1. Introduction
A DNA sequence holds the most fundamental information about an individual life form; thus, its analysis offers significant potential for a variety of studies related to biology and medicine [
1,
2,
3,
4,
5,
6,
7,
8,
9,
10,
11,
12,
13,
14]. Traditional DNA sequence analysis, such as high-resolution melting (HRM) [
3,
15], is widely used not only in lab-scale research but also in real-world applications for pathogen diagnostics. During the recent coronavirus disease of 2019 (COVID-19) pandemic, real-time polymerase chain reaction (PCR), which is based on HRM, has been utilized as the standard diagnostic tool since it is the most powerful and reliable technique available.
However, laser-based DNA analysis schemes have been introduced in the last few years, challenging the traditional PCR technique in simplicity, sample consumption, and particularly in detection speed [
16,
17,
18,
19,
20,
21,
22]. Since a laser is a nonlinear optical phenomenon with a laser threshold, DNA sequences can be analyzed with a single laser emission. When compared with traditional DNA analysis, which takes at least a few hours, laser-based analysis has a huge advantage in rapid, high-throughput DNA sequential detection.
DNA detecting lasers that have been described mostly utilize optofluidic platforms, and laser gain has been obtained using dyes labeled with DNA or intercalating dyes. Similarly to traditional fluorescence-based analyses, dye-labeled DNA molecules, such as molecular beacons, have allowed for efficient DNA detecting lasers; however, they include complex DNA labeling sample preparation processes [
16,
18,
20]. The use of intercalating dyes rendered laser-based DNA analysis simple and cost-effective, and lasers were obtained with several different laser cavities/schemes [
17,
21]. While an optofluidic platform with a Fabry–Perot cavity implemented laser-based DNA detection [
22], a few different ring resonator cavities exhibited superior laser characteristics owing to an intrinsically higher Q factor. The very first form of the optofluidic ring resonator (OFRR)-based DNA laser was introduced in 2012 [
17]. This technique utilizes a glass capillary filled with a DNA sample mixed with intercalating dye, demonstrating that DNA sequences can be simply and rapidly detected using DNA laser emission. However, since the non-target DNA still emits fluorescence, which does not contribute to the laser feedback, the analysis cannot be regarded as a “digital-like” scheme from an optical standpoint. DNA probe molecules functionalized on the surface of the OFRR enabled a truly digital-like optical detection of DNA sequences, suppressing the fluorescence background from the non-target DNA molecules [
21]. A drawback of this novel method is that pretreatment of the OFRR with a DNA probe hinders the performance of a rapid, high-throughput DNA analysis platform, which is the most desirable trait of laser-based DNA analysis to replace existing traditional DNA analysis.
In this study, we propose an OFRR laser scheme using microdroplets of a DNA/intercalating dye and investigate optically true, digital-like DNA sequence detection to yield a realistic, rapid, high-throughput DNA analysis platform. The nL-sized microdroplets filled with a simple mixture of DNA and intercalating dye are individual reaction chambers for high-throughput analysis, and the glass-capillary-based OFRR laser connected downstream detects the DNA laser by analyzing their sequences. The laser emission was subsequently detected by the fiber taper to reject the unwanted fluorescence background. This resulted in a truly digital optical signal similar to the DNA laser with the surface-functionalized OFRR.
2. Materials and Methods
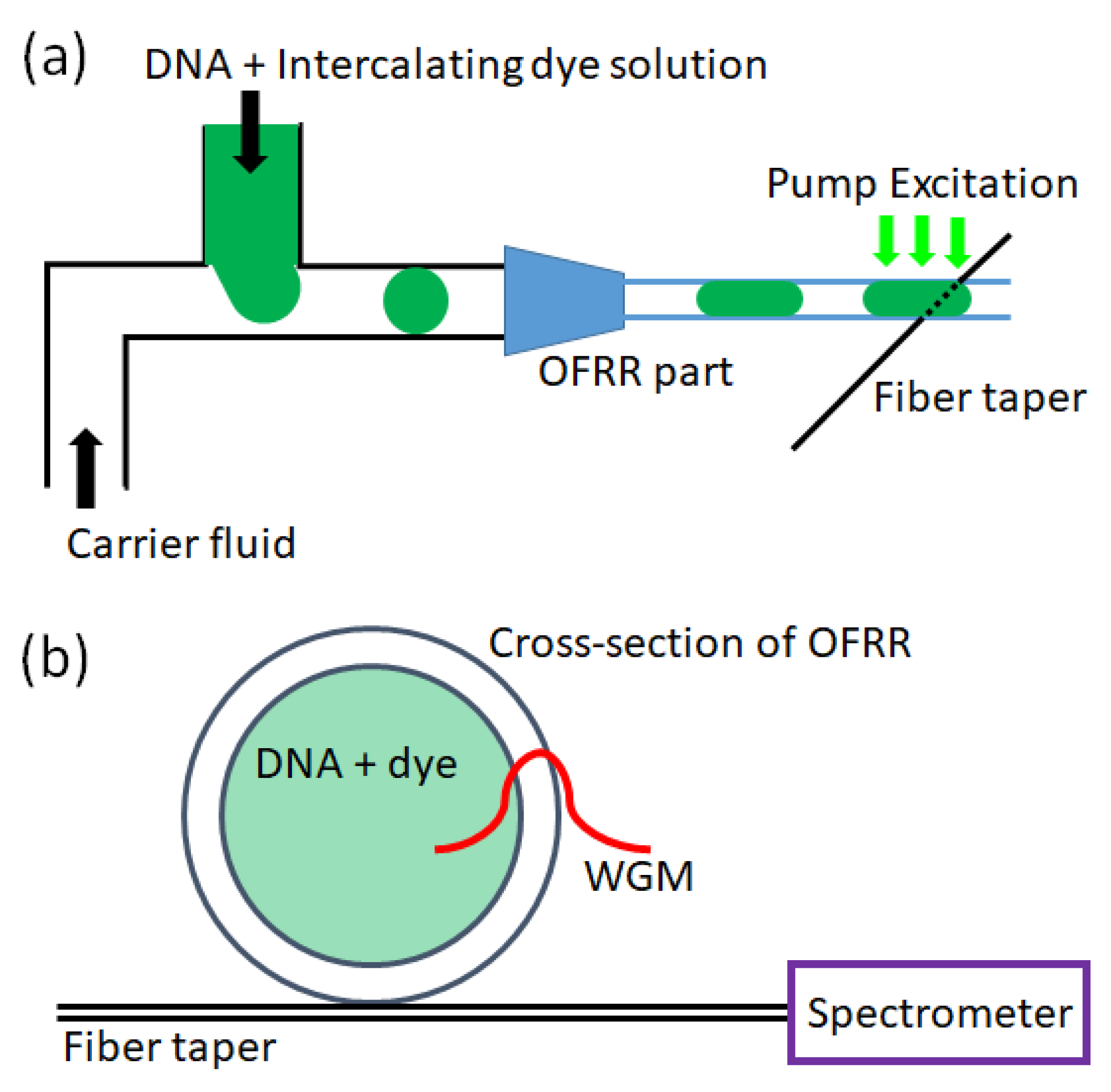
The microdroplets were generated with plastic tubing (Upchurch, P1476, inner diameter (ID) = 150 μm) and T-junctions (Upchurch, P-890), as schematically shown in
Figure 1a, in accordance with previous studies [
23,
24,
25]. Silicon oil was introduced upstream as a hydrophobic carrier fluid, and an aqueous solution of DNA and intercalating dye (SYTO
®-13 Green Fluorescent Nucleic Acid Stain, Life Technologies, Carlsbad, CA, USA) was injected into the T-junction. The flow rates of the carrier fluid and DNA samples were 10 μL/min and 3 μL/min, respectively. The immiscible phase forms individual microdroplets, of which approximately 125 nL can be derived from a generation rate of 0.4 Hz.
The fused silica glass capillary was fabricated using the existing drawing method with CO2 laser irradiation and an inner diameter and wall thickness of approximately 70 μm and 7 μm, respectively. The microdroplets of DNA/dye were infused downstream into the capillary OFRR. In a capillary with a smaller diameter, the DNA/dye microdroplets filled the entire cross-section, placing the DNA/dye samples in contact with the capillary wall. When the microdroplets passed a point of interest, the OFRR was optically pumped using an optical parametric oscillator (continuum, 488 nm wavelength, 5 ns pulse width, 20 Hz repetition rate). As the evanescent field of light confined in the OFRR whispering-gallery mode (WGM) interacted with the DNA/dye solution, a laser was emitted and coupled into the fiber taper in contact with the OFRR. Only the laser emission was carried into the spectrometer (iHR550, Horiba Jobin Yvon) connected to the distal end of the fiber taper, excluding the undesired fluorescence background.
The microdroplets can also serve as high-Q OFRR cavities because they are natural spheres with smooth surfaces. However, the relatively low refractive index of the aqueous DNA/dye solution (~1.33) renders it impossible to confine light when the droplets are in carrier fluid with a higher index. Microdroplets in a carrier fluid are free from evaporation issues and are individual reaction chambers that can be easily manipulated. Thus, this platform is ideal for high-throughput biosensors and/or molecular detectors. However, a low water index significantly limits the choices of the continuous phase for light confinement in the WGM. This is one of the main reasons for utilizing a glass capillary in the laser cavity. Although most of the WGM-confined light resides inside the glass capillary, there is enough of an evanescent field inside, resulting in sufficient laser feedback from the laser gain, which is the DNA/dye mixture in this experiment.
The arbitrary DNA sequences used in this study are listed in
Table 1. The target DNA, which was perfectly matched double-stranded DNA, had 21 base pairs. Sequences 1 and 2 of single-stranded DNA (ssDNA) forming the target double-stranded DNA were complementary pairs. In control experiments, base mismatches in the middle of the sequences were intentionally introduced to verify that our DNA analysis platform was capable of distinguishing the non-target DNA with only a single base mismatch. In this study, we classified the base mismatched DNA described above using the suggested microdroplet OFRR laser system. Therefore, our DNA detection platform could distinguish any kind of non-target DNA with a thermodynamic difference larger than a single base mismatch.
Both samples were mixed with SYTO 13 nucleic acid stain (Invitrogen), and the DNA intercalating dye was originally dissolved in dimethyl sulfoxide (DMSO) at a concentration of 5 mM. The final concentrations of DNA and dye were 250 µM when dissolved in a buffer solution (Tris-acetate ethylenediaminetetraacetic acid [EDTA] buffer, pH = 8.3).
3. Results
The laser emission spectra from the target DNA assay and base mismatched DNA samples are shown in
Figure 2. While microdroplets of each DNA sample were generated in separate fluidic tubing systems, both droplet samples were connected to the same OFRR and optically pumped with an energy density of 2.1 mJ/mm
2. The curves were shifted vertically for clarity. The target DNA samples exhibited a clear laser emission with typical multi-mode lasing from the OFRR cavity. However, the base mismatched DNA samples presented virtually zero optical signals. Although a free-space coupled fluorescence emission from the base mismatched samples can be observed with the naked eye, there is no observable optical signal from the detection portion. This is because our system, with a fiber taper coupler, only detects the laser emission, excluding the undesired fluorescence background. The pump energy density of 2.1 mJ/mm
2 is above the lasing threshold of the target DNA laser and below that of the base mismatched DNA laser. Thus, the target DNA was distinguishable from non-target DNA using only a single pulse of laser emission.
To further investigate the lasing threshold difference between the target and base mismatches, the laser emission intensities were measured by varying the pump energy density.
Figure 3 illustrates the spectrally integrated laser emission intensity as a function of pump energy density. The lasing threshold derived from the linear fit of the laser intensity was approximately 0.62 mJ/mm
2. This is slightly higher than that of the DNA laser with a continuous phase but comparable to previous reports. When microdroplets of the DNA sample solution are inserted into the OFRR portion, the droplets are elongated and the DNA solution comes into contact with the capillary OFRR wall. This is because the hydrophilic phase of the aqueous solution expels the hydrophobic continuous phase (silicon oil). Thus, this is likely not the main reason for the slight increase in the laser threshold. In fact, laser characteristics can be affected by several factors, such as the intrinsic Q factor of the cavity.
The most important caveat of this experiment is that lasing from the base mismatched DNA (non-target DNA) is not measurable, even at a high pump energy density. However, the laser threshold of the base mismatched DNA is significantly higher than that of the target at room temperature. Therefore, distinguishing the 21-base-pair target DNA from non-target samples is a straightforward process with our laser-based DNA sequence analysis system.
The most distinctive advantage of our DNA analysis system Is that it can rapidly detect the target DNA in different samples. To prove this concept, each microdroplet should be analyzed separately; thus, they can be regarded as individual reaction chambers with small volumes. We generated alternating microdroplets of the target DNA and base-mismatched DNA with two T-junctions in the fluidic tubing portion, as previously described.
Figure 4 shows the optical signals observed using the spectrometer at the distal end of the fiber taper as a function of time. When the target DNA sample microdroplets passed the analysis point of the OFRR, clear laser emission was observed, and there was no optical signal when the base-mismatched microdroplets were analyzed. This directly proves that our OFRR DNA detection platform is suitable for rapidly analyzing multiple microdroplets with different DNA sequences.
4. Discussion
Using advanced microfluidic systems, microdroplets of various DNA samples can be simultaneously generated and manipulated, and each individual microdroplet can be read if the target DNA is present with a single laser emission. We propose an optofluidic system that generates microdroplets of DNA samples and detects the target DNA with only a single laser emission. This novel DNA analysis platform is based on a glass capillary OFRR laser cavity, which is capable of lasing in an aqueous DNA solution with a relatively low index. In addition, only the laser signal was observed in the detection portion despite the unwanted fluorescence background from the DNA/dye solution, which does not contribute to the laser emission. This is because the optical signal is coupled into the fiber taper and measured. These data suggest that our DNA detection scheme is truly digital from an optical perspective.
More importantly, our DNA analysis platform is capable of rapidly detecting the target DNA from multiple microdroplets because the DNA laser can be measured separately using the suggested OFRR system. Recent advances in microfluidics enabled the generation and manipulation of many individual microdroplets. Combined with this, our laser system is a promising, novel technology for high-throughput, rapid DNA sequence detection platforms, resolving the bottleneck of time-consuming traditional DNA analysis.
Author Contributions
Conceptualization, W.L.; methodology, W.L.; investigation, C.S.J. and W.L.; writing—original draft preparation, C.S.J. and W.L.; writing—review and editing, W.L.; visualization, W.L.; supervision, W.L.; project administration, W.L. All authors have read and agreed to the published version of the manuscript.
Funding
This work was supported by the R&D program funded by the Korea Institute of Science and Technology (2E31781).
Institutional Review Board Statement
Not applicable.
Informed Consent Statement
Not applicable.
Data Availability Statement
Not applicable.
Conflicts of Interest
The authors declare no conflict of interest.
References
- Canales, R.D.; Luo, Y.; Willey, J.C.; Austermiller, B.; Barbacioru, C.C.; Boysen, C.; Hunkapiller, K.; Jensen, R.V.; Knight, C.R.; Lee, K.Y.; et al. Evaluation of DNA microarray results with quantitative gene expression platforms. Nat. Biotechnol. 2006, 24, 1115–1122. [Google Scholar] [CrossRef]
- Chen, Z.; Qian, S.; Chen, X.; Chen, J.; Lin, Y.; Liu, J. Label. Label-free detection of target DNA sequence and single-base mismatch in hepatitis C virus corresponding to oligonucleotide by resonance light scattering technique. RSC Adv. 2012, 2, 2562. [Google Scholar] [CrossRef]
- Christopoulos, T.K. Nucleic acid analysis. Anal. Chem. 1999, 71, 425–438. [Google Scholar] [CrossRef]
- Hawkins, R.D.; Hon, G.C.; Ren, B. Next-generation genomics: An integrative approach. Nat. Rev. Genet. 2010, 11, 476–486. [Google Scholar] [CrossRef] [PubMed]
- Higuchi, R.; von Beroldingen, C.H.v.; Sensabaugh, G.F.; Erlich, H.A. DNA typing from single hairs. Nature. 1988, 332, 543–546. [Google Scholar] [CrossRef]
- Jackson, S.P.; Bartek, J. The DNA-damage response in human biology and disease. Nature. 2009, 461, 1071–1078. [Google Scholar] [CrossRef] [Green Version]
- Jones, P.A.; Baylin, S.B. The fundamental role of epigenetic events in cancer. Nat. Rev. Genet. 2002, 3, 415–428. [Google Scholar] [CrossRef]
- Kapitonov, V.V.; Jurka, J. Molecular paleontology of transposable elements in the Drosophila melanogaster genome. Proc. Natl Acad. Sci. USA 2003, 100, 6569–6574. [Google Scholar] [CrossRef] [Green Version]
- Lupski, J.R. Genomic disorders: Structural features of the genome can lead to DNA rearrangements and human disease traits. Trends Genet. 1998, 14, 417–422. [Google Scholar] [CrossRef]
- Pollack, J.R.; Perou, C.M.; Alizadeh, A.A.; Eisen, M.B.; Pergamenschikov, A.; Williams, C.F.; Jeffrey, S.S.; Botstein, D.; Brown, P.O. Genome-wide analysis of DNA copy-number changes using cDNA microarrays. Nat. Genet. 1999, 23, 41–46. [Google Scholar] [CrossRef]
- Robertson, K.D. DNA methylation and human disease. Nat. Rev. Genet. 2005, 6, 597–610. [Google Scholar] [CrossRef] [PubMed]
- Shastry, B.S. Pharmacogenetics and the concept of individualized medicine. Pharm. J. 2006, 6, 16–21. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Taylor, R.W.; Turnbull, D.M. Mitochondrial DNA mutations in human disease. Nat. Rev. Genet. 2005, 6, 389–402. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wang, D.G.; Fan, J.B.; Siao, C.J.; Berno, A.; Young, P.; Sapolsky, R.; Ghandour, G.; Perkins, N.; Winchester, E.; Spencer, J.; et al. Large-scale identification, mapping, and genotyping of single-nucleotide polymorphisms in the human genome. Science. 1998, 280, 1077–1082. [Google Scholar] [CrossRef] [Green Version]
- Montgomery, J.; Wittwer, C.T.; Palais, R.; Zhou, L. Simultaneous mutation scanning and genotyping by high-resolution DNA melting analysis. Nat. Protoc. 2007, 2, 59–66. [Google Scholar] [CrossRef]
- Sun, Y.; Shopova, S.I.; Wu, C.S.; Arnold, S.; Fan, X. Bioinspired optofluidic FRET lasers via DNA scaffolds. Proc. Natl Acad. Sci. USA 2010, 107, 16039–16042. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lee, W.; Fan, X. Intracavity DNA melting analysis with optofluidic lasers. Anal. Chem. 2012, 84, 9558–9563. [Google Scholar] [CrossRef]
- Sun, Y.; Fan, X. Distinguishing DNA by analog-to-digital-like conversion by using optofluidic lasers. Angew. Chem. Int Ed Engl. 2012, 51, 1236–1239. [Google Scholar] [CrossRef] [Green Version]
- Chen, Q.; Liu, H.; Lee, W.; Sun, Y.; Zhu, D.; Pei, H.; Fan, C.; Fan, X. Self-assembled DNA tetrahedral optofluidic lasers with precise and tunable gain control. Lab Chip 2013, 13, 3351–3354. [Google Scholar] [CrossRef]
- Fan, X.; Yun, S.H. The potential of optofluidic biolasers. Nat. Methods 2014, 11, 141–147. [Google Scholar] [CrossRef] [PubMed]
- Lee, W.; Chen, Q.; Fan, X.; Yoon, D.K. Digital DNA detection based on a compact optofluidic laser with ultra-low sample consumption. Lab Chip 2016, 16, 4770–4776. [Google Scholar] [CrossRef] [Green Version]
- Hou, M.; Liang, X.; Zhang, T.; Qiu, C.; Chen, J.; Liu, S.; Wang, W.; Fan, X.D.N.A. Melting analysis with optofluidic lasers based on Fabry-Perot microcavity. ACS Sens. 2018, 3, 1750–1755. [Google Scholar] [CrossRef]
- Trivedi, V.; Doshi, A.; Kurup, G.K.; Ereifej, E.; Vandevord, P.J.; Basu, A.S. A modular approach for the generation, storage, mixing, and detection of droplet libraries for high throughput screening. Lab Chip 2010, 10, 2433–2442. [Google Scholar] [CrossRef]
- Lee, W.; Luo, Y.; Zhu, Q.; Fan, X. Versatile optofluidic ring resonator lasers based on microdroplets. Opt. Express. 2011, 19, 19668–19674. [Google Scholar] [CrossRef]
- Saqib, M.; Şahinoğlu, O.B.; Erdem, E.Y. Alternating droplet formation by using tapered channel geometry. Sci. Rep. 2018, 8, 1606. [Google Scholar] [CrossRef]
| Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).