Simple Summary
Yak is the dominant animal species in China’s plateau regions. The development of the yak industry is conducive to protecting the ecosystem of the Qinghai–Tibet Plateau and promoting local economic development. However, the slow growth and development of yak have seriously affected the development of the yak industry. Structural variation (SV) has been widely applied in livestock breeding for growth traits. Therefore, it is of great significance to use SV to improve yak growth traits in yak breeding. Meanwhile, the GHR gene plays an important role in the formation and normal development of bones. This study associated the correlation between the yak GHR SV gene and growth traits and confirmed that GHR-SV can be used as a molecular marker for the early reproduction of yaks. This study provides a theoretical basis for the early growth and development of yaks.
Abstract
The growth hormone receptor (GHR) is a member of the cytokine/hematopoietic factor receptor superfamily, which plays an important role in the growth and development, immunity, and metabolism of animals. This study identified a 246 bp deletion variant in the intronic region of the GHR gene, and three genotypes, including type II, type ID, and type DD, were observed. Genotype analysis of structural variation (SV) was performed on 585 individuals from 14 yak breeds, and it was found that 246 bp deletion was present in each breed. The II genotype was dominant in all yak breeds except for SB yak. The association analysis of gene polymorphisms and growth traits in the ASD yak population showed that the 246 bp SV was significantly associated with body length at 6 months (p < 0.05). GHR messenger RNA (mRNA) was expressed in all the tested tissues, with significantly higher levels in the liver, muscle, and fat than in other organs. The results of transcription activity showed that the luciferase activity of the pGL4.10-DD vector was significantly higher than that of the pGL4.10-II vector (p < 0.05). Additionally, the transcription-factor binding prediction results showed that the SV in the runt-related transcription factor 1 (Runx1) transcription-factor binding site may affect the transcriptional activity of the GHR gene, regulating yak growth and development. This study showed that the novel SV of the GHR gene could be used as a candidate molecular marker for the selection of the early growth trait in ASD yak.
1. Introduction
Yak (Bos grunniens) mainly lives in the Qinghai–Tibet Plateau at an altitude of 3000–5500 m [1]. In China, there are about 16 million yaks, accounting for about 95% of the world’s population [2]. It is an important and dominant animal species in plateau animal husbandry, which can provide meat, milk, fur, etc. [3]. Among them, yak meat is one of the main resources of the plateau animal husbandry economy [4]. Compared with cattle beef, yak meat is high in protein and energy, low in fat, and rich in amino acids, and the habitat of yak is least affected by humans. Therefore, yak meat is considered a natural green food, which meets people’s demand for green and healthy meat quality [5]. As a plateau variety, yak can protect the Qinghai–Tibet Plateau ecosystem and promote local economic development. Compared with cattle, the growth rate of yak is slow due to the lack of an efficient yak breeding program to improve growth traits, which affects the yield of yak meat. Therefore, it is of great significance to study the growth traits of yak.
The growth hormone receptor (GHR) gene is an essential growth-related candidate gene that plays a critical role in the early stages of animal growth and development [6]. The growth hormone (GH)–insulin-like growth factor (IGF) growth axis is crucial for the growth and differentiation of skeletal muscle [7]. The GHR gene combines with GH to initiate the intracellular signal transduction mechanism, increase the expression of insulin-like growth factor 1 (IGF-1), promote cell proliferation, and affect the growth and development of animal skeletal muscle [8]. In mice, GHR knockouts exhibit impaired skeletal muscle development characterized by a reduced number and area of muscle fibers and concomitant functional defects. GHR gene variants are associated with growth and development in livestock. The GHR gene is considered to be a molecular genetic marker of growth performance in fertile pigs [9]. In sheep, the GHR gene variation is associated with growth traits [10]. According to recent studies, the presence of genetic variations in the GHR gene is associated with economic traits, such as cattle growth and production [11,12]. These studies suggest that genetic variation in the GHR gene plays an important role in regulating animal growth and development.
Structural variation (SV) is an important and abundant source of genetic and phenotypic variation, including insertions/deletions (Indel), duplications, translocations, and copy-number variations (CNVs) [13]. Compared with a single nucleotide variant (SNV), SVs could have a greater impact on the function and expression of genes [14]. Up to now, many SVs are found to be associated with economic traits in domestic animals. For example, a deletion of 110 kb in the MER1 repeat containing the imprinted transcript 1 (MIMT1) gene is associated with abortion and stillbirth in cattle [15]. The lysine acetyltransferase 6A (KAT6A) gene is associated with the development of different body size traits in sheep [16]. In addition, a 225 bp deletion in the glutaminyl–peptide cyclotransferase-like (QPCTL) gene was associated with body weight and carcass traits in chickens [17]. Upadhyay et al. [18] detected an SV in the regulatory region of polypeptide N-acetylgalactosaminyltransferase-like 6 (GALNTL6) associated with feed efficiency and growth traits in cattle. However, there is no detailed study on SVs of the yak GHR gene.
In this study, we found a 246 bp sequence deletion SV of the yak GHR gene. In order to identify the role of the GHR gene SV in yak populations in China, we analyzed the association between GHR 246 bp deletion variation and phenotypic traits to illustrate the genetic effect of GHR 246 bp deletion variation in yak breeds. This study provides useful information for the genetic protection and improvement of Chinese yak.
2. Materials and Methods
2.1. Ethics Statement
This work has been approved by the Lanzhou Institute of Husbandry and Pharmaceutical Sciences; the grant number is No. LIHPS-CAAS-2017-115. In addition, we collected all blood samples and analyzed the data strictly following the guidelines for the Care and Use of Laboratory Animals.
2.2. Sample Collection
Blood samples were collected from a total of 585 individuals of 14 yak breeds. The numbers of samples from each breed were as follows: Datong yak (DT, n = 22); Xueduo yak (XD, n = 21); Huanhu yak (HH, n = 21); Gannan yak (GN, n = 21); Tianzhu White yak (TZB, n =21); Niangya yak (NY, n = 18); Leiwuqi yak (LWQ, n = 21); Sibu yak (SB, n = 18); Muli yak (ML, n = 23); Jiulong yak (JL, n = 21); Maiwa yak (MW, n = 21); Zhongdian yak (ZD, n = 21); Bazhou yak (BZ, n = 21); and Ashidan yak (ASD, n = 315). We tracked the phenotypic data of these Ashidan yaks (i.e., body weight (BW, kg), withers height (WH, cm), body length (BL, cm), and chest girth (CG, cm)) at four age groups of growth (i.e., 6 months old, 12 months old, 18 months old, and 30 months old). The phenotype was measured using the standard method of measurement [19]. Tissues, including the heart, muscle, liver, spleen, kidney, lung, and brain, were collected for expression profiling.
2.3. Genomic DNA (gDNA) Extraction and Polymerase Chain Reaction (PCR)
Yak blood gDNA was extracted using the Tiangen whole-genome blood extraction kit (Beijing, China), and the DNA quality was detected using an ultra-microspectrophotometer (Thermo Fisher Scientific, USA). The gDNA was stored at −20 °C for further analysis. The primers for gene structure variation were designed by the National Center for Biotechnology Information (NCBI) online primer design software Primer-BLAST and synthesized by Qingke Zexi (Xi’an) Biotechnology company (Xi’an, Shaanxi, China) (Table 1). PCR was then performed. A total volume of 10 μL PCR mixture contained 1 μL of gDNA (50 ng/μL), 1 μL of each primer (10 μmol/L), 5 μL of Go Taq®Green Master Mix (Promega, Madison, WI, USA), and 2 μL of ddH2O. The PCR conditions were as follows: 95 °C for 3 min, 95 °C for 5 s, 58 °C for 30 s, 72 °C for 1 min for 30 cycles, and 72 °C for 5 min. The products were identified via 2% agarose gel electrophoresis. Six PCR products were selected and sequenced by Qingke Zexi (Xi’an) Biotechnology Company (Xi’an, Shaanxi, China).

Table 1.
The information of primer sequence.
2.4. RNA Extraction and Quantitative PCR Identification
The total RNA of yak tissue was extracted using the Trizol method, and the integrity of RNA was identified with 1% agarose gel electrophoresis. cDNA was synthesized via reverse transcription using a translator first-strand cDNA synthesis kit (Roche, Shanghai, China) and stored at −20 °C for further analysis. Quantitative real-time polymerase chain reaction (RT-qPCR) was performed using Go Taq®qPCR and RT-qPCR Systems (Promega, Madison, WI, USA) to investigate the expression levels of the GHR messenger RNA (mRNA) in each tissue. Three replicates were selected for each sample. The primer information is provided in Supplementary Table S1. The RT-qPCR conditions were as follows: 95 °C for 3 min, 95 °C for 5 s, 64 °C for 20 s for a total of 40 cycles, and 72 °C for 30 s. The results were analyzed using the 2−ΔΔCT method [20].
2.5. Cell Culture and Cell Transfection
The 293T cell line was purchased from the Cell Bank of the Chinese Academy of Sciences (Shanghai, China). High-sugar DMEM (Hyclone, Logan, UT, USA), fetal bovine serum (Gibco, Waltham, MA, USA), and 1% streptomycin/penicillin (Invitrogen, Waltham, MA, USA) were used. Plasmids were transfected into cells using ViaFectTMTransfection Reagent (Promega, Madison, WI, USA) according to the manufacturer’s instructions.
2.6. Plasmid Construction and Prediction of Transcription-Factor Binding Sites (TFBSs)
Based on the yak GHR 246 bp deletion locus, the luciferase reporter vector pGL4.10 was selected, and the pGL4.10-II (324 bp, containing the 246 bp deletion) and pGL4.10-DD (78 bp, lacking the 246 bp deletion) vectors were constructed to detect the transcriptional activity of GHR 246 bp deletion.
In this study, the online software AnimalTFDB 3.0 (http://bioinfo.life.hust.edu.cn/AnimalTFDB/#!/) (accessed on 10 February 2022) and JASPAR (http://jaspar.genereg.net/) (accessed on 10 February 2022) were used for the prediction of TFBSs at the GHR 246 bp deletion locus. The runt-related transcription factor 1 (Runx1) binding sequence (TAAATGCAAA) was identified on the II genotype, and the pGL4.10-KO-Runx1 dual-luciferase reporter vector (324 bp) was constructed. The vector pGL4.75 (Promega, Madison, WI, USA) was used as an internal reference vector for determining the luciferase reporter system.
2.7. Statistical Analysis
Gene frequency and genotype frequency were calculated using the following equations:
Genotype frequency (FAiAj) = AiAj number of individuals/total number of samples;
Allele frequency (FAi) = FAiAj + 1/2 FAiAj.
The online website SHEsis was used to calculate Hardy–Weinberg equilibrium, heterozygosity, polymorphism information, and genetic variation content of GHR gene SV in the population. Meanwhile, we analyzed the association between the SV type of the GHR gene and the growth traits of Ashidan yak by using a one-way analysis of variance (ANOVA) and frequency-distribution histogram test. A mixed linear model used in the analysis is as follows: Yij = μ + Gij + eij, where Yij is the observed growth trait; μ is the population average of each trait; Gij is the fixed effect of SV genotype of the GHR gene; and eij is the random residual. The difference between the mean values was evaluated by using one-way ANOVA with a post-event least significant digit (LSD) multiple-comparison test. The software used for the statistical analyses was the IBM SPSS Statistics software (Version 23.0). Data are expressed as mean± standard error (SE).
3. Results
3.1. Identification of SV of Yak GHR Gene
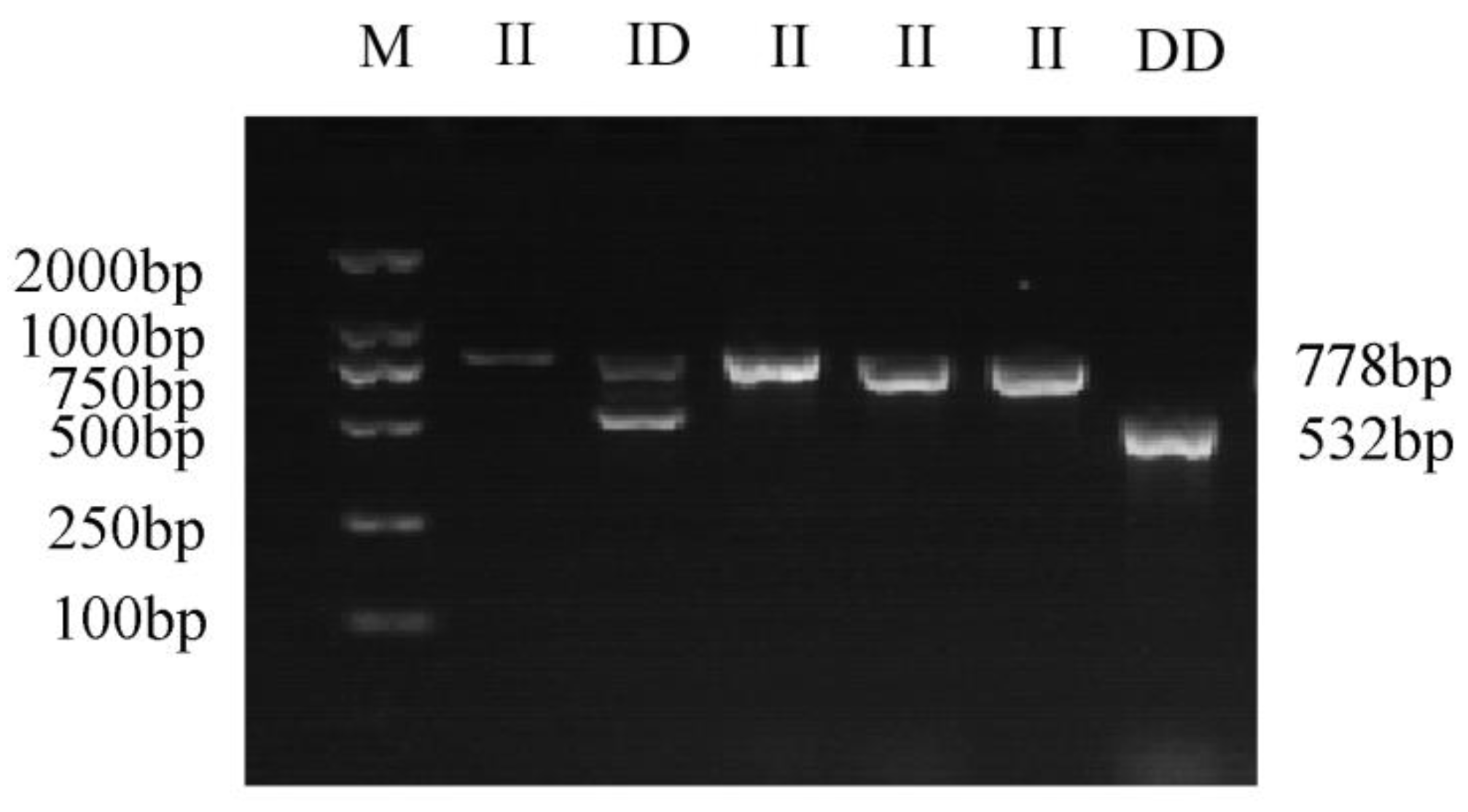
A 246 bp deletion variation was identified in intron 6 of the GHR gene (NW_005393449.1: 3052434-3052680). Based on the designed primers (F1, R1), GHR-SV246 was amplified with PCR using yak gDNA as the template. The GHR-SV246 polymorphism was analyzed via PCR amplification and agarose gel electrophoresis of the product (Figure 1). The product length matched the expectation, and three genotypes were identified, namely II (778 bp), ID (778 bp and 532 bp), and DD (532 bp). The sequencing analysis of the amplified products showed that the sequencing results were consistent with the reference genome sequence.
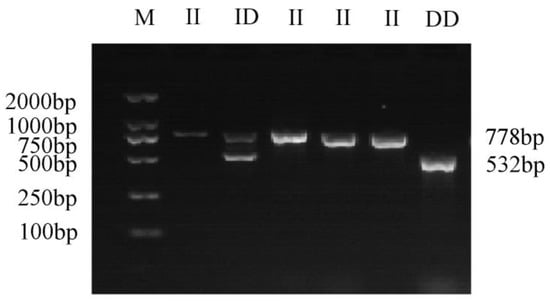
Figure 1.
The gel electrophoresis of different genotypes of GHR-SV246; M, GL DNA marker 2000; II, homozygous insertion type; ID, heterozygous type; DD, homozygous deletion type.
3.2. Identification of SV of Yak GHR Gene
The genotype frequencies and allele frequencies of GHR-SV246 in different yak breeds are shown in Table 2. The frequencies of different genotypes in Chinese yak populations were 0.723, 0.202, and 0.075, respectively. The II genotype was the dominant genotype in different populations except for SB yak; the allele frequencies were 0.824 and 0.176, respectively. Among them, the frequency of the I allele was the highest in the ML yak population (0.979) and the lowest in the SB yak population (0.500) (Table 2). The Ho ranged from 0.500 to 0.959, indicating high heterozygosity within different yak breeds, and the Ne ranged from 1.04 to 2.00. Classification based on the polymorphism information content (PIC value < 0.25, low polymorphism; 0.25 < PIC value < 0.5, medium polymorphism; PIC value > 0.5, high polymorphism) showed that the 246 bp deletion of the GHR gene exhibited relatively low polymorphism in ZD, ML, LWQ, NN, GN, and XD yak populations, while moderate polymorphism was observed in other yak populations (Table 3).

Table 2.
Genotype and allele frequencies of GHR-SV246 in different Yaks.

Table 3.
The genetic parameters of Yak GHR-SV246.
3.3. The Association of the 246 bp Deletion of GHR with Growth Traits of Yak
In this study, the normal distribution of yak growth traits (Supplementary Figure S1–S4) and the average value of the growth parameters were statistically analyzed (Supplementary Table S1). We also analyzed the association of GHR gene polymorphisms with the growth traits in yaks using a linear model. As shown in Table 4, the polymorphisms in the GHR gene were significantly associated with body length in 6-month-old ASD yaks (p < 0.05), while there was no significant relationship with other growth traits (p > 0.05).

Table 4.
The association analysis between different body size traits and different GHR genotypes of Yaks.
3.4. Identification of Fluorescence Activity of GHR Gene Polymorphism in Yak
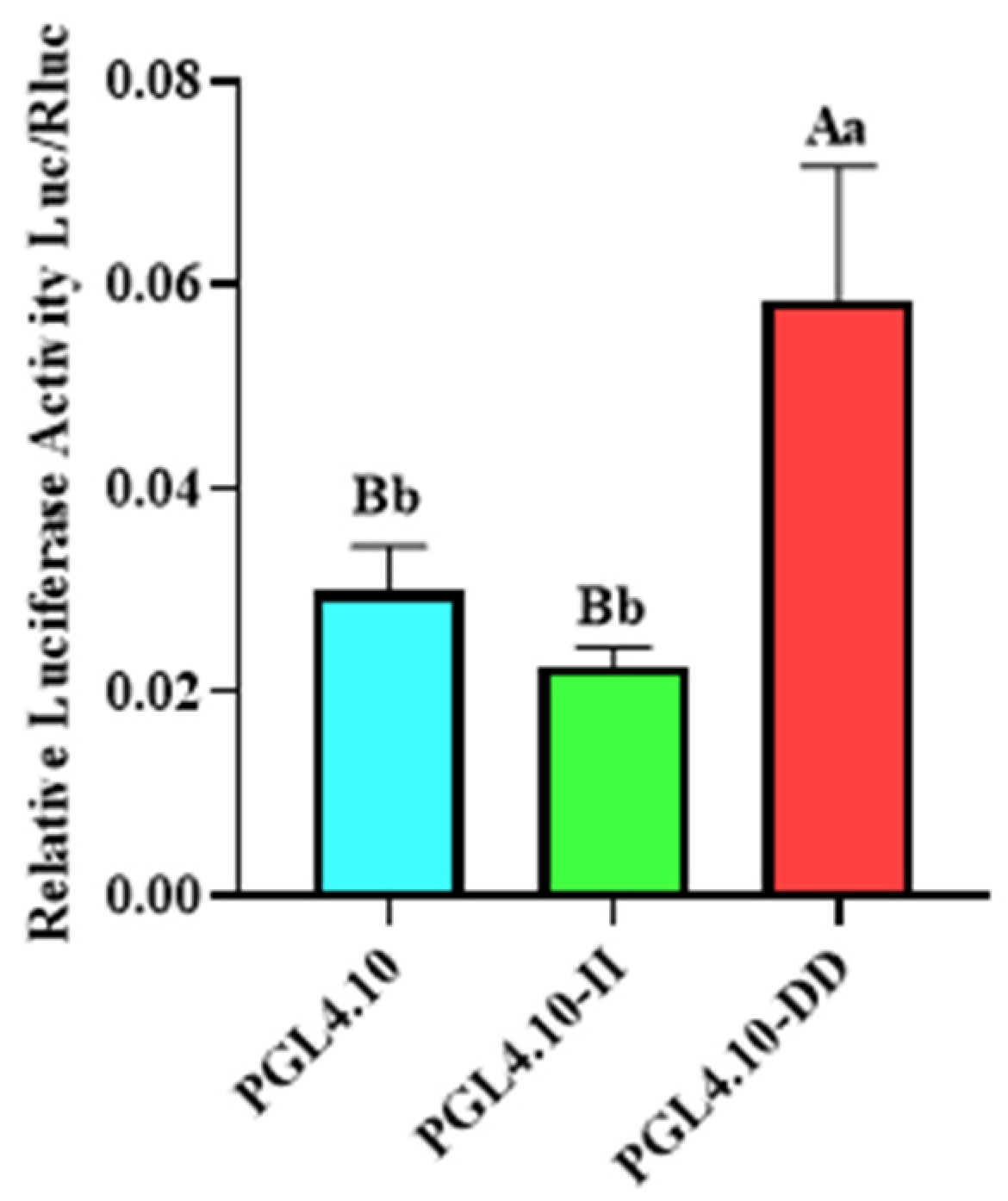
To detect the transcriptional activity of different GHR-SV246 genotypes in cells, pGL4.10, pGL4.10-II, and pGL4.10-DD vectors were co-transfected into 293T cells with the internal reference plasmid pGL4.75. The fluorescence activity of the pGL4.10-II vector was significantly different from that of the blank control group pGL4.10 (p < 0.05) and was significantly lower than that of the pGL4.10-DD vector (p < 0.05) (Figure 2).
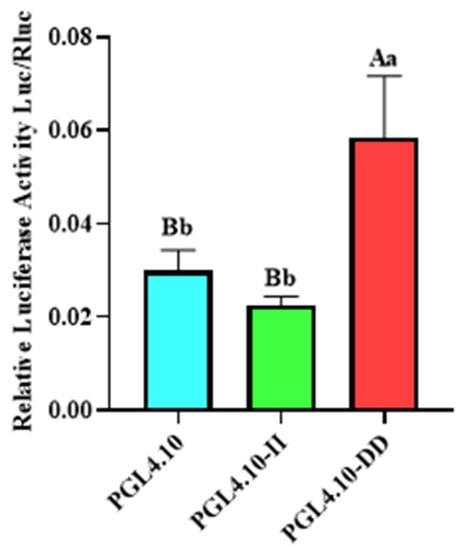
Figure 2.
The dual-luciferase activity of GHR-SV246 mutant recombinant plasmid in 293T cells; the same letter means no significant difference (p > 0.05); different letters indicate significant differences (p < 0.05).
3.5. Effect of Transcription Factors on the Transcriptional Activity of Mutant Recombinant Plasmids
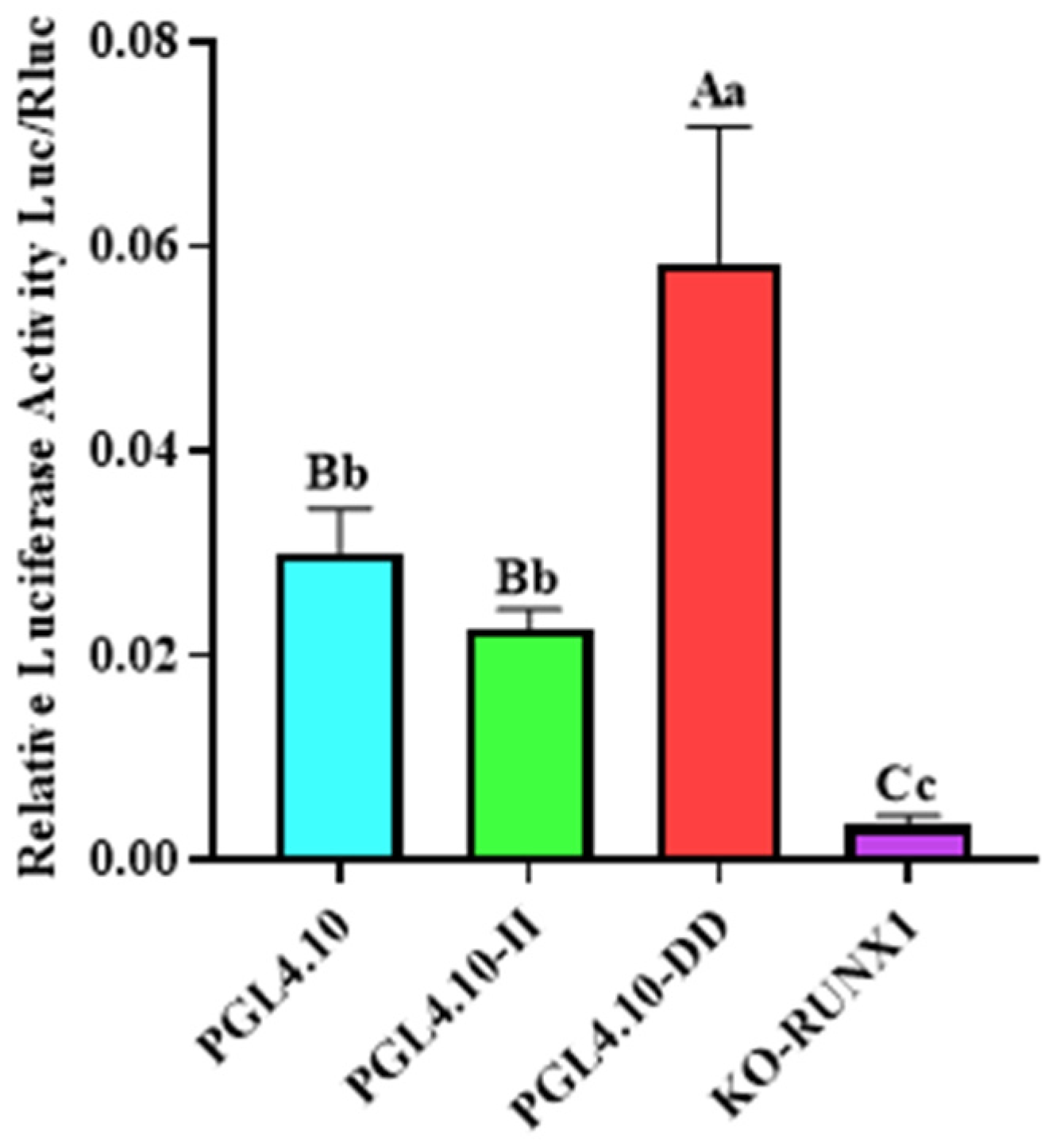
In order to determine the effect of Runx1 transcription-factor binding with the 246 bp deletion on transcriptional activity in cells, the pGL4.10-KO-Runx1 luciferase reporter vector with mutant Runx1 binding site was constructed, and its fluorescence activity was identified. The luciferase activity of vector pGL4.10-KO-Runx1 was significantly lower than that of vector pGL4.10-II (Figure 3; p < 0.05), indicating that the transcription factor Runx1 could be combined with the II genotype sequence, thus increasing its transcriptional activity in cells.
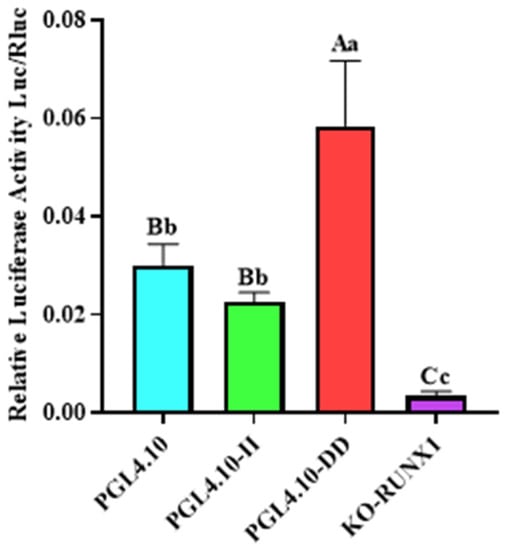
Figure 3.
The dual-luciferase activity of GHR-SV246 recombinant plasmid in 293T cells.
3.6. mRNA Expression Profile of the GHR Gene in Yak
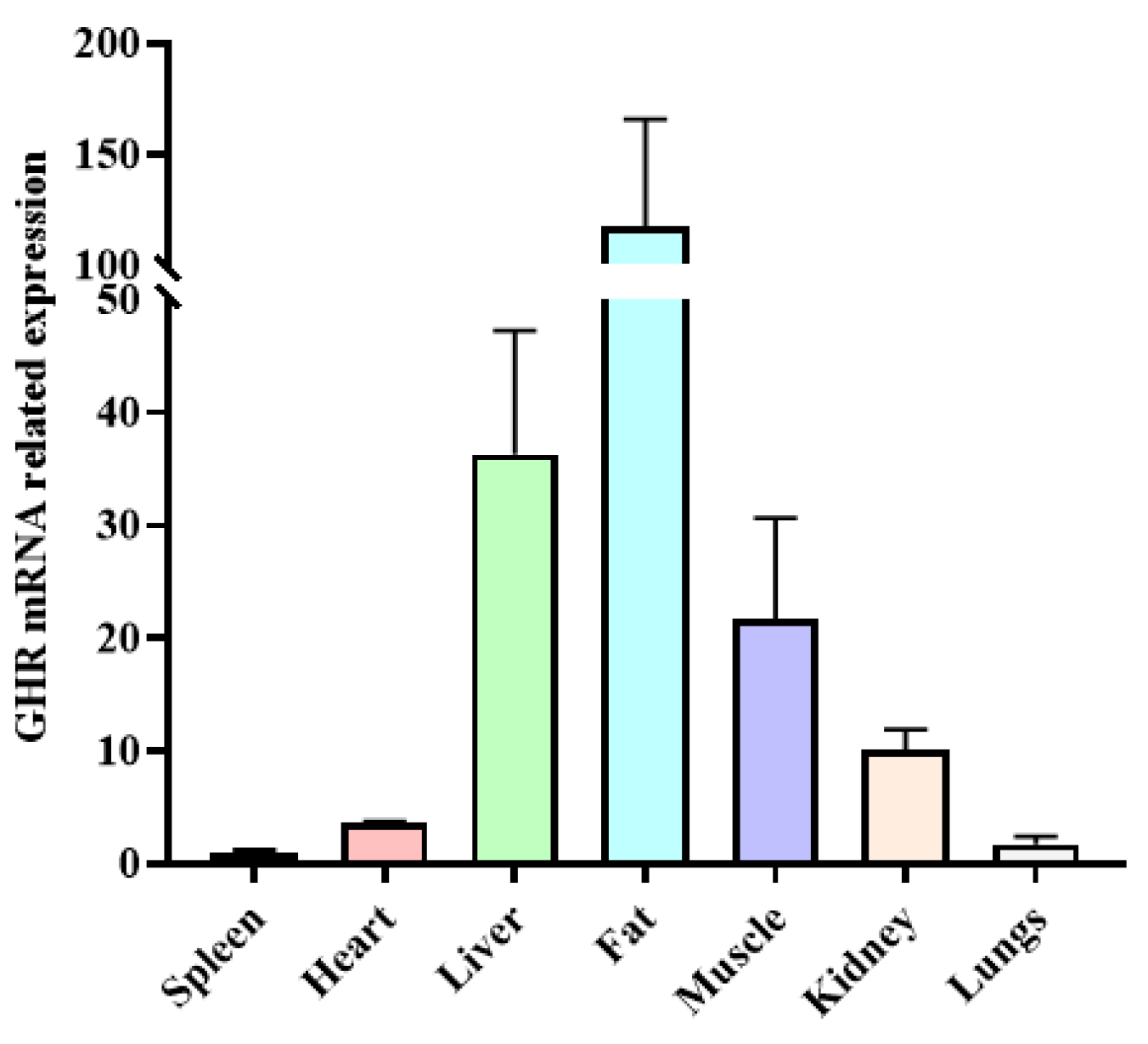
The expression levels of the GHR gene in different tissues were identified in yaks. There were significant differences in the expression of the GHR gene in various tissues of yaks, with the highest expression in fat, followed by the liver, muscle, and kidney tissues, and the lowest expression in the heart, spleen, and lung tissues (Figure 4).
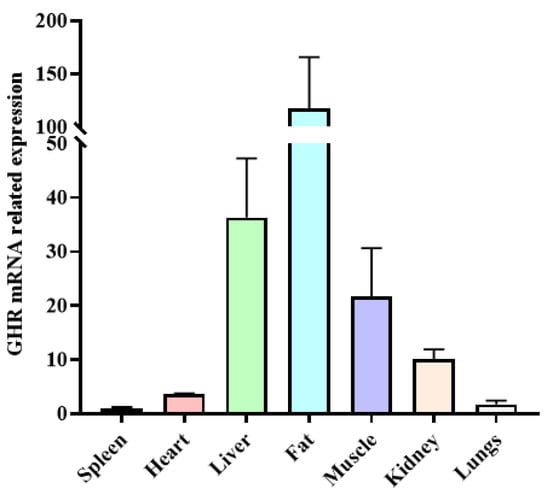
Figure 4.
The expression level of GHR gene mRNA in different tissues of yak.
4. Discussion
SV is a major determinant of the phenotypic diversity of animals. Studies on chickens [21], sheep [22], and cattle [10] have reported that SVs play a significant role in their genetic diversity. In poultry, the SVs of prolactin receptor (PRLR) [23] and Lpin1 [24] were significantly associated with chicken growth, carcass characteristics, and other economic traits. In cattle, the SVs of SIRT4 and NPM1 were significantly associated with bovine growth traits and meat quality [25,26]. These studies show that SV is an important class of molecular genetic markers, which is of great value in revealing the genetic mechanism of the economic traits of livestock and poultry.
The GHR gene is a member of the cytokine receptor superfamily [27]. Some studies have pointed out that the GHR gene can regulate the growth of skeletal muscle by interacting with the growth hormone (GH) [22,28]. In this study, we analyzed the expression of the GHR gene in different tissues of yak and found that the GHR gene was expressed in different tissues, with high expression in adipose tissue and moderate expression in the muscle. This is consistent with previous studies on GHR expression profiles [29]. GHR mRNA expression levels in muscle and fat have a positive effect on post-weaning weight gain in cattle, while GHR mRNA expression in the liver has a negative effect on post-weaning weight gain in cattle [30]. A study reported that there is a natural antisense transcript (GHR-AS) on the GHR gene of chickens, and its overexpression can promote the expression of myogenic differentiation 1 (MyoD) and myosin heavy chain (MyHC) and the differentiation of myoblasts, thus enhancing the differentiation of myoblasts [31].
A previous study found associations between GHR gene polymorphisms and the growth traits in cattle at different ages [16], showing that GHR gene variation can affect the growth and production traits of cattle. However, few studies have reported an association between GHR gene variation and yak growth and production traits. In this study, we found a 246 bp structural variation site in intron 6 of the yak GHR gene. The genotyping results showed that the three genotypes of GHR genes existed in different yak breeds, and the analysis of genetic parameters suggested that the 246 bp deletion was in Hardy–Weinberg equilibrium in different yak breeds, which indicates that it is relatively stable in yak population. In ASD yak, this SV belongs to a moderate polymorphism, indicating that the degree of genetic variation is relatively large, and it has certain breeding potential. We found that individuals with the DD genotype showed higher body length at 6 months of age in ASD yak breeds. These data suggest that SVs in the GHR gene might influence the early development of yak.
Introns are noncoding segments of genes that are removed via splicing during gene transcription and ultimately do not exist in mature RNA molecules [32]. Introns play an important role in the regulation of gene expression networks. Ostrovsky et al. [33] found that the intron 9 of the heparanase (HPSE) gene has a regulatory effect on its expression, and the existence of mutation sites in this intron has a significant impact on the regulatory effect of the intron. In this study, the transcriptional activity of the deletion sequence in the yak GHR gene was analyzed, and the luciferase activity of the pGL4.10-DD vector was significantly higher than that of the pGL4.10-II vector (p < 0.05). This finding indicates that there may be repressive TFBSs in the II genotype sequence, which interferes with the expression of the GHR gene by binding to transcription factors and affects the growth and development of yaks. Genomic variation in introns can affect gene expression through cis-element targets or by binding to other functional genes [27]. Boriushkin et al. [34] found a Krüppel-like factor 4 (KLF4) transcription-factor binding site located in the intron of the delta-like canonical Notch ligand 4 (DLL4) gene, which is a key regulator of angiogenesis. They also found that KLF4 can inhibit the expression of DLL4 via this binding site. The appearance of gene variant sites may introduce novel transcription-factor binding sites or destroy existing binding sites [35,36]. In this study, in the II genotype sequence, we detected a Runx1 transcription-factor binding site, which was first discovered in acute myeloid leukemia [37]. Runx1 has an important regulatory role in the growth and development of animal muscles. In mouse muscle cells, the overexpression of Runx1 inhibits myogenic differentiation but promotes myoblast proliferation [38]. When skeletal muscle is injured, the expression of Runx1 is upregulated in skeletal muscle, thereby inhibiting the premature differentiation of primary myoblasts and promoting muscle regeneration [39]. Runx1 inhibits Spi1 gene expression by interacting with Spil introns [29]. In this study, we hypothesized that Runx1 has an effect on the transcriptional regulation of the GHR gene II genotype in yaks. This study found that changes in Runx1-TFBS reduced the fluorescence activity of the II genotype vector, indicating that the Runx1 could bind to the II genotype sequence and increase the transcriptional activity of the GHR gene. The luciferase activity of the pGL4.10-II vector was not significantly different from the pGL4.10 empty-vector fluorescence activity (p > 0.05). There may also be a binding site of an inhibitory transcription factor in the II genotype sequence.
5. Conclusions
We detected and validated the SV of the GHR gene in yaks for the first time. The results indicated that SV of the GHR gene was significantly correlated with body length at 6 months of age in ASD yaks. Moreover, the transcriptional activity assay found significant differences among different genotypes, and transcription factor Runx1 promoted transcriptional activity in the II genotype. Our study provides primary evidence for the role of the GHR gene, which may be a molecular marker for early yak breeding in the future.
Supplementary Materials
The following supporting information can be downloaded at: https://www.mdpi.com/article/10.3390/ani13050851/s1, Figure S1: Histogram of normal distribution of body weight frequency of yaks at different ages, Figure S2: Histogram of normal distribution of yak body length frequency at different months, Figure S3: Histogram of normal distribution of yak body height frequency at different months, Figure S4: Histogram of normal distribution of chest girth frequency of yaks at different ages, Table S1: The overall average of the growth parameters.
Author Contributions
Conceptualization, F.W., X.W. and P.Y.; methodology, F.W. and X.W.; software, X.M., Q.Z. and Q.B.; validation, F.W., C.L. and P.Y.; formal analysis, F.W., X.W. and Q.B.; investigation, F.W. and Q.Z.; resources, C.L., M.C. and X.G.; data curation, F.W., X.W. and C.L.; writing—original draft preparation, F.W.; writing—review and editing, C.L., X.G. and X.W.; visualization, P.Y.; supervision, P.Y.; project administration, P.Y.; funding acquisition, P.Y. All authors have read and agreed to the published version of the manuscript.
Funding
This study was supported by the National Key R&D Program of China (2021YFD1600200), Central Public-interest Scientific Institution Basal Research Fund (Y2022LM10), Agricultural Science and Technology Innovation Program (25-LZIHPS-01), China Agriculture Research System of MOF and MARA (CARS-37) and Foundation for Innovation Groups of Basic Research in Gansu Province (20JR5RA580).
Institutional Review Board Statement
This work has been approved by the Lanzhou Institute of Husbandry and Pharmaceutical Sciences, the grant number is No. LIHPS-CAAS-2017-115.
Data Availability Statement
This study did not report any data.
Conflicts of Interest
The authors declare no conflict of interest.
References
- Li, A.; Wang, Y.; Pei, L.; Mehmood, K.; Li, K.; Qamar, H.; Iqbal, M.; Waqas, M.; Liu, J.; Li, J. Influence of dietary supplementation with Bacillus velezensis on intestinal microbial diversity of mice. Microb. Pathog. 2019, 136, 103671. [Google Scholar] [CrossRef] [PubMed]
- Ruan, C.-M.; Wang, J.; Yang, Y.-X.; Hu, J.-J.; Ma, Y.-J.; Zhang, Y.; Zhao, X.-X. Proteomic analysis of Tianzhu White Yak (Bos grunniens) testis at different sexual developmental stages. Anim. Sci. J. 2019, 90, 333–343. [Google Scholar] [CrossRef] [PubMed]
- Ding, L.; Wang, Y.; Kreuzer, M.; Guo, X.; Mi, J.; Gou, Y.; Shang, Z.; Zhang, Y.; Zhou, J.; Wang, H.; et al. Seasonal variations in the fatty acid profile of milk from yaks grazing on the Qinghai-Tibetan plateau. J. Dairy Res. 2013, 80, 410–417. [Google Scholar] [CrossRef]
- Ma, Z.; Xu, J.; Zhong, J.; Dou, Q.; Sun, Y.; Mu, Y. Structural features of the 5’ flanking region of the Yak (Bos grunniens) growth hormone receptor (GHR) gene (Brief Report). Arch. Anim. Breed. 2010, 53, 372–376. [Google Scholar] [CrossRef]
- Guo, Z.; Ge, X.; Yang, L.; Ma, G.; Ma, J.; Yu, Q.-L.; Han, L. Ultrasound-assisted thawing of frozen white yak meat: Effects on thawing rate, meat quality, nutrients, and microstructure. Ultrason. Sonochem. 2021, 70, 105345. [Google Scholar] [CrossRef]
- Etherton, T.D.; Bauman, D.E. Biology of Somatotropin in Growth and Lactation of Domestic Animals. Physiol. Rev. 1998, 78, 745–761. [Google Scholar] [CrossRef]
- Li, W.-Y.; Liu, Y.; Gao, C.-F.; Lan, X.-Y.; Wu, X.-F. A novel duplicated insertion/deletion (InDel) of the CPT1a gene and its effects on growth traits in goat. Anim. Biotechnol. 2021, 32, 343–351. [Google Scholar] [CrossRef]
- Yang, S.; Liu, Z.; Yan, Z.; Zhao, Z.; Zhang, C.; Gong, Q.; Du, X.; Wu, J.; Feng, Y.; Du, J.; et al. Improvement of skeletal muscle growth by GH/IGF growth-axis contributes to growth performance in commercial fleshy sturgeon. Aquaculture 2021, 543, 736929. [Google Scholar] [CrossRef]
- Slifierz, M.J.; Friendship, R.; de Lange, C.F.; Rudar, M.; Farzan, A. An epidemiological investigation into the association between biomarkers and growth performance in nursery pigs. BMC Veter. Res. 2013, 9, 247. [Google Scholar] [CrossRef]
- Yang, Q.; Yan, H.; Li, J.; Xu, H.; Wang, K.; Zhu, H.; Chen, H.; Qu, L.; Lan, X. A novel 14-bp duplicated deletion within goat GHR gene is significantly associated with growth traits and litter size. Anim. Genet. 2017, 48, 499–500. [Google Scholar] [CrossRef]
- Blott, S.; Kim, J.-J.; Moisio, S.; Schmidt-Küntzel, A.; Cornet, A.; Berzi, P.; Cambisano, N.; Ford, C.; Grisart, B.; Johnson, D.; et al. Molecular dissection of a quantitative trait locus: A phenylalanine-to-tyrosine substitution in the transmembrane domain of the bovine growth hormone receptor is associated with a major effect on milk yield and composition. Genetics 2003, 163, 253–266. [Google Scholar] [CrossRef]
- Waters, S.M.; McCabe, M.S.; Howard, D.J.; Giblin, L.; Magee, D.A.; MacHugh, D.E.; Berry, D.P. Associations between newly discovered polymorphisms in the Bos taurus growth hormone receptor gene and performance traits in Holstein-Friesian dairy cattle. Anim. Genet. 2011, 42, 39–49. [Google Scholar] [CrossRef]
- Shen, R.; Wang, L.; Liu, X.; Wu, J.; Jin, W.; Zhao, X.; Xie, X.; Zhu, Q.; Tang, H.; Li, Q.; et al. Genomic structural variation-mediated allelic suppression causes hybrid male sterility in rice. Nat. Commun. 2017, 8, 1310. [Google Scholar] [CrossRef]
- Gong, J.-Y.; Wen, C.-J.; Tang, M.-L.; Duan, R.-F.; Chen, J.-N.; Zhang, J.-Y.; Zheng, K.-W.; He, Y.-D.; Hao, Y.-H.; Yu, Q.; et al. G-quadruplex structural variations in human genome associated with single-nucleotide variations and their impact on gene activity. Proc. Natl. Acad. Sci. USA 2021, 118, e2013230118. [Google Scholar] [CrossRef]
- Flisikowski, K.; Venhoranta, H.; Nowacka-Woszuk, J.; McKay, S.; Flyckt, A.; Taponen, J.; Schnabel, R.; Schwarzenbacher, H.; Szczerbal, I.; Lohi, H.; et al. A Novel Mutation in the Maternally Imprinted PEG3 Domain Results in a Loss of MIMT1 Expression and Causes Abortions and Stillbirths in Cattle (Bos taurus). PLoS ONE 2010, 5, e15116. [Google Scholar] [CrossRef]
- Sakar, Ç.M.; Zülkadir, U. Determination of the relationship between Anatolian black cattle growth properties and myostatin, GHR and Pit-1 gene. Anim. Biotechnol. 2022, 33, 536–545. [Google Scholar] [CrossRef] [PubMed]
- Ren, T.; Li, W.; Liu, D.; Liang, K.; Wang, X.; Li, H.; Jiang, R.; Tian, Y.; Kang, X.; Li, Z. Two insertion/deletion variants in the promoter region of the QPCTL gene are significantly associated with body weight and carcass traits in chickens. Anim. Genet. 2019, 50, 279–282. [Google Scholar] [CrossRef] [PubMed]
- Upadhyay, M.; Derks, M.F.; Andersson, G.; Medugorac, I.; Groenen, M.A.; Crooijmans, R.P. Introgression contributes to distribution of structural variations in cattle. Genomics 2021, 113, 3092–3102. [Google Scholar] [CrossRef] [PubMed]
- Gilbert, R.P.; Bailey, D.R.; Shannon, N.H. Linear body measurements of cattle before and after 20 years of selection for postweaning gain when fed two different diets2. J. Anim. Sci. 1993, 71, 1712–1720. [Google Scholar] [CrossRef]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef]
- Florini, J.R.; Ewton, D.Z.; Coolican, S.A. Growth hormone and the insulin-like growth factor system in myogenesis. Endocr. Rev. 1996, 17, 481–517. [Google Scholar] [PubMed]
- Rehfeldt, C.; Nissen, P.M.; Kuhn, G.; Vestergaard, M.; Ender, K.; Oksbjerg, N. Effects of maternal nutrition and porcine growth hormone (pGH) treatment during gestation on endocrine and metabolic factors in sows, fetuses and pigs, skeletal muscle development, and postnatal growth. Domest. Anim. Endocrinol. 2004, 27, 267–285. [Google Scholar] [CrossRef] [PubMed]
- Liang, K.; Wang, X.; Tian, X.; Geng, R.; Li, W.; Jing, Z.; Han, R.; Tian, Y.; Liu, X.; Kang, X.; et al. Molecular characterization and an 80-bp indel polymorphism within the prolactin receptor (PRLR) gene and its associations with chicken growth and carcass traits. 3 Biotech 2019, 9, 296. [Google Scholar] [CrossRef] [PubMed]
- Wang, R.; Wang, T.; Lu, W.; Zhang, W.; Chen, W.; Kang, X.; Huang, Y. Three indel variants in chicken LPIN1 exon 6/flanking region are associated with performance and carcass traits. Br. Poult. Sci. 2015, 56, 621–630. [Google Scholar] [CrossRef] [PubMed]
- Huang, Y.Z.; He, H.; Wang, J.; Li, Z.J.; Lan, X.Y.; Lei, C.Z.; Zhang, E.P.; Zhang, C.L.; Shen, Q.W.; Chen, H. Sequence variants in the bovine nucleophosmin 1 gene, their linkage and their associations with body weight in native cattle breeds in China. Anim. Genet. 2011, 42, 556–559. [Google Scholar] [CrossRef]
- Zhang, Q.; Jin, Y.; Jiang, F.; Cheng, H.; Wang, Y.; Lan, X.; Song, E. Relationship between an indel mutation within the SIRT4 gene and growth traits in Chinese cattle. Anim. Biotechnol. 2019, 30, 352–357. [Google Scholar] [CrossRef] [PubMed]
- Kim, B.-Y.; Park, J.H.; Jo, H.-Y.; Koo, S.K.; Park, M.-H. Optimized detection of insertions/deletions (INDELs) in whole-exome sequencing data. PLoS ONE 2017, 12, e0182272. [Google Scholar] [CrossRef] [PubMed]
- Stewart, C.E.; Rotwein, P.; Vélez, E.J.; Azizi, S.; Millán-Cubillo, A.; Fernández-Borràs, J.; Blasco, J.; Chan, S.J.; Calduch-Giner, J.A.; Pérez-Sánchez, J.; et al. Growth, differentiation, and survival: Multiple physiological functions for insulin-like growth factors. Physiol. Rev. 1996, 76, 1005–1026. [Google Scholar] [CrossRef] [PubMed]
- Hosokawa, H.; Koizumi, M.; Masuhara, K.; Romero-Wolf, M.; Tanaka, T.; Nakayama, T.; Rothenberg, E.V. Stage-specific action of Runx1 and GATA3 controls silencing of PU.1 expression in mouse pro–T cells. J. Exp. Med. 2021, 218, e20202648. [Google Scholar] [CrossRef]
- Zheng, W.; Leng, X.; Vinsky, M.; Li, C.; Jiang, H. Association of body weight gain with muscle, fat, and liver expression levels of growth hormone receptor, insulin-like growth factor I, and beta-adrenergic receptor mRNAs in steers. Domest. Anim. Endocrinol. 2018, 64, 31–37. [Google Scholar] [CrossRef]
- Xu, H.; Li, T.; Wang, Z.; Adu-Asiamah, P.; Leng, Q.; Zheng, J.; Zhao, Z.; An, L.; Zhang, X.; Zhang, L. Roles of chicken growth hormone receptor antisense transcript in chicken muscle development and myoblast differentiation. Poult. Sci. 2019, 98, 6980–6988. [Google Scholar] [CrossRef] [PubMed]
- Pena, V.; Rozov, A.; Fabrizio, P.; Lührmann, R.; Wahl, M.C. Structure and function of an RNase H domain at the heart of the spliceosome. EMBO J. 2008, 27, 2929–2940. [Google Scholar] [CrossRef] [PubMed]
- Ostrovsky, O.; Baryakh, P.; Morgulis, Y.; Mayorov, M.; Bloom, N.; Beider, K.; Shimoni, A.; Vlodavsky, I.; Nagler, A. The HPSE Gene Insulator-A Novel Regulatory Element That Affects Heparanase Expression, Stem Cell Mobilization, and the Risk of Acute Graft versus Host Disease. Cells 2021, 10, 2523. [Google Scholar] [CrossRef]
- Boriushkin, E.; Zhang, H.; Becker, M.; Peachey, J.; Shatat, M.A.; Adams, R.H.; Hamik, A. Kruppel-like factor 4 regulates developmental angiogenesis through disruption of the RBP-J–NICD–MAML complex in intron 3 of Dll4. Angiogenesis 2019, 22, 295–309. [Google Scholar] [CrossRef]
- Su, K.; Chen, S.; Ye, J.; Kuang, L.; Zhang, T.; Wang, H.; Yang, X. A functional indel polymorphism rs34396413 in TFAP2A intron-5 significantly increases female encephalocele risk in Han Chinese population. Child’s Nerv. Syst. 2019, 35, 965–972. [Google Scholar] [CrossRef]
- Lin, W.; Ren, T.; Li, W.; Liu, M.; He, D.; Liang, S.; Luo, W.; Zhang, X. Novel 61-bp Indel of RIN2 Is Associated With Fat and Hatching Weight Traits in Chickens. Front. Genet. 2021, 12, 672888. [Google Scholar] [CrossRef]
- Morita, K.; Maeda, S.; Suzuki, K.; Kiyose, H.; Taniguchi, J.; Liu, P.P.; Sugiyama, H.; Adachi, S.; Kamikubo, Y. Paradoxical enhancement of leukemogenesis in acute myeloid leukemia with moderately attenuated RUNX1 expressions. Blood Adv. 2017, 1, 1440–1451. [Google Scholar] [CrossRef]
- Bao, M.; Liu, S.; Yu, X.-Y.; Wu, C.; Chen, Q.; Ding, H.; Shen, C.; Wang, B.; Wang, S.; Song, Y.-H.; et al. Runx1 promotes satellite cell proliferation during ischemia—Induced muscle regeneration. Biochem. Biophys. Res. Commun. 2018, 503, 2993–2997. [Google Scholar] [CrossRef]
- Umansky, K.B.; Gruenbaum-Cohen, Y.; Tsoory, M.; Feldmesser, E.; Goldenberg, D.; Brenner, O.; Groner, Y. Runx1 Transcription Factor Is Required for Myoblasts Proliferation during Muscle Regeneration. PLoS Genet. 2015, 11, e1005457. [Google Scholar] [CrossRef] [PubMed]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).