1. Introduction
Contemporary digital workflows have overtaken histopathology’s traditional diagnostic and analysis protocols [
1]. It is standard for pathologists to use digitalized images within virtual microscope software. Such software visualizes images obtained from a scanning microscope. Typically, an image consists of an entire glass slide of tissue stained to highlight relevant biological features. A microscope illuminates a given slide and allows the digitization of the sample. Any given staining approach has affiliated interactions with specific biological components in the slide [
2]. If a histological slide is not stained, it remains transparent and will be masked by the illuminance source of the microscope penetrating through the tissue. In a standard, red, green, and blue (RGB) channel format, the stain will absorb a distinct amount of light in each channel [
2]. The resulting images will thus vary due to laboratory changes in stain manufacturer, general storing conditions, microscope/slide scanner configuration, and dye-application methods [
1,
2,
3,
4].
In diagnostic applications, hematoxylin and eosin (H&E) is the default staining method for revealing salient histologic features [
5]. Hematoxylin stains histologic cell nuclei to a purple-blue hue, while eosin stains cytoplasm and extracellular matrixes to a pink-red hue. Like other staining methods, the results of an H&E stain may vary between different laboratories or even within the same laboratory [
2]. Such variance negatively impacts the time-to-insight for histopathologists and can even affect the accuracy of computer-aided diagnostic software (CAD) [
4,
6,
7,
8]. It is therefore imperative to effectively normalize the staining variance between histological slides.
Various methods have been developed for stain normalization [
2,
9,
10,
11,
12,
13,
14]. Often, methods require a target image to imitate by artificially altering an input image [
15,
16]. Essentially, the staining variance of one laboratory is used to alter out-of-laboratory stain samples. Studies such as [
17] highlight the efficacy of structure-preserving normalization [
18] with a target. In work [
19], it was noted that the Food and Drugs Administration recommended adopting target-based normalization. Overall, the variance between laboratories can be essential in organizing available targets for more effective normalization. As noted in [
19], there is no ‘gold standard’ when it comes to color-management issues.
Regarding classification effects, prior investigations show that color normalization within samples of the same laboratory did not positively affect classification accuracy [
20]. In addition, the same study showed inconsistent positive effects across all its datasets. Another study [
21] showed that color pre-processing reduced accuracy in cancer classification, especially when coupled with color related features. In that regard, another work showed similar effects for kidney tissue classification [
22]. In [
8], the authors showed that color transformations were more effective than color normalization in four different applications. In contrast positive accuracy effects of color normalization were previously shown for diagnostic tasks such as colorectal cancer classification [
6]. All these results strongly highlighted a potential task-related positive or negative impact of color normalization approaches.
We saw that stain normalization approaches have been investigated in various ways. To date, the underlying structure of inter-laboratory stain variance as a normalization target remains unclear. How can it be systematically exploited for task specific positive normalization effects? When considering this variance, we first consider the following questions. How many laboratories have distinct staining results? Are staining effects consistent between tissue types? Beginning to answer these questions can help digital pathologists and the research community fine-tune their assumptions about target and normalization choices, ultimately in an effort to improve time-to-insight and classification accuracy in CAD applications.
This study analyzed H&E staining results from skin, kidney, and liver tissues between 66 laboratories from 11 countries. The slides were distributed to the central Finland biobank via the Lab quality (Helsinki) company. After standard preprocessing, we used a principal-component-analysis-based RGB image representation; this image representation solely focused on color-related components and minimized the interference of morphological changes between the images. We used unsupervised machine learning to discover clusters of laboratory staining outcomes. The k-means algorithm was set to investigate groupings of stains up to 24 clusters. We furthered our investigation with supervised learning, designating laboratories as the ground truth. In the unsupervised approach, we used the rand index metric to compare cluster assignments between laboratories and tissue types. In the supervised approach we compared classification accuracy between tissue types. These approaches allowed us to infer a rough estimate of distinct outcomes from the set of laboratories and whether tissue type also affected the results.
4. Discussion
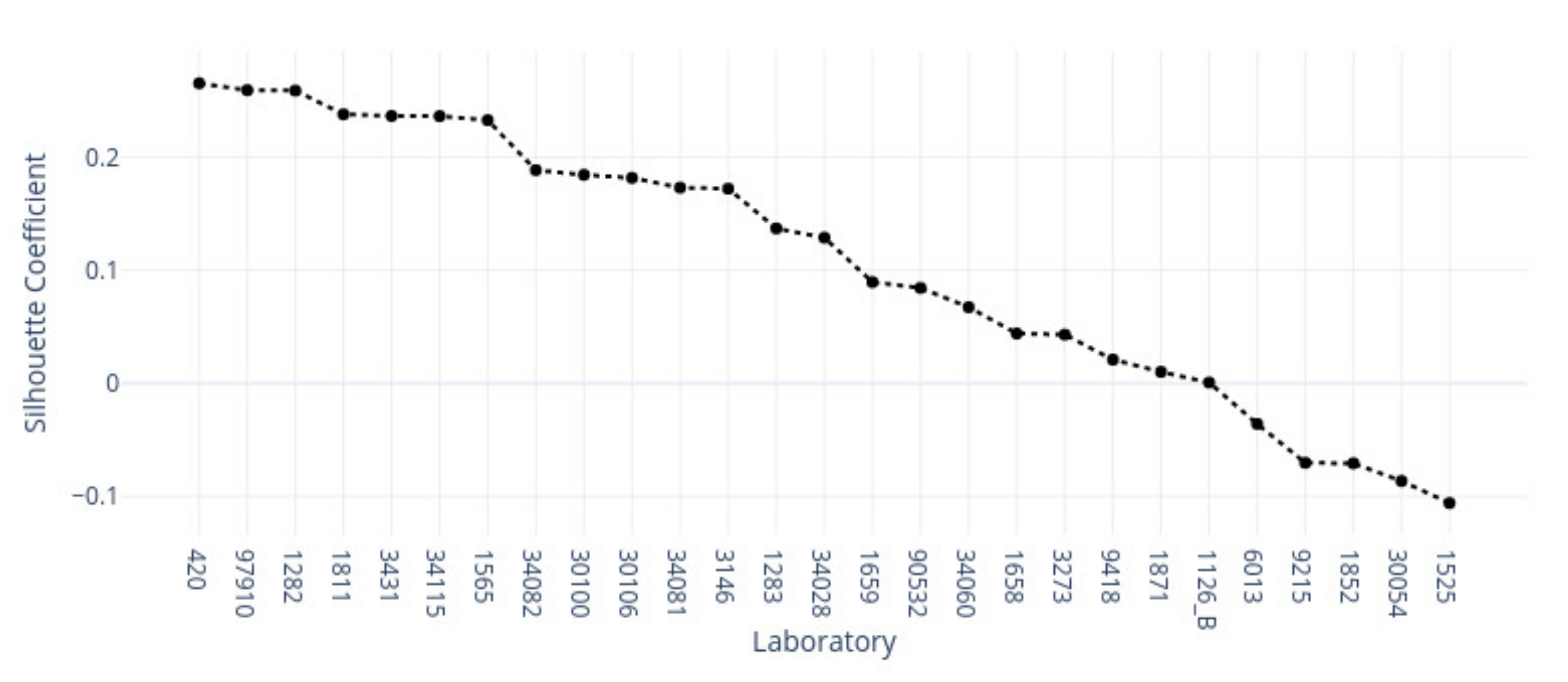
This study demonstrates the first results of the underlying structure between multiple laboratory stainings. The clustering section showcased that the stainings formed two primary clusters within different tissue types. The laboratories in these clusters maintain a similarity that steadily decreases as we approach each cluster’s edges. Although clusters did form according to the best silhouette scores, the clustering effects were not as strong as expected, with average silhouette scores below 0.3. It is important to note that we only had a single example per tissue type per laboratory. Such limitations can make it more challenging to analyze the staining variance, since more examples from each laboratory may be needed to characterize any given laboratory more accurately.
In contrast, the supervised approach showed good separability between laboratories. This result showed that laboratories have an effect, part of which can be predicted. We demonstrated that the tissue type affected the supervised results and cluster assignment. The large accuracy differences between tissue types were shown in the supervised approach. The same effect was shown in the unsupervised results where the rand index showed that laboratory assignment within clusters and between tissue types did not match, i.e., were slightly above the 50% chance level. It is important to note that we simultaneously analyzed the hematoxylin and eosin components in both machine learning approaches. Individual variation may exist in the two staining components whose impact and contribution to these results are yet undetermined. It therefore remains unclear if this effect would persist if the clustering and learning were to occur separately for hematoxylin and eosin.
This novel study was designed as a proof-of-concept approach and an initial investigation into inter-laboratory staining variation. Consequently, additional limitations include the limited resolution, a fixed zoom level, and one region of interest per input sample. If these parameters are updated in future studies, we might obtain an even more adequate margin of error and resolution for the numerical results. Additionally, we note that the features and preprocessing pipeline focused on color-related information, not morphological information. Such an approach has the benefit of highlighting non-structural features but does not consider tissue morphology. Another approach could use active learning such that clustering labels are used as ground truth to a supervised learning set-up; in this way, it might be possible to investigate which features play a more critical role in the clustering result by modeling the clustering result itself with a classification algorithm.
In conclusion, we investigated the impact and relationships of multi-laboratory H&E staining variance on different tissue types with machine learning methods. We hypothesized that individual laboratory stainings are consistent between tissue types and that different staining approaches form subgroups with similar features. Our results showed that multi-laboratory staining varies between tissue types. Multiple laboratories did not appear to form strong clusters within each tissue type but were relatively well separated in the supervised approach. From a clinical perspective, we showed that only some of the laboratories were identified. According to the result, not all laboratory samples were distinct and were confused with other laboratories. The results suggest value in considering tissue-specific normalization targets. This might require changing assumptions for the efficacy of a laboratory given incompatible tissue-type examples. Ultimately, tissue type appears to be an additional driver of the formed clusters and classification results. Thus, tissue type should also be considered when color-normalizing digital pathology slides between laboratories.