1. Introduction
The gut microbiota are among the most important and active components of intestinal micro-ecology. Accordingly, this structure has recently become a research hotspot in various scientific fields. More than 1000 bacterial species are estimated to reside in the human intestine, and these organisms harbour up to 100 times as many genes as the human genome [
1]. The gut microbiota plays a very important role in host health. First, the microbiota colonises the intestinal mucosa and thus constitutes a natural protective barrier over the host intestine [
2]. Second, the microbiota digests food nutrients, particularly some non-digestible polysaccharides by the host, to provide energy to the host [
3]. Third, the microbiota inhibits the invasion and infection of pathogenic microorganisms by competing for intestinal nutrients and niches, as well as secreting bacteriostatic and antimicrobial substances [
4,
5]. Fourth, the microbiota synthesises beneficial metabolites, such as short-chain fatty acids (SCFAs) [
6], vitamin B12 and folic acid [
7]. Finally, the microbiota regulates the host immune system and promotes the maturation of immune cells [
8]. The composition of the gut microbiota is determined by the mutual adaptation of microorganisms and their hosts, and these interactions maintain the functional stability of the intestinal micro-ecological system [
9]. This system also involves metabolites produced by the gut microbiota, including SCFAs such as acetate, propionate and butyrate [
6]. These SCFAs can activate G protein-coupled receptors on the surfaces of host cells to regulate the growth, metabolism and functions of a variety of cells, including intestinal epithelial cells, adipocytes and leukocytes [
10,
11].
‘Lactic acid bacteria’ (LAB) refers to a group of bacteria that degrade carbohydrates to produce lactic acid, which has antimicrobial properties. Accordingly, humans have used LAB in the fermentation and preservation of foods and eaten for thousands of years. These traditional uses have led to the perception of most LAB as organisms generally regarded as safe (GRAS). Other studies have shown that some LAB species provide benefits to the host and are therefore considered probiotic. Most probiotic species exhibit resistance to acids, bile salts and enzymes in the gastrointestinal (GI) environment and can adhere to the intestinal lining to colonise the human GI tract [
12,
13].
Bacteriocins are small antimicrobial peptides produced by bacteria which may be subject to posttranslational modification. The production of bacteriocin is usually considered to be a probiotic trait. Studies have identified bacteriocin-producing LAB in the GI tracts of hosts [
14,
15,
16]. However, very few studies have investigated whether the production of bacteriocins by probiotic strains in the GI tract affect host health [
17,
18,
19], other than studies in which pathogenic bacteria were antagonised.
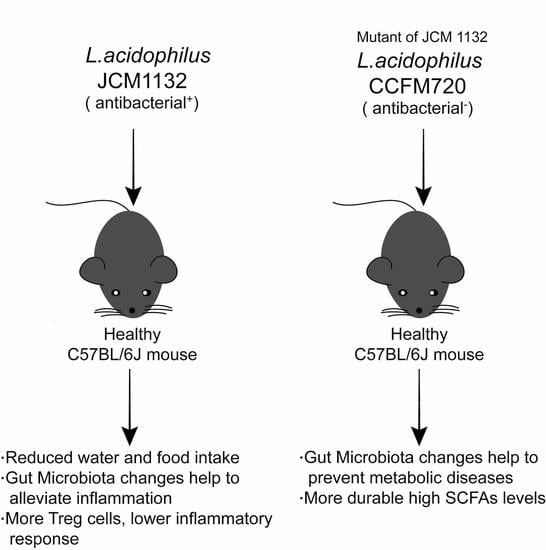
In this study, two strains of Lactobacillus acidophilus that share similar physiological characteristics but differ in bacteriocin production, JCM 1132 and CCFM 720, were investigated to assess their effects on the gut microbiota and metabolite composition in healthy mice. L. acidophilus CCFM 720 is a spontaneous mutant strain with no antimicrobial activity, obtained from the bacteriocin-producing strain L. acidophilus JCM 1132 after multiple passages. These two strains enabled the study of the different effects of phenotypically different strains of the same LAB species on the gut microbiota and metabolite composition.
4. Discussion
Some LAB may contribute to intestinal micro-ecological homeostasis and can improve host health [
31,
32,
33]. Bacteriocins are antimicrobial peptides produced by some LAB, which play important roles as probiotics. In recent years, increased research into LAB bacteriocins has led to the continuous development of bacteriocin-producing strains for use in food and manufacturing processes as a means of controlling microbiota communities in food systems and inhibiting the growth of pathogenic bacteria [
34,
35]. Moreover, these bacteriocin-producing LAB have been proposed as alternatives to antibiotics for the treatment of infections caused by pathogenic bacteria, including some drug-resistant bacteria. These LAB have a narrow spectrum and thus have little effect on the host gut microbiota [
36,
37]. However, few studies have evaluated the effects of bacteriocin-producing LAB on the healthy gut microbiota and metabolite composition. Only a few studies have mentioned the transient effects of LAB bacteriocin on the gut microbiota. For instance, neither Abp118-producing nor non-producing strains of
L. salivarius had significant effects on the gut microbiota at the phylum level in the pig and mouse, other than a significant decrease in
Spirochaete [
18]. Plantaricin, which is produced by
L. plantarum P-8, also induced a shift in the human faecal bacterial structure [
19]. In this study, we evaluate the effects of the bacteriocin-producing
L. acidophilus strain JCM 1132 and its bacteriocin-free spontaneous mutant CCFM 720 on the gut microbiota and metabolism in healthy mice. Although these strains have not previously been reported to have significant effects on body weight [
18,
38], the bacteriocin-producing
L. acidophilus strain significantly affected the food and water intake of the mice in our study, as well as some physiological and immunological indicators. An analysis of the gut microbiota revealed that treatment with
L. acidophilus JCM 1132 and CCFM 720 induced significant differences in the levels of several genera in the gut microbiota of healthy mice, as well as changes in SCFA production.
The bacteriocins synthesized by LAB can be roughly divided into three categories: Class I, Class II and Class III [
39,
40,
41]. The bacteriocin identified by BAGEL4 in the
L. acidophilus JCM 1132 genome was Acidocin J1132, a well-known pore-forming bacteriocin. Acidocin J1132 is a two-component Class IIb bacteriocin, formed by a subunit α and β. Acidocin J1132 has a narrow antimicrobial spectrum, with inhibitory effects only against some species of lactobacilli [
30]. Although the effect of Acidocin J1132 on the gut microbiota has not been studied, the effects of other Class II bacteriocins on the gut microbiota have been reported. In a previous study, five Class II bacteriocins with different antimicrobial spectra had little effect on the gut microbiota at the phylum level, whereas wide-spectrum bacteriocins induced transient changes at the genus level which were generally considered to be beneficial [
42]. In our study, despite the narrow antibacterial spectrum of bacteriocin from JCM 1132, the different effects of the
L. acidophilus JCM 1132 and CCFM 720 strains on certain genera in the gut microbiota persisted even one week after gavage withdrawal. Therefore, the effects of these LAB on gut microbiota may be more persistent than expected, possibly because of the formation of a new steady-state via competition within the gut microbiota. It is also necessary to consider the duration for which the LAB resides in the intestinal tract. For a potentially intestine-colonizing bacterium, the production of bacteriocin may determine the effect on the gut microbiota in situ. Although the generation time and adherence index to the Caco-2 cell line of two strains are similar in vitro, it is conceivable that the lack of antibacterial activity may affect the competitive growth of bacteria in the intestine and colonisation. The lack of antibacterial activity may have an all-round effect on the growth of
L. acidophilus in the intestine. Therefore, it can not be ruled out that some of the changes may be driven by altered growth/colonisation ability, but we believe that the altered growth/colonisation ability between the two strains, if any, may be largely driven by the loss of antibacterial activity.
In general, our results indicate that neither
L. acidophilus JCM 1132 nor CCFM 720 significantly affects the α diversity of the gut microbiota. This finding appears advantageous, as it suggests that probiotics do not induce large structural changes in the host gut microbiota. In contrast, the results of β diversity analyses revealed significant differences in the gut microbiota between the
L. acidophilus JCM 1132 and CCFM 720 groups. These differences were confirmed by changes at the phylum level, as the two strains differently affected the abundance of Actinobacteria, Verrucomicrobia and Tenericutes, and these changes had significantly different durations. Among them, the effect of
L. acidophilus on Actinobacteria may have been mainly due to its effect on
Bifidobacterium. Previously,
L. acidophilus was shown to increase the abundance of
Bifidobacterium in BALB/c mice [
43]. The presence of
Bifidobacterium, a typical beneficial bacteria, in the host intestinal tract often indicates an improvement in the intestinal environment [
44]. However, our results indicate that only treatment with
L. acidophilus CCFM 720 led to a significant increase in and the stable maintenance of the abundance of
Bifidobacterium in the mouse intestine. This observation suggests that the increased abundance of
Bifidobacterium induced by
L. acidophilus may be offset by bacteriocin. Specifically, this offset may be due not directly to the inhibition of
Bifidobacterium by bacteriocin, but rather to feedback from the inhibition of other intestinal bacteria. The beneficial bacteria
Akkermansia was also shown to be differentially affected by
L. acidophilus.
A. muciniphila, a member of Verrucomicrobia, is an important physiological regulator and next-generation probiotic that is beneficial to humans [
45,
46,
47]. In previous studies, a close relationship was observed between the abundance of
A. muciniphila and improved metabolic signs in mice, and supplementation with this strain could regulate high-fat diet-related glucose tolerance and relieve adipose tissue inflammation [
48,
49,
50]. Our results showed that gavage with
L. acidophilus JCM 1132 or CCFM 720 did not affect the slight increase in the abundance of
Akkermansia. Interestingly, the abundance of
Akkermansia in the JCM 1132 group increased significantly after a 1-week withdrawal period. This large increase may have been caused by a re-organization of the microbiota structure after
L. acidophilus administration. The effect of
L. acidophilus JCM 1132 on
Akkermansia may also guide approaches to the alleviation of metabolic diseases such as obesity and diabetes.
The intake of
L. acidophilus also reduced the Firmicutes/Bacteroidetes (F/B) ratio and the abundance of Tenericutes. Studies have shown that obesity might be linked to the gut microbiota composition in the host. The relative abundance of Firmicutes and Bacteroides in obese mice was shown to significantly increase and decrease, respectively, resulting in an increase in the harvesting of energy in the host [
51,
52]. A similar gut microbiota structure has also been observed in patients with type 2 diabetes and metabolic syndrome [
53]. The abundance of Tenericutes has been linked to obesity or type 2 diabetes-associated metabolic parameters [
54,
55]. Although the effects of reducing the F/B ratio did not appear to be directly related to the presence or absence of bacteriocin during gavage in our study, treatment with
L. acidophilus JCM 1132 had a delayed reducing effect on the F/B ratio. The difference in the reduction in Tenericutes abundance between the groups treated with the two
L. acidophilus strains only manifested after a 1-week withdrawal period. Moreover, the decreased abundance of Tenericutes associated with
L. acidophilus JCM 1132 treatment disappeared rapidly, indicating that this phenomenon might not be the result of direct inhibition by the
L. acidophilus JCM 1132. Regarding the mechanism by which
L. acidophilus JCM 1132 treatment affects
Akkermansia, we can speculate that bacteriocin induces changes in the gut microbiota by killing target bacteria directly. The subsequent disappearance of bacteriocin may then cause a rapid over-correction. The ability of
L. acidophilus to control/balance the population of beneficial gut microbiota that related to obesity, metabolic syndrome, diabetes etc. suggest that these two strains may help to maintain intestinal health and metabolic balance in the host. However, the targeted microbiota and ways of adjustment were different between these two strains, although the two strains did not seem to have significantly different effects on indicators of lipid metabolism.
Acidocin J1132 exhibits a narrow antibacterial spectrum against some
Lactobacillus species in vitro [
30]. In this study, the intrinsic
Lactobacillus strains in the gut were inhibited and substituted by
L. acidophilus JCM 1132 during the gavage period but rebounded slightly after a 1-week withdrawal period. This finding suggests that the administration of
L. acidophilus JCM 1132 triggered or replaced the growth of intrinsic
Lactobacillus, as suggested previously [
56]. The rapid decrease in the abundance of
Lactobacillus in the
L. acidophilus CCFM 720 group also indicated that the abundance of this species was maintained within a relatively stable range in the healthy gut.
L. acidophilus JCM 1132 does not manifest an outstanding capacity for colonization [
57], although this situation may be more prominent in different host species [
58]. Non-mouse-derived, non-bacteriocin-producing
L. acidophilus strains face a disadvantage with respect to the colonisation and replacement of native lactobacilli in the mouse intestine. Even in the presence of bacteriocin, a proportion of the intrinsic lactobacilli may be killed, but the abundance of
Lactobacillus will not change greatly. This characteristic is beneficial for maintaining homeostasis in the host gut microbiota.
Our analysis of other genera revealed that the effect of
L. acidophilus on the relative abundance of Bacteroidetes was mainly due to an increase in the abundance of
S24-7. However, the effects of
L. acidophilus on
S24-7,
Bacteroides and
Parabacteroides are unlikely to be related directly to the effect of bacteriocin. Previously, a combination of polysaccharide with probiotics (
L. acidophilus NCFM and
B. lactis Bi-07) was shown to decrease the proportion of
Parabacteroides in gut microbiota significantly in weaned rats [
59]. Antibacterial substances other than bacteriocins produced by LAB, such as organic acids and hydrogen peroxide, can also influence the intestinal environment [
60,
61]. However, we observed a considerable difference in the regulation of the relative
Rikenella abundance by
L. acidophilus between the
L. acidophilus JCM 1132 and CCFM 720 groups after a 1-week withdrawal period. Previous studies have revealed a negative correlation between the abundance of
Rikenella and body weight or fat mass [
62,
63]. This increase in the abundance of
Rikenella also confirms the role of
L. acidophilus in maintaining metabolic health in the host. As with the mechanism used to regulate the abundance of Tenericutes, treatment with
L. acidophilus CCFM 720 appeared to better support a lasting beneficial effect on host metabolism.
Additionally, gavage with the different bacterial strains led to differential regulation of the abundance of the Firmicutes genera
Turicibacter,
Oscillospira, rc4-4 and
Dorea. In particular,
L. acidophilus CCFM 720 more strongly inhibited
rc4-4 and
Dorea, and had a more persistent inhibitory effect on the latter genus. Moreover, after a 1-week withdrawal period, treatment with
L. acidophilus CCFM 720 was associated with an increase in the abundance of
Oscillospira, whereas treatment with JCM 1132 had a stronger inhibitory effect on
Ruminococcus. An increased abundance of
rc4-4 was shown to be associated with high-fat diet-induced obesity [
64,
65]. Moreover, several reports have described associations between a high abundance of
Dorea and some diseases [
66,
67,
68].
Ruminococcus is thought to be associated with inflammatory bowel disease, while
Oscillospira has been reported to improve weight loss [
69,
70]. As with the F/B ratio, Tenericutes and
Rikenella, the altered patterns of
rc4-4, Dorea, and
Oscillospira suggest that treatment with
L. acidophilus CCFM 720 may have a more lasting beneficial effect on metabolism. The effects of the two
L. acidophilus strains on the levels of beneficial SCFAs after a 1-week withdrawal period also supported this conjecture.
In contrast to the above conclusions, however, treatment with either JCM 1132 or CCFM 720 led to a decreased abundance of
Anaeroplasma, which was reported to be correlated negatively with obesity and hypercholesterolemia [
71,
72]. Additionally, CCFM 720 had a more persistent inhibitory effect. In our experiments, only
L. acidophilus JCM 1132 had a beneficial regulatory effect on the abundance of
Turicibacter which was probably due to the reversal of the inhibitory effect of bacteriocin on
Turicibacter.
Turicibacter has been reported to alleviate the symptoms of hyperlipidaemia and relieve inflammation in the body [
73,
74]. Treatment with
L. acidophilus JCM 1132 also more strongly inhibited species of the pro-inflammatory genus
Ruminococcus [
69]. The ability of JCM 1132 treatment to regulate the abundance of
Turicibacter and
Ruminococcus may also explain why this strain more effectively promoted an immunosuppressive response.
Our results suggest that
L. acidophilus JCM 1132 and CCFM 720 have different effects on the gut microbiota of healthy mice, as well as some physiological indicators, and that some of these differences persist even one week after treatment withdrawal. Although previous studies have suggested that the differential effects of bacteria with or without bacteriocins on the gut microbiota would disappear at the end of the experimental process, our results indicate persistent effects on some phyla or genera, despite significant effects on the gut microbiota diversity. High doses of bacteriocin may pose a health risk to the host [
75]. Therefore, although LAB are GRAS, our research suggests that further studies of the effects of bacteriocin associated with LAB applications are needed. However, it must be acknowledged that, because of the currently unavailable on genetic modification on
L. acidophilus JCM 1132 or CCFM 720 in this research, there is a potential possibility that CCFM 720 is not a perfect bacteriocin non-producing control for JCM 1132. Although no obvious differences in the other physiological characteristics of the two strains have been found in addition to the loss of antibacterial ability and putative β peptide of Acidocin J1132, we cannot rule out the potential impacts of the other three missing putative proteins on the physiological characteristics of
L. acidophilus JCM1132. The underlying factors which impacted mice gut microbiota other than bacteriocin cannot be completely ruled out, such as colonisation and metabolite production. Therefore, further attempts to genetically modify the Acidocin encoding loci in JCM1132 or complement CCFM 720 with this locus from JCM1132 will help us to more convincingly determine the presence or absence of bacteriocin on the effects by
L. acidophilus on healthy mice. In addition, further work would be needed to assess which of the observed outcomes are due to the direct effect of different doses of the bacteriocin, once purified, and which are indirect effects linked to modifications on the gut microbiota structure.