Haplotype Diversity of NADPH-Cytochrome P450 Reductase Gene of Ophiocordyceps sinensis and the Effect on Fungal Infection in Host Insects
Abstract
:1. Introduction
2. Materials and Methods
2.1. Sampling
2.2. DNA Extraction and Sequencing
2.3. Data Analyses
3. Results
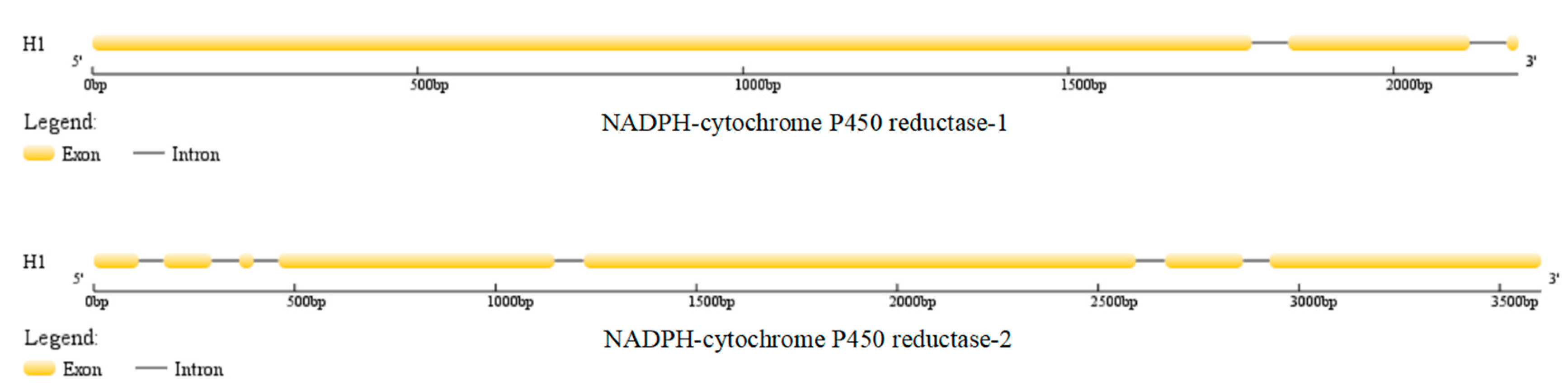
3.1. Gene Sequence Characteristics and Protein Structures
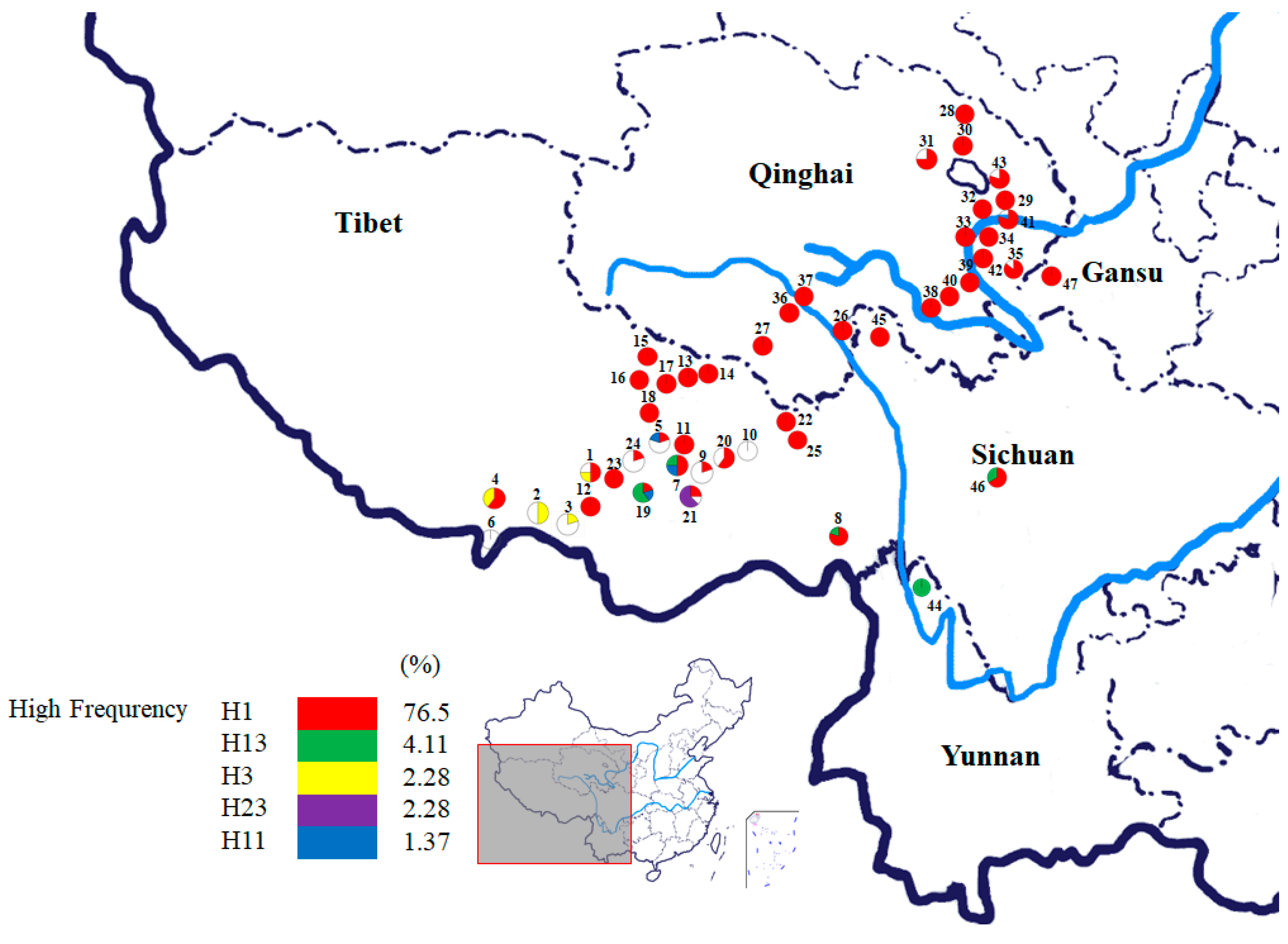
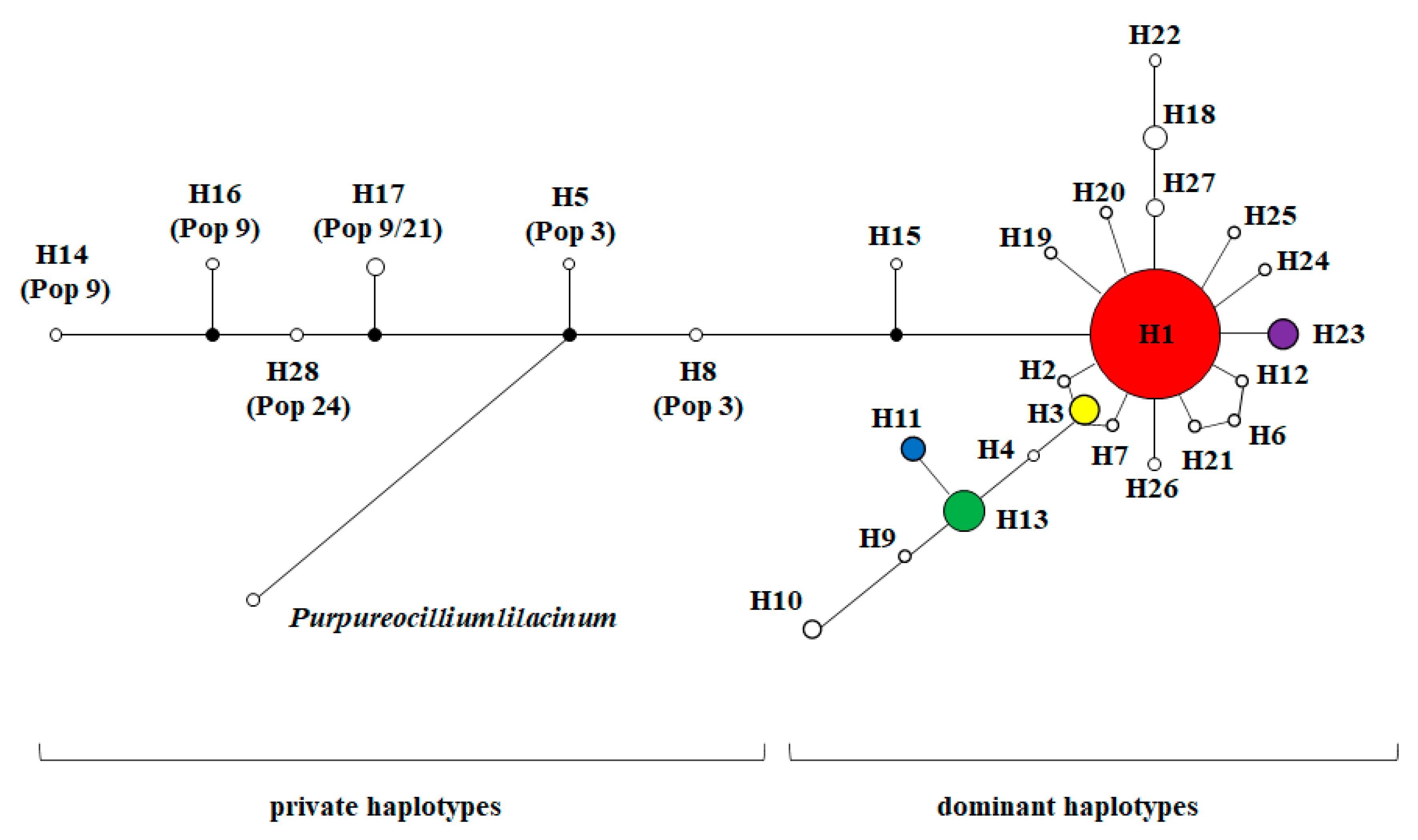
3.2. Haplotypes Diversity of NADPH CPR Genes
3.3. Functional Analysis of NADPH CPR
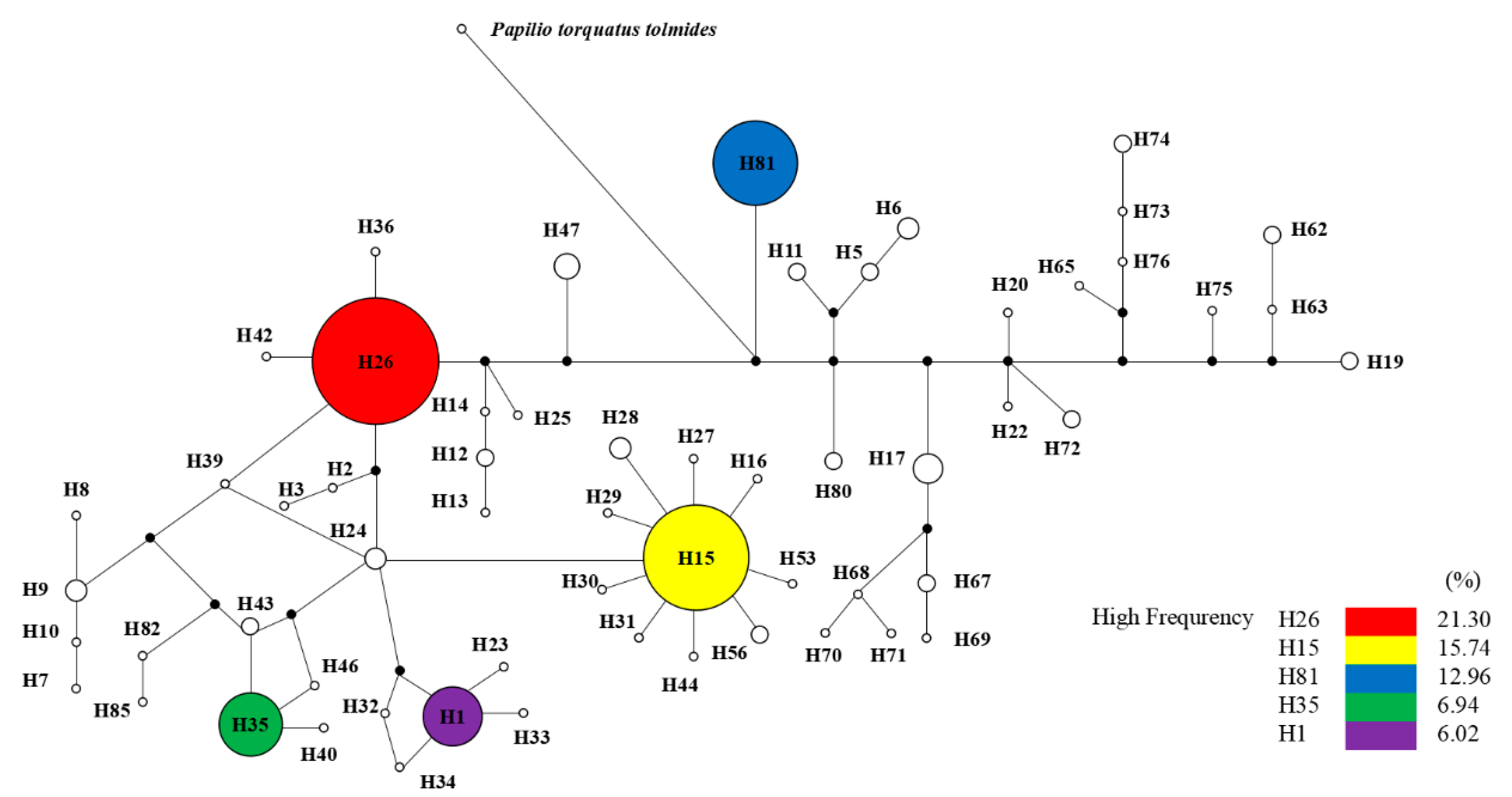
3.4. Correspondence between O. sinensis and Its Host Insects
4. Discussion
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Zhu, J.S.; Halpern, G.M.; Jones, K. The scientific rediscovery of an ancient Chinese herbal medicine: Cordyceps sinensis Part I. J. Altern. Complement. Med. 1998, 4, 289–303. [Google Scholar] [CrossRef] [PubMed]
- Sung, G.H.; Hyweljones, N.L.; Sung, J.M.; Luangsaard, J.J.; Shrestha, B.; Spatafora, J.W. Phylogenetic classification of Cordyceps and the clavicipitaceous fungi. Stud. Mycol. 2007, 57, 5–59. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hu, X.; Zhang, Y.J.; Xiao, G.H.; Zheng, P.; Xia, Y.L.; Zhang, X.Y.; St Leger, R.J.; Liu, X.Z.; Wang, C.S. Genome survey uncovers the secrets of sex and lifestyle in caterpillar fungus. Chin. Sci. Bull. 2013, 58, 2846–2854. [Google Scholar] [CrossRef] [Green Version]
- Yang, D.R.; Yang, Y.X.; Shen, F.R.; Dong, D.Z.; Yu, R.Q.; Lu, Z.; Chun, S.S. Studied on Hepialid larvae infection by Cordyceps sinensis. In Proceedings of the First National Symposium on Entomogenous Fungi, Gongzhuling, Jilin, China, 26–29 August 1986; Fungal Society of China, Botanical Society of China: Bejing, China, 1986; pp. 248–252. [Google Scholar]
- Zeng, W.; Yin, D.H.; Li, Q.S.; Li, L. The growth of Cordyceps sinensis (Berk.) Sacc. in the infection and parasitic phases. Mycosystema 2006, 25, 646–650. [Google Scholar] [CrossRef]
- Zhong, X.; Peng, Q.Y.; Song, L.S.; Chen, H.; Sun, H.X.; Zhang, G.R.; Liu, X. Detection of Ophiocordyceps sinensis in the roots of plants in alpine meadows by nested-touchdown polymerase chain reaction. Fungal Biol. 2014, 118, 359–363. [Google Scholar] [CrossRef]
- Guo, L.X.; Hong, Y.H.; Zhou, Q.Z.; Zhu, Q.; Xu, X.M.; Wang, J.H. Fungus-larva relation in the formation of Cordyceps sinensis as revealed by stable carbon isotope analysis. Sci. Rep. 2017, 7, 7789:1–7789:10. [Google Scholar] [CrossRef] [Green Version]
- Lei, W.; Zhang, G.R.; Peng, Q.Y.; Liu, X. Development of Ophiocordyceps sinensis through plant-mediated interkingdom host colonization. Int. J. Mol. Sci. 2015, 16, 17482–17493. [Google Scholar] [CrossRef]
- Yang, Y.X.; Yang, D.R.; Shen, F.R.; Dong, D.Z. Studies on hepialid larvae for being infected by Chinese “insect herb” fungus (Cordyceps sinensis). Zool. Res. 1989, 10, 227–231. [Google Scholar]
- Tu, Y.Q.; Zhang, D.L.; Zeng, W.; Chen, S.J.; Yin, D.H. An experiment of infecting hepialus larvae with Cordyceps sinensis. Edible Fungi 2010, 32, 16–17. [Google Scholar] [CrossRef]
- Guo, L.X.; Xu, X.M.; Liang, F.R.; Yuan, J.P.; Peng, J.; Wu, C.F.; Wang, J.H.; Rita, G. Morphological observations and fatty acid composition of indoor-cultivated Cordyceps sinensis at a high-altitude laboratory on Sejila mountain, Tibet. PLoS ONE 2015, 10, e0126095:1–e0126095:15. [Google Scholar] [CrossRef]
- Xia, E.H.; Yang, D.R.; Jiang, J.J.; Zhang, Q.J.; Liu, Y.; Liu, Y.L.; Zhang, Y.; Zhang, H.B.; Shi, C.; Tong, Y.; et al. The caterpillar fungus, Ophiocordyceps sinensis, genome provides insights into highland adaptation of fungal pathogenicity. Sci. Rep. 2017, 7, 1806:1–1806:11. [Google Scholar] [CrossRef]
- Alka, S.; Elizabeth, M.J.G.; Paul, V.B. Direct electrochemistry of human and rat NADPH cytochrome P450 reductase. Electrochem. Commun. 2006, 8, 1845–1849. [Google Scholar] [CrossRef]
- Esteves, F.; Campelo, D.; Gomes, B.C.; Urban, P.; Bozonnet, S.; Lautier, T.; Rueff, J.; Truan, G.; Kranendonk, M. The role of the FMN-domain of human cytochrome P450 oxidoreductase in its promiscuous interactions with structurally diverse redox partners. Front. Pharmacol. 2020, 11, 299:1–299:16. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Siewers, V.; Viaud, M.; Jimenez-Teja, D.; Collado, I.G.; Gronover, C.S.; Pradier, J.M.; Tudzynski, B.; Tudzynski, P. Functional analysis of the cytochrome P450 monooxygenase gene bcbot1 of Botrytis cinerea indicates that botrydial is a strain-specific virulence factor. Mol. Plant Microbe Interact. 2005, 18, 602–612. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Moraga, J.; Dalmais, B.; Izquierdo-Bueno, I.; Aleu, J.; Hanson, J.R.; Hernandez-Galan, R.; Viaud, M.; Collado, I.G. Genetic and molecular basis of botrydial biosynthesis: Connecting cytochrome P450-encoding genes to biosynthetic intermediates. ACS Chem. Biol. 2016, 11, 2838–2846. [Google Scholar] [CrossRef] [Green Version]
- Xie, J.; Li, S.; Mo, C.; Xiao, X.; Peng, D.; Wang, G.; Xiao, Y. Genome and transcriptome sequences reveal the specific parasitism of the nematophagous Purpureocillium lilacinum 36-1. Front. Microbiol. 2016, 7, 1084:1–1084:32. [Google Scholar] [CrossRef]
- Liang, H.H.; Cheng, Z.; Yang, X.L.; Shan, L.; Zhou, T.S.; Zhang, W.J.; Chen, J.K. Genetic variation and aiffinity of Cordyceps sinensis in Qinghai province based on analysis of morphologic characters and inter-simple sequence repeat markers. Chin. Tradit. Herb. Drugs 2005, 12, 1859–1864. [Google Scholar]
- Quan, Q.M.; Chen, L.L.; Wang, X.; Li, S.; Yang, X.L.; Zhu, Y.G.; Wang, M.; Cheng, Z. Genetic diversity and distribution patterns of host insects of caterpillar fungus Ophiocordyceps sinensis in the Qinghai-Tibet Plateau. PLoS ONE 2014, 9, e92293:1–e92293:10. [Google Scholar] [CrossRef] [Green Version]
- Kumar, S.; Stecher, G.; Tamura, K. MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol. Biol. Evol. 2016, 33, 1870–1874. [Google Scholar] [CrossRef] [Green Version]
- Rozas, J.; Ferrer-Mata, A.; Sanchez-DelBarrio, J.C.; Guirao-Rico, S.; Librado, P.; Ramos-Onsins, S.E.; Sanchez-Gracia, A. DnaSP 6: DNA sequence polymorphism analysis of large data sets. Mol. Biol. Evol. 2017, 34, 3299–3302. [Google Scholar] [CrossRef]
- Bandelt, H.J.; Forster, P.; Rohl, A. Median-joining networks for inferring intraspecific phylogenies. Mol. Biol. Evol. 1999, 16, 37–48. [Google Scholar] [CrossRef] [PubMed]
- Peakall, R.; Smouse, P.E. GenAlEx 6.5: genetic analysis in Excel. Population genetic software for teaching and research-an update. Bioinformatics 2012, 28, 2537–2539. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mantel, N. The detection of disease clustering and a generalized regression approach. Cancer Res. 1967, 27, 209–220. [Google Scholar] [PubMed]
- Schuh, A.; Becq, J.; Humphray, S.; Alexa, A.; Burns, A.; Clifford, R.; Feller, S.M.; Grocock, R.; Henderson, S.; Khrebtukova, I.; et al. Monitoring chronic lymphocytic leukemia progression by whole genome sequencing reveals heterogeneous clonal evolution patterns. Blood 2012, 120, 4191–4196. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Venter, J.C.; Adams, M.D.; Myers, E.W. The sequence of the human genome. Science 2001, 291, 1304–1351. [Google Scholar] [CrossRef] [Green Version]
- Yu, J.; Hu, S.; Wang, J.; Wong, G.; Li, S.; Liu, B.; Deng, Y.; Dai, L.; Zhou, Y.; Zhang, X. A draft sequence of the rice genome (Oryza sativa L. ssp. indica). Science 2002, 296, 92–100. [Google Scholar] [CrossRef]
- Zhang, Y.J.; Zhang, S.; Li, Y.L.; Ma, S.L.; Wang, C.S.; Xiang, M.C.; Liu, X.; An, Z.Q.; Xu, J.P.; Liu, X.Z. Phylogeography and evolution of a fungal-insect association on the Tibetan Plateau. Mol. Ecol. 2014, 23, 5337–5355. [Google Scholar] [CrossRef]
- Quan, Q.M.; Wang, Q.X.; Zhou, X.L.; Li, S.; Yang, X.L.; Zhu, Y.G.; Cheng, Z. Comparative phylogenetic relationships and genetic structure of the caterpillar fungus Ophiocordyceps sinensis and its host insects inferred from multiple gene sequences. J. Microbiol. 2014, 52, 99–105. [Google Scholar] [CrossRef]
- Shen, A.L.; Porter, T.D.; Wilson, T.E.; Kasper, C.B. Structural analysis of the FMN binding domain of NADPH-cytochrome P450 oxidoreductase by site-directed mutagenesis. J. Biol. Chem. 1989, 264, 7584–7589. [Google Scholar]
- Hertz, E.P.T.; Kruse, T.; Davey, N.E.; López-Méndez, B.; Sigurðsson, J.O.; Montoya, G.; Olsen, J.V.; Nilsson, J. A conserved motif provides binding specificity to the PP2A-B56 phosphatase. Mol. Cell 2016, 63, 686–695. [Google Scholar] [CrossRef] [Green Version]
- Kushiro, T.; Okamoto, M.; Nakabayashi, K.; Yamagishi, K.; Kitamura, S.; Asami, T.; Hirai, N.; Koshiba, T.; Kamiya, Y.; Nambara, E. The Arabidopsis cytochrome P450 CYP707A encodes ABA 8’-hydroxylases: Key enzymes in ABA catabolism. EMBO J. 2004, 23, 1647–1656. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wang, M.; Roberts, D.L.; Paschke, R.; Shea, T.M.; Masters, B.S.; Kim, J.J. Three-dimensional structure of NADPH-cytochrome P450 reductase: prototype for FMN- and FAD-containing enzymes. Proc. Natl. Acad. Sci. USA 1997, 94, 8411–8416. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Suwanchaichinda, C.; Brattsten, L.B. Genomic and bioinformatic analysis of NADPH-cytochrome P450 reductase in Anopheles stephensi (Diptera: Culicidae). J. Insect Sci. 2014, 14, 165:1–165:8. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chen, Y.Q.; Wang, N.; Qu, L.; Li, T.; Zhang, W. Determination of the anamorph of Cordyceps sinensis inferred from the analysis of the ribosomal DNA internal transcribed spacers and 5.8S rDNA. Biochem. Syst. Ecol. 2001, 29, 597–607. [Google Scholar] [CrossRef]
- Xiao, W.; Yang, J.L.; Ping, Z.; Di, C.K.; Xia, H.H.; Xin, Z.H.; Wang, Q. Non-support of species complex hypothesis of Cordyceps sinensis by targeted rDNA-ITS sequence analysis. Mycosystema 2009, 28, 724–730. [Google Scholar] [CrossRef]
- Zhang, P.; Cui, S.H.; Liu, B.; Ren, X.; Kang, S.; Wei, F.; Ma, S.C. Discriminatory power evaluation of nuclear ribosomal rna barcoding sequences through Ophiocordyceps sinensis related samples. Front. Microbiol. 2018, 9, 2498:1–2498:8. [Google Scholar] [CrossRef] [Green Version]
- Wang, X.L.; Yao, Y.J. Host insect species of Ophiocordyceps sinensis: A review. ZooKeys 2011, 127, 43–49. [Google Scholar] [CrossRef] [Green Version]
- Li, S.P.; Tsim, K.W.K. The biological and pharmacological properties of Cordyceps sinensis, a traditional Chinese medicine that has broad clinical applications. In Herbal and Traditional Medicine: Molecular Aspects of Health; Wachtel-Galor, S., Ed.; CRC Press: New York, NY, USA, 2004; pp. 657–686. [Google Scholar]
- Yang, D.R.; Li, C.D.; Shu, C.; Yang, Y.X. Studies on the Chinese species of the genus Hepialus and their geographical distribution. Acta Entomol. Sin. 1996, 39, 413–422. [Google Scholar] [CrossRef]
- St. Leger, R.J. Polydnavirus Genome Organization. In Parasites and Pathogens of Insects; Beckage, N.E., Thompson, S.N., Federici, B.A., Eds.; Academic Press: San Diego, CA, USA, 1993; Volume 2, pp. 211–229. [Google Scholar]
- Duplessis, S.; Cuomo, C.A.; Lin, Y.C.; Aerts, A.; Tisserant, E.; Veneault-Fourrey, C.; Joly, D.L.; Francis Martina, F. Obligate biotrophy features unraveled by the genomic analysis of rust fungi. Proc. Natl. Acad. Sci. USA 2011, 108, 9166–9171. [Google Scholar] [CrossRef] [Green Version]
- Cresnar, B.; Petric, S. Cytochrome P450 enzymes in the fungal kingdom. Biochim. Biophys. Acta 2011, 1814, 29–35. [Google Scholar] [CrossRef]



| Position | H1 | H5 | H8 | H14 | H16 | H17 | H28 |
|---|---|---|---|---|---|---|---|
| 335 | T | T | - | T | - | T | T |
| 397 | T | - | - | T | - | - | - |
| 401 | S | - | - | S | - | - | - |
| 425 | - | S | S | - | S | S | S |
| 443 | - | T | T | - | T | T | T |
| 454 | - | S | S | - | S | S | S |
| 636 | T | T | T | - | - | T | - |
| Position | H1 | H4 | H10 | H12 | H29 |
|---|---|---|---|---|---|
| 23 | T | - | - | - | - |
| 24 | T | - | - | - | - |
| 25 | S | - | - | - | - |
| 189 | - | S | S | S | S |
| 252 | S | - | - | - | - |
| 257 | - | T | T | T | T |
| 437 | - | Y | Y | Y | - |
| 539 | - | Y | Y | Y | - |
| 777 | S | - | S | S | S |
| 881 | S | S | - | - | S |
| Functional Site | H1 | H5 | H8 | H14 | H16 | H17/H28 | ||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Position | Sequence | Position | Sequence | Position | Sequence | Position | Sequence | Position | Sequence | Position | Sequence | |
| N-gly | 167–170 | NNTY | 167–170 | NNTY | 167–170 | NNTY | 167–170 | NNTY | 167–170 | NNTY | 167–170 | NNTY |
| 244–247 | NLTR | 244–247 | NLTR | 244–247 | NLTR | 244–247 | NLTR | 244–247 | NLTR | 244–247 | NLTR | |
| 305–308 | NLTY | 305–308 | NLTY | 305–308 | NLTY | 305–308 | NLTY | 305–308 | NLTY | 305–308 | NLTY | |
| cA/GMP | 461–464 | KKIS | 461–464 | KKIS | 461–464 | KKIS | 461–464 | KKIS | 461–464 | KKIS | 461–464 | KKIS |
| 581–584 | RKRT | 581–584 | RKRT | 581–584 | RKRT | 581–584 | RKRT | 581–584 | RKRT | 581–584 | RKRT | |
| PK C | 61–63 | SGK | 61–63 | SGK | 61–63 | SGK | 61–63 | SGK | 61–63 | SGK | 61–63 | SGK |
| 113–115 | SDK | 113–115 | SDK | 113–115 | SDK | 113–115 | SDK | 113–115 | SDK | 113–115 | SDK | |
| 265–267 | TAK | 265–267 | TAK | 265–267 | TAK | 265–267 | TAK | 265–267 | TAK | 265–267 | TAK | |
| 335–337 | TAK | 335–337 | TAK | 335–337 | TAK | 335–337 | TAK | 335–337 | TAK | |||
| 343–345 | SVK | 343–345 | SVK | 343–345 | SVK | 343–345 | SVK | 343–345 | SVK | 343–345 | SVK | |
| 350–352 | TAK | 350–352 | TAK | 350–352 | TAK | 350–352 | TAK | 350–352 | TAK | 350–352 | TAK | |
| 401–403 | SDK | 401–403 | SDK | |||||||||
| 615–617 | SKK | 615–617 | SKK | 615–617 | SKK | 615–617 | SKK | 615–617 | SKK | 615–617 | SKK | |
| 636–638 | TQK | 636–638 | TQK | 636–638 | TQK | |||||||
| CK II | 75–78 | TAED | 75–78 | TAED | 75–78 | TAED | 75–78 | TAED | 75–78 | TAED | 75–78 | TAED |
| 123–126 | TYGE | 123–126 | TYGE | 123–126 | TYGE | 123–126 | TYGE | 123–126 | TYGE | 123–126 | TYGE | |
| 141–144 | TADD | 141–144 | TADD | 141–144 | TADD | 141–144 | TADD | 141–144 | TADD | 141–144 | TADD | |
| 204–207 | TMEE | 204–207 | TMEE | 204–207 | TMEE | 204–207 | TMEE | 204–207 | TMEE | 204–207 | TMEE | |
| 359–362 | TTFD | 359–362 | TTFD | 359–362 | TTFD | 359–362 | TTFD | 359–362 | TTFD | 359–362 | TTFD | |
| 401–404 | SDKD | 401–404 | SDKD | |||||||||
| 591–594 | SEWE | 591–594 | SEWE | 591–594 | SEWE | 591–594 | SEWE | 591–594 | SEWE | 591–594 | SEWE | |
| 425–428 | SKGE | 425–428 | SKGE | 425–428 | SKGE | 425–428 | SKGE | |||||
| TK 1 | 582–589 | KRTEDFLY | 582–589 | KRTEDFLY | 582–589 | KRTEDFLY | 582–589 | KRTEDFLY | 582–589 | KRTEDFLY | 582–589 | KRTEDFLY |
| N-myr | 26–31 | GTYWGV | 26–31 | GTYWGV | 26–31 | GTYWGV | 26–31 | GTYWGV | 26–31 | GTYWGV | 26–31 | GTYWGV |
| 45–50 | GVKAGR | 45–50 | GVKAGR | 45–50 | GVKAGR | 45–50 | GVKAGR | 45–50 | GVKAGR | 45–50 | GVKAGR | |
| 70–75 | GSQTGT | 70–75 | GSQTGT | 70–75 | GSQTGT | 70–75 | GSQTGT | 70–75 | GSQTGT | 70–75 | GSQTGT | |
| 150–155 | GNDPAL | 150–155 | GNDPAL | 150–155 | GNDPAL | 150–155 | GNDPAL | 150–155 | GNDPAL | 150–155 | GNDPAL | |
| 164–169 | GLGNNT | 164–169 | GLGNNT | 164–169 | GLGNNT | 164–169 | GLGNNT | 164–169 | GLGNNT | 164–169 | GLGNNT | |
| 303–308 | GSNLTY | 303–308 | GSNLTY | 303–308 | GSNLTY | 303–308 | GSNLTY | 303–308 | GSNLTY | 303–308 | GSNLTY | |
| 483–488 | GVATNY | 483–488 | GVATNY | 483–488 | GVATNY | 483–488 | GVATNY | 483–488 | GVATNY | 483–488 | GVATNY | |
| 568–573 | GLDVGR | 568–573 | GLDVGR | 568–573 | GLDVGR | 568–573 | GLDVGR | 568–573 | GLDVGR | 568–573 | GLDVGR | |
| TK 2 | 398–405 | RLGSDKDY | 398–405 | RLGNDKDY | 398–405 | RLGNDKDY | 398–405 | RLGSDKDY | 398–405 | RLGNDKDY | 398–405 | RLGNDKDY |
| 582–589 | KRTEDFLY | 582–589 | KRTEDFLY | 582–589 | KRTEDFLY | 582–589 | KRTEDFLY | 582–589 | KRTEDFLY | 582–589 | KRTEDFLY | |
| Functional Site | H1 | H4 | H10/H12 | H29 | ||||
|---|---|---|---|---|---|---|---|---|
| Position | Sequence | Position | Sequence | Position | Sequence | Position | Sequence | |
| N-gly | 255–258 | NITD | 255–258 | NITD | 255–258 | NITD | 255–258 | NITD |
| 429–432 | NFSL | 429–432 | NFSL | 429–432 | NFSL | 429–432 | NFSL | |
| 734–737 | NLSW | 734–737 | NLSW | 734–737 | NLSW | 734–737 | NLSW | |
| cA/GMP | 190–193 | KRPS | 190–193 | KRPS | 190–193 | KRPS | 190–193 | KRPS |
| 225–228 | RKES | 225–228 | RKES | 225–228 | RKES | 225–228 | RKES | |
| 670–673 | RKVT | 670–673 | RKVT | 670–673 | RKVT | 670–673 | RKVT | |
| 809–812 | KRTS | 809–812 | KRTS | 809–812 | KRTS | 809–812 | KRTS | |
| PK C | 112–114 | SIR | 112–114 | SIR | 112–114 | SIR | 112–114 | SIR |
| 230–232 | SGR | 230–232 | SGR | 230–232 | SGR | 230–232 | SGR | |
| 323–325 | TLR | 323–325 | TLR | 323–325 | TLR | 323–325 | TLR | |
| 446–448 | TIK | 446–448 | TIK | 446–448 | TIK | 446–448 | TIK | |
| 457–459 | SLR | 457–459 | SLR | 457–459 | SLR | 457–459 | SLR | |
| 673–675 | TLR | 673–675 | TLR | 673–675 | TLR | 673–675 | TLR | |
| 685–687 | SAR | 685–687 | SAR | 685–687 | SAR | 685–687 | SAR | |
| 694–696 | SVK | 694–696 | SVK | 694–696 | SVK | 694–696 | SVK | |
| 708–710 | TYR | 708–710 | TYR | 708–710 | TYR | 708–710 | TYR | |
| 740–742 | TLK | 740–742 | TLK | 740–742 | TLK | 740–742 | TLK | |
| 745–747 | SDR | 745–747 | SDR | 745–747 | SDR | 745–747 | SDR | |
| 774–776 | TKR | 774–776 | TKR | 774–776 | TKR | 774–776 | TKR | |
| 822–824 | SIK | 822–824 | SIK | 822–824 | SIK | 822–824 | SIK | |
| 189–191 | SKR | 189–191 | SKR | 189–191 | SKR | |||
| CK II | 54–57 | STNE | 54–57 | STNE | 54–57 | STNE | 54–57 | STNE |
| 89–92 | TAYD | 89–92 | TAYD | 89–92 | TAYD | 89–92 | TAYD | |
| 246–249 | TTGD | 246–249 | TTGD | 246–249 | TTGD | 246–249 | TTGD | |
| 431–434 | SLDD | 431–434 | SLDD | 431–434 | SLDD | 431–434 | SLDD | |
| 463–466 | TPTE | 463–466 | TPTE | 463–466 | TPTE | 463–466 | TPTE | |
| 476–479 | TAAE | 476–479 | TAAE | 476–479 | TAAE | 476–479 | TAAE | |
| 568–471 | SWME | 568–471 | SWME | 568–471 | SWME | 568–471 | SWME | |
| 631–634 | SAFE | 631–634 | SAFE | 631–634 | SAFE | 631–634 | SAFE | |
| 657–660 | SGND | 657–660 | SGND | 657–660 | SGND | 657–660 | SGND | |
| 758–761 | SASD | 758–761 | SASD | 758–761 | SASD | 758–761 | SASD | |
| 812–815 | SILD | 812–815 | SILD | 812–815 | SILD | 812–815 | SILD | |
| 859–862 | SVLE | 859–862 | SVLE | 859–862 | SVLE | 859–862 | SVLE | |
| 881–884 | SGLE | 881–884 | SGLE | 881–884 | SGLE | |||
| 950–953 | SPDE | 950–953 | SPDE | 950–953 | SPDE | 950–953 | SPDE | |
| N-myr | 82–87 | GVHDGL | 82–87 | GVHDGL | 82–87 | GVHDGL | 82–87 | GVHDGL |
| 242–247 | GVDPTT | 242–247 | GVDPTT | 242–247 | GVDPTT | 242–247 | GVDPTT | |
| 461–466 | GMTPTE | 461–466 | GMTPTE | |||||
| 491–496 | GIAHTR | 491–496 | GIAHTR | 491–496 | GIAHTR | 491–496 | GIAHTR | |
| 505–510 | GSNSGT | 505–510 | GSNSGT | 505–510 | GSNSGT | 505–510 | GSNSGT | |
| 557–562 | GQPPSN | 557–562 | GQPPSN | 557–562 | GQPPSN | 557–562 | GQPPSN | |
| 580–585 | GVSYAV | 580–585 | GVSYAV | 580–585 | GVSYAV | 580–585 | GVSYAV | |
| 650–655 | GTEDSS | 650–655 | GTEDSS | 650–655 | GTEDSS | 650–655 | GTEDSS | |
| 693–698 | GSVKNH | 693–698 | GSVKNH | 693–698 | GSVKNH | 693–698 | GSVKNH | |
| 828–833 | GSYLGM | 828–833 | GSYLGM | 828–833 | GSYLGM | 828–833 | GSYLGM | |
| 874–879 | GVATSF | 874–879 | GVATSF | 874–879 | GVATSF | 874–879 | GVATSF | |
| 882–887 | GLEPGE | 882–887 | GLEPGE | 882–887 | GLVPGE | 882–887 | GLEPGE | |
| 1035–1040 | GCKQGW | 1035–1040 | GCKQGW | 1035–1040 | GCKQGW | 1035–1040 | GCKQGW | |
| Ami | 185–188 | CGKR | 185–188 | CGKR | 185–188 | CGKR | 185–188 | CGKR |
| 230–233 | SGRK | 230–233 | SGRK | 230–233 | SGRK | 230–233 | SGRK | |
| 402–405 | NGKR | 402–405 | NGKR | 402–405 | NGKR | 402–405 | NGKR | |
| 935–938 | AGRK | 935–938 | AGRK | 935–938 | AGRK | 935–938 | AGRK | |
| CYP450 | 400–409 | FGNGKRACIG | 400–409 | FGNGKRACIG | 400–409 | FGNGKRACIG | ||
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Xu, Z.; Zhu, Y.; Xuan, L.; Li, S.; Cheng, Z. Haplotype Diversity of NADPH-Cytochrome P450 Reductase Gene of Ophiocordyceps sinensis and the Effect on Fungal Infection in Host Insects. Microorganisms 2020, 8, 968. https://doi.org/10.3390/microorganisms8070968
Xu Z, Zhu Y, Xuan L, Li S, Cheng Z. Haplotype Diversity of NADPH-Cytochrome P450 Reductase Gene of Ophiocordyceps sinensis and the Effect on Fungal Infection in Host Insects. Microorganisms. 2020; 8(7):968. https://doi.org/10.3390/microorganisms8070968
Chicago/Turabian StyleXu, Zixian, Yunguo Zhu, Lingyan Xuan, Shan Li, and Zhou Cheng. 2020. "Haplotype Diversity of NADPH-Cytochrome P450 Reductase Gene of Ophiocordyceps sinensis and the Effect on Fungal Infection in Host Insects" Microorganisms 8, no. 7: 968. https://doi.org/10.3390/microorganisms8070968





