Transcriptional Regulation of the Outer Membrane Protein A in Acinetobacter baumannii
Abstract
:1. Introduction
2. Materials and Methods
2.1. Bacterial Strains, Plasmids, and Culture Conditions
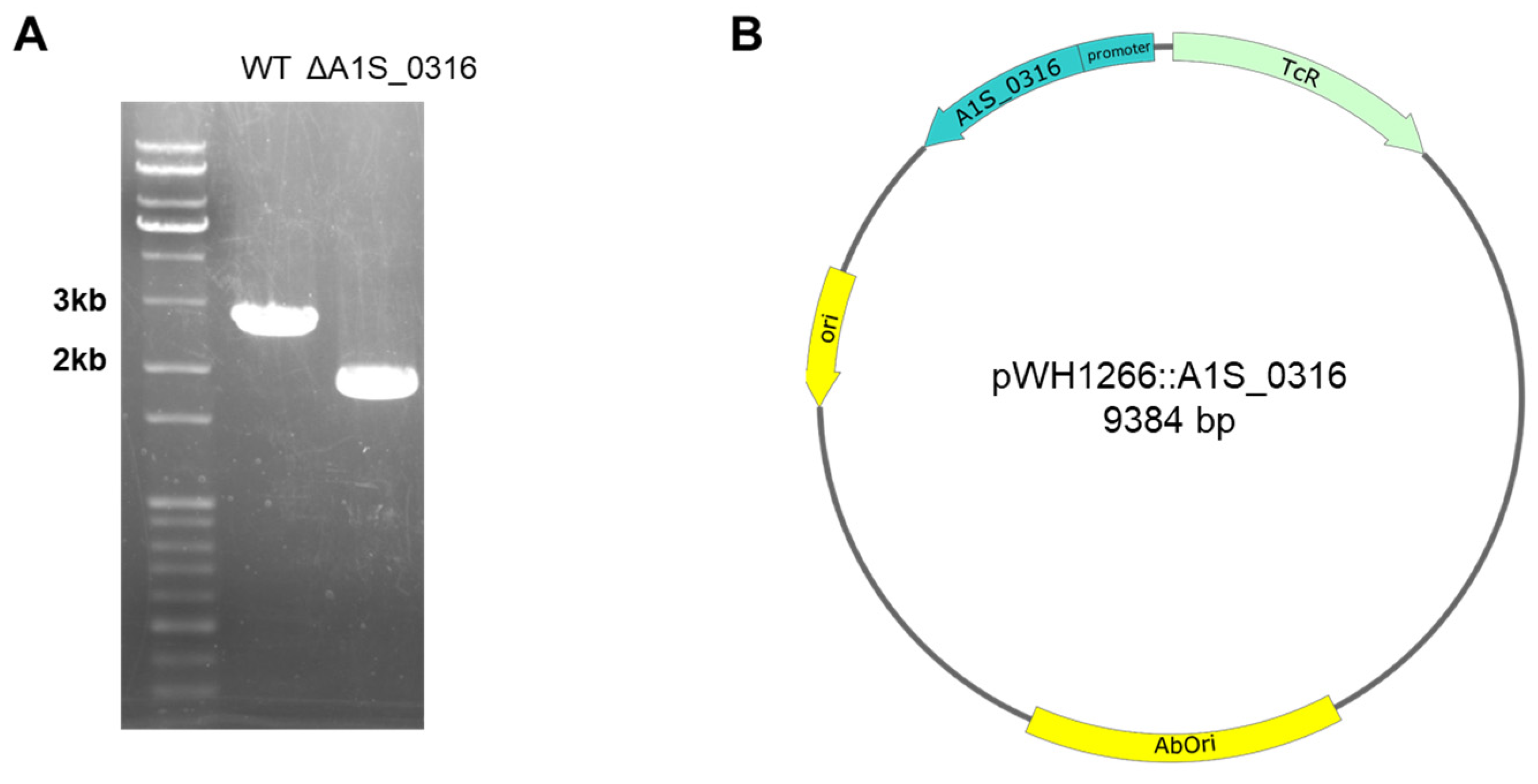
2.2. Construction of the ΔA1S_0316 Mutant Strain
2.3. Construction of a ΔA1S_0316 Complementary Strain Using pWH1266:A1S_0316
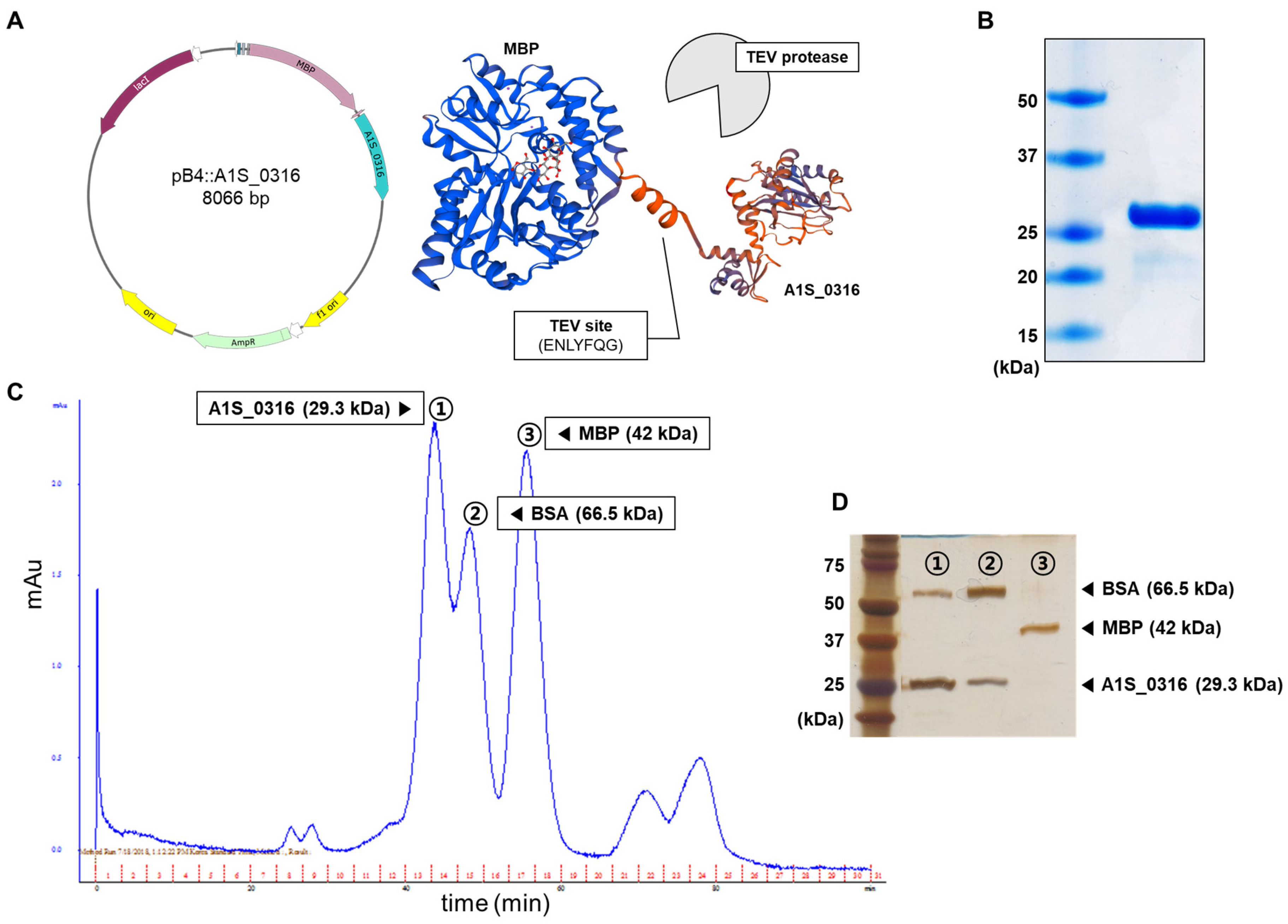
2.4. Construction of pB4:A1S_0316
2.5. Construction of pSA508:AbOmpAp for the In Vitro Transcription Assay
2.6. Primer Extension Assay
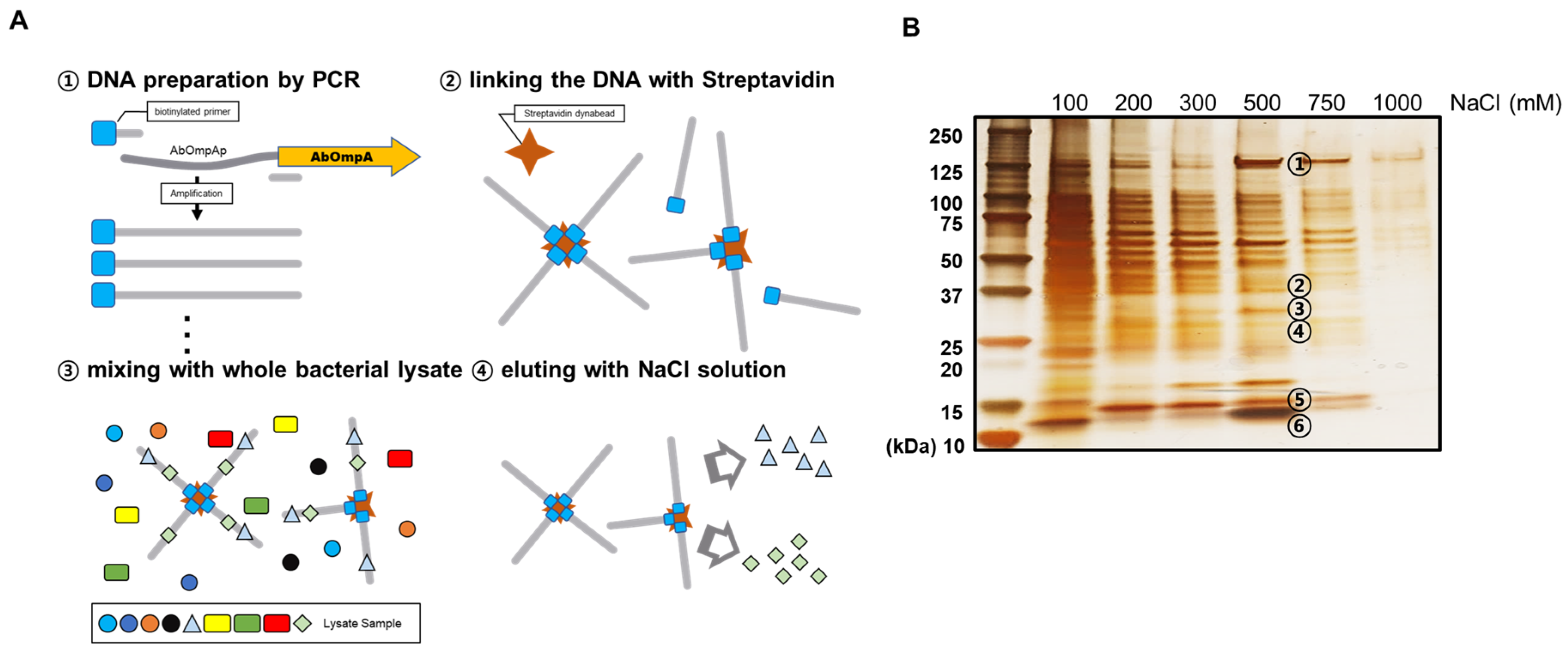
2.7. DNA Affinity Chromatography
2.8. Purification of A1S_0316
2.9. Electrophoretic Mobility Shift Assay (EMSA)
2.10. Real-Time qPCR
2.11. Western Blotting of the Bacterial Membrane Fraction
2.12. In Vitro Transcription Assay
3. Results
3.1. Characterization of the OmpA Promoter Region in A. baumannii
3.2. Identification of Proteins that Bind to the OmpA Promoter
3.3. Molecular Features of A1S_0316
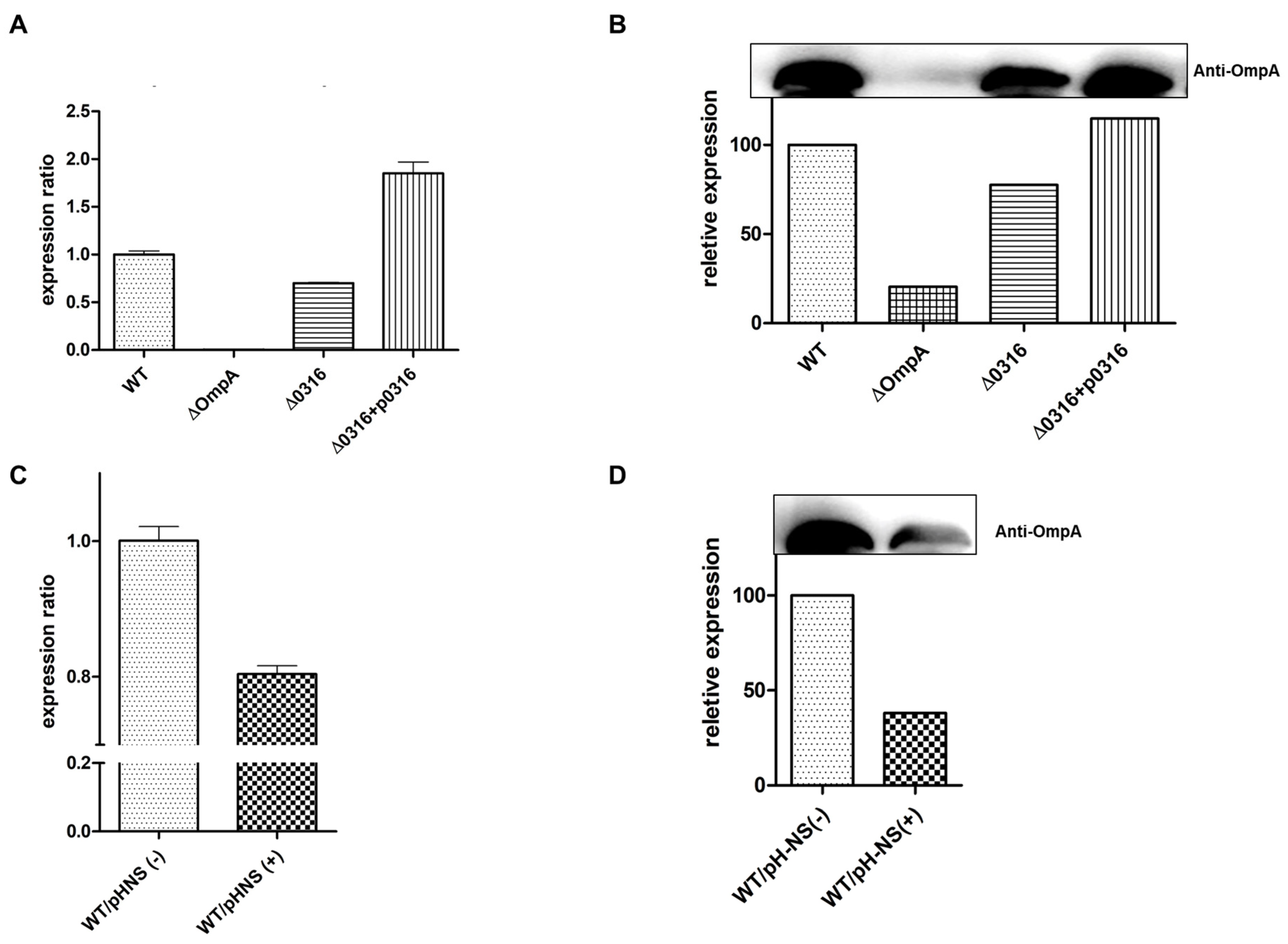
3.4. Effect of A1S_0316 and AbH-NS on OmpA Expression In Vivo
3.5. Effect of A1S_0316 and AbH-NS on OmpA Expression In Vitro
4. Discussion
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Doughari, H.J.; Ndakidemi, P.A.; Human, I.S.; Benade, S. The Ecology, Biology and Pathogenesis of Acinetobacter spp.: An Overview. Microbes Environ. 2011, 26, 101–112. [Google Scholar] [CrossRef] [Green Version]
- Dent, L.L.; Marshall, D.R.; Pratap, S.; Hulette, R.B. Multidrug resistant Acinetobacter baumannii: A descriptive study in a city hospital. BMC Infect. Dis. 2010, 10, 196. [Google Scholar] [CrossRef] [Green Version]
- Rahal, J.J. Novel Antibiotic Combinations against Infections with Almost Completely Resistant Pseudomonas aeruginosa and Acinetobacter Species. Clin. Infect. Dis. 2006, 43, S95–S99. [Google Scholar] [CrossRef] [Green Version]
- Boucher, H.W.; Talbot, G.H.; Bradley, J.S.; Edwards, J.E.; Gilbert, D.; Rice, L.B.; Scheld, M.; Spellberg, B.; Bartlett, J. Bad Bugs, No Drugs: No ESKAPE! An Update from the Infectious Diseases Society of America. Clin. Infect. Dis. 2009, 48, 1–12. [Google Scholar] [CrossRef] [Green Version]
- Koebnik, R.; Locher, K.P.; Van Gelder, P. Structure and function of bacterial outer membrane proteins: Barrels in a nutshell. Mol. Microbiol. 2000, 37, 239–253. [Google Scholar] [CrossRef] [PubMed]
- Smith, S.G.J.; Mahon, V.; Lambert, M.A.; Fagan, R.P. A molecular Swiss army knife: OmpA structure, function and expression. FEMS Microbiol. Lett. 2007, 273, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Nilsson, G.; Belasco, J.G.; Cohen, S.N.; von Gabain, A. Growth-rate dependent regulation of mRNA stability in Escherichia coli. Nature 1984, 312, 75–77. [Google Scholar] [CrossRef] [PubMed]
- Rasmussen, A.A.; Eriksen, M.; Gilany, K.; Udesen, C.; Franch, T.; Petersen, C.; Valentin-Hansen, P. Regulation of OmpA mRNA stability: The role of a small regulatory RNA in growth phase-dependent control. Mol. Microbiol. 2005, 58, 1421–1429. [Google Scholar] [CrossRef] [PubMed]
- Udekwu, K.I.; Darfeuille, F.; Vogel, J.; Reimegard, J.; Holmqvist, E.; Wagner, E.G. Hfq-dependent regulation of OmpA synthesis is mediated by an antisense RNA. Genes Dev. 2005, 19, 2355–2366. [Google Scholar] [CrossRef] [Green Version]
- Vytvytska, O.; Moll, I.; Kaberdin, V.R.; von Gabain, A.; Blasi, U. Hfq (HF1) stimulates OmpA mRNA decay by interfering with ribosome binding. Genes Dev. 2000, 14, 1109–1118. [Google Scholar]
- Murphy, E.R.; Rossmanith, J.; Sieg, J.; Fris, M.E.; Hussein, H.; Kouse, A.B.; Gross, K.; Zeng, C.; Hines, J.V.; Narberhaus, F.; et al. Regulation of OmpA Translation and Shigella dysenteriae Virulence by an RNA Thermometer. Infect. Immun. 2020, 88, e00871-19. [Google Scholar] [CrossRef]
- Oh, M.H.; Lee, J.C.; Kim, J.; Choi, C.H.; Han, K. Simple Method for Markerless Gene Deletion in Multidrug-Resistant Acinetobacter baumannii. Appl. Environ. Microbiol. 2015, 81, 3357–3368. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Aslanidis, C.; de Jong, P.J. Ligation-independent cloning of PCR products (LIC-PCR). Nucleic Acids Res. 1990, 18, 6069–6074. [Google Scholar] [CrossRef] [PubMed]
- Sun, J.; Jeon, J.H.; Shin, M.; Shin, H.C.; Oh, B.H.; Kim, J.S. Crystal structure and CRISPR RNA-binding site of the Cmr1 subunit of the Cmr interference complex. Acta Crystallogr. D Biol. Crystallogr. 2014, 70, 535–543. [Google Scholar] [CrossRef]
- Choy, H.E.; Adhya, S. RNA polymerase idling and clearance in gal promoters: Use of supercoiled minicircle DNA template made in vivo. Proc. Natl. Acad. Sci. USA 1993, 90, 472–476. [Google Scholar] [CrossRef] [Green Version]
- Carey, M.F.; Peterson, C.L.; Smale, S.T. The Primer Extension Assay. Cold Spring Harb. Protoc. 2013, 2013. [Google Scholar] [CrossRef] [Green Version]
- Kim, B.N.; Shin, M.; Ha, S.C.; Park, S.Y.; Seo, P.W.; Hofmann, A.; Kim, J.S. Crystal structure of an ASCH protein from Zymomonas mobilis and its ribonuclease activity specific for single-stranded RNA. Sci. Rep. 2017, 7, 12303. [Google Scholar] [CrossRef] [Green Version]
- Shin, M. The mechanism underlying Ler-mediated alleviation of gene repression by H-NS. Biochem. Biophys. Res. Commun. 2017, 483, 392–396. [Google Scholar] [CrossRef]
- Navarre, W.W. The Impact of Gene Silencing on Horizontal Gene Transfer and Bacterial Evolution. Adv. Microb. Physiol. 2016, 69, 157–186. [Google Scholar] [CrossRef]
- Herrou, J.; Czyz, D.M.; Fiebig, A.; Willett, J.W.; Kim, Y.; Wu, R.; Babnigg, G.; Crosson, S. Molecular control of gene expression by Brucella BaaR, an IclR-type transcriptional repressor. J. Biol. Chem. 2018, 293, 7437–7456. [Google Scholar] [CrossRef] [Green Version]
- Mortensen, B.L.; Fuller, J.R.; Taft-Benz, S.; Kijek, T.M.; Miller, C.N.; Huang, M.T.; Kawula, T.H. Effects of the putative transcriptional regulator IclR on Francisella tularensis pathogenesis. Infect. Immun. 2010, 78, 5022–5032. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Stoebel, D.M.; Free, A.; Dorman, C.J. Anti-silencing: Overcoming H-NS-mediated repression of transcription in Gram-negative enteric bacteria. Microbiology 2008, 154, 2533–2545. [Google Scholar] [CrossRef] [PubMed]
- Perez, J.C.; Latifi, T.; Groisman, E.A. Overcoming H-NS-mediated transcriptional silencing of horizontally acquired genes by the PhoP and SlyA proteins in Salmonella enterica. J. Biol. Chem. 2008, 283, 10773–10783. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kim, S.W.; Oh, M.H.; Jun, S.H.; Jeon, H.; Kim, S.I.; Kim, K.; Lee, Y.C.; Lee, J.C. Outer membrane Protein A plays a role in pathogenesis of Acinetobacter nosocomialis. Virulence 2016, 7, 413–426. [Google Scholar] [CrossRef] [Green Version]
- Kwon, H.I.; Kim, S.; Oh, M.H.; Na, S.H.; Kim, Y.J.; Jeon, Y.H.; Lee, J.C. Outer membrane protein A contributes to antimicrobial resistance of Acinetobacter baumannii through the OmpA-like domain. J. Antimicrob. Chemother. 2017, 72, 3012–3015. [Google Scholar] [CrossRef]
- Nie, D.; Hu, Y.; Chen, Z.; Li, M.; Hou, Z.; Luo, X.; Mao, X.; Xue, X. Outer membrane protein A (OmpA) as a potential therapeutic target for Acinetobacter baumannii infection. J. Biomed. Sci. 2020, 27, 26. [Google Scholar] [CrossRef] [Green Version]




© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Oh, K.-W.; Kim, K.; Islam, M.M.; Jung, H.-W.; Lim, D.; Lee, J.C.; Shin, M. Transcriptional Regulation of the Outer Membrane Protein A in Acinetobacter baumannii. Microorganisms 2020, 8, 706. https://doi.org/10.3390/microorganisms8050706
Oh K-W, Kim K, Islam MM, Jung H-W, Lim D, Lee JC, Shin M. Transcriptional Regulation of the Outer Membrane Protein A in Acinetobacter baumannii. Microorganisms. 2020; 8(5):706. https://doi.org/10.3390/microorganisms8050706
Chicago/Turabian StyleOh, Kyu-Wan, Kyeongmin Kim, Md. Maidul Islam, Hye-Won Jung, Daejin Lim, Je Chul Lee, and Minsang Shin. 2020. "Transcriptional Regulation of the Outer Membrane Protein A in Acinetobacter baumannii" Microorganisms 8, no. 5: 706. https://doi.org/10.3390/microorganisms8050706





