Genetic Association of APOA5 and AKT3 Genes with Milk Production Traits in Chinese Holstein Cows
Abstract
:1. Introduction
2. Materials and Methods
2.1. Animals and Phenotypic Data
2.2. DNA Extraction and Quality Control
2.3. SNP Identification and Genotyping
2.4. Association Analyses
2.5. Functional Prediction of Mutation Sites
3. Results
3.1. SNP Identification
3.2. Association Analyses between SNPs and Five Milk Production Traits
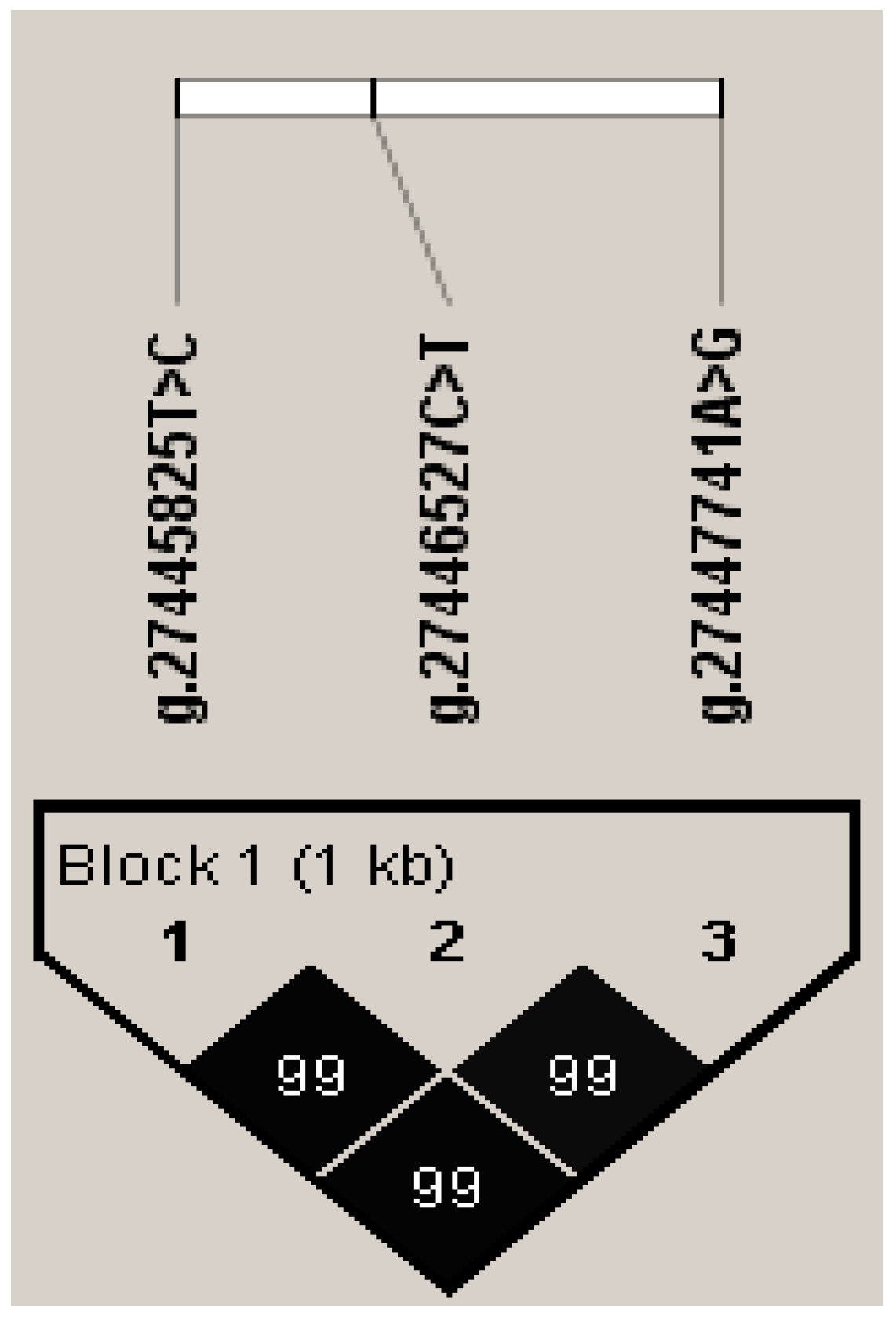
3.3. Association between Haplotype Block and Five Milk Production Traits
3.4. Changes in Transcription Factor Binding Sites Caused by SNPs in 5′ Region
3.5. Changes in mRNA and Protein Structure and Function by Missense Mutation
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Rumbold, P.; McCullogh, N.; Boldon, R.; Haskell-Ramsay, C.; James, L.; Stevenson, E.; Green, B. The potential nutrition-, physical- and health-related benefits of cow’s milk for primary-school-aged children. Nutr. Res. Rev. 2022, 35, 50–69. [Google Scholar] [CrossRef] [PubMed]
- Scholz-Ahrens, K.E.; Ahrens, F.; Barth, C.A. Nutritional and health attributes of milk and milk imitations. Eur. J. Nutr. 2020, 59, 19–34. [Google Scholar] [CrossRef] [PubMed]
- Dai, J.; Yin, T.; Cao, L. Dairy consumption and liver cancer risk: A meta-analysis of observational studies. Oncol. Lett. 2024, 27, 108. [Google Scholar] [CrossRef] [PubMed]
- Pereira, P.C. Milk nutritional composition and its role in human health. Nutrition 2014, 30, 619–627. [Google Scholar] [CrossRef] [PubMed]
- Tunick, M.H.; Van Hekken, D.L. Dairy Products and Health: Recent Insights. J. Agric. Food Chem. 2015, 63, 9381–9388. [Google Scholar] [CrossRef] [PubMed]
- Yang, S.; Bhargava, N.; Connor, A.O.; Gibney, E.R.; Feeney, E.L. Dairy consumption in adults in China: A systematic review. BMC Nutr. 2023, 9, 116. [Google Scholar] [CrossRef] [PubMed]
- Guinan, F.L.; Wiggans, G.R.; Norman, H.D.; Dürr, J.W.; Cole, J.B.; Van Tassell, C.P.; Misztal, I.; Lourenco, D. Changes in genetic trends in US dairy cattle since the implementation of genomic selection. J. Dairy Sci. 2023, 106, 1110–1129. [Google Scholar] [CrossRef]
- König, S.; Simianer, H.; Willam, A. Economic evaluation of genomic breeding programs. J. Dairy Sci. 2009, 92, 382–391. [Google Scholar] [CrossRef]
- Schaeffer, L.R. Strategy for applying genome-wide selection in dairy cattle. J. Anim. Breed. Genet. 2006, 123, 218–223. [Google Scholar] [CrossRef]
- Ding, X.; Zhang, Z.; Li, X.; Wang, S.; Wu, X.; Sun, D.; Yu, Y.; Liu, J.; Wang, Y.; Zhang, Y.; et al. Accuracy of genomic prediction for milk production traits in the Chinese Holstein population using a reference population consisting of cows. J. Dairy Sci. 2013, 96, 5315–5323. [Google Scholar] [CrossRef]
- Wiggans, G.R.; Cole, J.B.; Hubbard, S.M.; Sonstegard, T.S. Genomic Selection in Dairy Cattle: The USDA Experience. Annu. Rev. Anim. Biosci. 2017, 5, 309–327. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Z.; Ober, U.; Erbe, M.; Zhang, H.; Gao, N.; He, J.; Li, J.; Simianer, H. Improving the accuracy of whole genome prediction for complex traits using the results of genome wide association studies. PLoS ONE 2014, 9, e93017. [Google Scholar] [CrossRef] [PubMed]
- de Las, H.S.; Lopez, B.I.; Moghaddar, N.; Park, W.; Park, J.E.; Chung, K.Y.; Lim, D.; Lee, S.H.; Shin, D.; van der Werf, J. Use of gene expression and whole-genome sequence information to improve the accuracy of genomic prediction for carcass traits in Hanwoo cattle. Genet. Sel. Evol. 2020, 52, 54. [Google Scholar] [CrossRef] [PubMed]
- Han, B.; Yuan, Y.; Liang, R.; Li, Y.; Liu, L.; Sun, D. Genetic Effects of LPIN1 Polymorphisms on Milk Production Traits in Dairy Cattle. Genes 2019, 10, 265. [Google Scholar] [CrossRef] [PubMed]
- Khan, K.; Suhail, S.M.; Khan, R.; Ahmed, I.; Khan, F.A.; Khan, M.J. Genetic polymorphism of Β-casein gene and its association with milk production and composition in Azi-Kheli buffalo. Trop. Anim. Health Prod. 2023, 55, 94. [Google Scholar] [CrossRef] [PubMed]
- Sun, Y.; Wu, X.; Ma, Y.; Liu, D.; Lu, X.; Zhao, T.; Yang, Z. Molecular Marker-Assisted Selection of ABCG2, CD44, SPP1 Genes Contribute to Milk Production Traits of Chinese Holstein. Animals 2023, 13, 89. [Google Scholar] [CrossRef] [PubMed]
- Worku, D.; Gowane, G.; Verma, A. Genetic variation in promoter region of the bovine LAP3 gene associated with estimated breeding values of milk production traits and clinical mastitis in dairy cattle. PLoS ONE 2023, 18, e277156. [Google Scholar] [CrossRef]
- Fu, Y.; Jia, R.; Xu, L.; Su, D.; Li, Y.; Liu, L.; Ma, Z.; Sun, D.; Han, B. Fatty acid desaturase 2 affects the milk-production traits in Chinese Holsteins. Anim. Genet. 2022, 53, 422–426. [Google Scholar] [CrossRef]
- Du, A.; Zhao, F.; Liu, Y.; Xu, L.; Chen, K.; Sun, D.; Han, B. Genetic polymorphisms of PKLR gene and their associations with milk production traits in Chinese Holstein cows. Front. Genet. 2022, 13, 1002706. [Google Scholar] [CrossRef]
- Xu, L.; Shi, L.; Liu, L.; Liang, R.; Li, Q.; Li, J.; Han, B.; Sun, D. Analysis of Liver Proteome and Identification of Critical Proteins Affecting Milk Fat, Protein, and Lactose Metabolism in Dariy Cattle with iTRAQ. Proteomics 2019, 19, 1800387. [Google Scholar] [CrossRef]
- Coleman, D.N.; Vailati-Riboni, M.; Elolimy, A.A.; Cardoso, F.C.; Rodriguez-Zas, S.L.; Miura, M.; Pan, Y.; Loor, J.J. Hepatic betaine-homocysteine methyltransferase and methionine synthase activity and intermediates of the methionine cycle are altered by choline supply during negative energy balance in Holstein cows. J. Dairy Sci. 2019, 102, 8305–8318. [Google Scholar] [CrossRef] [PubMed]
- Fruchart-Najib, J.; Baugé, E.; Niculescu, L.; Pham, T.; Thomas, B.; Rommens, C.; Majd, Z.; Brewer, B.; Pennacchio, L.A.; Fruchart, J. Mechanism of triglyceride lowering in mice expressing human apolipoprotein A5. Biochem. Biophys. Res. Commun. 2004, 319, 397–404. [Google Scholar] [CrossRef] [PubMed]
- Pennacchio, L.A.; Olivier, M.; Hubacek, J.A.; Cohen, J.C.; Cox, D.R.; Fruchart, J.C.; Krauss, R.M.; Rubin, E.M. An apolipoprotein influencing triglycerides in humans and mice revealed by comparative sequencing. Science 2001, 294, 169–173. [Google Scholar] [CrossRef] [PubMed]
- Khan, M.J.; Jacometo, C.B.; Graugnard, D.E.; Correa, M.N.; Schmitt, E.; Cardoso, F.; Loor, J.J. Overfeeding Dairy Cattle During Late-Pregnancy Alters Hepatic PPARalpha-Regulated Pathways Including Hepatokines: Impact on Metabolism and Peripheral Insulin Sensitivity. Gene Regul. Syst. Biol. 2014, 8, 97–111. [Google Scholar] [CrossRef] [PubMed]
- Xu, T.; Cardoso, F.C.; Pineda, A.; Trevisi, E.; Shen, X.; Rosa, F.; Osorio, J.S.; Loor, J.J. Grain challenge affects systemic and hepatic molecular biomarkers of inflammation, stress, and metabolic responses to a greater extent in Holstein than Jersey cows. J. Dairy Sci. 2017, 100, 9153–9162. [Google Scholar] [CrossRef] [PubMed]
- Coffer, P.J.; Jin, J.; Woodgett, J.R. Protein kinase B (c-Akt): A multifunctional mediator of phosphatidylinositol 3-kinase activation. Biochem. J. 1998, 335 Pt 1, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Rivière, J.; Mirzaa, G.M.; O’Roak, B.J.; Beddaoui, M.; Alcantara, D.; Conway, R.L.; St-Onge, J.; Schwartzentruber, J.A.; Gripp, K.W.; Nikkel, S.M.; et al. De novo germline and postzygotic mutations in AKT3, PIK3R2 and PIK3CA cause a spectrum of related megalencephaly syndromes. Nat. Genet. 2012, 44, 934–940. [Google Scholar] [CrossRef]
- Bionaz, M.; Loor, J.J. mTOR, AMPK, and insulin receptor signaling networks in the bovine mammary gland during the lactation cycle. FASEB J. 2007, 21, A1109. [Google Scholar]
- Meredith, B.K.; Kearney, F.J.; Finlay, E.K.; Bradley, D.G.; Fahey, A.G.; Berry, D.P.; Lynn, D.J. Genome-wide associations for milk production and somatic cell score in Holstein-Friesian cattle in Ireland. BMC Genet. 2012, 13, 21. [Google Scholar] [CrossRef]
- Cole, J.B.; Wiggans, G.R.; Ma, L.; Sonstegard, T.S.; Lawlor, T.J.; Crooker, B.A.; Van Tassell, C.P.; Yang, J.; Wang, S.; Matukumalli, L.K.; et al. Genome-wide association analysis of thirty one production, health, reproduction and body conformation traits in contemporary U.S. Holstein cows. BMC Genom. 2011, 12, 408. [Google Scholar] [CrossRef]
- Nayeri, S.; Sargolzaei, M.; Abo-Ismail, M.K.; May, N.; Miller, S.P.; Schenkel, F.; Moore, S.S.; Stothard, P. Genome-wide association for milk production and female fertility traits in Canadian dairy Holstein cattle. BMC Genet. 2016, 17, 75. [Google Scholar] [CrossRef] [PubMed]
- Olsen, H.G.; Knutsen, T.M.; Kohler, A.; Svendsen, M.; Gidskehaug, L.; Grove, H.; Nome, T.; Sodeland, M.; Sundsaasen, K.K.; Kent, M.P.; et al. Genome-wide association mapping for milk fat composition and fine mapping of a QTL for de novo synthesis of milk fatty acids on bovine chromosome 13. Genet. Sel. Evol. 2017, 49, 20. [Google Scholar] [CrossRef] [PubMed]
- Li, C.; Sun, D.; Zhang, S.; Wang, S.; Wu, X.; Zhang, Q.; Liu, L.; Li, Y.; Qiao, L. Genome wide association study identifies 20 novel promising genes associated with milk fatty acid traits in Chinese Holstein. PLoS ONE 2014, 9, e96186. [Google Scholar] [CrossRef] [PubMed]
- Brøndum, R.F.; Su, G.; Janss, L.; Sahana, G.; Guldbrandtsen, B.; Boichard, D.; Lund, M.S. Quantitative trait loci markers derived from whole genome sequence data increases the reliability of genomic prediction. J. Dairy Sci. 2015, 98, 4107–4116. [Google Scholar] [CrossRef] [PubMed]
- Georgakopoulos-Soares, I.; Deng, C.; Agarwal, V.; Chan, C.S.Y.; Zhao, J.; Inoue, F.; Ahituv, N. Transcription factor binding site orientation and order are major drivers of gene regulatory activity. Nat. Commun. 2023, 14, 2333. [Google Scholar] [CrossRef] [PubMed]
- Calkhoven, C.F.; Ab, G. Multiple steps in the regulation of transcription-factor level and activity. Biochem. J. 1996, 317 Pt 2, 329–342. [Google Scholar] [CrossRef] [PubMed]
- Degtyareva, A.O.; Antontseva, E.V.; Merkulova, T.I. Regulatory SNPs: Altered Transcription Factor Binding Sites Implicated in Complex Traits and Diseases. Int. J. Mol. Sci. 2021, 22, 6454. [Google Scholar] [CrossRef] [PubMed]
- Zheng, W.; Zhao, H.; Mancera, E.; Steinmetz, L.M.; Snyder, M. Genetic analysis of variation in transcription factor binding in yeast. Nature 2010, 464, 1187–1191. [Google Scholar] [CrossRef] [PubMed]
- Huang, P.; Peslak, S.A.; Ren, R.; Khandros, E.; Qin, K.; Keller, C.A.; Giardine, B.; Bell, H.W.; Lan, X.; Sharma, M.; et al. HIC2 controls developmental hemoglobin switching by repressing BCL11A transcription. Nat. Genet. 2022, 54, 1417–1426. [Google Scholar] [CrossRef]
- Somasundaram, R.; Jensen, C.T.; Tingvall-Gustafsson, J.; Ahsberg, J.; Okuyama, K.; Prasad, M.; Hagman, J.R.; Wang, X.; Soneji, S.; Strid, T.; et al. EBF1 and PAX5 control pro-B cell expansion via opposing regulation of the Myc gene. Blood 2021, 137, 3037–3049. [Google Scholar] [CrossRef]
- Huerga Encabo, H.; Traveset, L.; Argilaguet, J.; Angulo, A.; Nistal-Villán, E.; Jaiswal, R.; Escalante, C.R.; Gekas, C.; Meyerhans, A.; Aramburu, J.; et al. The transcription factor NFAT5 limits infection-induced type I interferon responses. J. Exp. Med. 2020, 217, e20190449. [Google Scholar] [CrossRef]
- Grassmeyer, J.; Mukherjee, M.; DeRiso, J.; Hettinger, C.; Bailey, M.; Sinha, S.; Visvader, J.E.; Zhao, H.; Fogarty, E.; Surendran, K. Elf5 is a principal cell lineage specific transcription factor in the kidney that contributes to Aqp 2 and Avpr 2 gene expression. Dev. Biol. 2017, 424, 77–89. [Google Scholar] [CrossRef]
- Liu, Y.; Peng, L.; Chen, J.; Chen, L.; Wu, Y.; Cheng, M.; Chen, M.; Ye, X.; Jin, Y. EIF5A2 specifically regulates the transcription of aging-related genes in human neuroblastoma cells. BMC Geriatr. 2023, 23, 83. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Z.; Xiong, L.; Wu, Z.; Jiang, L.; Li, Y.; Li, Z.; Peng, Y.; Ning, K.; Zou, X.; Liu, Z.; et al. Nkx2.8 promotes chemosensitivity in bladder urothelial carcinoma via transcriptional repression of MDR1. Cell Death Dis. 2022, 13, 492. [Google Scholar] [CrossRef]
- Yu, C.; Liu, Z.; Chen, Q.; Li, Y.; Jiang, L.; Zhang, Z.; Zhou, F. Nkx2.8 Inhibits Epithelial–Mesenchymal Transition in Bladder Urothelial Carcinoma via Transcriptional Repression ofTwist1. Cancer Res. 2018, 78, 1241–1252. [Google Scholar] [CrossRef] [PubMed]
- Fang, X.; Cai, Y.; Liu, J.; Wang, Z.; Wu, Q.; Zhang, Z.; Yang, C.J.; Yuan, L.; Ouyang, G. Twist2 contributes to breast cancer progression by promoting an epithelial-mesenchymal transition and cancer stem-like cell self-renewal. Oncogene 2011, 30, 4707–4720. [Google Scholar] [CrossRef] [PubMed]
- Silva, P.R.B.; Weber, W.J.; Crooker, B.A.; Collier, R.J.; Thatcher, W.W.; Chebel, R.C. Hepatic mRNA expression for genes related to somatotropic axis, glucose and lipid metabolisms, and inflammatory response of periparturient dairy cows treated with recombinant bovine somatotropin. J. Dairy Sci. 2017, 100, 3983–3999. [Google Scholar] [CrossRef]
- Jiao, B.L.; Zhang, X.L.; Wang, S.H.; Wang, L.X.; Luo, Z.X.; Zhao, H.B.; Khatib, H.; Wang, X. MicroRNA-221 regulates proliferation of bovine mammary gland epithelial cells by targeting the STAT5a and IRS1 genes. J. Dairy Sci. 2019, 102, 426–435. [Google Scholar] [CrossRef] [PubMed]
- Li, S.S.; Loor, J.J.; Liu, H.Y.; Liu, L.; Hosseini, A.; Zhao, W.S.; Liu, J.X. Optimal ratios of essential amino acids stimulate β-casein synthesis via activation of the mammalian target of rapamycin signaling pathway in MAC-T cells and bovine mammary tissue explants. J. Dairy Sci. 2017, 100, 6676–6688. [Google Scholar] [CrossRef]
- Chiaruttini, C.; Guillier, M. On the role of mRNA secondary structure in bacterial translation. Wiley Interdiscip. Rev. RNA 2020, 11, e1579. [Google Scholar] [CrossRef]
- Faure, G.; Ogurtsov, A.Y.; Shabalina, S.A.; Koonin, E.V. Role of mRNA structure in the control of protein folding. Nucleic Acids Res. 2016, 44, 10898–10911. [Google Scholar] [CrossRef] [PubMed]
- Bucher, M.; Niebling, S.; Han, Y.; Molodenskiy, D.; Hassani Nia, F.; Kreienkamp, H.; Svergun, D.; Kim, E.; Kostyukova, A.S.; Kreutz, M.R.; et al. Autism-associated SHANK3 missense point mutations impact conformational fluctuations and protein turnover at synapses. eLife 2021, 10, e66165. [Google Scholar] [CrossRef] [PubMed]

| Gene | SNP Name | GenBank No. | Location | Genotype | Genotypic Frequency | Allele | Allelic Frequency |
|---|---|---|---|---|---|---|---|
| APOA5 | 15:g.27447741A>G | rs41755770 | 5′ regulatory region | AA | 0.0975 | A | 0.3173 |
| AG | 0.4396 | G | 0.6827 | ||||
| GG | 0.4629 | ||||||
| 15:g.27446527C>T | rs41755767 | 5′ regulatory region | CC | 0.1070 | C | 0.3332 | |
| CT | 0.4523 | T | 0.6668 | ||||
| TT | 0.4407 | ||||||
| 15:g.27445825T>C (missense mutation) | rs41755766 | exon 1 | CC | 0.4523 | C | 0.6748 | |
| CT | 0.4449 | T | 0.3252 | ||||
| TT | 0.1028 | ||||||
| AKT3 | 16:g.33367767T>C | rs208316642 | 5′ regulatory region | CC | 0.1631 | C | 0.4115 |
| CT | 0.4968 | T | 0.5885 | ||||
| TT | 0.3400 | ||||||
| 16:g.33417238C>T | rs41798799 | intron 1 | CC | 0.3612 | C | 0.5990 | |
| CT | 0.4756 | T | 0.4010 | ||||
| TT | 0.1631 | ||||||
| 16:g.33551706T>C | rs209739552 | intron 6 | CC | 0.0191 | C | 0.1563 | |
| CT | 0.2744 | T | 0.8438 | ||||
| TT | 0.7066 |
| Lactation | Haplotype Combination | Milk Yield (kg) | Fat Yield (kg) | Fat Percentage (%) | Protein Yield (kg) | Protein Percentage (%) |
|---|---|---|---|---|---|---|
| 1 | H1H1 (415) | 10,439 Aa ± 64.96 | 345.89 a ± 2.87 | 3.34 ± 0.027 | 308.56 a ± 2.09 | 2.97 ± 0.02 |
| H1H2 (405) | 10,295 ABb ± 63.30 | 341.90 ab ± 2.80 | 3.33 ± 0.026 | 304.47 b ± 2.04 | 2.97 ± 0.02 | |
| H2H2 (91) | 10,163 Bab ± 94.21 | 337.46 b ± 3.97 | 3.33 ± 0.038 | 301.73 b ± 2.89 | 2.99 ± 0.03 | |
| p | 0.0011 | 0.0259 | 0.9935 | 0.0044 | 0.7444 | |
| 2 | H1H1 (268) | 10,893 Aa ± 71.27 | 392.86 A ± 3.12 | 3.63 ± 0.029 | 321.60 A ± 2.27 | 2.96 ± 0.02 |
| H1H2 (279) | 10,649 Bab ± 67.19 | 382.75 B ± 2.97 | 3.61 ± 0.028 | 313.79 B ± 2.16 | 2.96 ± 0.02 | |
| H2H2 (64) | 10,591 ABb ± 110.92 | 385.54 AB ± 4.63 | 3.65 ± 0.045 | 315.26 AB ± 3.38 | 2.99 ± 0.03 | |
| p | 0.0005 | 0.0012 | 0.6972 | 0.0005 | 0.5671 |
| Gene | SNP Name | Allele | Transcription Factor | Relative Score (≥0.85) | Predicted Core Binding Site Sequence |
|---|---|---|---|---|---|
| APOA5 | 15:g.27447741A>G | A | NKX2.8 | 0.86 | GCACCTCAG |
| G | EBF1 | 0.87 | ACCCCAGGAA | ||
| ELF5 | 0.86 | CCCAGGAAGAGA | |||
| HIC2 | 0.88 | GTGCACCCC | |||
| 15:g.27446527C>T | C | TWIST2 | 0.86 | CAGAGCTGGG | |
| RHOXF1 | 0.88 | CAGAGCTG | |||
| T | - | - | - | ||
| AKT3 | 16:g.33367767T>C | T | - | - | - |
| C | NFAT5 | 0.87 | ATTTTCTTTT |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Guo, Z.; Du, A.; Han, B.; Li, H.; Tian, R.; Sun, W.; Zhao, G.; Tian, J.; Bao, X.; Zhang, J.; et al. Genetic Association of APOA5 and AKT3 Genes with Milk Production Traits in Chinese Holstein Cows. Agriculture 2024, 14, 869. https://doi.org/10.3390/agriculture14060869
Guo Z, Du A, Han B, Li H, Tian R, Sun W, Zhao G, Tian J, Bao X, Zhang J, et al. Genetic Association of APOA5 and AKT3 Genes with Milk Production Traits in Chinese Holstein Cows. Agriculture. 2024; 14(6):869. https://doi.org/10.3390/agriculture14060869
Chicago/Turabian StyleGuo, Zijiao, Aixia Du, Bo Han, Hui Li, Rugang Tian, Wei Sun, Gaoping Zhao, Jing Tian, Xiangnan Bao, Jixin Zhang, and et al. 2024. "Genetic Association of APOA5 and AKT3 Genes with Milk Production Traits in Chinese Holstein Cows" Agriculture 14, no. 6: 869. https://doi.org/10.3390/agriculture14060869





