Altered Envelope Structure and Nanomechanical Properties of a C-Terminal Protease A-Deficient Rhizobium leguminosarum
Abstract
1. Introduction
2. Materials and Methods
2.1. Bacterial Strains, Plasmids, and Media
2.2. Isolation of Peptidoglycan
2.3. Muropeptide Structural Analysis
2.4. Transmission Electron Microscopy (TEM) Imaging
2.5. AFM Imaging and Force Data Acquisition
2.6. AFM Creep Experiments
3. Results
3.1. Cell Envelope Ultrastructure
3.2. Muropeptide Composition
3.3. R. leguminosarum Cell Envelope Elasticity and Spring Constant
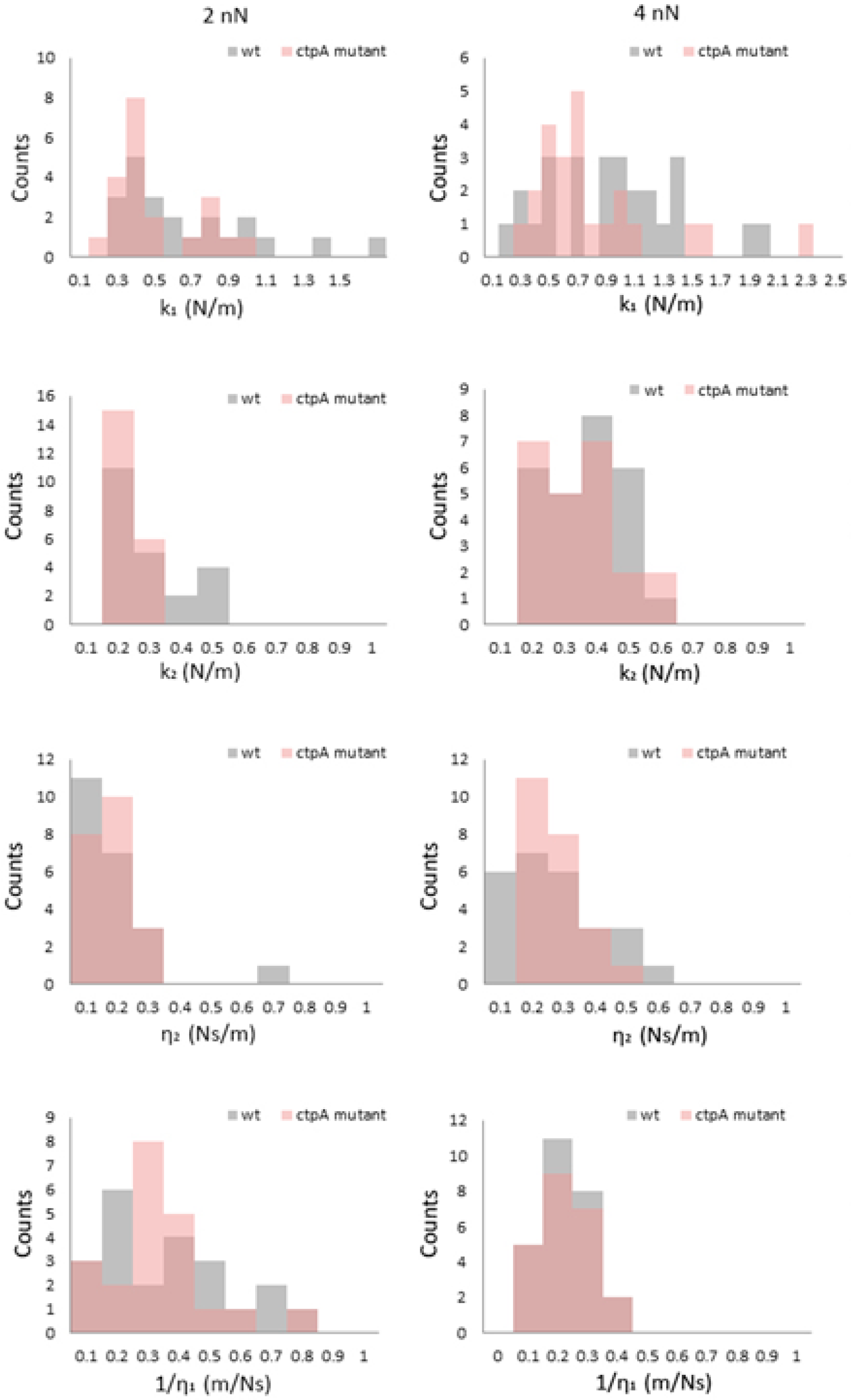
3.4. Viscoelasticity of the Cell Envelope
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Auer, G.K.; Weibel, D.B. Bacterial cell mechanics. Biochemistry 2017, 56, 3710–3724. [Google Scholar] [CrossRef]
- Findley, W.N.; Lai, J.S.; Onaran, K. Creep and Relaxation of Nonlinear Viscoelastic Materials with an Introduction to Linear Viscoelasticity; Dover Publications Inc.: New York, NY, USA, 1989. [Google Scholar]
- Doyle, R.J.; Marquis, R.E. Elastic, flexible peptidoglycan and bacterial cell wall properties. Trends Microbiol. 1994, 2, 57–60. [Google Scholar] [CrossRef]
- Vadillo-Rodriguez, V.; Dutcher, J.R. Viscoelasticity of the cell wall. Soft Matter 2011, 7, 4101–4110. [Google Scholar] [CrossRef]
- Bhat, S.V.; Jun, D.; Paul, B.C.; Dahms, T.E.S. Viscoelasticity in Biological Systems: A Special Focus on Microbes. In Viscoelasticity—From Theory to Biological Applications; de Vicente, J., Ed.; InTech: Rijeka, Croatia, 2012; Volume 6, pp. 123–156. [Google Scholar]
- Vadillo-Rodriguez, V.; Beveridge, T.J.; Dutcher, J.R. Surface viscoelasticity of individual gram-negative bacterial cells measured using atomic force microscopy. J. Bacteriol. 2008, 190, 4225–4232. [Google Scholar] [CrossRef]
- Glauert, A.M.; Thornley, M.J. The topography of the bacterial cell wall. Annu. Rev. Microbiol. 1969, 23, 159–198. [Google Scholar] [CrossRef]
- Silhavy, T.J.; Kahne, D.; Walker, S. The bacterial cell envelope. Cold Spring Harb. Perspect. Biol. 2010, 2, a000414. [Google Scholar] [CrossRef]
- Papanastasiou, M.; Orfanoudaki, G.; Koukaki, M.; Kountourakis, N.; Sardis, M.F.; Aivaliotis, M.; Karamanou, S.; Economou, A. The Escherichia coli peripheral inner membrane proteome. Mol. Cell. Proteom. 2013, 12, 599–610. [Google Scholar] [CrossRef]
- Schlegel, S.; Hjelm, A.; Baumgarten, T.; Vikström, D.; Gier, J.-W. Bacterial-based membrane protein production. Biochim. Biophys. Acta 2014, 1843, 1739–1749. [Google Scholar] [CrossRef]
- Mullineaux, C.W.; Nenninger, A.; Ray, N.; Robinson, C. Diffusion of green fluorescent protein in three cell environments in Escherichia coli. J. Bacteriol. 2006, 188, 3442–3448. [Google Scholar] [CrossRef]
- Van Wielink, J.E.; Duine, J.A. How big is the periplasmic space? Trends Biochem. Sci. 1990, 15, 136–137. [Google Scholar] [CrossRef]
- Vollmer, W.; Blanot, D.; De Pedro, M.A. Peptidoglycan structure and architecture. FEMS Microbiol. Rev. 2008, 32, 149–167. [Google Scholar] [CrossRef] [PubMed]
- Raetz, C.R.; Guan, Z.; Ingram, B.O.; Six, D.A.; Song, F.; Wang, X.; Zhao, J. Discovery of new biosynthetic pathways: The lipid A story. J. Lipid Res. 2009, 50, S103–S108. [Google Scholar] [CrossRef]
- Raetz, C.R.; Whitfield, C. Lipopolysaccharide endotoxins. Annu. Rev. Biochem. 2002, 71, 635–700. [Google Scholar] [CrossRef]
- Lerouge, I.; Vanderleyden, J. O-antigen structural variation: Mechanisms and possible roles in animal/plant–microbe interactions. FEMS Microbiol. Rev. 2002, 26, 17–47. [Google Scholar] [CrossRef]
- Breedveld, M.W.; Yoo, J.S.; Reinhold, V.N.; Miller, K.J. Synthesis of glycerophosphorylated cyclic beta-(1,2)-glucans by Rhizobium meliloti ndv mutants. J. Bacteriol. 1994, 176, 1047–1051. [Google Scholar] [CrossRef]
- Bohin, J.P. Osmoregulated periplasmic glucans in Proteobacteria. FEMS Microbiol. Lett. 2000, 186, 11–19. [Google Scholar] [CrossRef]
- Lee., S.; Cho, E.; Jung, S. Periplasmic glucans isolated from Proteobacteria. BMB Rep. 2009, 42, 769–775. [Google Scholar] [CrossRef]
- Koch, A.L.; Woeste, S. Elasticity of the sacculus of Escherichia coli. J. Bacteriol. 1992, 174, 4811–4819. [Google Scholar] [CrossRef]
- Chen, N.; Xiang, X.; Saha, R.; Bagley, S.T.; Heiden, P.A. Copolymerization of bacterial cell wall materials to enhance the stability of polyhydroxyalkanoate. Macromol. Chem. Phys. 2012, 313, 2647–2652. [Google Scholar] [CrossRef]
- Vadillo-Rodriguez, V.; Schooling, S.R.; Dutcher, J.R. In situ characterization of differences in the viscoelastic response of individual gram-negative and gram-positive bacterial cells. J. Bacteriol. 2009, 191, 5518–5525. [Google Scholar] [CrossRef]
- Garcia-Manyes, S.; Sanz, F. Nanomechanics of lipid bilayers by force spectroscopy with AFM: A perspective. Biochim. Biophys. Acta 2010, 1798, 741–749. [Google Scholar] [CrossRef] [PubMed]
- Espinosa, G.; Lopez-Montero, I.; Monroy, F.; Langevin, D. Shear rheology of lipid monolayers and insights on membrane fluidity. Proc. Natl. Acad. Sci. USA 2011, 108, 6008–6013. [Google Scholar] [CrossRef]
- Picas., L.; Rico, F.; Scheuring, S. Direct measurement of the mechanical properties of lipid phases in supported bilayers. Biophys. J. 2012, 102, L01–L03. [Google Scholar] [CrossRef]
- Gambin, Y.; Lopez-Esparza, R.; Reffay, M.; Sierecki, E.; Gov, N.S.; Genest, M.; Hodges, R.S.; Urbach, W. Corrections to the Saffman-Delbruck mobility for membrane bound proteins. Proc. Natl. Acad. Sci. USA 2006, 103, 2098–2102. [Google Scholar] [CrossRef]
- Jeon, J.; Voth, G.A. The dynamic stress responses to area change in planar lipid bilayer membranes. Biophys. J. 2005, 88, 1104–1119. [Google Scholar] [CrossRef]
- Jauffred, L.; Callisen, T.H..; Oddershede, L.B. Visco-elastic membrane tethers extracted from Escherichia coli by optical tweezers. Biophys. J. 2007, 93, 4068–4075. [Google Scholar] [CrossRef]
- Touhami, A.; Nysten, B.; Dufrêne, Y.F. Nanoscale mapping of the elasticity of microbial cells by atomic force microscopy. Langmuir 2003, 19, 4539–4543. [Google Scholar] [CrossRef]
- Lu., S.; Walters, G.; Parg, R.; Dutcher, J.R. Nanomechanical response of bacterial cells to cationic antimicrobial peptides. Soft Matter 2014, 10, 1806–1815. [Google Scholar] [CrossRef]
- Zahran, H.H. Rhizobium-legume symbiosis and nitrogen fixation under severe conditions and in an arid climate. Microbiol. Mol. Biol. Rev. 1999, 63, 968–989. [Google Scholar] [CrossRef]
- Vriezen, J.A.C.; de Bruijn, F.J.; Nüsslein, K. Responses of rhizobia to desiccation in relation to osmotic stress, oxygen, and temperature. Appl. Environ. Microbiol. 2007, 73, 3451–3459. [Google Scholar] [CrossRef]
- Long, S.R. Rhizobium-legume nodulation: Life together in the underground. Cell 1989, 56, 203–214. [Google Scholar] [CrossRef]
- Hara, H.; Yamamoto, Y.; Higashitani, A.; Suzuki, H.; Nishimura, Y. Cloning, mapping, and characterization of the Escherichia coli prc gene, which is involved in C-terminal processing of penicillin-binding protein 3. J. Bacteriol. 1991, 173, 4799–4813. [Google Scholar] [CrossRef] [PubMed]
- Seoane, A.; Sabbaj, A.; McMurry, L.M.; Levy, S.B. Multiple antibiotic susceptibility associated with inactivation of the prc gene. J. Bacteriol. 1992, 174, 7844–7847. [Google Scholar] [CrossRef][Green Version]
- Wang, C.Y.; Wang, S.W.; Huang, W.C.; Kim, K.S.; Chang, N.S.; Wang, Y.H.; Wu, M.H.; Teng, C.H. Prc contributes to Escherichia coli evasion of classical complement-mediated serum killing. Infect. Immun. 2012, 80, 3399–3409. [Google Scholar] [CrossRef]
- Gilbert, K.B.; Vanderlinde, E.M.; Yost, C.K. Mutagenesis of the carboxy terminal protease CtpA decreases desiccation tolerance in Rhizobium leguminosarum. FEMS Microbiol. Lett. 2007, 272, 65–74. [Google Scholar] [CrossRef]
- Dong, J.; Signo, K.S.L.; Vanderlinde, E.M.; Yost, C.K.; Dahms, T.E.S. Atomic force microscopy of a ctpA mutant in Rhizobium leguminosarum reveals surface defects linking CtpA function to biofilm formation. Microbiology 2011, 157, 3049–3058. [Google Scholar] [CrossRef]
- Jun, D.; Minic, Z.; Vanderlinde, E.M.; Bhat, S.V.; Yost, C.K.; Babu, M.; Dahms, T.E.S. Metabolic adaptation of a C-terminal protease A-deficient Rhizobium leguminosarum in response to loss of nutrient transport. Front. Microbiol. 2018, 8, 2617. [Google Scholar] [CrossRef]
- Johnston, A.W.; Beringer, J.E. Identification of the rhizobium strains in pea root nodules using genetic markers. J. Gen. Microbiol. 1975, 87, 343–350. [Google Scholar] [CrossRef]
- Vincent, J.M. A Manual for the Practical Study of Root-Nodule Bacteria; Blackwell Scientific: Oxford, UK, 1970. [Google Scholar]
- Pisabarro, A.G.; de Pedro, M.A.; Vazquez, D. Structural modifications in the peptidoglycan of Escherichia coli associated with changes in the state of growth of the culture. J. Bacteriol. 1985, 161, 238–242. [Google Scholar] [CrossRef]
- Zhang, H.; Olson, D.J.H.; Van, D.; Purves, R.W.; Smith, M.A. Rapid identification of 5 triacylglycerol-estolides in plant and fungal oils. Ind. Crops Prod. 2012, 37, 186–194. [Google Scholar] [CrossRef]
- Paul, B.C.; Ma, H.; Snook, L.A.; Dahms, T.E.S. High spatial resolution surface imaging and analysis of fungal cells using AFM and force spectroscopy. In Laboratory Protocols in Fungal Biology; Gupta, V.K., Tuohy, M.G., Ayyachamy, M., Turner, K.M., O’Donovan, A., Eds.; Springer: New York, NY, USA, 2013; pp. 151–160. [Google Scholar]
- Cleveland, J.P.; Manne, S.; Bocek, D.; Hansma, P.K. A nondestructive method for determining the spring constant of cantilevers for scanning force microscopy. Rev. Sci. Instrum. 1993, 64, 403–405. [Google Scholar] [CrossRef]
- Lévy, R.; Maaloum, M. Measuring the spring constant of atomic force microscope cantilevers: Thermal fluctuations and other methods. Nanotechnology 2001, 13, 33–37. [Google Scholar] [CrossRef]
- Geiss, P. Creep load conditions. In Handbook of Adhesion Technology; da Silva, L.M., Ãchsner, A., Adams, R., Eds.; Springer: Berlin/Heidelberg, Germany, 2011; Volume 8, pp. 875–902. [Google Scholar]
- Baldwin, W.W.; Sheu, M.J.; Bankston, P.W.; Woldringh, C.L. Changes in buoyant density and cell size of Escherichia coli in response to osmotic shocks. J. Bacteriol. 1988, 170, 452–455. [Google Scholar] [CrossRef] [PubMed]
- Koch, A.L. Shrinkage of growing Escherichia coli cells by osmotic challenge. J. Bacteriol. 1984, 159, 919–924. [Google Scholar] [CrossRef] [PubMed]
- Sonntag, I.; Schwarz, H.; Hirota, Y.; Henning, U. Cell envelope and shape of Escherichia coli: Multiple mutants missing the outer membrane lipoprotein and other major outer membrane proteins. J. Bacteriol. 1978, 136, 280–285. [Google Scholar] [CrossRef] [PubMed]
- Kanehisa, M.; Furumichi, M.; Tanabe, M.; Sato, Y.; Morishima, K. KEGG: New perspectives on genomes, pathways, diseases and drugs. Nucleic Acids Res. 2017, 45, D353–D361. [Google Scholar] [CrossRef]
- Keiler, K.C.; Sauer, R.T. Identification of Active Site Residues of the Tsp Protease. J. Biol. Chem. 1995, 270, 28864–28868. [Google Scholar] [CrossRef]
- Altschul, S.F.; Gish, W.; Miller, W.; Myers, E.W.; Lipman, D.J. Basic local alignment search tool. J. Mol. Biol. 1990, 215, 403–410. [Google Scholar] [CrossRef]
- Gaboriaud, F.; Bailet, S.; Dague, E.; Jorand, F. Surface structure and nanomechanical properties of Shewanella putrefaciens bacteria at two pH values (4 and 10) determined by atomic force microscopy. J. Bacteriol. 2005, 187, 3864–3868. [Google Scholar] [CrossRef]
- Yao, X.; Jericho, M.; Pink, D.; Beveridge, T. Thickness and elasticity of gram-negative murein sacculi measured by atomic force microscopy. J. Bacteriol. 1999, 181, 6865–6875. [Google Scholar] [CrossRef]
- Barnickel, G.; Labischinski, H.; Bradaczek, H.; Giesbrecht, P. Conformational energy calculation on the peptide part of murein. Eur. J. Biochem. 1979, 95, 157–165. [Google Scholar] [CrossRef]
- Labischinski, H.; Barnickel, G.; Bradaczek, H.; Giesbrecht, P. On the secondary and tertiary structure of murein. Low and medium-angle X-ray evidence against chitin-based conformations of bacterial peptidoglycan. Eur. J. Biochem. 1979, 95, 147–155. [Google Scholar] [CrossRef] [PubMed]
- Mayer, F. Cytoskeletons in prokaryotes. Cell Biol. Int. 2003, 27, 429–438. [Google Scholar] [CrossRef]
- Smolyakov, G.; Formosa-Dague, C.; Severac, C.; Duval, R.E.; Dague, E. High speed indentation measures by FV, QI and QNM introduce a new understanding of bionanomechanical experiments. Micron 2016, 85, 8–14. [Google Scholar] [CrossRef] [PubMed]



| Peak | Proposed Structure | Observed m/z | Calculated [M+Na]+ | % of All Peaks | |
|---|---|---|---|---|---|
| Wild Type | ctpA Mutant | ||||
| 2 | GlcNAc-MurNAc-Ala-Glu-meso-DAP | 893.4 | 893.8 | 2.0 | 2.7 |
| 3 | GlcNAc-MurNAc-Ala-Glu-meso-DAP-Gly | 950.4 | 950.9 | 1.6 | 1.6 |
| 6 | GlcNAc-MurNAc-Ala-Glu-meso-DAP-Ala | 964.5 | 964.9 | 6.2 | 8.3 |
| 10 | GlcNAc-MurNAc-Ala-Glu-meso-DAP-Ala-meso-DAP-Glu-Ala-MurNAc-GlcNAc | 1816.9 | 1816.8 | 1.3 | 2.4 |
| 11 | MurNAc-Ala-Glu-meso-DAP-Ala-meso-DAP-Glu-Ala-MurNAc | 1408.7 | 1408.4 | 1.1 | 2.2 |
| 12 | GlcNAc-MurNAc-Ala-Glu-meso-DAP-Ala-Ala-meso-DAP-Glu-Ala-MurNAc-GlcNAc | 1888.0 | 1887.9 | 2.9 | 3.0 |
| Force (nN) | k1 (N/m) | k2 (N/m) | η2 (N•s/m) | η1 (N•s/m) | ||||
|---|---|---|---|---|---|---|---|---|
| wt | ctpA | wt | ctpA | wt | ctpA | wt | ctpA | |
| 2 | 0.52 ± 0.37 n = 22 | 0.45 ± 0.22 n = 21 | 0.22 ± 0.12 n = 22 | 0.18 ± 0.06 n = 21 | 0.14 ± 0.12 n = 22 | 0.13 ± 0.06 n = 21 | 5.27 ± 4.38 n = 22 | 5.04 ± 4.24 n = 21 |
| 4 | 0.89 ± 0.48 n = 26 | 0.758 ± 0.46 n = 23 | 0.31 ± 0.13 n = 26 | 0.30 ± 0.13 n = 23 | 0.23 ± 0.15 n = 26 | 0.23 ± 0.10 n = 23 | 8.22 ± 6.26 n = 26 | 7.51 ± 5.58 n = 23 |
| 6 | 0.97 ± 0.56 * n = 18 | 1.39 ± 0.78 * n = 23 | 0.37 ± 0.12 n = 18 | 0.41 ± 0.16 n = 23 | 0.27 ± 0.16 n = 18 | 0.33 ± 0.23 n = 23 | 9.95 ± 5.66 * n = 18 | 7.04 ± 4.50 * n = 23 |
| 10 | 1.56 ± 0.59 n = 21 | 1.71 ± 0.92 n = 27 | 0.57 ± 0.15 * n = 21 | 0.46 ± 0.21 * n = 27 | 0.52 ± 0.36 n = 21 | 0.48 ± 0.21 n = 27 | 12.5 ± 5.97 † n = 21 | 6.78 ± 3.87 † n = 27 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Jun, D.; Idem, U.; Dahms, T.E.S. Altered Envelope Structure and Nanomechanical Properties of a C-Terminal Protease A-Deficient Rhizobium leguminosarum. Microorganisms 2020, 8, 1421. https://doi.org/10.3390/microorganisms8091421
Jun D, Idem U, Dahms TES. Altered Envelope Structure and Nanomechanical Properties of a C-Terminal Protease A-Deficient Rhizobium leguminosarum. Microorganisms. 2020; 8(9):1421. https://doi.org/10.3390/microorganisms8091421
Chicago/Turabian StyleJun, Dong, Ubong Idem, and Tanya E. S. Dahms. 2020. "Altered Envelope Structure and Nanomechanical Properties of a C-Terminal Protease A-Deficient Rhizobium leguminosarum" Microorganisms 8, no. 9: 1421. https://doi.org/10.3390/microorganisms8091421
APA StyleJun, D., Idem, U., & Dahms, T. E. S. (2020). Altered Envelope Structure and Nanomechanical Properties of a C-Terminal Protease A-Deficient Rhizobium leguminosarum. Microorganisms, 8(9), 1421. https://doi.org/10.3390/microorganisms8091421







