Peripheral Pulmonary Lesions Classification Using Endobronchial Ultrasonography Images Based on Bagging Ensemble Learning and Down-Sampling Technique
Abstract
1. Introduction
- The CAD system implements a down-sampling technique that alleviates repercussions from imbalanced data to improve performance. Every benign case in the training set was used during the model’s training phase, whereas malignant cases were down-sampled and averaged out.
- If only malignant cases are down-sampled, the training dataset will not be fully utilized, and data will be wasted; therefore, the CAD system harnesses an ensemble learning technique. All trained models are integrated to combine all benign and malignant dataset features for the final classification.
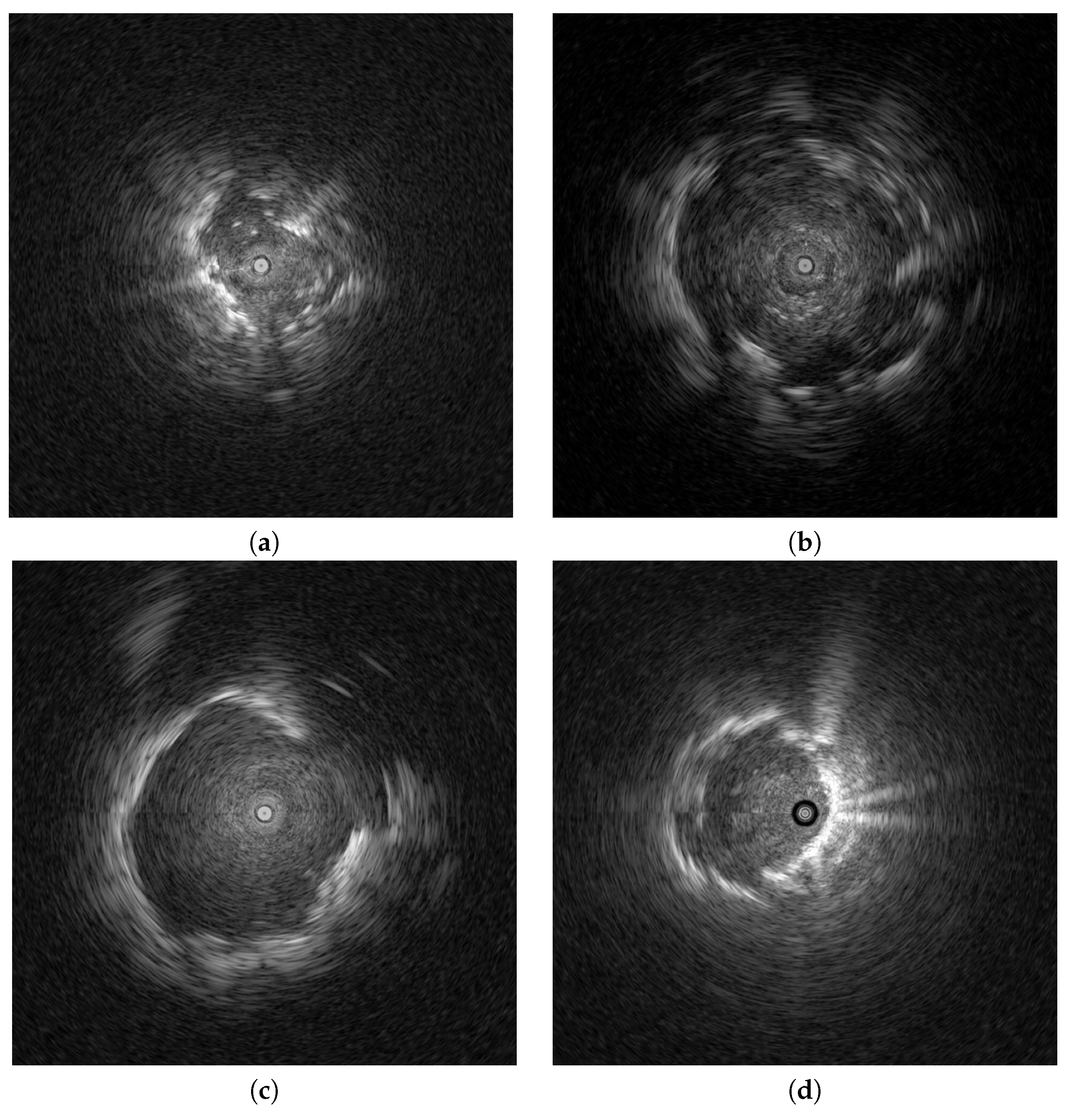
2. Experimental Materials
3. Proposed Methods
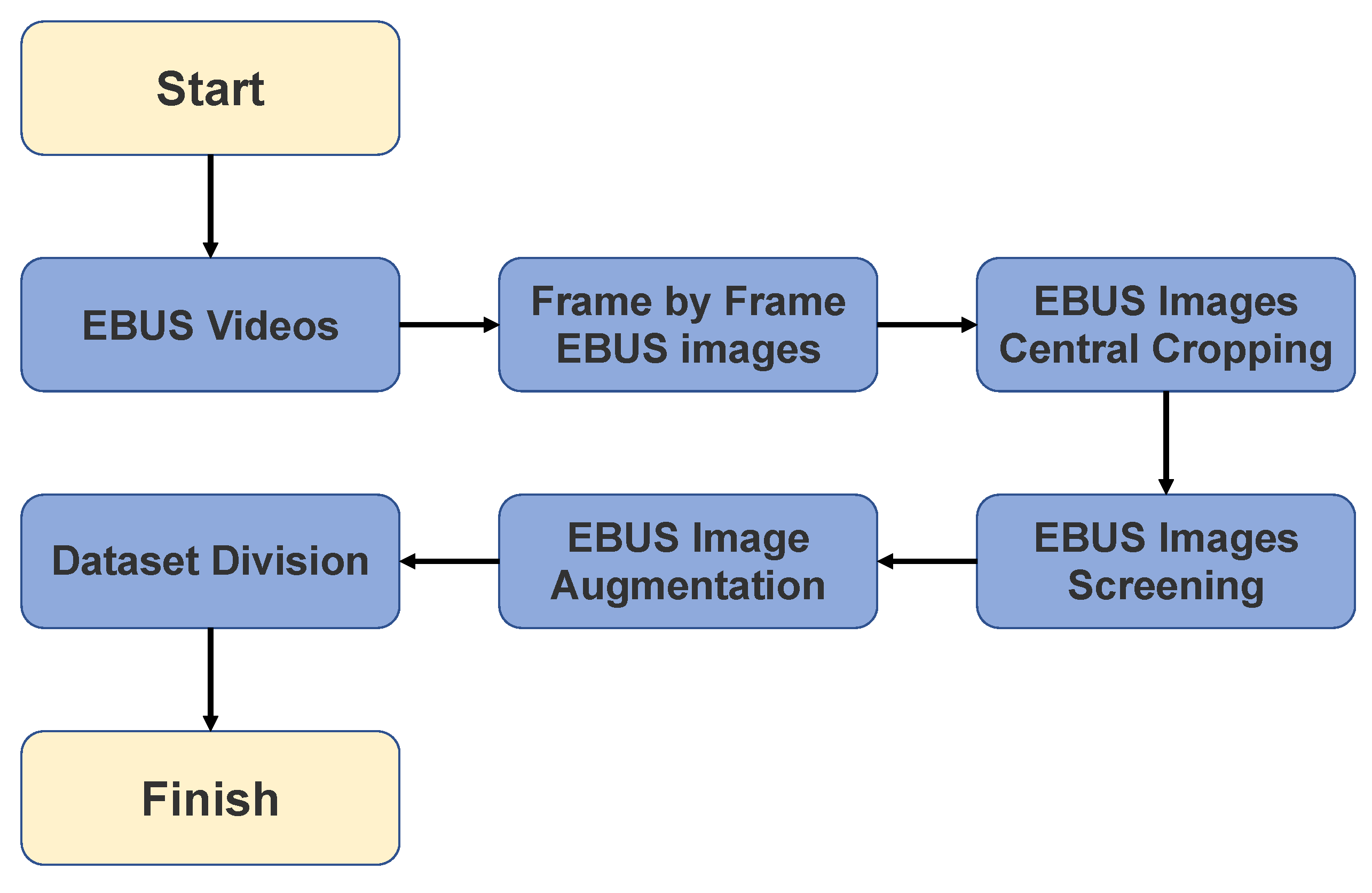
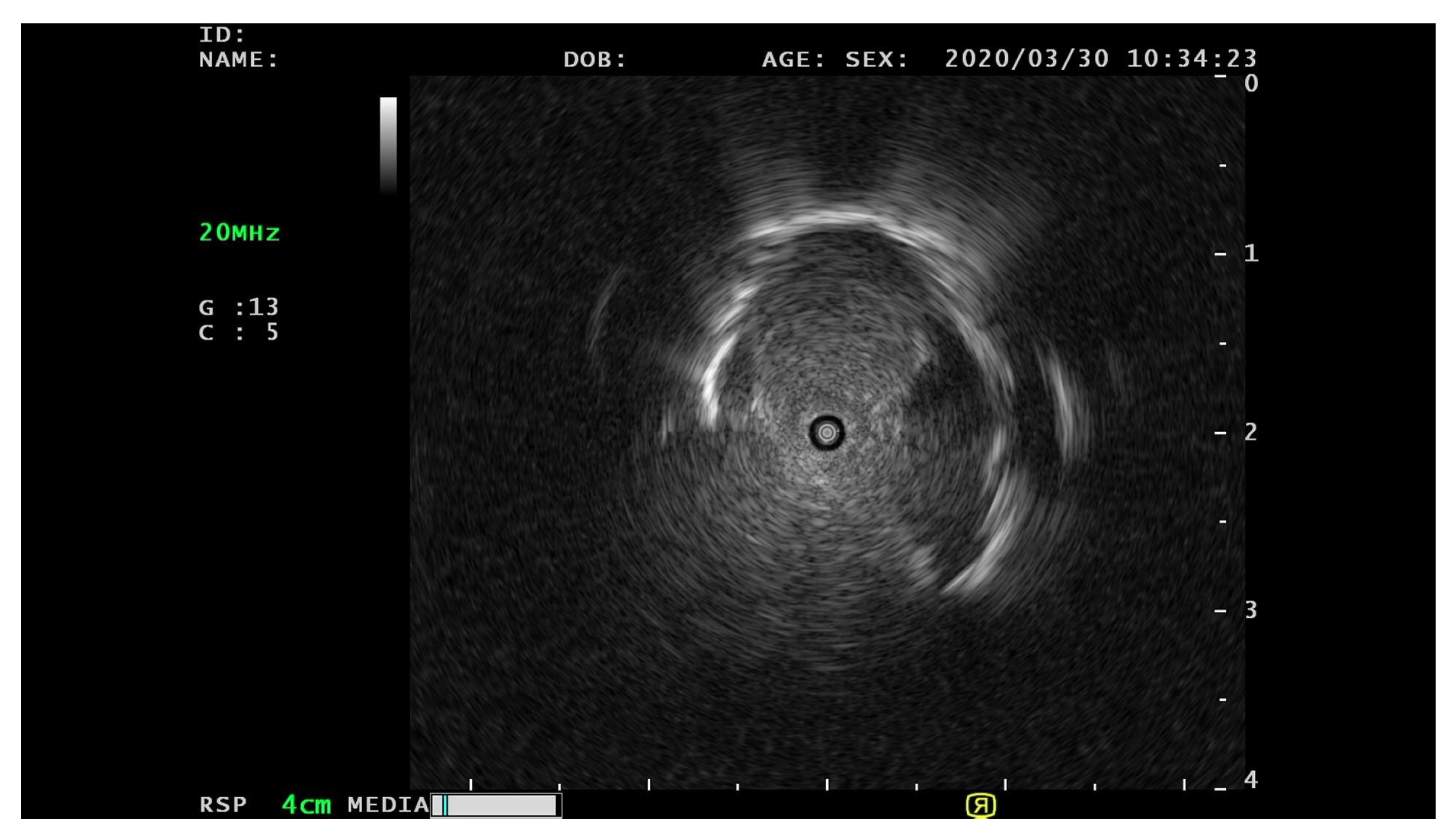
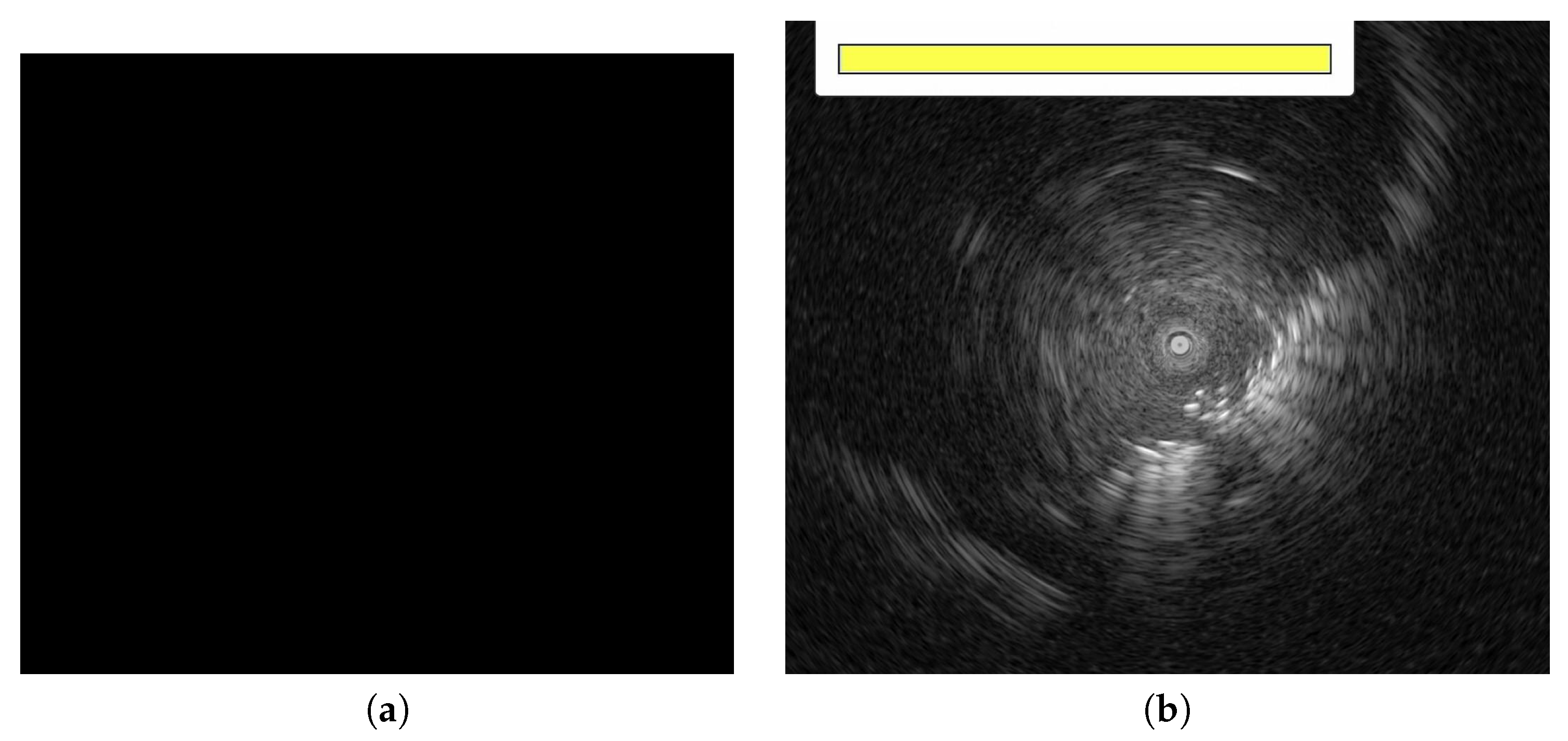
3.1. Data Pre-Processing
3.2. Data Balancing
3.3. The CAD Framework
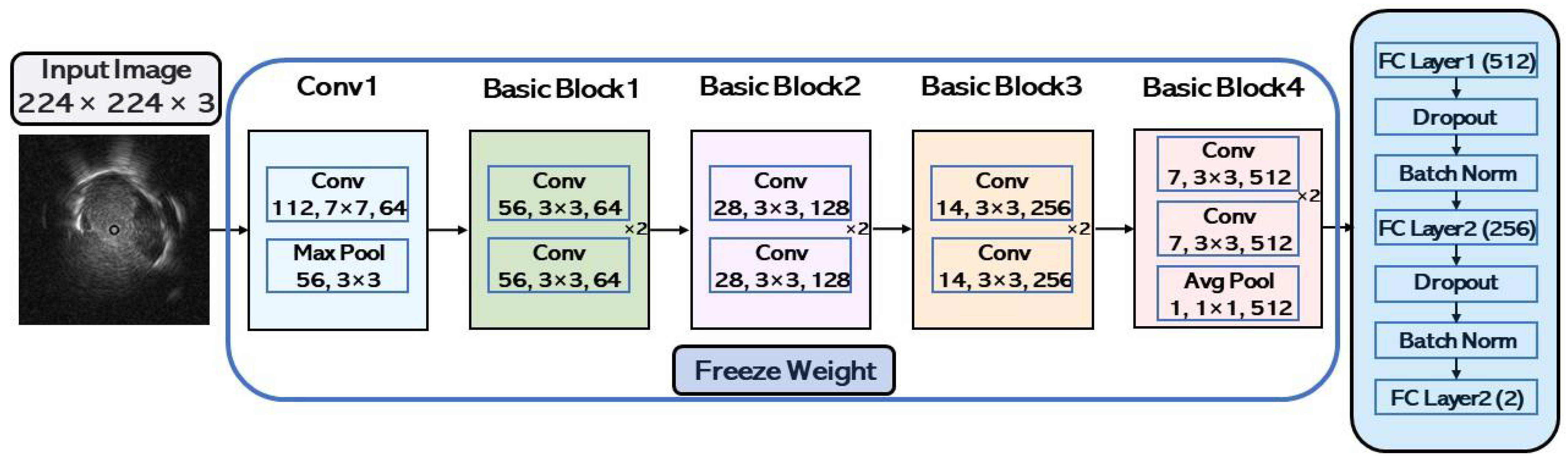
3.3.1. CNN Model Architecture
3.3.2. Down-Sampling Malignant Cases
3.3.3. Bagging Ensemble
3.4. Performance Evaluation
4. Experimental Results
4.1. Experimental Setup
4.2. Classification Results
4.2.1. Baseline Classification Results
4.2.2. Bagging Ensemble Classification Results
5. Discussion and Conclusions
5.1. Discussion
5.2. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Pizzato, M.; Li, M.; Vignat, J.; Laversanne, M.; Singh, D.; La Vecchia, C.; Vaccarella, S. The epidemiological landscape of thyroid cancer worldwide: GLOBOCAN estimates for incidence and mortality rates in 2020. Lancet Diabetes Endocrinol. 2022, 10, 264–272. [Google Scholar] [CrossRef] [PubMed]
- Alberg, A.J.; Brock, M.V.; Ford, J.G.; Samet, J.M.; Spivack, S.D. Epidemiology of Lung Cancer: Diagnosis and Management of Lung Cancer, 3rd ed: American College of Chest Physicians Evidence-Based Clinical Practice Guidelines. Chest 2013, 143, e1S–e29S. [Google Scholar] [CrossRef]
- Khomkham, B.; Lipikorn, R. Pulmonary Lesion Classification Framework Using the Weighted Ensemble Classification with Random Forest and CNN Models for EBUS Images. Diagnostics 2022, 12, 1552. [Google Scholar] [CrossRef]
- Chen, C.H.; Lee, Y.W.; Huang, Y.S.; Lan, W.R.; Chang, R.F.; Tu, C.Y.; Chen, C.Y.; Liao, W.C. Computer-aided diagnosis of endobronchial ultrasound images using convolutional neural network. Comput. Methods Programs Biomed. 2019, 177, 175–182. [Google Scholar] [CrossRef]
- Zhan, P.; Zhu, Q.Q.; Miu, Y.Y.; Liu, Y.F.; Wang, X.X.; Zhou, Z.J.; Jin, J.J.; Li, Q.; Sasada, S.; Izumo, T.; et al. Comparison between endobronchial ultrasound-guided transbronchial biopsy and CT-guided transthoracic lung biopsy for the diagnosis of peripheral lung cancer: A systematic review and meta-analysis. Transl. Lung Cancer Res. 2017, 6, 23. [Google Scholar] [CrossRef] [PubMed]
- Lou, L.; Huang, X.; Tu, J.; Xu, Z. Endobronchial ultrasound-guided transbronchial needle aspiration in peripheral pulmonary lesions: A systematic review and meta-analysis. Clin. Exp. Metastasis 2022, 40, 45–52. [Google Scholar] [CrossRef]
- Hearst, M.A.; Dumais, S.T.; Osuna, E.; Platt, J.; Scholkopf, B. Support vector machines. IEEE Intell. Syst. Their Appl. 1998, 13, 18–28. [Google Scholar] [CrossRef]
- Breiman, L. Random forests. Mach. Learn. 2001, 45, 5–32. [Google Scholar] [CrossRef]
- Kuruvilla, J.; Gunavathi, K. Lung cancer classification using neural networks for CT images. Comput. Methods Programs Biomed. 2014, 113, 202–209. [Google Scholar] [CrossRef] [PubMed]
- Alakwaa, W.; Nassef, M.; Badr, A. Lung cancer detection and classification with 3D convolutional neural network (3D-CNN). Int. J. Adv. Comput. Sci. Appl. 2017, 8, 409–417. [Google Scholar] [CrossRef]
- Chaunzwa, T.L.; Hosny, A.; Xu, Y.; Shafer, A.; Diao, N.; Lanuti, M.; Christiani, D.C.; Mak, R.H.; Aerts, H.J. Deep learning classification of lung cancer histology using CT images. Sci. Rep. 2021, 11, 5471. [Google Scholar] [CrossRef]
- Nasrullah, N.; Sang, J.; Alam, M.S.; Mateen, M.; Cai, B.; Hu, H. Automated lung nodule detection and classification using deep learning combined with multiple strategies. Sensors 2019, 19, 3722. [Google Scholar] [CrossRef]
- Chen, J.; Zeng, H.; Zhang, C.; Shi, Z.; Dekker, A.; Wee, L.; Bermejo, I. Lung cancer diagnosis using deep attention-based multiple instance learning and radiomics. Med. Phys. 2022, 49, 3134–3143. [Google Scholar] [CrossRef]
- He, K.; Zhang, X.; Ren, S.; Sun, J. Deep residual learning for image recognition. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Las Vegas, NV, USA, 27–30 June 2016; pp. 770–778. [Google Scholar]
- Huang, G.; Liu, Z.; Van Der Maaten, L.; Weinberger, K.Q. Densely connected convolutional networks. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Honolulu, HI, USA, 21–26 July 2017; pp. 4700–4708. [Google Scholar]
- Sandler, M.; Howard, A.; Zhu, M.; Zhmoginov, A.; Chen, L.C. Mobilenetv2: Inverted residuals and linear bottlenecks. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Salt Lake City, UT, USA, 18–23 June 2018; pp. 4510–4520. [Google Scholar]
- Ma, N.; Zhang, X.; Zheng, H.T.; Sun, J. Shufflenet v2: Practical guidelines for efficient cnn architecture design. In Proceedings of the European Conference on Computer Vision (ECCV), Munich, Germany, 8–14 September 2018; pp. 116–131. [Google Scholar]
- Hinton, G.E.; Srivastava, N.; Krizhevsky, A.; Sutskever, I.; Salakhutdinov, R.R. Improving neural networks by preventing co-adaptation of feature detectors. arXiv 2012, arXiv:1207.0580. [Google Scholar]
- Ioffe, S.; Szegedy, C. Batch normalization: Accelerating deep network training by reducing internal covariate shift. Proc. Mach. Learn. Res. 2015, 37, 448–456. [Google Scholar]
- Kingma, D.P.; Ba, J. Adam: A method for stochastic optimization. arXiv 2014, arXiv:1412.6980. [Google Scholar]
- Tibshirani, R. Regression shrinkage and selection via the lasso. J. R. Stat. Soc. Ser. B Methodol. 1996, 58, 267–288. [Google Scholar] [CrossRef]
- Vidaurre, D.; Bielza, C.; Larranaga, P. A survey of L1 regression. Int. Stat. Rev. 2013, 81, 361–387. [Google Scholar] [CrossRef]
- Van Laarhoven, T. L2 regularization versus batch and weight normalization. arXiv 2017, arXiv:1706.05350. [Google Scholar]
- He, H.; Garcia, E.A. Learning from imbalanced data. IEEE Trans. Knowl. Data Eng. 2009, 21, 1263–1284. [Google Scholar]
- Buda, M.; Maki, A.; Mazurowski, M.A. A systematic study of the class imbalance problem in convolutional neural networks. Neural Netw. 2018, 106, 249–259. [Google Scholar] [CrossRef]
- Haixiang, G.; Yijing, L.; Shang, J.; Mingyun, G.; Yuanyue, H.; Bing, G. Learning from class-imbalanced data: Review of methods and applications. Expert Syst. Appl. 2017, 73, 220–239. [Google Scholar] [CrossRef]
- Levi, G.; Hassner, T. Age and gender classification using convolutional neural networks. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition Workshops, Boston, MA, USA, 7–12 June 2015; pp. 34–42. [Google Scholar]
- Elkan, C. The foundations of cost-sensitive learning. In Proceedings of the International Joint Conference on Artificial Intelligence, Seattle, WA, USA, 4–10 August 2001; Volume 17, pp. 973–978. [Google Scholar]
- Rokach, L. Ensemble-based classifiers. Artif. Intell. Rev. 2010, 33, 1–39. [Google Scholar] [CrossRef]
- Breiman, L. Bagging predictors. Mach. Learn. 1996, 24, 123–140. [Google Scholar] [CrossRef]




| Model | Depth | Complexity FLOPs (G) | Total Parameters (M) |
|---|---|---|---|
| ResNet-18 | 18 | 58.36 | 11.34 |
| ResNet-34 | 34 | 117.71 | 21.45 |
| DenseNet-121 | 121 | 92.69 | 7.54 |
| DenseNet-169 | 169 | 109.93 | 13.40 |
| MobileNet-V2 | 54 | 10.45 | 2.58 |
| ShuffleNet-V2 | 56 | 1.40 | 0.64 |
| Hyper-Parameter | Value |
|---|---|
| Epochs | 50 |
| Batch size | 32 |
| Input size | 224 × 224 |
| Learning rate | 0.001 |
| Loss function | Cross-Entropy |
| Optimizer | Adam |
| Patience for early stopping | 5 |
| Regularization | L1 + L2 |
| Fold ID | Train Set | Validation Set | Test Set | |||
|---|---|---|---|---|---|---|
| Part ID | Case Num of B and M | Part ID | Case Num of B and M | Part ID | Case Num of B and M | |
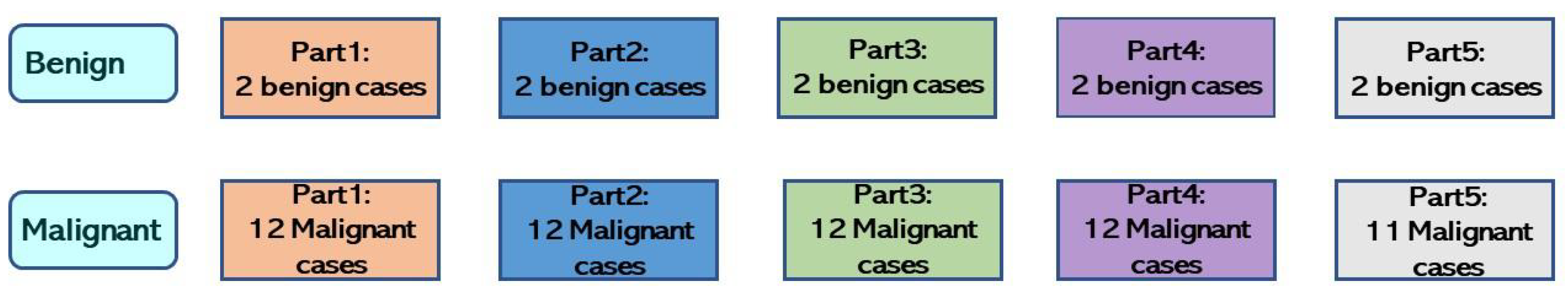
| Fold-1 | part2, part4, part5 | Benign: 6 Malignant: 35 | Part3 | Benign: 2 Malignant: 12 | Part1 | Benign: 2 Malignant: 12 |
| Fold-2 | part1, part3, part5 | Benign: 6 Malignant: 35 | Part4 | Benign: 2 Malignant: 12 | Part2 | Benign: 2 Malignant: 12 |
| Fold-3 | part1, part2, part4 | Benign: 6 Malignant: 36 | Part5 | Benign: 2 Malignant: 11 | Part3 | Benign: 2 Malignant: 12 |
| Fold-4 | part2, part3, part5 | Benign: 6 Malignant: 36 | Part1 | Benign: 2 Malignant: 12 | Part4 | Benign: 2 Malignant: 12 |
| Fold-5 | part1, part3, part4 | Benign: 6 Malignant: 36 | Part2 | Benign: 2 Malignant: 12 | Part5 | Benign: 2 Malignant: 11 |
| Model | Accuracy | F1-Score | AUC | PPV | NPV | Sensitivity | Specificity |
|---|---|---|---|---|---|---|---|
| ResNet-18 | 0.59 | 0.60 | 0.61 | 0.65 | 0.50 | 0.60 | 0.59 |
| ResNet-34 | 0.53 | 0.59 | 0.64 | 0.54 | 0.43 | 0.68 | 0.35 |
| DenseNet-121 | 0.62 | 0.66 | 0.66 | 0.68 | 0.54 | 0.68 | 0.54 |
| DenseNet-169 | 0.62 | 0.61 | 0.62 | 0.70 | 0.60 | 0.58 | 0.67 |
| MobileNet-V2 | 0.63 | 0.62 | 0.62 | 0.76 | 0.59 | 0.50 | 0.78 |
| ShuffleNet-V2 | 0.58 | 0.63 | 0.57 | 0.62 | 0.52 | 0.67 | 0.47 |
| Bagging Ensemble | Model Name | Model Numbers | Accuracy | F1-Score | AUC | PPV | NPV | Sensitivity | Specificity |
|---|---|---|---|---|---|---|---|---|---|
| Majority Voting | ResNet-18 | 3 | 0.62 | 0.56 | 0.62 | 0.76 | 0.57 | 0.51 | 0.73 |
| ResNet-34 | 3 | 0.65 | 0.61 | 0.65 | 0.78 | 0.62 | 0.57 | 0.73 | |
| DenseNet-121 | 3 | 0.62 | 0.60 | 0.62 | 0.67 | 0.61 | 0.59 | 0.64 | |
| ResNet-18+ | 3 | 0.67 | 0.61 | 0.67 | 0.83 | 0.64 | 0.55 | 0.79 | |
| DenseNet-169+ | |||||||||
| MobileNet-V2 | |||||||||
| ResNet-34+ | 3 | 0.67 | 0.60 | 0.67 | 0.78 | 0.65 | 0.54 | 0.80 | |
| ResNet-18+ | |||||||||
| ResNet-18 | |||||||||
| Mobilenet_V2+ | 3 | 0.66 | 0.62 | 0.66 | 0.77 | 0.66 | 0.58 | 0.74 | |
| DenseNet-121+ | |||||||||
| DenseNet-169 | |||||||||
| ResNet-34+ | 3 | 0.70 | 0.63 | 0.70 | 0.83 | 0.68 | 0.58 | 0.82 | |
| ResNet-18+ | |||||||||
| MobileNet-V2 |
| Bagging Ensemble | Model Name | Model Numbers | Accuracy | F1-Score | AUC | PPV | NPV | Sensitivity | Specificity |
|---|---|---|---|---|---|---|---|---|---|
| Output Average | ResNet-18 | 2 | 0.70 | 0.63 | 0.70 | 0.87 | 0.67 | 0.56 | 0.84 |
| ResNet-34 | 3 | 0.65 | 0.61 | 0.65 | 0.79 | 0.63 | 0.58 | 0.73 | |
| DenseNet-121 | 2 | 0.66 | 0.60 | 0.66 | 0.84 | 0.70 | 0.54 | 0.78 | |
| MobileNet-V2 | 3 | 0.64 | 0.57 | 0.64 | 0.75 | 0.61 | 0.51 | 0.77 | |
| ResNet-18+ | 3 | 0.67 | 0.61 | 0.67 | 0.84 | 0.64 | 0.55 | 0.80 | |
| DenseNet-169+ | |||||||||
| MobileNet-V2 | |||||||||
| ResNet-34+ | 3 | 0.68 | 0.60 | 0.68 | 0.81 | 0.66 | 0.53 | 0.84 | |
| Output | ResNet-18+ | ||||||||
| Average | ResNet-18 | ||||||||
| Mobilenet_V2+ | 3 | 0.65 | 0.59 | 0.65 | 0.77 | 0.64 | 0.55 | 0.75 | |
| DenseNet-121+ | |||||||||
| DenseNet-169 | |||||||||
| ResNet-34+ | 3 | 0.70 | 0.63 | 0.70 | 0.84 | 0.68 | 0.57 | 0.84 | |
| ResNet-18+ | |||||||||
| MobileNet-V2 |
| Bagging Ensemble | Model Name | Model Numbers | Accuracy | F1-Score | AUC | PPV | NPV | Sensitivity | Specificity |
|---|---|---|---|---|---|---|---|---|---|
| ResNet-18 | 3 | 0.68 | 0.56 | 0.67 | 0.87 | 0.63 | 0.46 | 0.91 | |
| ResNet-34 | 2 | 0.68 | 0.61 | 0.71 | 0.82 | 0.67 | 0.56 | 0.80 | |
| DenseNet-121 | 2 | 0.68 | 0.64 | 0.69 | 0.81 | 0.69 | 0.60 | 0.76 | |
| MobileNet-V2 | 2 | 0.67 | 0.58 | 0.69 | 0.87 | 0.62 | 0.49 | 0.86 | |
| ResNet-18+ | 3 | 0.67 | 0.61 | 0.73 | 0.83 | 0.63 | 0.54 | 0.80 | |
| DenseNet-169+ | |||||||||
| Performance | MobileNet-V2 | ||||||||
| Weighting | ResNet-34+ | 3 | 0.69 | 0.60 | 0.72 | 0.82 | 0.66 | 0.53 | 0.85 |
| ResNet-18+ | |||||||||
| ResNet-18 | |||||||||
| Mobilenet_V2+ | 3 | 0.65 | 0.60 | 0.68 | 0.76 | 0.64 | 0.56 | 0.74 | |
| DenseNet-121+ | |||||||||
| DenseNet-169 | |||||||||
| ResNet-34+ | 3 | 0.70 | 0.63 | 0.75 | 0.84 | 0.68 | 0.56 | 0.85 | |
| ResNet-18+ | |||||||||
| MobileNet-V2 |
| Model Name or Fusion Name | Accuracy | F1-Score | AUC | PPV | NPV | Sensitivity | Specificity | |
|---|---|---|---|---|---|---|---|---|
| Baseline | ResNet-18 | 0.59 | 0.60 | 0.61 | 0.65 | 0.50 | 0.60 | 0.59 |
| ResNet-34 | 0.53 | 0.59 | 0.64 | 0.54 | 0.43 | 0.68 | 0.35 | |
| DenseNet-121 | 0.62 | 0.66 | 0.66 | 0.68 | 0.54 | 0.68 | 0.54 | |
| DenseNet-169 | 0.62 | 0.61 | 0.62 | 0.70 | 0.60 | 0.58 | 0.67 | |
| MobileNet-V2 | 0.63 | 0.56 | 0.62 | 0.76 | 0.59 | 0.50 | 0.78 | |
| ShuffleNet-V2 | 0.58 | 0.63 | 0.57 | 0.62 | 0.52 | 0.67 | 0.47 | |
| Bagging Ensemble | Majority Voting | 0.70 | 0.63 | 0.70 | 0.83 | 0.68 | 0.58 | 0.82 |
| Output Average | 0.70 | 0.63 | 0.70 | 0.87 | 0.67 | 0.56 | 0.84 | |
| Performance Weighting | 0.70 | 0.63 | 0.75 | 0.84 | 0.68 | 0.56 | 0.85 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wang, H.; Shikano, K.; Nakajima, T.; Nomura, Y.; Nakaguchi, T. Peripheral Pulmonary Lesions Classification Using Endobronchial Ultrasonography Images Based on Bagging Ensemble Learning and Down-Sampling Technique. Appl. Sci. 2023, 13, 8403. https://doi.org/10.3390/app13148403
Wang H, Shikano K, Nakajima T, Nomura Y, Nakaguchi T. Peripheral Pulmonary Lesions Classification Using Endobronchial Ultrasonography Images Based on Bagging Ensemble Learning and Down-Sampling Technique. Applied Sciences. 2023; 13(14):8403. https://doi.org/10.3390/app13148403
Chicago/Turabian StyleWang, Huitao, Kohei Shikano, Takahiro Nakajima, Yukihiro Nomura, and Toshiya Nakaguchi. 2023. "Peripheral Pulmonary Lesions Classification Using Endobronchial Ultrasonography Images Based on Bagging Ensemble Learning and Down-Sampling Technique" Applied Sciences 13, no. 14: 8403. https://doi.org/10.3390/app13148403
APA StyleWang, H., Shikano, K., Nakajima, T., Nomura, Y., & Nakaguchi, T. (2023). Peripheral Pulmonary Lesions Classification Using Endobronchial Ultrasonography Images Based on Bagging Ensemble Learning and Down-Sampling Technique. Applied Sciences, 13(14), 8403. https://doi.org/10.3390/app13148403






