Genomics and Virulence of Klebsiella pneumoniae Kpnu95 ST1412 Harboring a Novel Incf Plasmid Encoding Blactx-M-15 and Qnrs1 Causing Community Urinary Tract Infection
Abstract
1. Introduction
2. Materials and Methods
2.1. K. pneumoniae UTI Strain, Growth Conditions and Antibiotic Susceptibility Testing
2.2. Purification of the ESBL-Encoding Plasmid and Transformation
2.3. Curing of pKpnU95 and Plasmid Reconstitution
2.4. Fitness Studies
2.5. Virulence Assessment in the Caenorhabditis Elegans Host–Pathogen Model
2.6. WGS of KpnU95
2.7. Complete Sequencing of Plasmid pKpnu95
2.8. Development of a Targeted PCR Screening for the Presence of pKpnu95
2.9. Bioinformatics Analysis
2.9.1. GenBank Submission
2.9.2. Analysis of pKpnU95-Closely Related Plasmids
2.9.3. Phylogenetic Analysis of pKpnU95-Closely Related Plasmids
3. Results
3.1. Kpnu95 Isolate and Generation of a Cured and a Reconstituted Cured Strain
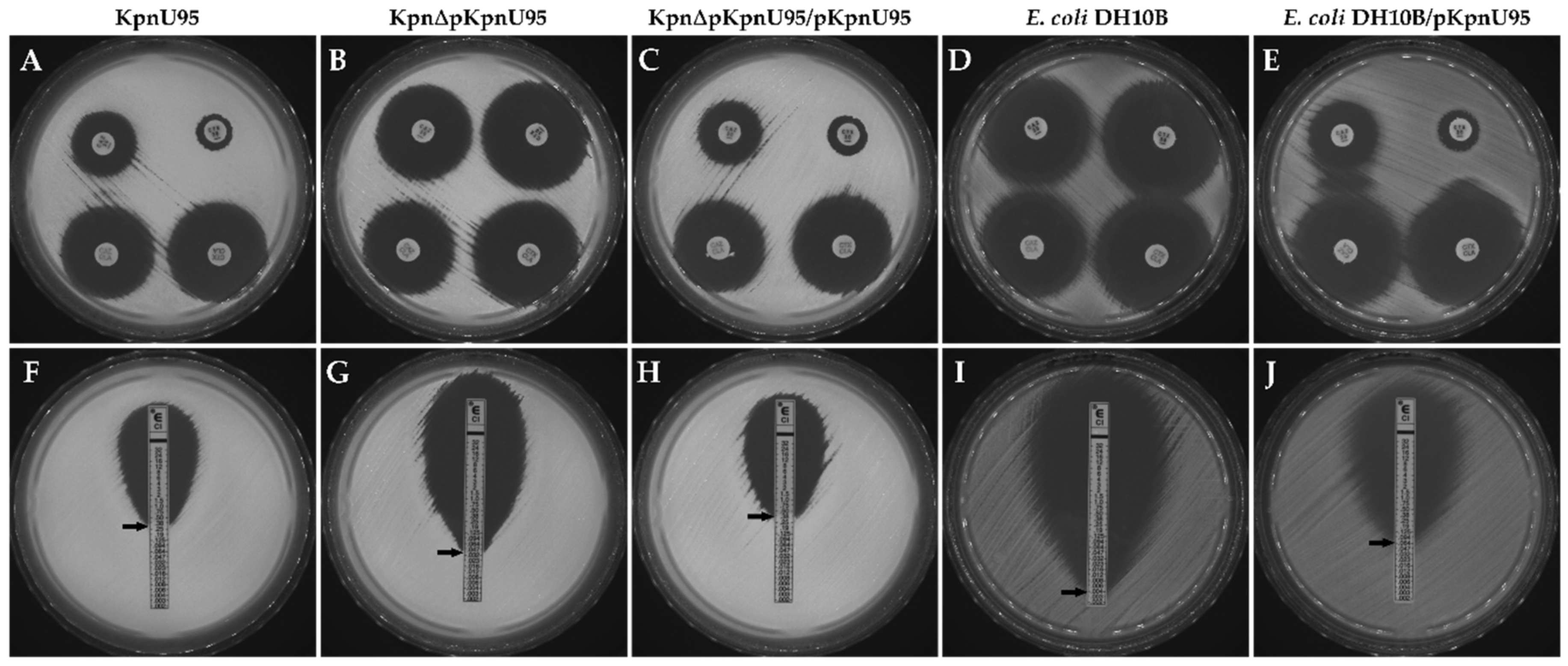
3.2. Antibiotic Susceptibility
3.3. Fitness and In Vivo Virulence of K. pneumoniae KpnU95 Strains
3.4. Genomic Characterization of KpnU95
3.4.1. KpnU95 Chromosome
3.4.2. Complete Sequence of pKpnU95
3.5. Analysis of pKpnU95 Closely Related Global Plasmids
3.6. Co-Evolution of pKpnU95-Related Plasmids and Their K. pneumoniae Clonal Lineages Hosts
4. Discussion
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Boucher, H.W.; Talbot, G.H.; Bradley, J.S.; Edwards, J.E.; Gilbert, D.; Rice, L.B.; Scheld, M.; Spellberg, B.; Bartlett, J. Bad Bugs, No Drugs: No ESKAPE! An Update from the Infectious Diseases Society of America. Clin. Infect. Dis. 2009, 48, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Willyard, C. Drug-resistant bacteria ranked. Nature 2017, 543, 15. [Google Scholar] [CrossRef]
- Wyres, K.L.; Lam, M.M.C.; Holt, K.E. Population genomics of Klebsiella pneumoniae. Nat. Rev. Microbiol. 2020, 18, 344–359. [Google Scholar] [CrossRef]
- Paczosa, M.K.; Mecsas, J. Klebsiella pneumoniae: Going on the Offense with a Strong Defense. Microbiol. Mol. Biol. Rev. 2016, 80, 629–661. [Google Scholar] [CrossRef]
- Navon-Venezia, S.; Kondratyeva, K.; Carattoli, A. Klebsiella pneumoniae: A major worldwide source and shuttle for antibiotic resistance. FEMS Microbiol. Rev. 2017, 41, 252–275. [Google Scholar] [CrossRef]
- Flores-Mireles, A.L.; Walker, J.N.; Caparon, M.; Hultgren, S.J. Urinary tract infections: Epidemiology, mechanisms of infection and treatment options. Nat. Rev. Microbiol. 2015, 13, 269–284. [Google Scholar] [CrossRef]
- Koksal, E.; Tulek, N.; Sonmezer, M.C.; Temocin, F.; Bulut, C.; Hatipoglu, C.; Erdinc, F.S.; Ertem, G. Investigation of risk factors for community-acquired urinary tract infections caused by extended-spectrum beta-lactamase Escherichia coli and Klebsiella species. Investig. Clin. Urol. 2019, 60, 46. [Google Scholar] [CrossRef]
- Xercavins, M.; Jiménez, E.; Padilla, E.; Riera, M.; Freixas, N.; Boix-Palop, L.; Pérez, J.; Calbo, E. High clonal diversity of ESBL-producing Klebsiella pneumoniae isolates from clinical samples in a non-outbreak situation. A cohort study. Antimicrob. Resist. Infect. Control 2020, 9, 5. [Google Scholar] [CrossRef]
- Zhang, J.; Zhou, K.; Zheng, B.; Zhao, L.; Shen, P.; Ji, J.; Wei, Z.; Li, L.; Zhou, J.; Xiao, Y. High prevalence of ESBL-producing klebsiella pneumoniae causing community-onset infections in China. Front. Microbiol. 2016, 7, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Russo, T.A.; Marr, C.M. Hypervirulent Klebsiella pneumoniae. Clin. Microbiol. Rev. 2019, 32, 1–42. [Google Scholar] [CrossRef] [PubMed]
- Choby, J.E.; Howard-Anderson, J.; Weiss, D.S. Hypervirulent Klebsiella pneumoniae—Clinical and molecular perspectives. J. Intern. Med. 2020, 287, 283–300. [Google Scholar] [CrossRef]
- Zhou, K.; Yu, W.; Shen, P.; Lu, H.; Wang, B.; Rossen, J.W.A.; Xiao, Y. A novel Tn1696-like composite transposon (Tn6404) harboring bla IMP-4 in a Klebsiella pneumoniae isolate carrying a rare ESBL gene bla SFO-1. Sci. Rep. 2017, 7, 1–7. [Google Scholar] [CrossRef] [PubMed]
- Shaidullina, E.; Shelenkov, A.; Yanushevich, Y.; Mikhaylova, Y.; Shagin, D.; Alexandrova, I.; Ershova, O.; Akimkin, V.; Kozlov, R.; Edelstein, M. Antimicrobial Resistance and Genomic Characterization of OXA-48- and CTX-M-15-Co-Producing Hypervirulent Klebsiella pneumoniae ST23 Recovered from Nosocomial Outbreak. Antibiotics 2020, 9, 862. [Google Scholar] [CrossRef] [PubMed]
- Brosh-Nissimov, T.; Navon-Venezia, S.; Keller, N.; Amit, S. Risk analysis of antimicrobial resistance in outpatient urinary tract infections of young healthy adults. J. Antimicrob. Chemother. 2019, 74, 499–502. [Google Scholar] [CrossRef] [PubMed]
- EUCAST Breakpoint Tables for Interpretation of MICs and Zone Diameters. 2019. Available online: http://www.eucast.org/fileadmin/src/media/PDFs/EUCAST_files/Breakpoint_tables/v_9.0_Breakpoint_Tables.pdf (accessed on 4 August 2019).
- Woodford, N.; Fagan, E.J.; Ellington, M.J. Multiplex PCR for rapid detection of genes encoding CTX-M extended-spectrum β-lactamases. J. Antimicrob. Chemother. 2006, 57, 154–155. [Google Scholar] [CrossRef]
- Leavitt, A.; Chmelnitsky, I.; Ofek, I.; Carmeli, Y.; Navon-Venezia, S. Plasmid pKpQIL encoding KPC-3 and TEM-1 confers carbapenem resistance in an extremely drug-resistant epidemic Klebsiella pneumoniae strain. J. Antimicrob. Chemother. 2009, 65, 243–248. [Google Scholar] [CrossRef] [PubMed]
- Versalovic, J.; Koeuth, T.; Lupski, J.R. Distribution of repetitive DNA sequences in eubacteria and application to fingerprinting of bacterial genomes. Nucleic Acids Res. 1991, 19, 6823–6831. [Google Scholar] [CrossRef]
- Cohen, D.; Mechold, U.; Nevenzal, H.; Yarmiyhu, Y.; Randall, T.E.; Bay, D.C.; Rich, J.D.; Parsek, M.R.; Kaever, V.; Harrison, J.J.; et al. Oligoribonuclease is a central feature of cyclic diguanylate signaling in Pseudomonas aeruginosa. Proc. Natl. Acad. Sci. USA 2015, 112, 11359–11364. [Google Scholar] [CrossRef]
- Chutipongtanate, S.; Thongboonkerd, V. Systematic comparisons of artificial urine formulas for in vitro cellular study. Anal. Biochem. 2010, 402, 110–112. [Google Scholar] [CrossRef]
- Teitzel, G.M.; Geddie, A.; De Long, S.K.; Kirisits, M.J.; Whiteley, M.; Parsek, M.R. Survival and growth in the presence of elevated copper: Transcriptional profiling of copper-stressed Pseudomonas aeruginosa. J. Bacteriol. 2006, 188, 7242–7256. [Google Scholar] [CrossRef]
- Solis, G.M.; Petrascheck, M. Measuring Caenorhabditis elegans Life Span in 96 Well Microtiter Plates. J. Vis. Exp. 2011, 18, 2496. [Google Scholar] [CrossRef]
- Schindelin, J.; Arganda-Carreras, I.; Frise, E.; Kaynig, V.; Longair, M.; Pietzsch, T.; Preibisch, S.; Rueden, C.; Saalfeld, S.; Schmid, B.; et al. Fiji: An open-source platform for biological-image analysis. Nat. Methods 2012, 9, 676–682. [Google Scholar] [CrossRef]
- Bolger, A.M.; Lohse, M.; Usadel, B. Trimmomatic: A flexible trimmer for Illumina sequence data. Bioinformatics 2014, 30, 2114–2120. [Google Scholar] [CrossRef] [PubMed]
- Bankevich, A.; Nurk, S.; Antipov, D.; Gurevich, A.A.; Dvorkin, M.; Kulikov, A.S.; Lesin, V.M.; Nikolenko, S.I.; Pham, S.; Prjibelski, A.D.; et al. SPAdes: A new genome assembly algorithm and its applications to single-cell sequencing. J. Comput. Biol. 2012, 19, 455–477. [Google Scholar] [CrossRef]
- Wick, R.R.; Judd, L.M.; Gorrie, C.L.; Holt, K.E. Unicycler: Resolving bacterial genome assemblies from short and long sequencing reads. PLoS Comput. Biol. 2017, 13, 1–22. [Google Scholar] [CrossRef] [PubMed]
- Tatusova, T.; Dicuccio, M.; Badretdin, A.; Chetvernin, V.; Nawrocki, E.P.; Zaslavsky, L.; Lomsadze, A.; Pruitt, K.D.; Borodovsky, M.; Ostell, J. NCBI prokaryotic genome annotation pipeline. Nucleic Acids Res. 2016, 44, 6614–6624. [Google Scholar] [CrossRef] [PubMed]
- Aziz, R.K.; Bartels, D.; Best, A.A.; DeJongh, M.; Disz, T.; Edwards, R.A.; Formsma, K.; Gerdes, S.; Glass, E.M.; Kubal, M.; et al. The RAST Server: Rapid annotations using subsystems technology. BMC Genom. 2008, 9, 75. [Google Scholar] [CrossRef]
- Bousquet, A.; Henquet, S.; Compain, F.; Genel, N.; Arlet, G.; Decré, D. Partition locus-based classification of selected plasmids in Klebsiella pneumoniae, Escherichia coli and Salmonella enterica spp.: An additional tool. J. Microbiol. Methods 2015, 110, 85–91. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Xie, Y.; Liu, M.; Tai, C.; Sun, J.; Deng, Z.; Ou, H.Y. OriTfinder: A web-based tool for the identification of origin of transfers in DNA sequences of bacterial mobile genetic elements. Nucleic Acids Res. 2018, 46, W229–W234. [Google Scholar] [CrossRef] [PubMed]
- Wick, R.R.; Heinz, E.; Holt, K.E.; Wyres, K.L. Kaptive web: User-Friendly capsule and lipopolysaccharide serotype prediction for Klebsiella genomes. J. Clin. Microbiol. 2018, 56, 1–12. [Google Scholar] [CrossRef]
- Alikhan, N.-F.; Petty, N.K.; Ben Zakour, N.L.; Beatson, S.A. BLAST Ring Image Generator (BRIG): Simple prokaryote genome comparisons. BMC Genom. 2011, 12, 402. [Google Scholar] [CrossRef]
- Croucher, N.J.; Page, A.J.; Connor, T.R.; Delaney, A.J.; Keane, J.A.; Bentley, S.D.; Parkhill, J.; Harris, S.R. Rapid phylogenetic analysis of large samples of recombinant bacterial whole genome sequences using Gubbins. Nucleic Acids Res. 2015, 43, e15. [Google Scholar] [CrossRef]
- Trifinopoulos, J.; Nguyen, L.T.; von Haeseler, A.; Minh, B.Q. W-IQ-TREE: A fast online phylogenetic tool for maximum likelihood analysis. Nucleic Acids Res. 2016, 44, W232–W235. [Google Scholar] [CrossRef]
- Letunic, I.; Bork, P. Interactive Tree of Life (iTOL) v4: Recent updates and new developments. Nucleic Acids Res. 2019, 47, W256–W259. [Google Scholar] [CrossRef]
- Weinstein, M.P.; Patel, J.B.; Bobenchik, A.M.; Campeau, S.; Cullen, S.K.; Galas, M.F.; Gold, H.; Humphries, R.M.; Kirn, T.J., Jr.; Lewis, J.S., II; et al. M100 Performance Standards for Antimicrobial Susceptibility Testing, 29th ed.; CLSI: Annapolis Junction, MD, USA, 2019; pp. 1–25. [Google Scholar]
- Zenno, S.; Koike, H.; Kumar, A.N.; Jayaraman, R.; Tanokura, M.; Saigo, K. Biochemical characterization of NfsA, the Escherichia coli major nitroreductase exhibiting a high amino acid sequence homology to Frp, a Vibrio harveyi flavin oxidoreductase. J. Bacteriol. 1996, 178, 4508–4514. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Zhang, H.; Ning, J.; Sajid, A.; Cheng, G.; Yuan, Z.; Hao, H. The nature and epidemiology of OqxAB, a multidrug efflux pump. Antimicrob. Resist. Infect. Control 2019, 8, 1–13. [Google Scholar] [CrossRef]
- Sahly, H.; Navon-Venezia, S.; Roesler, L.; Hay, A.; Carmeli, Y.; Podschun, R.; Hennequin, C.; Forestier, C.; Ofek, I. Extended-Spectrum β-Lactamase Production is Associated with an Increase in Cell Invasion and Expression of Fimbrial Adhesins in Klebsiella pneumoniae. Antimicrob. Agents Chemother. 2008, 52, 3029–3034. [Google Scholar] [CrossRef] [PubMed]
- Sichtig, H.; Minogue, T.; Yan, Y.; Stefan, C.; Hall, A.; Tallon, L.; Sadzewicz, L.; Nadendla, S.; Klimke, W.; Hatcher, E.; et al. FDA-ARGOS is a database with public quality-controlled reference genomes for diagnostic use and regulatory science. Nat. Commun. 2019, 10, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Long, S.W.; Olsen, R.J.; Eager, T.N.; Beres, S.B.; Zhao, P.; Davis, J.J.; Brettin, T.; Xia, F.; Musser, J.M. Population Genomic Analysis of 1777 Extended-Spectrum Beta-Lactamase-Producing Klebsiella pneumoniae Isolates, Houston, Texas: Unexpected Abundance of Clonal Group 307. mBio 2017, 8, 1–18. [Google Scholar] [CrossRef]
- Huang, X.; Cheng, X.; Sun, P.; Tang, C.; Ni, F.; Liu, G. Characteristics of NDM-1-producing Klebsiella pneumoniae ST234 and ST1412 isolates spread in a neonatal unit. BMC Microbiol. 2018, 18, 186. [Google Scholar] [CrossRef]
- Dziri, O.; Alonso, C.A.; Dziri, R.; Gharsa, H.; Maraoub, A.; Torres, C.; Chouchani, C. Metallo-β-lactamases and class D carbapenemases in south-east Tunisia: Implication of mobile genetic elements in their dissemination. Int. J. Antimicrob. Agents 2018, 52, 871–877. [Google Scholar] [CrossRef]
- El Fertas-Aissani, R.; Messai, Y.; Alouache, S.; Bakour, R. Virulence profiles and antibiotic susceptibility patterns of Klebsiella pneumoniae strains isolated from different clinical specimens. Pathol. Biol. 2013, 61, 209–216. [Google Scholar] [CrossRef]
- Russo, T.A.; Olson, R.; MacDonald, U.; Beanan, J.; Davidsona, B.A. Aerobactin, but not yersiniabactin, salmochelin, or enterobactin, enables the growth/survival of hypervirulent (hypermucoviscous) Klebsiella pneumoniae ex vivo and in vivo. Infect. Immun. 2015, 83, 3325–3333. [Google Scholar] [CrossRef] [PubMed]
- Mahren, S.; Schnell, H.; Braun, V. Occurrence and regulation of the ferric citrate transport system in Escherichia coli B, Klebsiella pneumoniae, Enterobacter aerogenes, and Photorhabdus luminescens. Arch. Microbiol. 2005, 184, 175–186. [Google Scholar] [CrossRef]
- Scott, E.; Holden-Dye, L.; O’Connor, V.; Wand, M.E. Intra Strain Variation of the Effects of Gram-Negative ESKAPE Pathogens on Intestinal Colonization, Host Viability, and Host Response in the Model Organism Caenorhabditis elegans. Front. Microbiol. 2020, 10, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Lavigne, J.-P.; Cuzon, G.; Combescure, C.; Bourg, G.; Sotto, A.; Nordmann, P. Virulence of Klebsiella pneumoniae Isolates Harboring blaKPC-2 Carbapenemase Gene in a Caenorhabditis elegans Model. PLoS ONE 2013, 8, e67847. [Google Scholar] [CrossRef] [PubMed]
- Kaplan, E.; Sela, N.; Doron-Faigenboim, A.; Navon-Venezia, S.; Jurkevitch, E.; Cytryn, E. Genomic and functional characterization of qnr-encoding plasmids from municipal wastewater biosolid Klebsiella pneumoniae isolates. Front. Microbiol. 2015, 6, 1–12. [Google Scholar] [CrossRef]
- Shanmugam, D.; Esak, S.B.; Narayanaswamy, A. Molecular characterisation of nfsA gene in nitrofurantoin resistant uropathogens. J. Clin. Diagn. Res. 2016, 10, DC05–DC09. [Google Scholar] [CrossRef]
- Mckinnell, J.A.; Stollenwerk, N.S.; Jung, C.W.; Miller, L.G. Nitrofurantoin compares favorably to recommended agents as empirical treatment of uncomplicated urinary tract infections in a decision and cost analysis. Mayo Clin. Proc. 2011, 86, 480–488. [Google Scholar] [CrossRef]
- Martin, R.M.; Bachman, M.A. Colonization, infection, and the accessory genome of Klebsiella pneumoniae. Front. Cell. Infect. Microbiol. 2018, 8, 1–15. [Google Scholar] [CrossRef] [PubMed]
- Hyre, A.N.; Kavanagh, K.; Kock, N.D.; Donati, G.L.; Subashchandrabose, S. Copper is a host effector mobilized to urine during urinary tract infection to impair bacterial colonization. Infect. Immun. 2017, 85, 1–14. [Google Scholar] [CrossRef] [PubMed]
- Osawa, M.; Hashimoto, K.; Kawasaki, H.; Ito, K.; Dohmae, N.; Ishitani, R. ATP-dependent modulation of MgtE in Mg2+ homeostasis. Nat. Commun. 2017, 8, 1–11. [Google Scholar]
- Ble, C.; Lagarde, F. RcnB Is a Periplasmic Protein Essential for Maintaining Intracellular Ni and Co Concentrations in Escherichia coli. J. Bacteriol. 2011, 193, 3785–3793. [Google Scholar]
- Bellotti, D.; Rowińska-Żyrek, M.; Maurizio, R. Novel insights into the metal binding ability of ZinT periplasmic protein from Escherichia coli and Salmonella enterica. R. Soc. Chem. 2020, 49, 9393–9403. [Google Scholar] [CrossRef] [PubMed]
- Ipe, D.S.; Horton, E.; Ulett, G.C. The Basics of Bacteriuria: Strategies of Microbes for Persistence in Urine. Front. Cell. Infect. Microbiol. 2016, 6, 1–12. [Google Scholar] [CrossRef]
- San Millan, A. Evolution of Plasmid-Mediated Antibiotic Resistance in the Clinical Context. Trends Microbiol. 2018, 26, 978–985. [Google Scholar] [CrossRef] [PubMed]
- Arredondo-Alonso, S.; Willems, R.J.; van Schaik, W.; Schürch, A.C. On the (im)possibility of reconstructing plasmids from whole-genome short-read sequencing data. Microb. Genom. 2017, 3, 1–8. [Google Scholar] [CrossRef]
- Gullberg, E.; Albrecht, L.M.; Karlsson, C.; Sandegren, L.; Andersson, D.I. Selection of a Multidrug Resistance Plasmid by Sublethal Levels of Antibiotics and Heavy Metals. mBio 2014, 5, 19–23. [Google Scholar] [CrossRef]
- Wu, F.; Ying, Y.; Yin, M.; Jiang, Y.; Wu, C.; Qian, C.; Chen, Q.; Shen, K.; Cheng, C.; Zhu, L.; et al. Molecular Characterization of a Multidrug-Resistant Klebsiella pneumoniae Strain R46 Isolated from a Rabbit. Int. J. Genom. 2019, 2019, 1–12. [Google Scholar] [CrossRef]



| Antibiotic Agent | MIC (μg/mL) | ||||
|---|---|---|---|---|---|
| KpnU95 | KpnΔpKpnU95 1 | KpnΔpKpnU95/pKpnU95 2 | E. coli DH10B | E. coli DH10B/pKpnU95 3 | |
| Ampicillin | >16 | >16 | >16 | <8 | >16 |
| Piperacillin | >64 | <16 | >64 | <16 | >64 |
| Cefazolin | >16 | 2 | >16 | 4 | >16 |
| Ceftazidime | 16 | <1 | 16 | <1 | 16 |
| Ceftriaxone | >32 | <0.5 | >32 | <0.5 | >32 |
| Ampicillin/Sulbactam | 16/8 | 8/4 | 16/8 | <4/2 | 16/8 |
| Piperacillin/Tazobactam | <16/4 | <8/4 | <8/4 | <8/4 | <8/4 |
| Ticarcillin/ Clavulanic acid | 32/2 | <8/2 | 32/2 | <8/2 | 32/2 |
| Aztreonam | >16 | <1 | >16 | <1 | 16 |
| Imipenem | <0.5 | <0.5 | <0.5 | <0.5 | <0.5 |
| Ertapenem | <0.25 | <0.25 | <0.25 | <0.25 | <0.25 |
| Meropenem | <0.5 | <0.5 | <0.5 | <0.5 | <0.5 |
| Doripenem | <0.5 | <0.5 | <0.5 | <0.5 | <0.5 |
| Amikacin | <8 | <8 | <8 | <8 | <8 |
| Tobramycin | <2 | <2 | <2 | <2 | <2 |
| Gentamicin | <2 | <2 | <2 | <2 | <2 |
| Ciprofloxacin 4 | 0.38 | 0.032 | 0.38 | 0.002 | 0.064 |
| Nitrofurantoin | 64 | 64 | 64 | <32 | <32 |
| Trimethoprim/ Sulfamethoxazole | >4/76 | <2/38 | >4/76 | <2/38 | >4/76 |
| Tetracycline | <4 | <4 | <4 | <4 | <4 |
| Minocycline | 2 | 2 | 2 | <1 | <1 |
| Tigecycline | <1 | <1 | <1 | <1 | <1 |
| Fosfomicin 5 | ≤16 | ≤16 | ≤16 | ≤16 | ≤16 |
| Category | Functional Group | Genes 1 | Function/Resistance to |
|---|---|---|---|
| Virulence | Fimbria adhesins | fimBEAICDFGHK | Type 1 fimbria |
| Fimbria adhesins and biofilm formation | mrkABCDFJIH | Type 3 fimbria | |
| ecpRABCDE | E. coli common pilus | ||
| pilABC, pilQONM, pilT | Type VI pilus | ||
| Secretion systems (SS) | gspCDEFGHIJLMN, pulS, pulO | Type II SS | |
| tssBCKDHIMAFGJ, tssE | Type VI SS | ||
| Toxins–antitoxin systems | doc-phd, hipAB, ratAB, RelE-like TA | Type II TA | |
| YkfI-YfjZ | Type IV TA | ||
| ecnAB | Lipoprotein toxin entericidin | ||
| Siderophores | entABCDEFHS2, fepABCDGS2 | Enterobactin | |
| Siderophore receptors | iutA, iroN | Aerobactin and Salmochelin receptors | |
| Iron-acquisition and transport | msrQ, hmuRS, foxA, cirA, fhuD, yfeX, feoABC, efeO | heme-binding subunit, iron-acquisition proteins, hemin transport proteins | |
| Outer membrane porins | ompA, ompC, ompF, ompN, ompW | Iron uptake | |
| Transcriptional regulators | fur | Ferric iron uptake transcriptional regulator | |
| rcsA, rscB | Regulators of capsule synthesis/iron uptake | ||
| Host-associated metabolism | ureDABCEFG, urtABCDE | Urease, urea ABC transporter | |
| hpxB, hpxDE, hpxR, hpxO hyuE, pucU, codB | Allantoin/purine metabolism and transport | ||
| Virulence factors | pgaABCD | Biofilm synthesis | |
| msgA | Intracellular survival | ||
| hha | Hemolysin expression modulator | ||
| mviMN, yihY | Putative virulence factors | ||
| Persistence | Heavy metal resistance and homeostasis | chrAB | Chromate resistance and transport |
| cpxRA, cusF, cutE, cutC, cutF, cueO, cueR, copA, copCD, scsABCD | Copper-binding response regulator systems, copper-binding protein, copper homeostasis, copper resistance | ||
| cusRS | Copper/silver response regulator system | ||
| zraPSR | Zinc/copper resistance | ||
| corA, corB, corC mgtA, mgtE | Magnesium-cobalt transporters Mg/Co/Ni transporter | ||
| rcnA, rcnB | Nickel/cobalt efflux Nickel/cobalt homeostasis | ||
| cbtJKL, cbiMNQO | Cobalt ABC transporter, cobalt ECF transporter | ||
| nikABCDER, dppBC, oppDF | Nickel ABC transporter | ||
| mntH, mntS, mntP, mntR sitABC, mtsC | Manganese efflux pump, transport regulator, manganese ABC transporters | ||
| znuABC, zupT, zntB, zitB, fieF, zur, zntR | Zinc transporters/uptake regulators | ||
| zntA, czcD | Zn(II)/Cd(II)/Pb(II) translocating P-type ATPase, Cobalt-zinc-cadmium resistance protein/cation transporter | ||
| Metalloid resistance | arsRBC, yffB | Arsenate reductase, regulator, arsenic efflux pump | |
| tehAB, ydcL, yebE | Tellurite resistance | ||
| tsgA | Putative selenite and tellurite transporter | ||
| UV | umuCD | Error-prone, lesion bypass DNA polymerase V | |
| Antibiotic resistance | ARG | blaSHV-1 | Penicillin |
| fosA | Fosfomycin | ||
| nfsA | Nitrofurantoin | ||
| Multidrug transporters | oqxAB | Quinolone resistance | |
| macAB | Macrolide resistance | ||
| mexB | Acriflavine resistance | ||
| eefABC | Tolerance response to inorganic acid | ||
| acrAB, acrD, acrEF, tolC | Resistance to beta-lactams, aminoglycosides, fluoroquinolones, tetracycline, chloramphenicol, acriflavine | ||
| mdtABCD, mdtG, mdtH, mdtJI, mdtK, mdtL, mdtM, mdtNOP, mdtQ | Transporters of wide range of multidrug and disinfectant components 3 | ||
| emrRAB, emrD | Resistance to nalidixic acid, thiolactomycin, novobiocin Participates in a low energy shock adaptive response | ||
| Transcriptional activators | marRAB, soxRS, ramA, rarA | Activation of both antibiotic resistance and oxidative stress genes |
| Category | Functional Group | Genes 1 | Function/Resistance to |
|---|---|---|---|
| Antibiotic resistance | ARG | blaCTX-M-15 | ESBL |
| mph(A) | Macrolides | ||
| strA, strB, aadA2, aph(3′)-Ia | Aminoglycosides | ||
| sul1, sul2 | Sulphonamides | ||
| qnrS1 | Quinolones | ||
| dfrA12 | Trimethoprim-sulfa | ||
| Multidrug transporter | qacEΔ1 (emrE) | Small multidrug resistance (SMR) efflux transporter | |
| Virulence | Iron | fecIRABCDE | Ferric citrate transport system |
| Persistence | Heavy metal resistance and homeostasis | chrA | Chromate transport |
| pcoABCDRS, copA, copG | Copper resistance Metal-binding protein | ||
| cusCFBA | Copper-silver efflux system proteins | ||
| cusRS, pcoE (silE), silP | Copper-silver response regulator system, copper-silver binding protein, Ag(+)/copper-translocating P-type ATPase | ||
| Metalloid resistance | arsRBC, arsH | Arsenate reductase, arsenic resistance, regulator, arsenic efflux pump | |
| Toxins-antitoxin | relB, yafO-like2 | Antitoxin and putative toxin | |
| Others | cld | Chlorite dismutase | |
| UV | umuCD | Error-prone, lesion bypass DNA polymerase V |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Gancz, A.; Kondratyeva, K.; Cohen-Eli, D.; Navon-Venezia, S. Genomics and Virulence of Klebsiella pneumoniae Kpnu95 ST1412 Harboring a Novel Incf Plasmid Encoding Blactx-M-15 and Qnrs1 Causing Community Urinary Tract Infection. Microorganisms 2021, 9, 1022. https://doi.org/10.3390/microorganisms9051022
Gancz A, Kondratyeva K, Cohen-Eli D, Navon-Venezia S. Genomics and Virulence of Klebsiella pneumoniae Kpnu95 ST1412 Harboring a Novel Incf Plasmid Encoding Blactx-M-15 and Qnrs1 Causing Community Urinary Tract Infection. Microorganisms. 2021; 9(5):1022. https://doi.org/10.3390/microorganisms9051022
Chicago/Turabian StyleGancz, Ayala, Kira Kondratyeva, Dorit Cohen-Eli, and Shiri Navon-Venezia. 2021. "Genomics and Virulence of Klebsiella pneumoniae Kpnu95 ST1412 Harboring a Novel Incf Plasmid Encoding Blactx-M-15 and Qnrs1 Causing Community Urinary Tract Infection" Microorganisms 9, no. 5: 1022. https://doi.org/10.3390/microorganisms9051022
APA StyleGancz, A., Kondratyeva, K., Cohen-Eli, D., & Navon-Venezia, S. (2021). Genomics and Virulence of Klebsiella pneumoniae Kpnu95 ST1412 Harboring a Novel Incf Plasmid Encoding Blactx-M-15 and Qnrs1 Causing Community Urinary Tract Infection. Microorganisms, 9(5), 1022. https://doi.org/10.3390/microorganisms9051022






