Forms of Bacterial Survival in Model Biofilms
Abstract
1. Introduction
2. Materials and Methods
2.1. Bacterial Cultures
2.2. Production of Planktonic Populations
2.3. Preparation of Experimental Biofilms
2.4. Determination of the Number of Viable Cells in Biofilms
2.5. Determination of the Proportion of Thermoresistant Cells in Planktonic Cultures and Biofilms
2.6. Determination of the Number of Antibiotic Tolerant Cells in Biofilms
2.7. Scanning Electron Microscopy
2.8. Transmission Electron Microscopy
3. Results
3.1. Dynamics of Surviving and Thermoresistant Cells in Developing Planktonic Bacterial Cultures
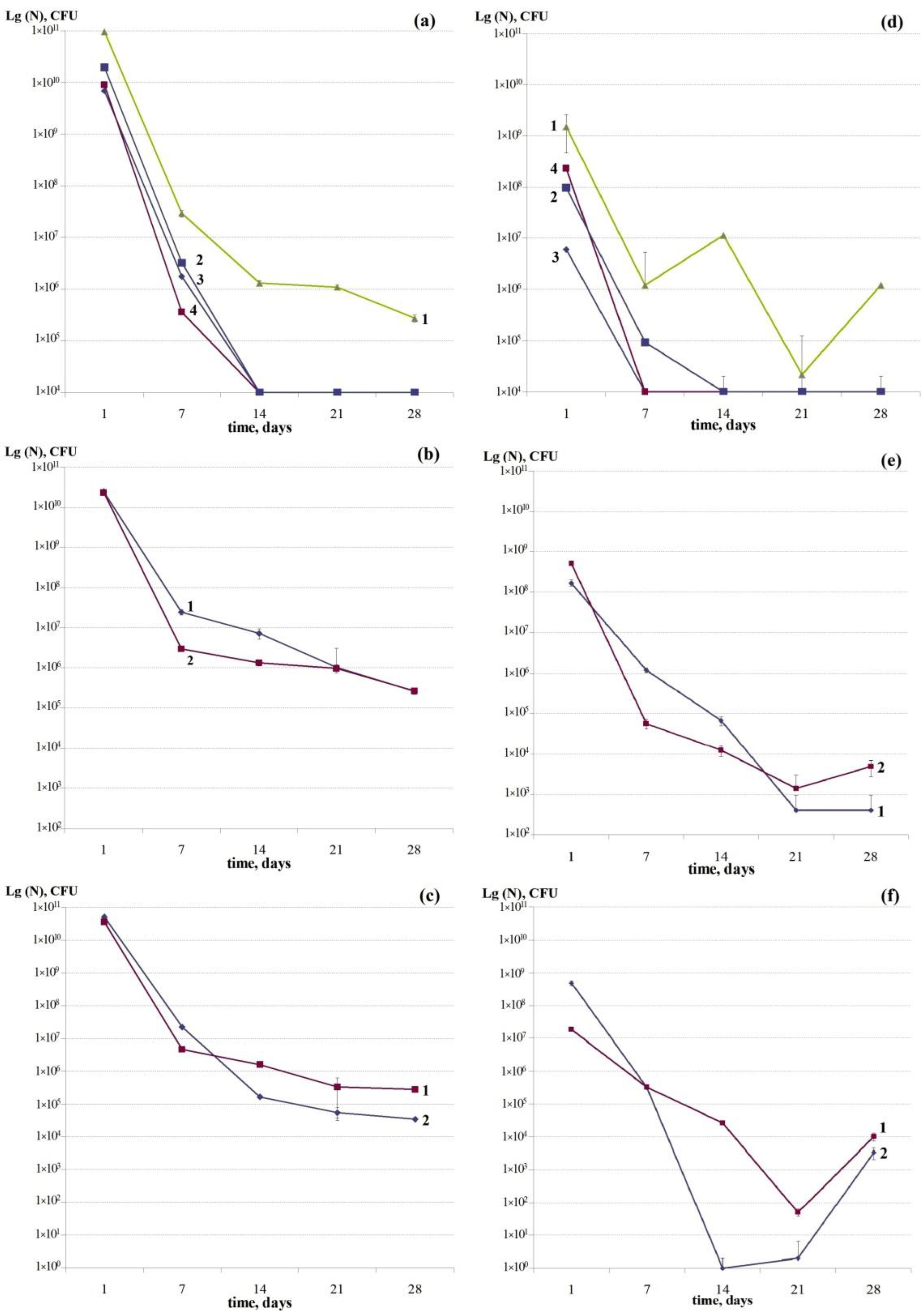
3.2. Dynamics of the Development of Mono-Species Biofilms of Bacteria under Study: Changes in the Number of Viable Cells, Antibiotic Tolerant Cells, and Thermoresistant Cells
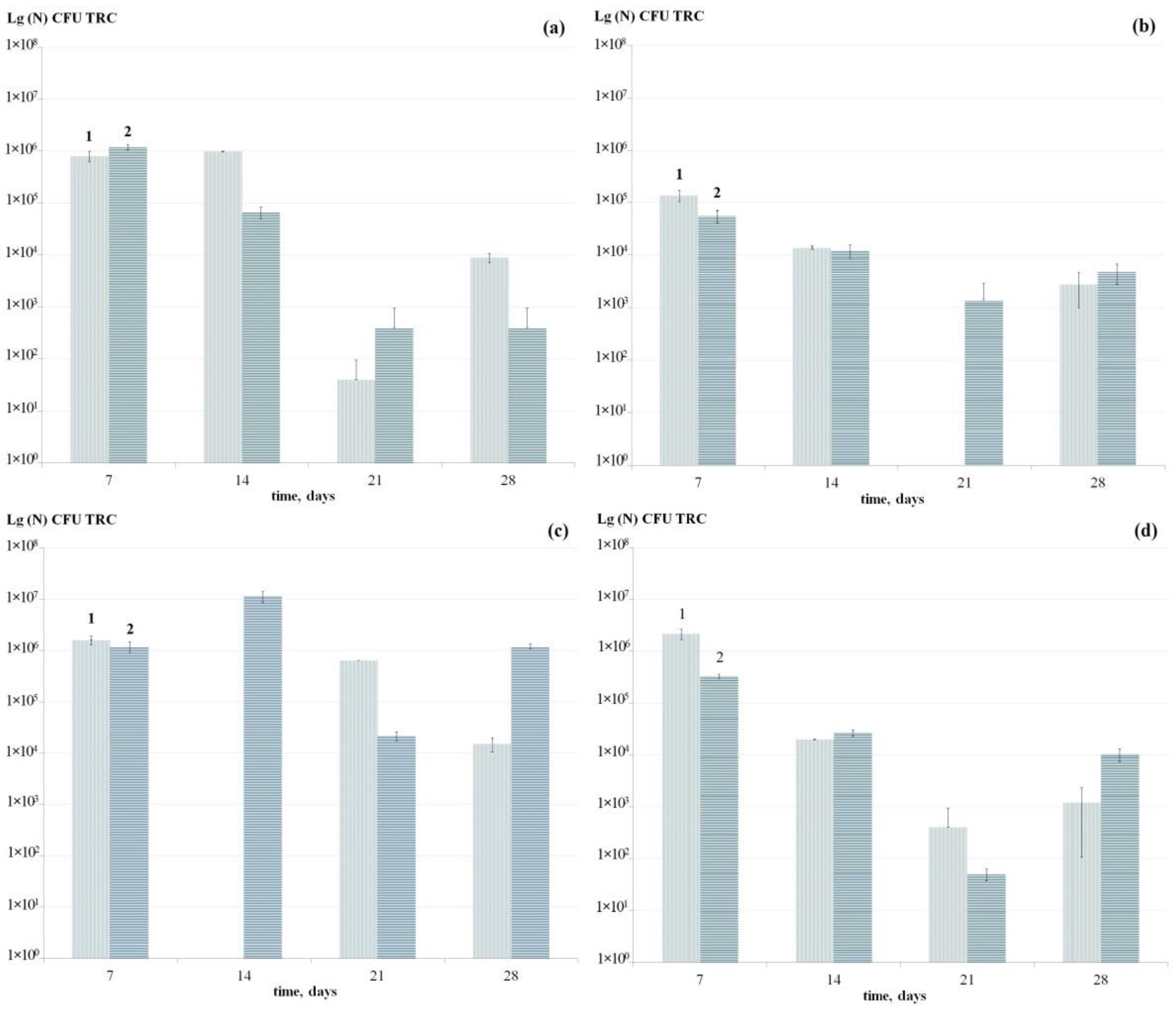
3.2.1. Dynamics of the Total Number of Cells of the Studied Bacteria
3.2.2. Dynamics of the Number of Antibiotic Tolerant Persister Cells Formed in Mono-Species Biofilms
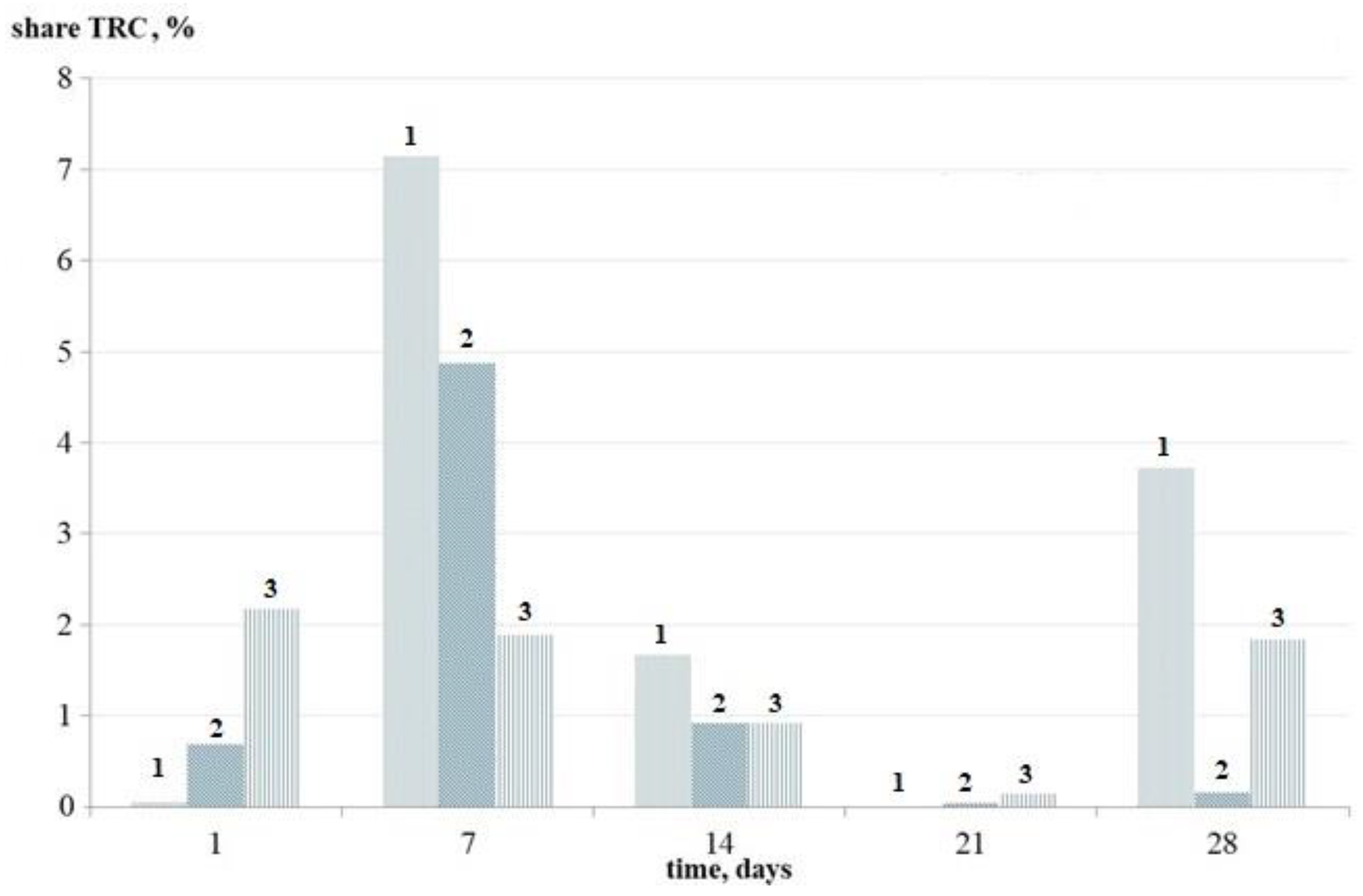
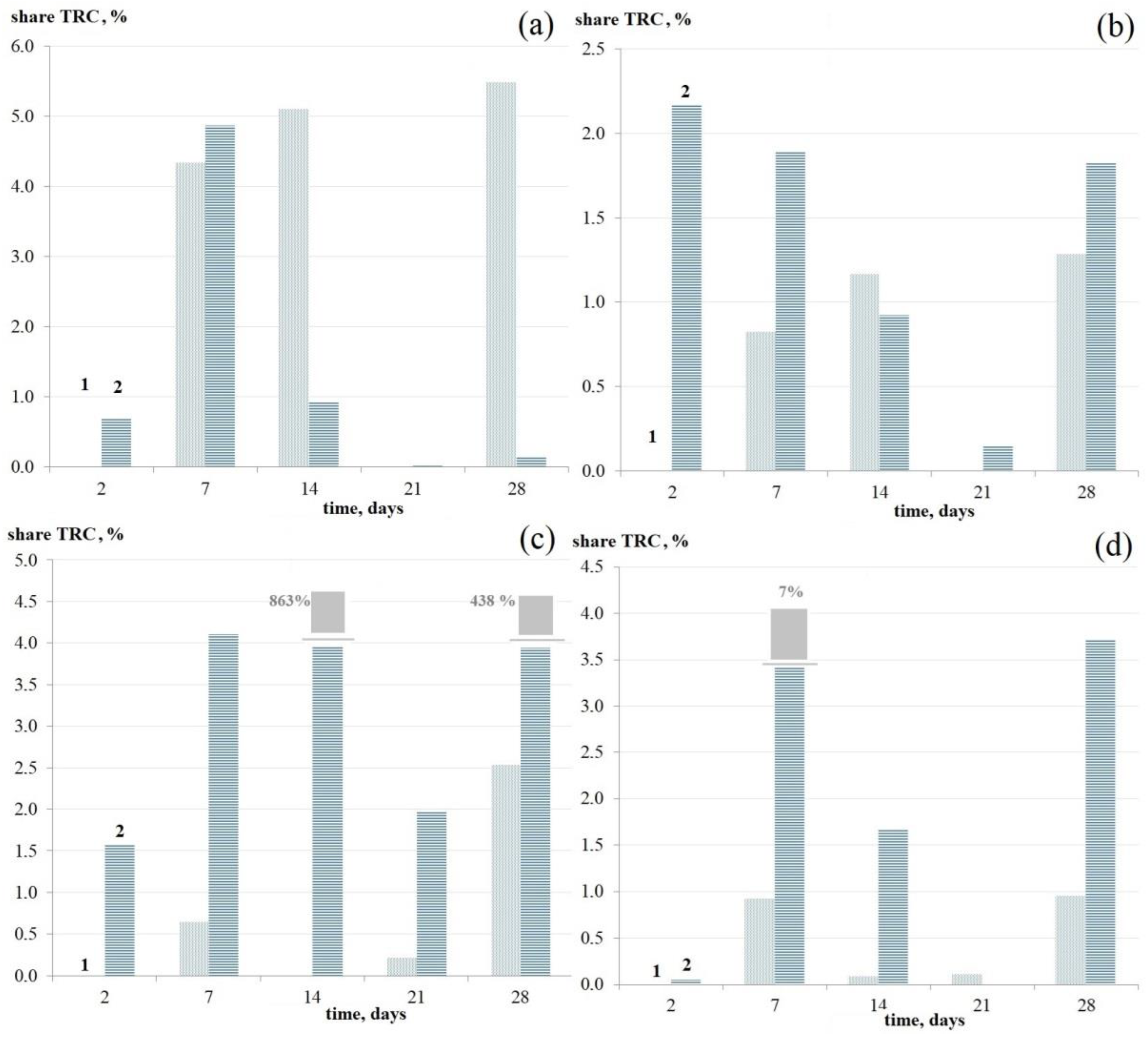
3.2.3. Dynamics of the Number of Thermoresistant Cells in Mono-Species Bacterial Biofilms
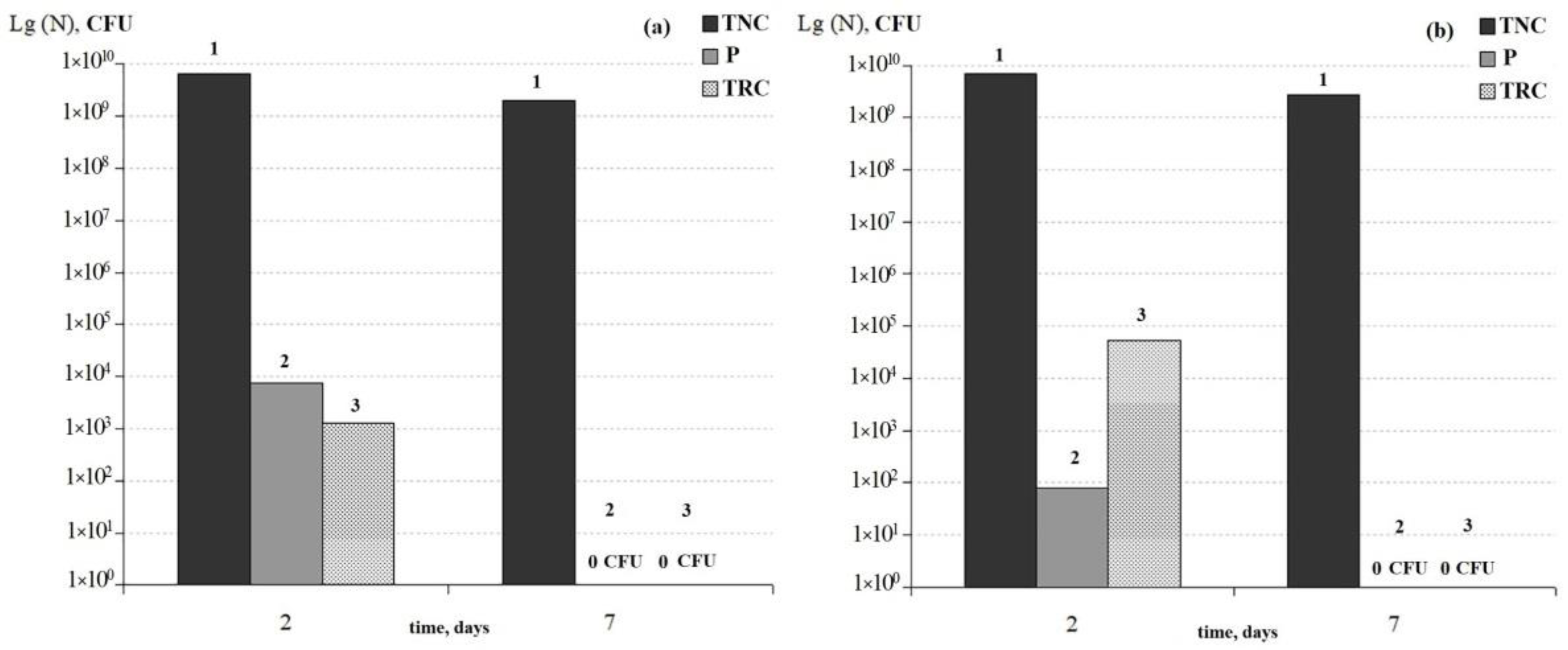
3.3. Comparison of the Dynamics of Thermoresistant Cell Formation in Planktonic and Biofilm Cultures
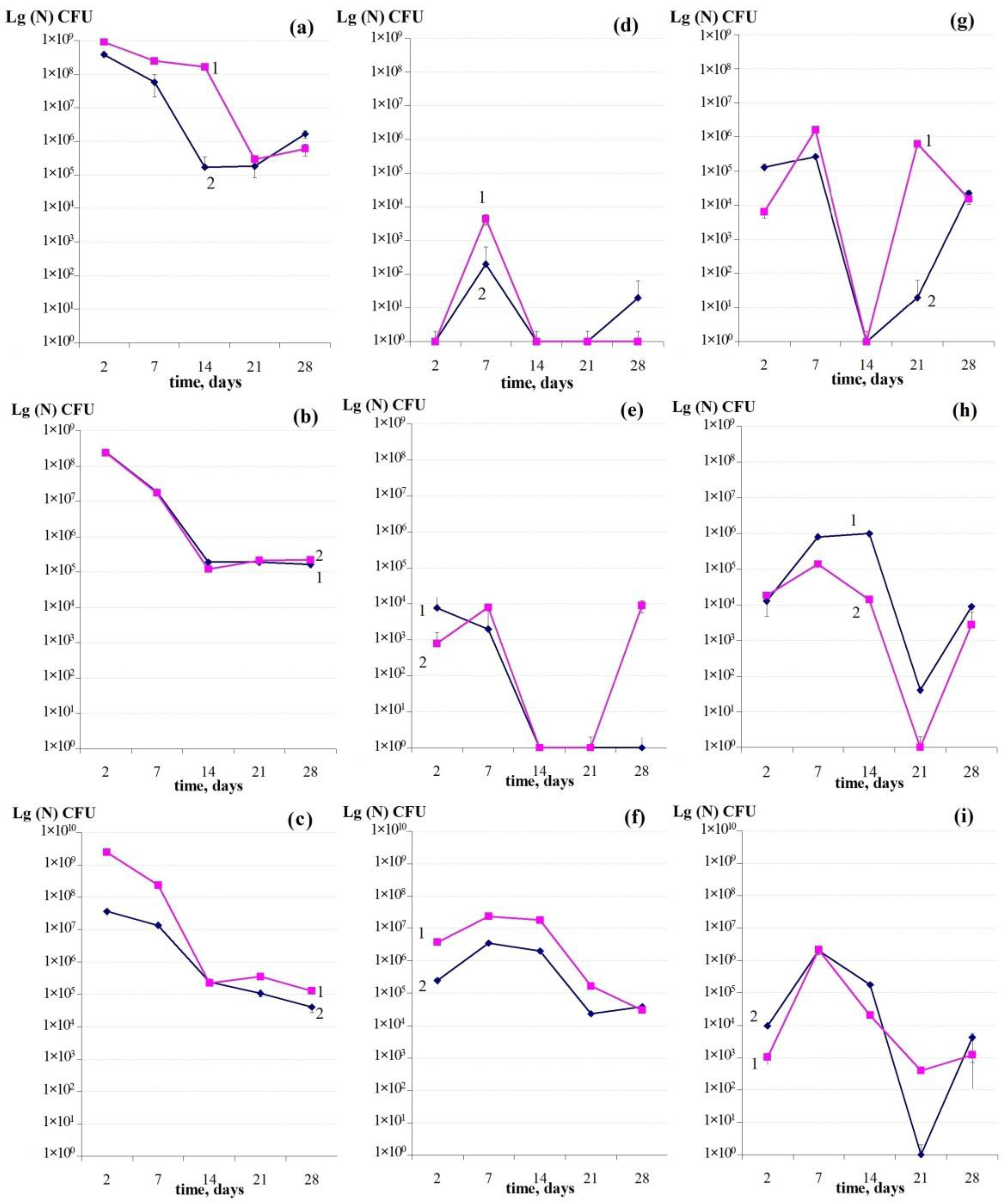
3.4. Binary Biofilm Development
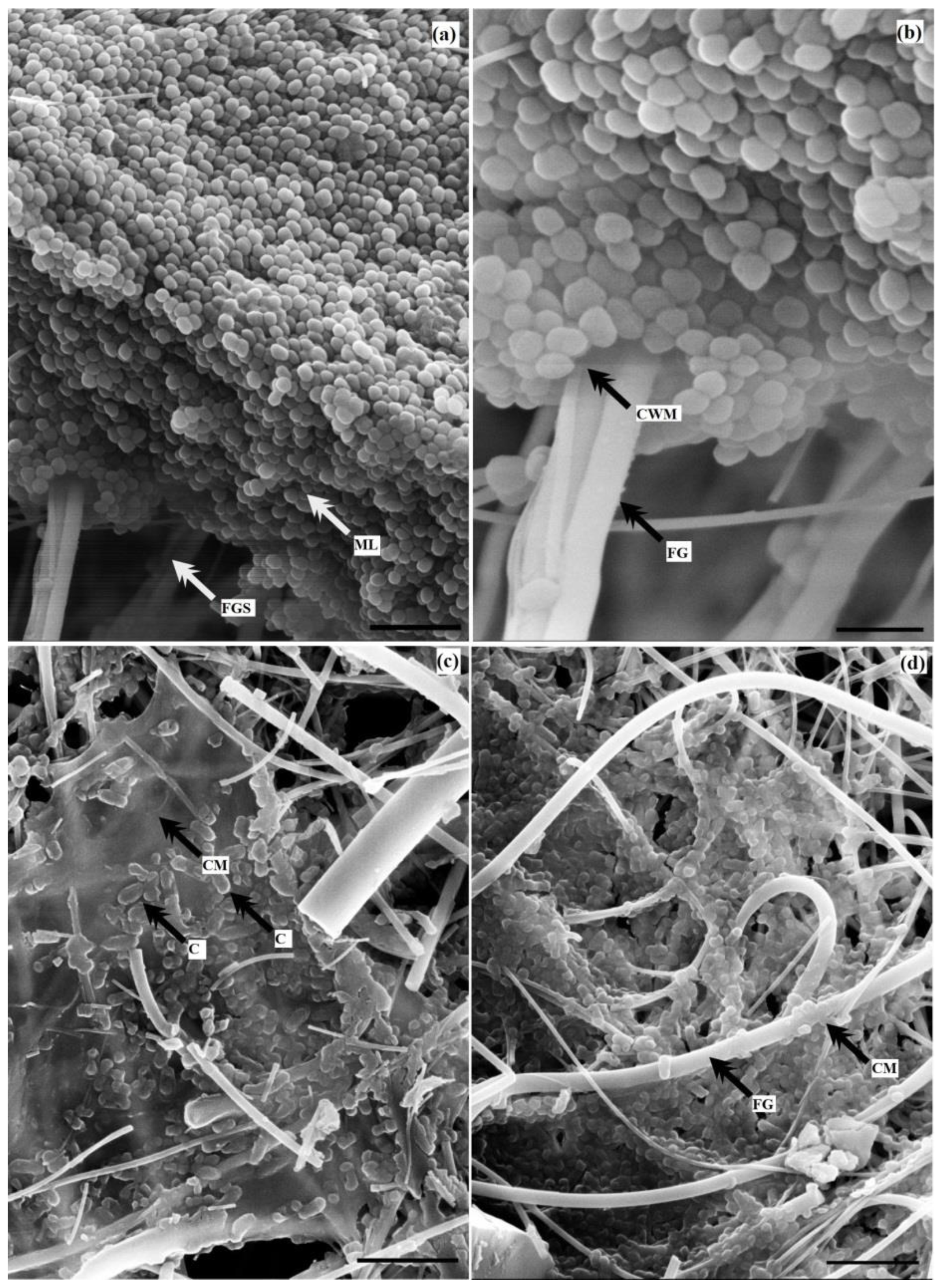
3.5. Electron Microscopic Studies of Mono-Species Biofilms
4. Discussion
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Satpathy, S.; Sen, S.K.; Pattanaik, S.; Raut, S. Review on bacterial biofilm: An universal cause of contamination. Biocatal. Agric. Biotechnol. 2016, 7, 56–66. [Google Scholar] [CrossRef]
- Plakunov, V.K.; Mart’yanov, S.V.; Teteneva, N.A.; Zhurina, M.V. Controlling of microbial biofilms formation: Anti- and probiofilm agents. Microbiology 2017, 86, 423–438. [Google Scholar] [CrossRef]
- Pandit, A.; Adholeya, A.; Cahill, D.; Brau, L.; Kochar, M. Microbial biofilms in nature: Unlocking their potential for agricultural applications. J. Appl. Microbiol. 2020, 129, 199–211. [Google Scholar] [CrossRef]
- González-Rivas, F.; Ripolles-Avila, C.; Fontecha-Umaña, F.; Ríos-Castillo, A.G.; Rodríguez-Jerez, J.J. Biofilms in the spotlight: Detection, quantification, and removal methods. Compr. Rev. Food Sci. Food Saf. 2018, 17, 1261–1276. [Google Scholar] [CrossRef]
- Sharahi, J.Y.; Azimi, T.; Shariati, A.; Safari, H.; Tehrani, M.K.; Hashemi, A. Advanced strategies for combating bacterial biofilms. J. Cell. Physiol. 2019, 234, 14689–14708. [Google Scholar] [CrossRef] [PubMed]
- Yuan, L.; Hansen, M.F.; Røder, H.L.; Wang, N.; Burmølle, M.; He, G. Mixed-species biofilms in the food industry: Current knowledge and novel control strategies. Crit. Rev. Food Sci. Nutr. 2020, 60, 2277–2293. [Google Scholar] [CrossRef] [PubMed]
- Yuan, L.; Wang, N.; Sadiq, F.A.; He, G. Interspecies interactions in dual-species biofilms formed by psychrotrophic bacteria and the tolerance of sessile communities to disinfectants. J. Food Prot. 2020, 83, 951–958. [Google Scholar] [CrossRef]
- Galié, S.; García-Gutiérrez, C.; Miguélez, E.M.; Villar, C.J.; Lombó, F. Biofilms in the food industry: Health aspects and control methods. Front. Microbiol. 2018, 9, 898. [Google Scholar] [CrossRef]
- Alvarez-Ordóñez, A.; Coughlan, L.M.; Briandet, R.; Cotter, P.D. Biofilms in food processing environments: Challenges and opportunities. Annu. Rev. Food. Sci. Technol. 2019, 10, 173–195. [Google Scholar] [CrossRef]
- Akinbobola, A.B.; Sherry, L.; Mckay, W.G.; Ramage, G.; Williams, C. Tolerance of Pseudomonas aeruginosa in in-vitro biofilms to high-level peracetic acid disinfection. J. Hosp. Infect. 2017, 97, 162–168. [Google Scholar] [CrossRef]
- Cai, L.; Li, Y.; Wang, H.; Xu, X.; Zhou, G. Biofilm formation by meat-borne Pseudomonas fluorescens on stainless steel and its resistance to disinfectants. Food Control. 2018, 91, 397–403. [Google Scholar] [CrossRef]
- Hall, C.W.; Mah, T.F. Molecular mechanisms of biofilm-based antibiotic resistance and tolerance in pathogenic bacteria. FEMS Microbiol. Rev. 2017, 41, 276–301. [Google Scholar] [CrossRef] [PubMed]
- Flemming, H.-C.; Wingender, J.; Szewzyk, U.; Steinberg, P.; Rice, S.A.; Kjelleberg, S. Biofilms: An emergent form of bacterial life. Nat. Rev. Microbiol. 2016, 14, 563–575. [Google Scholar] [CrossRef] [PubMed]
- Yuan, L.; Burmølle, M.; Sadiq, F.A.; Wang, N.; He, G. Interspecies variation in biofilm-forming capacity of psychrotrophic bacterial isolates from Chinese raw milk. Food Control 2018, 91, 47–57. [Google Scholar] [CrossRef]
- Stewart, P.; Franklin, M. Physiological heterogeneity in biofilms. Nat. Rev. Microbiol. 2008, 6, 199–210. [Google Scholar] [CrossRef] [PubMed]
- Wimpenny, J.; Manz, W.; Szewzyk, U. Heterogeneity in biofilms. FEMS Microbiol. Rev. 2000, 24, 661–671. [Google Scholar] [CrossRef]
- Zhurina, M.V.; Nikolaev, Y.A.; Plakunov, V.K. Role of the extracellular polymer matrix in azithromycin protection of Chromobacterium violaceum biofilms. Microbiology 2019, 88, 505–508. [Google Scholar] [CrossRef]
- Plakunov, V.K.; Nikolaev, Y.A.; Gannesen, A.V.; Chemaeva, D.S.; Zhurina, M.V. A new approach to detection of the protective effect of Escherichia coli on Gram-positive bacteria in binary biofilms in the presence of antibiotics. Microbiology 2019, 88, 275–281. [Google Scholar] [CrossRef]
- Bridier, A.; Sanchez-Vizuete, P.; Guilbaud, M.; Piard, J.; Naïtali, M. Biofilm-associated persistence of food-borne pathogens. Food Microbiol. 2015, 45, 167–178. [Google Scholar] [CrossRef]
- Wicaksono, W.A.; Erschen, S.; Krause, R.; Müller, H.; Cernava, T.; Berg, G. Enhanced survival of multi-species biofilms under stress is promoted by low-abundant but antimicrobial-resistant keystone species. J. Hazard Mater. 2022, 422, 126836. [Google Scholar] [CrossRef]
- Mirani, Z.A.; Urooj, S.; Zeesahn, F.; Khan, M.N.; Aziz, M.; Shaikh, I.A.; Khan, A.B. Role of phenotypic switching in stability and persistence of Pseudomonas aeruginosa biofilms. J. Microbiol. Infect. Dis. 2020, 10, 10–17. [Google Scholar] [CrossRef][Green Version]
- Mirani, Z.A.; Urooj, S.; Ullah, A.; Khan, M.N.; Rauf, N.; Mehmood, A.; Fenghuan, W.; Shaikh, I.A.; Khan, A.B. Phenotypic heterogeneity in biofilm consortia of E. coli. Microbiology 2021, 90, 237–246. [Google Scholar] [CrossRef]
- Nguyen, D.; Joshi-Datar, A.; Lepine, F.; Bauerle, E.; Olakanmi, O.; Beer, K.; McKay, G.; Siehnel, R.; Schafhauser, J.; Wang, Y.; et al. Active starvation responses mediate antibiotic tolerance in biofilms and nutrient-limited bacteria. Science 2011, 334, 982–986. [Google Scholar] [CrossRef] [PubMed]
- Miyaue, S.; Suzuki, E.; Komiyama, Y.; Kondo, Y.; Morikawa, M.; Maeda, S. Bacterial memory of persisters: Bacterial persister cells can retain their phenotype for days or weeks after withdrawal from colony-biofilm culture. Front. Microbiol. 2018, 9, 1396. [Google Scholar] [CrossRef] [PubMed]
- El-Registan, G.I.; Mulyukin, A.L.; Nikolaev, Y.A.; Suzina, N.E.; Gal’chenko, V.F.; Duda, V.I. Adaptogenic functions of extracellular autoregulators of microorganisms. Microbiology 2006, 75, 380–389. [Google Scholar] [CrossRef]
- Mulyukin, A.L.; Pogorelova, A.Y.; El’-Registan, G.I.; Suzina, N.E.; Duda, V.I.; Antonyuk, L.P. Diverse morphological types of dormant cells and conditions for their formation in Azospirillum brasilense. Microbiology 2009, 78, 33–41. [Google Scholar] [CrossRef]
- Mulyukin, A.L.; Suzina, N.E.; Mel’nikov, V.G.; Gal’chenko, V.F.; El’-Registan, G.I. Dormant state and phenotypic variability of Staphylococcus aureus and Corynebacterium pseudodiphtheriticum. Microbiology 2014, 83, 149–159. [Google Scholar] [CrossRef]
- Lewis, K. Persister cells. Annu. Rev. Microbiol. 2010, 64, 357–372. [Google Scholar] [CrossRef]
- Balaban, N.; Merrin, I.; Chait, R.; Kowalik, L.; Leibler, S. Bacterial persistence as a phenotypic switch. Science 2004, 305, 1622–1625. [Google Scholar] [CrossRef]
- Balaban, N.; Gerdes, K.; Lewis, K.; McKinney, J.D. A problem of persistence: Still more questions than answers? Nat. Rev. Microbial. 2013, 11, 587–591. [Google Scholar] [CrossRef]
- Kaplan, Y.; Reich, S.; Oster, E.; Maoz, S.; Levin-Reisman, I.; Ronin, I.; Gefen, O.; Agam, O.; Balaban, N.Q. Observation of universal ageing dynamics in antibiotic persistence. Nature 2021, 600, 290–294. [Google Scholar] [CrossRef]
- van den Bergh, B.; Fauvart, M.; Michiels, J. Formation, physiology, ecology, evolution and clinical importance of bacterial persisters. FEMS Microbiol. Rev. 2017, 41, 219–251. [Google Scholar] [CrossRef] [PubMed]
- Urbaniec, J.; Xu, Y.; Hu, Y.; Hingley-Wilson, S.; McFadden, J. Phenotypic heterogeneity in persisters: A novel ‘hunker’ theory of persistence. FEMS Microbiol. Rev. 2022, 46, fuab042. [Google Scholar] [CrossRef] [PubMed]
- Ayrapetyan, M.; Williams, T.C.; Oliver, J.D. Bridging the gap between viable but non-culturable and antibiotic persistent bacteria. Trends Microbiol. 2015, 23, 7–13. [Google Scholar] [CrossRef] [PubMed]
- Chen, H.; Green, A.; Martz, K.; Wu, X.; Alzahrani, A.; Warriner, K. The progress of type II persisters of Escherichia coli O157:H7 to a non-culturable state during prolonged exposure to antibiotic stress with revival being aided through acid-shock treatment and provision of methyl pyruvate. Can. J. Microbiol. 2021, 67, 518–528. [Google Scholar] [CrossRef]
- Kim, J.S.; Chowdhury, N.; Yamasaki, R.; Wood, T.K. Viable but non-culturable and persistence describe the same bacterial stress state. Environ Microbiol. 2018, 20, 2038–2048. [Google Scholar] [CrossRef]
- Song, S.; Wood, T.K. ‘Viable but non-culturable cells’ are dead. Environ. Microbiol. 2021, 23, 2335–2338. [Google Scholar] [CrossRef]
- Wang, J.; Wang, H.; Cai, K.; Yu, P.; Liu, Y.; Zhao, G.; Chen, R.; Xu, R.; Yu, M. Evaluation of three sample preparation methods for the identification of clinical strains by using two MALDI-TOF MS systems. J. Mass Spectrom. 2021, 56, e4696. [Google Scholar] [CrossRef]
- Bradley, D.; Jett, B.D.; Hatter, K.L.; Huycke, M.M.; Gilmore, M.S. Simplified agar plate method for quantifying viable bacteria. BioTechniques 1997, 23, 648–650. [Google Scholar] [CrossRef]
- Plakunov, V.K.; Mart’yanov, S.V.; Teteneva, N.A.; Teteneva, N.A.; Zhurina, M.V. A universal method for quantitative characterization of growth and metabolic activity of microbial biofilms in static models. Microbiology 2016, 85, 509–513. [Google Scholar] [CrossRef]
- Balaban, N.Q.; Helaine, S.; Lewis, K.; Ackermann, M.; Aldridge, B.; Andersson, D.I.; Brynildsen, M.P.; Bumann, D.; Camilli, A.; Collins, J.J.; et al. Definitions and guidelines for research on antibiotic persistence. Nat. Rev. Microbiol. 2019, 17, 441–448. [Google Scholar] [CrossRef] [PubMed]
- Nikolaev, Y.; Yushina, Y.; Mardanov, A.; Gruzdev, E.; Tikhonova, E.; El-Registan, G.; Beletskiy, A.; Semenova, A.; Zaiko, E.; Bataeva, D.; et al. Microbial biofilms at meat-processing plant as possible places of bacteria survival. Microorganisms 2022, 10, 1583. [Google Scholar] [CrossRef] [PubMed]
- Sussman, A.S.; Halvorson, H.O. Spores, Their Dormancy and Germination; Harper & Row: New York, NY, USA, 1966. [Google Scholar]
- Mulyukin, A.L.; Sorokin, V.V.; Loyko, N.G.; Suzina, N.E.; Duda, V.I.; Vorob’eva, E.A.; El’-Registan, G.I. Comparative study of the elemental composition of vegetative and dormant microbial cells. Microbiology 2002, 71, 37–48. [Google Scholar] [CrossRef]
- Nicholson, W.L.; Munakata, N.; Horneck, G.; Melosh, H.J.; Setlow, P. Resistance of Bacillus endospores to extreme terrestrial and extraterrestrial environments. Microbiol. Mol. Biol. Rev. 2000, 64, 548–572. [Google Scholar] [CrossRef] [PubMed]
- Nicholson, W.L.; Fajardo-Cavazos, P.; Rebeil, R.; Slieman, T.A.; Riesenman, P.J.; Law, J.F.; Xue, Y. Bacterial endospores and their significance in stress resistance. Antonie Van Leeuwenhoek 2002, 81, 27–32. [Google Scholar] [CrossRef]
- Parry, B.R.; Surovtsev, I.V.; Cabeen, M.T.; O’Hern, C.S.; Dufresne, E.R.; Jacobs-Wagner, C. The bacterial cytoplasm has glass-like properties and is fluidized by metabolic activity. Cell 2014, 156, 183–194. [Google Scholar] [CrossRef] [PubMed]
- Demkina, E.V.; Loiko, N.G.; Mulyukin, A.L.; Smirnova, T.A.; Gaponov, A.M.; Pisarev, V.M.; Tutel’yan, A.V.; Nikolaev, Y.A.; El’-Registan, G.I. Effect of inherent immunity factors on development of antibiotic tolerance and survival of bacterial populations under antibiotic attack. Microbiology 2015, 84, 764–774. [Google Scholar] [CrossRef]
- Nikolaev, Y.A.; Pankratov, T.A.; Gannesen, A.V.; Kolganova, T.V.; Suzina, N.E.; Demkina, E.V.; El’-Registan, G.I. Formation and properties of persister cells of Staphylococcus capitis and Staphylococcus epidermidis, bacteria inhabiting human skin. Microbiology 2020, 89, 425–434. [Google Scholar] [CrossRef]
- Lewis, K. Persister cells and the riddle of biofilm survival. Biochemistry 2005, 70, 267–274. [Google Scholar] [CrossRef]
- Zambrano, M.; Siegele, D.; Almiron, M.; Tormo, A.; Kolter, R. Microbial competition: Escherichia coli mutants that take over stationary phase cultures. Science 1993, 259, 1757–1760. [Google Scholar] [CrossRef]
- Kaldalu, N.; Hauryliuk, V.; Turnbull, K.J.; La Mensa, A.; Putrinš, M.; Tenson, T. In vitro studies of persister cells. Microbiol. Mol. Biol. Rev. 2020, 84, e00070-20. [Google Scholar] [CrossRef]
- Mulyukin, A.L.; Demkina, E.V.; Kryazhevskikh, N.A.; Suzina, N.E.; Vorob’Eva, L.I.; Duda, V.I.; Galchenko, V.F.; El-Registan, G.I. Dormant forms of Micrococcus luteus and Arthrobacter globiformis not platable on standard media. Microbiology 2009, 78, 407–418. [Google Scholar] [CrossRef]
- Chebotar’, I.V.; Emelyanova, M.A.; Bocharova, J.A.; Mayansky, N.A.; Kopantseva, E.E.; Mikhailovich, V.M. The classification of bacterial survival strategies in the presence of antimicrobials. Microb. Pathog. 2021, 155, e104901. [Google Scholar] [CrossRef] [PubMed]
- Gray, D.A.; Dugar, G.; Gamba, P.; Strahl, H.; Jonker, M.J.; Hamoen, L.W. Extreme slow growth as alternative strategy to survive deep starvation in bacteria. Nat. Commun. 2019, 10, 890. [Google Scholar] [CrossRef] [PubMed]
- Brauner, A.; Shoresh, N.; Fridman, O.; Balaban, N.Q. An experimental framework for quantifying bacterial tolerance. Biophys. J. 2017, 112, 2664–2671. [Google Scholar] [CrossRef] [PubMed]
- Colwell, R.R. Viable but nonculturable bacteria: A survival strategy. J. Infect. Chemother. 2000, 6, 121–125. [Google Scholar] [CrossRef] [PubMed]
- Kaprelyants, A.S.; Kell, D.B. Dormancy in stationary-phase cultures of Micrococcus luteus: Flow cytometric analysis of starvation and resuscitation. Appl. Environ. Microbiol. 1993, 59, 3187–3196. [Google Scholar] [CrossRef]
- Kaprelyants, A.S.; Mukamolova, G.V.; Davey, H.M.; Kell, D.B. Quantitative analysis of the physiological heterogeneity within starved cultures of Micrococcus luteus by flow cytometry and cell sorting. Appl. Environ. Microbiol. 1996, 62, 1311–1316. [Google Scholar] [CrossRef]
- Bamford, R.A.; Smith, A.; Metz, J.; Glover, G.; Titball, R.W.; Pagliara, S. Investigating the physiology of viable but non-culturable bacteria by microfluidics and time-lapse microscopy. BMC Biol. 2017, 15, 121. [Google Scholar] [CrossRef]
- Minsky, A.; Shimoni, E.; Frenkiel-Krispin, D. Stress, order and survival. Nat. Rev. Mol. Cell Biol. 2002, 3, 50–60. [Google Scholar] [CrossRef]
- Lennon, J.T.; Jones, S.E. Microbial seed banks: The ecological and evolutionary implications of dormancy. Nat. Rev. Microbiol. 2011, 9, 119–130. [Google Scholar] [CrossRef] [PubMed]
- Hunter, G.L.; Weeks, E.R. The physics of the colloidal glass transition. Rep. Prog. Phys. 2012, 75, e066501. [Google Scholar] [CrossRef] [PubMed]
- Wood, T.K.; Knabel, S.J.; Kwan, B.W. Bacterial persister cell formation and dormancy. Appl. Environ. Microbiol. 2013, 79, 7116–7121. [Google Scholar] [CrossRef] [PubMed]

| Strain | 18 h | 7 Day | 14 Day | 21 Day | 28 Day |
|---|---|---|---|---|---|
| Pseudomonas aeruginosa S481 | 1.56 | 4.11 | 863.64 | 1.96 | 437.96 |
| Pseudomonas extremorientalis S85 | 0.49 | 2.90 | 0.00 | 0.00 | 0.00 |
| Escherichia coli K12 | 0.69 | 4.88 | 0.92 | 0.04 | 0.16 |
| Salmonella typhimurium TA 1535 | 2.17 | 1.89 | 0.92 | 0.15 | 1.83 |
| Kocuria rhizophila S155 | 0.95 | 1.44 | 0.00 | 0.00 | 10.00 |
| Staphylococcus aureus ATCC6538 | 0.05 | 7.15 | 1.67 | 0.02 | 3.72 |
| Test Culture | Correlation Coefficient (Cp) | ||
|---|---|---|---|
| Strain | TNC/TRC * | TNC/P | TRC/P |
| Pseudomonas aeruginosa S481 | −0.19 | −0.08 | 0.92 |
| Pseudomonas extremorientalis S85 | 0.37 | −0.10 | 0.88 |
| Escherichia coli K12 | −0.32 | 0.98 | −0.28 |
| Salmonella typhimurium TA 1535 | −0.10 | −0.33 | 0.50 |
| Staphylococcus aureus ATCC6538 | −0.18 | −0.17 | 0.75 |
| Kocuria rhizophila S155 | −0.02 | 0.19 | 0.88 |
| Variants of Combinations | P, %, 7th Day | P, %, 14th Day |
|---|---|---|
| Mono-biofilm E. coli K12 | 0.01 | 0 |
| Binary biofilm E. coli K12 | 0.46 | 0.1 |
| Mono-biofilm K. rhizophila S155 | 25.97 | 83.47 |
| Binary biofilm K. rhizophila S155 | 8.15 | 4.76 |
| Mono-biofilm S. typhimurium TA1535 | 0.05 | 0 |
| Binary biofilm S. typhimurium TA1535 | 0.1 | 0 |
| Mono-biofilm S. aureus ATCC6538 | 10.25 | 81.08 |
| Binary biofilm S. aureus ATCC6538 | 3.67 | 20.42 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Pankratov, T.A.; Nikolaev, Y.A.; Yushina, Y.K.; Tikhonova, E.N.; El-Registan, G.I. Forms of Bacterial Survival in Model Biofilms. Coatings 2022, 12, 1913. https://doi.org/10.3390/coatings12121913
Pankratov TA, Nikolaev YA, Yushina YK, Tikhonova EN, El-Registan GI. Forms of Bacterial Survival in Model Biofilms. Coatings. 2022; 12(12):1913. https://doi.org/10.3390/coatings12121913
Chicago/Turabian StylePankratov, Timofei A., Yuri A. Nikolaev, Yulia K. Yushina, Ekaterina N. Tikhonova, and Galina I. El-Registan. 2022. "Forms of Bacterial Survival in Model Biofilms" Coatings 12, no. 12: 1913. https://doi.org/10.3390/coatings12121913
APA StylePankratov, T. A., Nikolaev, Y. A., Yushina, Y. K., Tikhonova, E. N., & El-Registan, G. I. (2022). Forms of Bacterial Survival in Model Biofilms. Coatings, 12(12), 1913. https://doi.org/10.3390/coatings12121913






