1. Introduction
Photoheterotrophs are obligately heterotrophic bacteria capable of using light energy for ATP generation without fixing inorganic carbon and producing molecular oxygen. Photoheterotrophs include rhodopsin-containing bacteria [
1] and aerobic anoxygenic phototrophs (AAPs) [
1,
2], using bacteriochlorophyll as a light-harvesting pigment. Photoheterotrophs lack RuBisCo, a key enzyme of a tricarbonic acid cycle, and therefore cannot fix CO
2 for subsequent synthesis of organics [
2,
3,
4,
5]. AAPs and rhodopsin-containing bacteria are considered to play a crucial role in carbon cycling and energy flux in both marine and freshwater ecosystems [
5,
6,
7,
8].
These two types of photoheterotrophic bacteria have fundamentally different mechanisms of conversion of light energy to chemical energy. Photosynthetic apparatus of rhodopsin-containing bacteria is the simplest of all known, lacking an electron transport chain. It consists of a transmembrane protein, opsin, which is a light-driven proton pump, a chromophore (retinal), and ATP synthase [
9].
Chlorophyll systems of AAPs are complex, being composed of multiple proteins, pigments, and cofactors forming reaction centers. The energy is passed along an electron transport chain [
5,
10]. AAPs likely evolved from the purple non-sulfur bacteria to fill an environmental niche, carrying out anoxygenic photosynthesis aerobically [
4].
The first aerobic species of anoxygenic phototrophs were discovered in the 1970s (Aburatsubo Inlet, Pacific Ocean) [
11]. The first freshwater AAP species,
Erythrobacter (now
Sandaracinobacter)
sibiricus and
Erythromicrobium (now
Erythromonas)
ursincola, were isolated from the surface of alkaline cyanobacterial mats growing in an alkaline thermal spring nearby Lake Baikal [
12,
13].
To date, the AAPs detected in different freshwater lakes and rivers belong to the class Alphaproteobacteria, orders Rhodospirillales, Rhodobacterales, Sphingomonadales, Rhizobiales, and Caulobacterales [
10,
14,
15,
16,
17,
18,
19,
20], as well as the class Betaproteobacteria, including genera
Limnohabitans [
16,
18,
19,
21] and
Polynucleobacter [
22]. Additionally, the bacteria of phyla Gemmatimonadetes [
23] and Firmicutes (Lactobacillales) [
18] were reported. In contrast to marine ecosystems, Gammaproteobacteria are rare in freshwater bodies [
10,
22].
AAP bacterial cells are on average two-fold larger than the average bacterial cell size [
24]. The use of light energy allows them to store more carbon (which would otherwise be respired) in the form of biomass [
5]. Thus, AAPs likely have an ecological advantage over heterotrophs under conditions when bacterial growth is energy or carbon limited [
25]. On the other hand, the larger size possibly makes AAP bacteria more subjected to protist grazing. AAPs can break down highly complex organics due to the activity of a broad range of proteins and enzymes. This makes them indispensable in carbon cycling in both saline and freshwater systems, as carbon available naturally is generally very complex both structurally and chemically for digestion by other microbes [
24]. The AAPs in freshwater bodies comprised on average 1–7% or 37% maximum of total bacterial abundance [
10,
24], whereas in marine ecosystems, their share was 1–10%, up to 18% [
10,
26].
The recognition of microbial rhodopsins as light-driven ion pumps began in the 1970s when the photopigment named bacteriorhodopsin was found in
Halobacterium salinarum (Archaea) [
27]. Metagenomic studies revealed the presence of a new family of rhodopsins in marine bacteria, which was named proteorhodopsin [
6]. Rhodopsin genes have been found in seawater, freshwater, and brackish water [
22,
28,
29,
30,
31,
32], and they are one of the most highly expressed and widely distributed proteins in marine bacterial communities [
8]. The amount of rhodopsin-containing bacteria was 13–70% in marine ecosystems [
9,
31,
33,
34], and these were the dominant fraction among photoheterotrophs in freshwater lakes Damariscotta, Mendota, and Sparkling (USA) [
22].
The diversity of rhodopsin-containing bacteria in freshwater ecosystems is poorly studied. The known dominant phylum is Actinobacteria [
22,
35,
36]. Rhodopsins are also found in freshwater Flavobacteria, Beta-, Gamma- and Deltaproteobacteria, Verrucomicrobia, Sphingobacteria, and Alphaproteobacteria, including the SAR11 freshwater cluster [
8,
22,
30].
The diversity of AAPs and rhodopsin containing bacteria in Lake Baikal has never been previously studied; this is the first work dedicated to this theme. Lake Baikal is of unique natural significance, being the oldest lake in the world with an age of ca. 25 million years. Taking into account that photoheterotrophs are a very active part of bacterial community playing an important role in carbon cycling and energy flux in freshwater ecosystems [
5,
6,
7,
8], it is essential to investigate those microorganisms for better understanding of microbial and biochemical processes occurring in Lake Baikal. The aim of the present work was to examine the diversity of aerobic anoxygenic phototrophs and rhodopsin-containing bacteria in different biotopes of Lake Baikal. We expected that photoheterotrophs community composition in Baikal water should be similar to other freshwater bodies, which was confirmed. We also hypothesized that photoheterotrophs community composition could differ in water and biofilms due to distinct living conditions, but those discrepancies eventually appeared to be mostly on the phylotype level.
2. Materials and Methods
Baikal is a dimictic oligotrophic lake with a surface area of 31,500 km
2 and a water volume of 23,000 km
3. Lake Baikal is conventionally divided into three basins: southern, central, and northern with maximum depths of about 1400, 1600, and 800 m, respectively. According to the content of major ions, water of the lake belongs to the bicarbonate class of the calcium group. The total dissolved solids content of Lake Baikal water is about 150 mg/L, which is very low [
37].
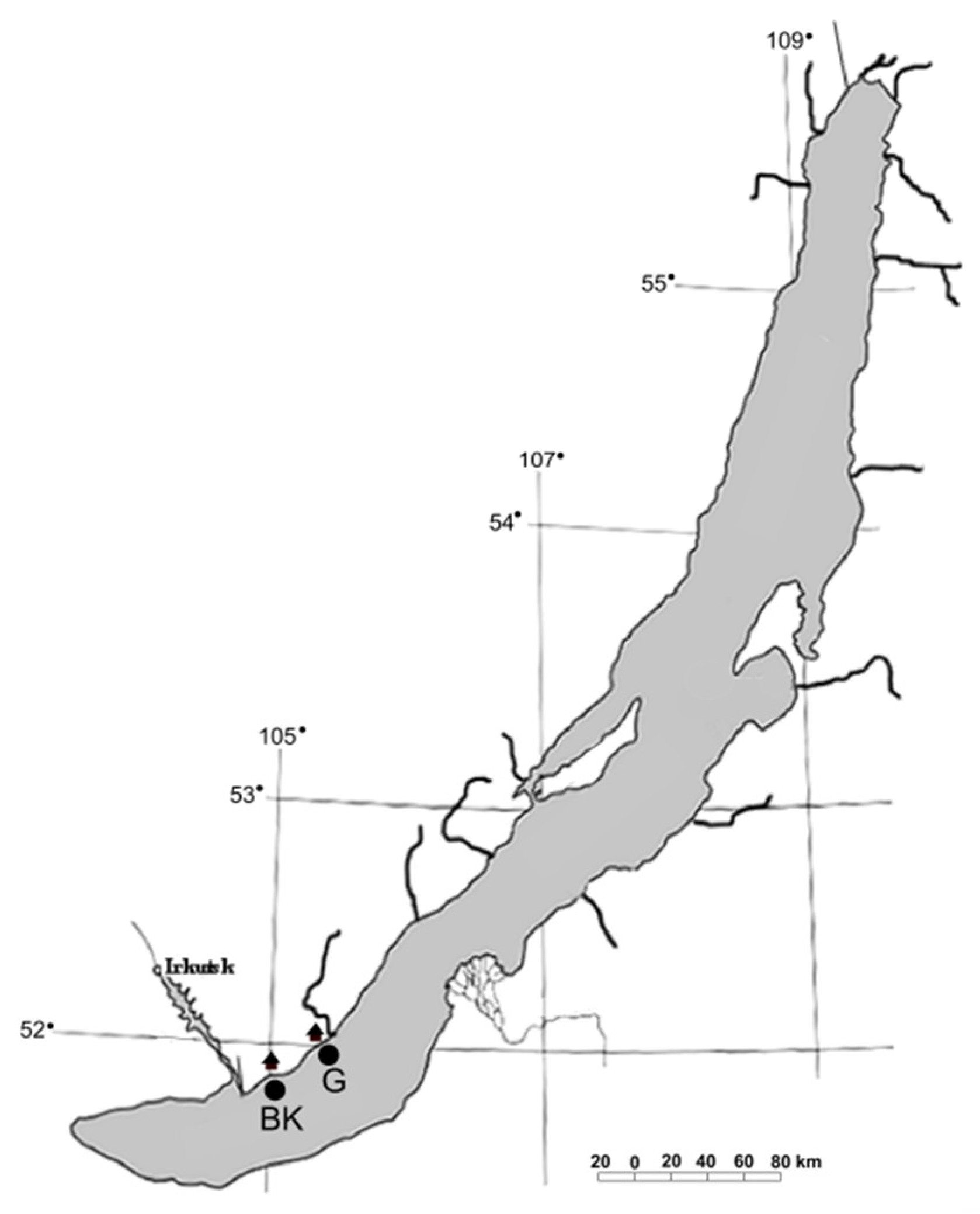
The samples of surface microlayer (BK1, G1), water column (BK2, G2), bottom water (BK4, G4), and epilithic biofilms (BK51, BK53, G51, G52) were taken in the littoral zone of the southern basin of Lake Baikal off the Bol’shiye Koty settlement (51°53′93.0″ N; 105°03′83.8″ E) (BK) and off the Bol’shoye Goloustnoye settlement (52°01′36.5″ N; 105°24′11.8″ E) (G) in August 2019 (
Figure 1). Surface microlayer was taken from a boat as described previously [
38]. A metal mesh screen (26.5 cm in diameter) was horizontally submerged into the water and then horizontally lifted. After several seconds, the screen was bent and tilted to drain the water from the cells of the screen into a sterile container. Water column samples were taken from a depth of 5 and 10 m with a bathometer, then integrated. Stones with epilithic biofilms (two stones from each site) and samples of bottom water were taken by scuba divers off the Bol’shiye Koty settlement from a depth of 17 m and off the Bol’shoye Goloustnoye settlement from a depth of 12.3 m. The stones were placed into sterile plastic containers fulfilled with surrounding water to prevent drying. Bottom water was collected into sterile plastic bottles.
1 cm
2 of epilithic biofilms from each stone was scraped off with a sterile scalpel and freezed until further analysis. Samples of water (1 L) were filtered through polycarbonate membrane filters (pore size 0.22 μm) (Millipore, Burlington, MA, USA). Total DNA was extracted using the phenol-chlorophorm method [
39]. Extracted DNA samples were equimolarly mixed and used as a template for the analysis.
PO
43−, NO
3−, NO
2−, and NH
4+ were analyzed in water samples filtered through mixed cellulose ester membrane filters (Advantec, Tokyo, Japan, pore diameter 0.45 µm). The content of total organic carbon (TOC) was determined in unfiltered water. Concentration of dissolved nutrients was determined using a photoelectric colorimeter (KFK-3-01-ZOM3, Zagorskii optiko-mekhanicheskii zavod [Zagorskii optical mechanics factory], Sergiev Posad, Russia) according to Wetzel and Likens [
40]. PO
43− was identified by the Denigès–Atkins method in modification with tin chloride. NH
4+ was detected by the indophenol method [
41]. NO
3− and NO
2− content was measured by high performance liquid chromatography (EcoNova, Novosibirsk, Russia) with UV detection on an inverse-phase column modified with octadecyltrimethylammonium bromide [
42]. Total organic carbon was determined by a total carbon/nitrogen analyzer (Vario TOC cube, Elementar, Langenselbold, Germany).
The samples of epilithic biofilms were scraped off with a sterile scalpel from each stone and dried before further analysis. Biofilms taken in the same sampling site were integrated (BK51 + BK53 and G51 + G52, respectively). TOC and N total were determined by element analyzer (Flash EA 1112 CHNS, Thermo Fisher Scientific, Waltham, MA, USA) in Baikal Analytical Collective Instrumental Center (A.E. Favorsky Irkutsk Institute of Chemistry SB RAS). P total was determined by the persulfate oxidation method [
43,
44,
45].
Physical and chemical characteristics of water samples and biofilms are provided in
Table 1 and
Table 2.
Fragments of the 16S rRNA gene containing two variable regions V3–V4 were amplified using eubacterial primers 343F (forward) (CTCCTACGGRRSGCAG) and 806RB (reverse) (GGACTACNVGGGTWTCTAAT) [
46,
47]. BioMaster HS-Taq PCR-Color (2x) kit (Biolabmix, Russia) was used to prepare the reaction mix. PCR conditions were as follows: an initial denaturation step at 94 °C for 9 min 50 s and 25 cycles at 94 °C (30 s), 58 °C (30 s), 72 °C (45 s), and a final elongation step at 72 °C for 8 min. The majority of environmental studies employ the
pufM gene (encoding the M subunit of the bacterial reaction center) as a marker for anoxygenic phototrophs harboring type-2 reaction centers [
1,
48,
49]. Fragments of the
pufM gene (~220 bp fragments) were amplified using primers
pufM_uniFfresh (forward) (GGNAAYYTGTTYTAYAACC) and
pufM_uniRfresh (reverse) (CCCATSGTCCANCKCCARAA) [
22,
50]. PCR conditions were as follows: an initial denaturation step at 95 °C for 5 min and 40 cycles at 94 °C (20 s), 50 °C (60 s), 72 °C (60 s), and a final elongation step at 72 °C for 10 min. Fragments of the rhodopsin gene (KEGG ID: "K04250") (~345 bp fragments) were amplified using primers PRpf2 (forward) (TAYCGYTAYGTNGAYTGG) and PRpr2 (reverse) (ATYGGRTANACRCCCCA) [
22,
36]. PCR conditions were as follows: an initial denaturation step at 95 °C for 5 min and 40 cycles at 94 °C (20 s), 46 °C (60 s), 72 °C (45 s), and a final elongation step at 72 °C for 10 min. Amplicon libraries were generated with gene-specific primers, prepared for sequencing with Nextera XT kit (Illumina, San Diego, CA, USA) and sequenced using MiSeq (Illumina, USA) (SB RAS Genomics Core Facility, Novosibirsk, Russia). Results were uploaded into NCBI SRA (PRJNA613961).
Assessment of amplicon libraries quality was performed with FastQC software (
http://www.bioinformatics.babraham.ac.uk/projects/fastqc Last access date: 14 April 2021), primers and spurious sequences were trimmed using cutadapt v1.14 [
51]. Metagenomic data for the G52 sample (16S rRNA gene amplicon library) were of poor quality and were not taken into further analysis. Metagenomic data were processed using DADA2 software (parameters: maxN = 0, maxEE = c(2,4)), including alignment, filtering for chimeras, short and spurious sequences, and exact sequence variants (ESVs) clustering [
52]. 16S rRNA gene fragments were aligned and taxonomically assigned using SILVA v.132 database with a confidence threshold of 80% [
53] and clustered to operational taxonomic units (OTUs) at 0.03 distance with mothur v.1.40.0 [
54].
Nucleotide sequences of functional genes fragments were translated into amino acid sequences, then aligned and taxonomically assigned using BLASTp algorithm against NCBI-nr database. After that, sequences were clustered at 100% of amino acid sequence similarity for further analysis.
Maximum Likelihood tree for 16S rRNA gene fragments alignment was built using MEGA X software tool [
55] with bootstrap sampling and using K2P + G + I substitution model based on jmodeltest tool results [
56]. Phylogenetic trees for functional gene alignments were computed following the Bayesian Markov chain Monte Carlo (MCMC) method using BEAST v1.10.4 [
57] with HKY + G + I substitution model. MCMC was run for 2 million steps. The output was analyzed in Tracer v1.7.1 [
58] and burn-in was adjusted to attain an appropriate effective sample size (ESS) more than 200.
Statistical analysis was performed with the vegan package [
59] using the R language [
60]. The rarefaction curves were plotted to evaluate the sufficiency of the sequencing depth (
Supplementary Materials Figure S1). We used sub-sampled by smallest value data set for further analysis of alpha- and beta-diversity. Alpha-diversity was analyzed using the Chao1, Shannon, and Simpson indices. Beta-diversity was analyzed using unweighted pair group method with arithmetic mean (UPGMA) and non-metric multidimensional scaling (NMDS) methods based on Bray-Curtis dissimilarity metrics. Analysis of variance (
PERMANOVA/“
adonis” function from
vegan package) was used to compare samples by sampling site and biotope (
Table S1).
4. Discussion
The main goal of our research was to assess the diversity of AAPs and rhodopsin-containing bacteria in Lake Baikal as it has never been done before. The method of sequencing of functional genes amplicons which was applied allowed us to get the first insight into the diversity of Baikal photoheterotrophs and to compare it with other freshwater bodies studied. Interesting data was obtained regarding differences between epilithon and water photoheterotrophic bacterial communities; the diversity of AAPs and rhodopsin-containing bacteria inhabiting those biotopes has not been compared, to the best of our knowledge.
16S rRNA gene diversity of Lake Baikal bacterioneuston and bacterioplankton communities was similar to that described by Galach’yants et al. (2017), Mikhailov et al. (2015, 2019), and Kurilkina et al. (2016) [
62,
63,
64,
65]. The dominant phyla in Lake Baikal neuston and plankton were Cyanobacteria, Actinobacteria, Proteobacteria, Bacteroidetes, and Verrucomicrobia. In the current work, Firmicutes and Fusobacteria were also referred to as the major phyla. The other phyla detected were Planctomycetes, Acidobacteria, Armatimonadetes, Chloroflexi, Deinococcus-Thermus, Gemmatimonadetes, Nitrospirae, and Firmicutes [
62,
63,
64,
65]. These phyla are known to be common in all freshwater bodies [
66]. 16S rRNA gene sequences of Lake Baikal bacteria had high homology with the sequences of bacteria inhabiting other freshwater bodies all over the world. This fact confirms the similarity of microbial communities of freshwater ecosystems [
67].
Biofilms are major sites of carbon cycling and ecosystem productivity in freshwater ecosystems, even in the world’s largest lakes [
68,
69]. Taxonomic composition of Lake Baikal epilithic biofilms was observed by Parfenova et al. (2013) and Sorokovikova et al. (2013) [
70,
71]. Phyla diversity was comparable to that described in the current work. Cyanobacteria, Proteobacteria, and Bacteroidetes dominated; Actinobacteria, Verrucomicrobia, Planctomycetes, Acidobacteria, Chloroflexi, Gemmatimonadetes, Nitrospirae, and Firmicutes were present as well. Similar composition of bacterial phyla in epilithic biofilms of oligotrophic mountain lakes was shown by Bartrons et al. (2012) [
72]. Recently, community structure of river biofilms was estimated by Romero et al. (2020) [
73]. Phyla composition turned out to be much the same as previously described. The most represented phyla were Proteobacteria, Bacteroidetes, Cyanobacteria, and Firmicutes.
The proportion of Cyanobacteria in the total number of sequences was big and averaged 23% in neuston and plankton and 17% in the epilithon. Cyanobacteria are a big part of Lake Baikal autotrophic picoplankton (more than 90% of its abundance), playing a key role in freshwater oligotrophic ecosystems as a considerable source of primary production [
74,
75]. Cyanobacteria convert carbon dioxide and water into organic matter during photosynthesis and release oxygen, making the existence of heterotrophic organisms that consume organic substances and aerobic organisms that require oxygen possible. Picoplankton species of the cluster
Synechococcus/
Cyanobium and
Dolichospermum lemmermannii mainly represented cyanobacteria in neuston and plankton. Benthic and periphyton species,
Synechococcus sp.,
Calothrix sp.,
Tychonema sp., and
Pseudanabaena sp., were abundant in the biofilms. One of the dominant picoplankton species was
Dolichospermum lemmermannii known to produce paralytic mollusc toxins (saxitoxins) harmful for human beings and mammals [
76].
Actinobacteria comprised 19% of all sequences in neuston and plankton and 22% in epilithon. Actinobacteria are ubiquitous in the epilimnion of freshwater bodies [
66]. These bacteria are chemo-organoheterotrophic microorganisms, at the same time possessing rhodopsin pigment allowing them to acquire the supplementary ATP from the solar light [
66]. Actinobacteria are free living bacteria with a small size of cells, which helps them to escape grazing. All these features enable dominating of Actinobacteria in different water bodies. In neuston and plankton, OTUs 8, 12, 17, 18, 23, and 25 (Acidobacteriales, Frankiales, and Microtrichales not identified at the genus level) prevailed, whereas OTUs 11, 20, 32, and 33 (Microtrichales, Propionibacteriales, and Frankiales not identified at the genus level) dominated epilithon.
Proteobacteria was the most represented phylum in all the biotopes, which included 24% of neuston and plankton and 33% of epilithon sequences. Proteobacteria are ubiquitous microorganisms, but freshwater bodies are usually dominated by Betaproteobacteria [
66]. These are copiotrophs growing fast in the excess of organics. In Lake Baikal, the most represented Betaproteobacteria genus was
Limnohabitans; it was detected exclusively in neuston and plankon. The genus
Limnohabitans (Burkholderiales, Betaproteobacteria) is a common and highly active component of freshwater bacterioplanktonic communities [
66].
Limnohabitans is capable of consuming phytoplankton-derived DOC and, thus, plays an important role in the carbon cycle in freshwater bodies [
77]. In the biofilms, dominant phylotypes were assigned as
Rhodoferax,
Methylotenera, as well as Burkholderiaceae not identified at the genus level.
Rhodoferax are purple non-sulfur bacteria common in biofilms capable of both living with or without oxygen acting as photoautotrophs [
78].
Methylotenera are representatives of methylotrophs, microbes capable of utilizing single C
1 compounds as sole sources of energy and carbon [
79]. These bacteria are also effective degraders of complex organic compounds [
79].
Among Alphaproteobacteria, the most represented OTU was assigned as the SAR11 cluster (Pelagibacterales). It was detected only in neuston and plankton. These bacteria are widely distributed in oligotrophic water bodies and are one of the most abundant microorganisms on the Earth [
66,
80]. In the epilithon, OTU 6 (
Tabrizicola) and OTU 63 (
Polymorphobacter) were the most represented phylotypes. Some species of the genera
Tabrizicola and
Polymorphobacter produce bacteriochlorophyll
a under aerobic heterotrophic conditions and possess
pufLM photosynthesis-related genes [
20,
81].
Bacteroidetes comprise a considerable part of bacterial community in the epilimnion of lakes [
66]. In our work, 10% of neuston and plankton sequences and 4% of epilithon sequences belonged to that phylum. These bacteria can attach to the particles and play an important role in the degradation of complex biopolymers. In water bacterial communities, the most represented genera were
Flavobacterium and
Algoriphagus. In the epilithon,
Flavobacterium dominated as well. Bacteria of the genus
Flavobacterium are one of the most numerous Bacteroidetes representatives in freshwater bodies acting as copiotrophs [
66]. Members of the genus
Algoriphagus are saccharolytic bacteria initially isolated from algal-rich biotopes [
82].
Verrucomicrobia are also presented in all freshwater lakes. These bacteria have been observed in both surface and hypolimnetic waters, suggesting a variety of metabolic strategies within the group [
83,
84]. In Lake Baikal, they were presented mainly in the biofilms (9% of sequences). In neuston and plankton, sequences belonging to that phylum comprised 5%. In water bacterial communities, OTU 9 (
Luteolibacter) was the most represented; the same OTU prevailed in the epilithon, but its proportion was higher compared to neuston and plankton.
Luteolibacter strains are chemo-organoheterotrophic bacteria showing a nutritional preference for simple sugars and complex protein substrates [
85].
In our study, Firmicutes was the major phylum as well. It was mostly represented by allochthonous microorganisms belonging to the genera
Lactobacillus and
Enterococcus, typical representatives of gut microbiome [
86], showing fecal contamination of water. This was probably due to the localization of sampling stations off the settlements not equipped by sewage treatment plants. Ships significantly contribute to the fecal contamination as well [
87].
Representatives of the phylum Fusobacteria were abundant in our samples as well. These are also members of gut microbiome [
86] and confirm fecal contamination of the water.
Thus, the members of Lake Baikal water and epilithic biofilms bacterial communities are active participants of carbon cycle and energy flux in the lake being essential to the maintenance of ecosystem functioning. Cyanobacteria perform primary production converting inorganic carbon into organic compounds using solar energy along with phytoplankton. Other members of the community are active degraders of complex organic matter being aerobic chemo-organoheterotrophs. Some bacteria are well adapted to the oligotrophic conditions due to possessing additional mechanism of energy harvesting, such as photoheterotrophs.
The taxonomic composition of AAP communities in Lake Baikal is similar to that in other freshwater bodies [
14,
15,
16,
18,
22,
88,
89]. As in Lake Baikal, Rhodobacterales and Burkholderiales dominated, and Rhizobiales, Rhodospirillales, and Sphingomonadales were detected almost in every studied water body.
pufM-containing Planctomycetes (class Phycisphaerae) were also detected in Lake Baikal. Planctomycetes are environmentally important bacteria that are key players in global carbon and nitrogen cycles. Planctomycetes seem to be associated primarily with particles, surfaces, microbial mats, and biofilms while they can be very abundant in other habitats as well [
90]. The closest
pufM homologue of Baikal representatives (MBC7770036, 98% similarity) was detected in a high arctic glacier in Northeast Greenland [
91].
pufM-containing Planctomycetes were also reported in the South China Sea [
92].
AAPs identified to the genus level belonged to the genera Tabrizicola, Erythrobacter, Blastomonas, and Sphingomonas.
The closest
pufM homologues of the Baikal representatives of
Erythrobacter,
Blastomonas and
Sphingomonas were detected in freshwater bodies.
Erythrobacter sp. HU12-14 (AGK27892, 98–100% similarity) was isolated from Lake Hulun (Inner Mongolia);
Sphingomonas sp. H151 (AGK27878, 100% similarity) and
Blastomonas sp. Y27 (AGK27877, 98–100% similarity) from Lake Yongxinghu (Inner Mongolia). These genera are known to include aerobic anoxygenic phototrophs [
93,
94,
95]. In Lake Baikal, phototrophic
Erythrobacter,
Blastomonas, and
Sphingomonas were detected mainly in the epilithon.
Type species of the genus
Tabrizicola were isolated from different freshwater and saline ecosystems [
96,
97,
98]. Since this genus has been discovered not long ago, its ecology is poorly studied. Notably, the phototrophic
Tabrizicola were rather abundant in Lake Baikal (4% of all
pufM gene sequences) and were detected exclusively in epilithon. The closest homologue,
Tabrizicola sp. (QBQ34549, 98–100% similarity), was detected in a lake on Tibetan Plateau.
Given that
pufM gene sequences were identified predominantly only to the order level, we reviewed the 16S rRNA diversity of hypothetically AAP bacteria in neuston, plankton, and epilithon (
Figure 8). The AAP genera,
Roseomonas (Acetobacterales),
Bradyrhizobium (Rhizobiales),
Tabrizicola (Rhodobacterales),
Blastomonas (Sphingomonadales),
Porphyrobacter (Sphingomonadales),
Sandarakinorhabdus (Sphingomonadales),
Sphingomonas (Sphingomonadales),
Polymorphobacter (Sphingomonadales),
Polynucleobacter (Burkholderiales), and
Limnohabitans (Burkholderiales), were detected in the studied biotopes of Lake Baikal (totally 10138 16S rRNA gene sequences, 6% of all obtained sequences).
Limnohabitans and
Tabrizicola dominated. Moreover, we detected the
Rhodobacter and
Rhodoferax representatives. These are purple non-sulfur bacteria capable of photolithoautotrophy in anaerobic conditions with the availability of light [
78]. In aerobic conditions, they function as chemoorganoheterotrophs.
Recently, the photosynthesis genes
pufLM and
bchY from the
Limnohabitans representatives were detected [
21]. Now it is known that
Limnohabitans comprises a big part of the AAP community in freshwater ecosystems [
21,
89]. A considerable amount (1.8%) of our 16S rRNA sequences was identified as
Limnohabitans. All of them were detected in the water biotopes: surface microlayer, water column, and bottom water. There were no
Limnohabitans bacteria in the epilithon. The closest homologue, betaproteobacterium SCGC AAA027-O07 (HQ663710, 99.53% similarity), possessed the
pufM gene; therefore, the Baikal representatives, probably, also had this gene (
Figure 8).
UPGMA and NMDS analysis indicated that biotope significantly biased the taxonomic composition of AAP bacteria. Water microbiomes and epilithon had distinct differences at the phylotype level (
Figure 4A,B). There were “typical epilithon phylotypes” and “typical water phylotypes”. AAPs belonging to Rhodobacterales dominated all biotopes. Rhizobiales and Burkholderiales were abundant in neuston and plankton in contrast to epilithon (
Figure 5). Phototrophic
Tabrizicola and
Erithrobacter were detected only in the epilithic biofilms, representing “typical epilithon phylotypes”.
The information about AAPs in epilithic biofilms is scarce; we managed to find only one paper dedicated to this theme. Hirose et al. (2016) investigated aerobic anoxygenic phototrophs in epilithic biofilms of the Tama River (Japan) [
17]. Rhodospirillales (
Roseomonas), Rhodobacterales (
Tabrizicola) and Sphingomonadales (
Polymorphobacter,
Sandarakinorhabdus,
Sphingomonas) were detected there [
17] as in the Baikal epilithic biofilms. Remarkably, the closest homologue of one Baikal
Tabrizicola phylotype,
Rhodobacteraceae bacterium W19 (LC094483, 100% similarity), was detected in epilithic biofilms of the Tama River [
17]. The AAP community of epilithic biofilms in Lake Baikal also included Burkholderiales and Gemmatimonadetes in contrast to the Tama River.
Rhodopsin-containing bacteria in freshwater bodies are poorly studied. The diversity of phyla of this phototrophic group in Lake Baikal is comparable to other investigated freshwater bodies. Based on BLASTp analysis, the rhodopsin-containing bacterial community of Lake Baikal included phylum Planctomycetes (
Figure 7). The closest homologue was detected in the sea water, off the coast of Alicante (Spain) (RZO64088, 72% similarity) [
99]. Rhodopsin-containing Planctomycetes were also detected in littoral microbial mats of high latitude freshwater lakes (Canada) [
100].
Thus, there were detected both
pufM-containing and rhodopsin-containing Planctomycetes in Lake Baikal. According to Zeng et al. (2020), “there is emerging genomic evidence that (bacterio-)chlorophyll- and proton-pumping rhodopsin-based phototrophic systems can coexist in a single bacterium” [
91]. At the same time, due to the fact that
pufM- and rhodopsin-containing Planctomycetes in Lake Baikal had different closest homologues, we can propose that the type of phototrophic system might depend on the distinct class, order, family, genus, or species affiliation of Planctomycetes strain, just like in Proteobacteria. In Proteobacteria, there are representatives of aerobic anoxygenic phototrophs [
16,
17,
18,
19,
20,
21,
22] as well as rhodopsin-containing bacteria [
29,
30,
100].
Actinobacteria are known to be the most abundant rhodopsin-containing bacteria in previously studied freshwater ecosystems [
22,
35,
36]. In Lake Baikal, they were not detected. Nevertheless, 16S rRNA diversity analysis revealed that Actinobacteria comprised 21% of all obtained sequences and were equal in epilithon, neuston, and plankton. Among them, 26% had close homologues (100% similarity) possessing the rhodopsin gene, especially, Actinobacterium SCGC AAA280-O03 (HQ663639, 100% similarity with OTU 10) (
Figure 8). Therefore, the Baikal representatives could also have this gene. Most likely, the Actinobacteria rhodopsin gene sequences were among those unidentified to the phylum level. According to the 16S rRNA data, Actinobacteria were a considerable part of the rhodopsin-containing phototrophic community of Lake Baikal.
Freshwater representatives of the SAR 11 cluster (Pelagibacterales) are also known to have rhodopsin [
29,
30]. 16S rRNA sequences of the SAR 11 bacteria were detected in neuston and plankton of Lake Baikal (1.1% of all sequences) but not in epilithon. The closest homologue was the rhodopsin-containing strain Alphaproteobacterium SCGC AAA 280-B11 (HQ663835, 100% similarity) [
22] (
Figure 8). We suppose that the Baikal SAR 11 bacteria also might possess a rhodopsin gene.
Data regarding the diversity of rhodopsin-containing bacteria in epilithic biofilms are scarce. Vigneron et al. (2018) studied genes coding for microbial rhodopsins in littoral microbial mats of high latitude freshwater lakes (Canada) [
100]. Rhodopsin genes were affiliated to Actinobacteria, Acidobacteria, Bacteroidetes, Cyanobacteria, Planctomycetes, and Alpha- and Betaproteobacteria. Epilithon of Lake Baikal included the same phyla detected in neuston and plankton: Bacteroidetes, Proteobacteria, Verrucomicrobia, and Planctomycetes. Nevertheless, UPGMA and NMDS analysis indicated significant differences between water microbiomes and epilithon at the phylotype level (
Figure 6A,B). Similar to the AAP bacterial communities, there were “typical epilithon phylotypes” and “typical water phylotypes”. For example, OTUs 1901, 2299 (Bacteroidetes, Cytophagales,
Aquirufa) and 1360 (Bacteroidetes, Cytophagales,
Runella) were exclusively in neuston and plankton, whereas OTU 678 (Bacteroidetes, Cytophagales,
Runella) only in epilithon (
Table 5).
The results showed that epilithon differed from neuston and plankton mainly at the genus and phylotype level; taxonomic composition of microbial communities on the higher levels was pretty similar. That was true for 16S rRNA gene as well as for
pufM and rhodopsin. It was an interesting finding needed to be explained. It is well known that there are free-living bacteria and attached forms, which need a substrate to adherence to. These two types of bacteria differ taxonomically at the phylotype level [
101]. The first type is obviously detected in plankton, and the second on the surfaces including stones. The second type takes part in the forming of epilithic biofilms. That is why epilithon and plankton in Lake Baikal were taxonomically distinct at the genus and phylotype level.