1. Introduction
Phospholipase is a class of enzymes responsible for the metabolism and synthesis of phospholipids in plants. They can catalyze the hydrolysis of phospholipids. According to the position where phospholipases hydrolyze phospholipids, they can be divided into five types: phospholipase A1, phospholipase A2, phospholipase B, phospholipase C, and phospholipase D [
1]. The most important kind of phospholipase in plants is phospholipase D, which can hydrolyze phospholipids to generate phosphatidic acid and free alcohol groups and is believed to play an important role in many cellular processes. In plants, PLD is involved in some physiological processes, such as cell membrane degradation, signal transduction, vesicle transport, and membrane metabolism during senescence, and abiotic stress damage during seed germination [
2,
3], including stomatal closure [
4], root elongation [
5], cold, drought and salt stress [
6,
7,
8]. The function of PLD is usually achieved through the enzymatic hydrolysis product of PA. PA is considered to be a ubiquitous lipid signal molecule, which keeps the cell membrane in a stable state and participates in regulating the dynamic balance of cell membrane lipid composition. Plant PLD is a complex gene family. As early as 1940, PLD was identified as an enzyme involved in lipid metabolism and membrane remodeling [
9]. Until 1994, the first PLD encoding cDNA was isolated from the castor bean [
10]. At present, the PLD gene family has been found in
Arabidopsis thaliana [
11],
Oryza sativa [
12],
Populus [
13],
Vitis vinifera [
13],
Cicer arietinum [
14],
Camellia sinensis [
15], and other plant species.
In dicotyledons and monocotyledons, the PLD family can be further divided into two subfamilies according to the composition of their N-terminal phospholipid-binding domains, namely, C2-PLD and PXPH-PLD subfamilies. The N-terminus of the C2-PLD subfamily contains the C2 domain, while the N-terminal of the PLD of the PXPH-PLD subfamily contains both Phox homology (Px) domain and Pleckstrin homology (PH) domain [
16]. All PLD genes include two HxKxxxxD motifs (motifs), namely HKD1 and HKD2 and C2 conserved domain at N-terminal. HKD is histidine (His), lysine (Lys) and aspartic acid (Asp) [
17]. These two motifs are the marker sequences of PLD. The HKD domain endows the enzyme with hydrolytic activity, and PLD in most plants contains a C2 domain in order to better bind to the cofactor Ca
2+ and enhance the affinity of the enzyme to its substrate [
18]. The difference is that in Arabidopsis and rice, it is found that some C2 domains in the PLD gene family are replaced by tandem PX-PH domains [
19]. The PX and PH domains of PXPH-PLD have shown a membrane targeting effect, which is closely related to the inositol polyphosphate signal [
20]. Protein domain analysis shows that in addition to C2-PLD and PXPH-PLD, there is a new subfamily in rice-SP-PLD. SP-PLD contains signal peptides at the N-terminal. Instead of C2 or PXPH domain [
12], its specific cellular localization may be related to plant growth and defense. The activity of PLD will increase under stress, and PA, the product of PLD, also plays an important role in the growth and development of plants [
21].
At present, the PLD gene has been found in many plants, 12 PLD genes have been identified in
Arabidopsis thaliana [
22], 17 PLD genes in
Oryza sativa [
12], 18 PLD genes in
Glycine max [
19], and 13 PLD genes in
Zea mays [
2]. According to the sequence characteristics, the PLD gene family can be divided into seven subtypes: PLDα, PLDβ, PLDγ, PLDδ, PLDε, PLDζ, and PLDφ, of which PLDα is the most common. In
Arabidopsis thaliana, PLDα1 is the most abundant PLD, which has been found to regulate salt tolerance and permeability, abscisic acid signal, and seed aging [
23]. PLDα1 is mainly responsible for the activity of PLDs [
24]. The study found that in Arabidopsis flowers, pollen tubes, pods, seeds, stems, leaves, roots, and other tissues, except for pollen tubes, AtPLDα1 showed higher expression levels than in other tissues. AtPLDα2, AtPLDβ1, AtPLDβ2, and AtPLDδ were highly expressed in pollen tubes, but low in other tissues. AtPLDγ2 was evenly expressed and not high in these tissues, which indicates the function of different PLD genes. The expression of some PLD genes will change only when they are stimulated by external stimuli [
25].
In this study, we identified the potato PLD gene family and obtained 16 potato PLD genes. The physical and chemical properties, gene structure, conservative motif, chromosome location, gene replication, phylogenetic evolution, cis-regulatory element prediction, GO annotation, interspecies collinearity, and qRT-PCR analysis of potato StPLD protein were studied by bioinformatics analysis, which provided some clues for further revealing the function of potato PLD gene family.
3. Results
3.1. Identification of Members of PLD Gene Family in Potato
Using the PLD gene family protein sequences of
Arabidopsis thaliana and rice, a Blastp search was carried out in the potato genome database and then screened by HMMER software according to the PLD gene domain (PF00614). Then the selected protein sequences were compared by SMART and Pfam, and the genes without the HxKxxxxD domain and only one HxKxxxxD domain were removed. Sixteen PLD genes were identified from the potato genome (
Table 1). Finally, in order to verify the correctness of the initial identification results, the InterproScan program was used to further screen to confirm the existence of two conserved HKD domains. The size of the ORF for StPLD proteins varied from 2268 bp to 3327 bp. The lengths of the proteins ranged from 755 to 1108 amino acids, and they possessed 86.96 kDa to 126.22 kDa molecular masses and predicted pI values of 5.40 to 8.52.
Combined with the classification of PLD gene families identified in
Arabidopsis thaliana and rice, the PLD genes in three plants are divided into six subfamilies: α, β/γ, δ, ζ, φ, κ. The subfamilies φ and κ only exist in rice, while PLD genes in potato and Arabidopsis are distributed in the other four families, and 16 PLD genes in potato are distributed in α, β, δ, ζ subfamilies. The number of genes was 6, 3, 5, and 2, respectively. The member domain of α, β, and δ family was C2, and the member domain of subfamily ζ was PH-PX (
Figure 1). The genetic relationship of PLD ζ was far from that of other groups, and the gene family distance of PLD φ was farther than that of other groups. The PLD family genes of potato, Arabidopsis and rice are mainly distributed in α, β and δ subfamilies, indicating that the three genes have high homology and may be similar in function. In addition, the gene distribution of potato PLD family was uneven, mainly in α and δ subfamilies, indicating that potato PLD gene family may have carried out genome replication in the process of evolution, so that the number and structure of genes among subgroups were different. By comparing the PLD protein sequence of potato and Arabidopsis, it is found that each gene contains HxKxxxxD conservative motifs at both ends of the sequence, which can indicate that the PLD protein is highly conserved among different species (
Figure 2).
3.2. Chromosomal Location and Gene Duplication of PLD Gene Family in Potato
According to the chromosome mapping, 16 potato PLD genes were randomly and unevenly distributed on chromosomes 1, 2, 3, 4, 6, 8, 10, and 12. There was only one potato PLD gene on chromosomes 4, 6, and 12, and four PLD genes on chromosome 1, which was the chromosome with the most PLD genes, followed by chromosome 8 with three PLD genes and two PLD genes on chromosomes 2, 3, and 10 (
Figure 3). There are many mechanisms of gene family amplification, including polyploidy, fragment replication, tandem replication, transposable elements, and so on [
33]. To study the genomic replication event of the potato PLD gene, according to the defined standard, two tandem repeat genes were found on chromosome 8 (StPLDα4/α5).
3.3. Gene Structure and Cis-Acting Elements Analysis and Conserved Motif Identification
In order to further study the conservation of potato PLD protein sequence and the difference in motif composition between potato proteins, MEME was used to further analyze the conservative motif of potato PLD protein sequence. Five conserved motifs of potato PLD were identified, which were between 34 and 50 amino acids in length (
Figure 4A,
Table 2). The results showed that the motifs of PLDα, PLDβ, and PLDδ subfamily genes were completely consistent, and they all contain five conserved motifs. The motif composition of PLDζ subfamily genes was also completely the same, all of which contained three conserved motifs. According to the results of Pfam website, Motif1 belongs to PLDc domain, which can also be called HKD domain (PF00614), Motif5 belongs to PLD_C (PF12357.8) domain, and other conserved motifs are predicted, such as Motif3 belongs to Pilin_GH domain (PF16734). All StPLD contained the HKD domain, C2-PLD had five motifs, PXPH-PLD had three motifs, in addition to StPLDζ1 and StPLDζ2, all contained Motif 2, Motif5; the deletion of two motifs of the ζ subfamily protein makes the domain of the ζ subfamily different from the other three subfamilies. Motif1, Motif3, and Motif4 were all distributed in the four subfamilies, which indicates that the StPLD protein sequence is conserved.
According to the structure of StPLD gene, the number of exons in the StPLD gene family members ranged from 3 to 20, and all genes had introns (
Figure 4B). In addition, it was also found that the gene structures of the same subfamily were similar. There were 10 exons and 9 introns in the subfamily PLDδ1~δ5, and 20 exons and 19 introns in both PLDζ1 and PLDζ2. The gene structure was completely the same. The number of exons in each gene in the same subfamily was basically the same, and there were great differences in the number of exons and introns among different subfamilies, indicating that the structure of StPLD gene is more complex.
In order to further study the regulation mechanism of StPLD gene under abiotic stress, according to the different functions of different cis-acting elements, 13 cis-acting elements related to growth and development, hormones and stress response were screened (
Figure 5). The promoter of StPLD gene contained many homeopathic regulatory elements, the most StPLDα3 had 16 cis-acting elements, and the least StPLDδ1 had three cis-acting elements. There are four stress response elements, namely drought-responsive (MBS), low temperature (LTR), anaerobic induction (ARE), and light response (G-box) elements, and five hormone response elements, namely abscisic acid (ABRE), methyl jasmonate (CGTCA-motif, TGACG-motif), salicylic acid (TCA-element), auxin (TGA-element), gibberellin (TATC-box, P-box). Two growth and development response elements were meristem expression regulation (CAT-box) and circadian rhythm regulation (circadian) (
Figure 5). Among the StPLD genes, G-box, ABRE, CGTCA-motif, and these three acting elements were the most, 28, 30, and 27, respectively, which were randomly distributed in each gene, indicating that these genes can participate in plant light response and respond to hormones, such as abscisic acid and methyl jasmonate. Cis-acting element analysis showed that StPLD gene was closely related to abiotic stress, growth and development and hormones in plants [
34].
3.4. Gene Ontology Annotations of StPLD Proteins
In order to further study the biological process of StPLD protein, GO annotation was carried out by Blast2GO software, and the role of genes in the biological process (BP), molecular function (MF), and cellular component (CC) was analyzed. Through GO annotation analysis, it was found that all 16 StPLD proteins participated in the process of phospholipid catabolism, and all had phospholipase D activity (
Figure 6).
In the biological process, 16 proteins were involved in the process of phospholipid catabolism. Except for StPLDζ1 and StPLDζ2, the other 14 proteins were involved in the metabolism of phosphatidylcholine. In addition, only the two proteins, StPLDζ1 and StPLDζ2, were involved in phosphatidic acid biosynthesis and inositol lipid-mediated signal transduction, and only the StPLDα6 protein was involved in water stress, salt stress, abscisic acid stress, and membrane lipid catabolism. Only the StPLDβ2 protein was involved in transcriptional regulation using DNA as a template. In molecular function, 16 StPLD proteins have phospholipase D activity, except StPLDζ1 and StPLDζ2, the other 14 proteins had a calcium ion binding function, which were distributed in StPLDα, StPLDβ, and StPLDδ, indicating that the C2 domain can better bind to cofactor Ca2+ and enhance the affinity of the enzyme to its substrate. Among the cellular components, 16 StPLD proteins were located in the plasma membrane, and only StPLDβ2 protein was located in the nucleus.
3.5. Collinear Analysis of PLD Gene in Potato and Arabidopsis thaliana
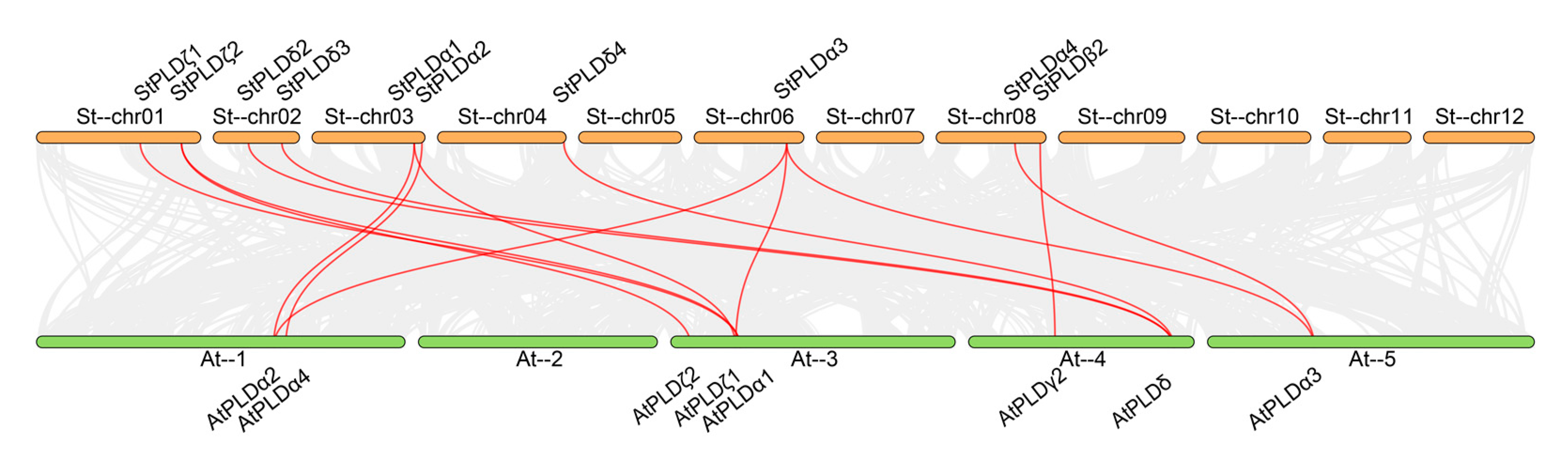
From the collinearity map of potato PLD and Arabidopsis PLD genes, it is known that there are 14 homologous gene pairs, including 14 potato PLD genes and 14 Arabidopsis PLD genes, which are distributed in four subfamilies: α, β, δ and ζ (
Figure 7) indicating that there is a close homologous evolution relationship between potato and Arabidopsis PLD gene family. Among them, StPLDα1, StPLDα3, and StPLDζ2 are collinear with at least two AtPLD genes, indicating that these genes may have similar functions and play an important role in the evolution of potato and Arabidopsis PLD gene family. We found that there was no collinear relationship between StPLDα5, StPLDα6, StPLDβ1, StPLDβ2, StPLDδ1, StPLDδ5, and the AtPLD gene, indicating that these genes may be specific to potato evolution. Among these genes, there was only one single gene pair, StPLDβ2 and AtPLDγ2, indicating that this pair of genes have a common origin before potato and
Arabidopsis thaliana gene differentiation.
3.6. Tissue Expression and Stress Treatment Expression Analysis of the Potato PLD Genes
In order to study the function of potato PLD gene in the process of growth and development, we obtained transcriptome data from potato genome database, found out the RNA-seq data of PLD gene in different tissues and under various stresses and drew heat maps (
Table A3 and
Table A4). The expression level of StPLD gene in leaves, roots, shoots, callus, stolons, tubers, flowers, petioles, petals, stamens, carpels, and sepals (
Figure 8A) and the expression levels of salt, mannitol, heat, Phytophthora infestans, β-aminobutyric acid, benzothiadiazole, abscisic acid, auxin, gibberellin, and benzo (a) pyrene (
Figure 8B) were revealed.
The results showed that StPLDα2, StPLDα6, and StPLDζ2 were not detected in all tissues, and StPLDα4, StPLDβ1, StPLDδ3, and StPLDζ1 were expressed in all tissues. Among them, StPLDα5 was expressed in stamens at a higher level, showing tissue-specific expression, StPLDα4, StPLDα5, StPLDβ1, StPLDδ3, StPLDζ1 were mainly expressed in roots and leaves, indicating that these genes may play a specific regulatory role in the development of roots and leaves. The expression of StPLDδ3 was significantly increased in petals, carpels, and sepals, and the expression of StPLDδ4 was significantly increased in stamens. Only StPLDα5 was not detected under P. infestans stress, while the other genes were upregulated under these 10 stresses, indicating that PLD gene family plays an important role in the regulation of potato growth and development. The gene expression of StPLDα5 increased significantly under NaCl, mannitol and 35 °C heat stress, indicating that StPLDα5 was involved in salt tolerance and drought resistance of potato, and was related to the heat tolerance mechanism of potato. StPLDα5 increased significantly under ABA hormone treatment.
3.7. Expression Analysis of StPLD Genes in Different Treatments
Based on the above bioinformatics analysis, the response of the potato PLD gene to biotic stress and abiotic stress was further studied. Potato Desiree materials were treated with NaCl, high temperature, drought, and ABA, and then the expression of StPLD gene was detected by real-time quantitative qRT-PCR to analyze the expression of 16 StPLD genes under NaCl, high temperature, drought, and ABA stress (
Figure 9). The results showed that the expression levels of 16 genes changed under the four treatments. Among them, the expression levels of the six genes, StPLDα1, StPLDα2, StPLDδ1, StPLDδ4, StPLDδ5, and StPLDζ2, were upregulated under NaCl, high temperature, drought, and ABA stress. The expression levels of the remaining genes were both upregulated and downregulated under these four treatments.
Under NaCl treatment, StPLDα1, StPLDα4, StPLDα5, StPLDδ1, StPLDδ2, StPLDδ3, StPLDδ4, StPLDδ5, StPLDζ2 were significantly upregulated, StPLDβ2, StPLDβ3 were downregulated, and the remaining genes were upregulated but not significantly (
Figure 9A), Under high temperature treatment, StPLDα1, StPLDα2, StPLDα4, StPLDα6, StPLDβ3, StPLDδ1, StPLDδ2, StPLDδ4, StPLDδ5, StPLDζ2 were significantly upregulated, and StPLDζ1 was significantly downregulated. (
Figure 9B), Under drought stress, StPLDα1, StPLDα2, StPLDα5, StPLDδ1, StPLDδ3, StPLDδ4, StPLDδ5, StPLDζ2 were significantly upregulated, and StPLDζ1 was significantly downregulated. (
Figure 9C). Under ABA treatment, StPLDα1, StPLDα2, StPLDβ3, StPLDδ1, StPLDδ4, StPLDδ5, StPLDζ2 were significantly upregulated, and StPLDα5, StPLDβ1, StPLDβ2, StPLDδ3, StPLDζ1 were significantly downregulated.(
Figure 9D).
Under NaCl treatment, the qRT-PCR results in the PLDα subfamily, StPLDδ subfamily, and StPLDζ subfamily are consistent with the results in the RNA-seq data, and the gene expression levels were all upregulated. The qRT-PCR results of β1 and β2 in the PLDβ subfamily were consistent with the results in the RNA-seq data. The gene expression of β1 increased, and the gene expression of β2 decreased. The qRT-PCR result of β3 was different from the result in the RNA-seq data, the qRT-PCR result was decreased, and the RNA-seq data were increased.
At 38 °C, the qRT-PCR results in the PLDβ subfamily were consistent with the results in the RNA-seq data. The gene expression of β1 and β3 increased, and the expression of the β2 gene decreased. In the PLDα subfamily, the qRT-PCR results of α1, α2, α4, and α5 were consistent with the results in the RNA-seq data, and the gene expression levels were all upregulated. The qRT-PCR results of α3 and α4 were different from the results in the RNA-seq data. The qRT-PCR results of α3 and α4 increased, but the RNA-seq data decreased. In the StPLDδ subfamily, the qRT-PCR results of δ1, δ2, δ4, and δ5 were consistent with the results in the RNA-seq data, and the gene expression levels were all upregulated. The qRT-PCR result of δ3 was different from the result in the RNA-seq data, the qRT-PCR result of δ3 decreased, and the result of the RNA-seq data increased. In the StPLDζ subfamily, the qRT-PCR results of ζ2 were consistent with the results in the RNA-seq data, and the gene expression increased. The qRT-PCR result of ζ1 was different from the result in the RNA-seq data. The qRT-PCR result of ζ1 decreased, and the result of RNA-seq increased.
Under drought treatment, the qRT-PCR results in the PLDβ subfamily were consistent with the results in the RNA-seq data. The gene expression of β1 and β3 increased, and the gene expression of β2 decreased. In the PLDα subfamily, the qRT-PCR results of α1, α2, α5, and α6 were consistent with the results in the RNA-seq data, and the gene expression levels were all upregulated. The qRT-PCR results of α3 and α4 were different from the results in RNA-seq data. The qRT-PCR results of α3 increased, the RNA-seq data decreased, the qRT-PCR result of α4 decreased, and the RNA-seq data increased. In the StPLDδ subfamily, the qRT-PCR results of δ1, δ2, δ4, and δ5 were consistent with the results in the RNA-seq data, and the gene expression levels were all upregulated. The qRT-PCR result of δ3 was different from the result in RNA-seq data. The qRT-PCR result of δ3 increased, but the result of RNA-seq decreased. In the StPLDζ subfamily, the qRT-PCR results of ζ2 were consistent with the results in the RNA-seq data, and the gene expression increased. The qRT-PCR result of ζ1 was different from the result in the RNA-seq data. The qRT-PCR result of ζ1 decreased, and the result of the RNA-seq data increased.
Under ABA treatment, in the PLDα subfamily, the qRT-PCR results of α1, α2, α3, and α4 were consistent with the results in the RNA-seq data. The gene expression of α1, α2, and α4 increased, and the gene expression of α3 decreased. The qRT-PCR results of α5 and α6 were different from the results in the RNA-seq data. The qRT-PCR results of α5 and α6 decreased, and the results of the RNA-seq data increased. The qRT-PCR results in the PLDβ subfamily were consistent with the results in the RNA-seq data. The gene expression of β1 and β2 decreased, and the gene expression of β3 increased. In the StPLDδ subfamily, the qRT-PCR results of δ1, δ3, δ4, and δ5 were consistent with the results in the RNA-seq data. The qRT-PCR result of δ2 was different from the result in the RNA-seq data, the qRT-PCR result of δ2 decreased, and the result of the RNA-seq data increased. In the StPLDζ subfamily, the qRT-PCR results of ζ1 were different from the results in the RNA-seq data. The qRT-PCR results of ζ1 decreased, and the RNA-seq data increased. The qRT-PCR results of ζ2 were consistent with the results in the RNA-seq data.
4. Discussion
The PLD gene family has been identified in many species, including 12 in
Arabidopsis thaliana [
22], 17 in
Oryza sativa [
12], 19 in
Gossypium [
35], 13 in
Zea mays [
3], and 13 in
Cicer arietinum [
14]. This experiment identified 16 PLD genes in the potato genome, all of which contained HxKxxxxD conserved motifs, with molecular weights ranging from 86.96 kDa to 126.22 kDa (
Table 1). Compared with the above-mentioned species, the potato PLD gene has not expanded significantly, only two genes were tandem repeat genes. These 16 PLD gene family members were randomly distributed on potato chromosomes 1, 2, 3, 4, 6, 8, 10, and 12 (
Figure 3). Based on the phylogenetic analysis of
Arabidopsis thaliana and rice, it was found that 16 potato PLD were divided into four subfamilies: α, β, δ, and ζ (
Figure 1). PLD α was the most common in potato PLD gene family, which was similar to the subfamily classification of PLD gene in
Arabidopsis thaliana, rice, and maize.
The difference is that the phylogenetic analysis of the PLD gene family in rice shows that there are two more subfamilies than potato and
Arabidopsis thaliana, namely κ and φ, and two more subfamilies in cotton, φ and ε (
Figure 1), which is caused by the different N-terminal domains of PLD. According to the different domains, it can be divided into three subfamilies: C2-PLD, PX/PH-PLD, and SP-PLD. α, β, δ, γ, and ε were classified into the C2-PLD subfamily, PLDζ was classified into the PX/PH-PLD subfamily, and PLDφ was classified into the SP-PLD subfamily (
Figure 4) [
12,
13,
36]. Collinear analysis showed that there were 14 pairs of homologous genes between potato PLD and Arabidopsis PLD. Except for StPLDβ2 and AtPLDγ2, each pair of collinear genes were in the same subfamily (
Figure 7), indicating the conservatism of genes in the same group of StPLD. At present, there are few reports on the collinear genes of PLD gene family, and their functions need to be further studied.
The difference in gene structure plays an important role in the evolution of gene families. Based on the analysis of the gene structure of potato and
Arabidopsis thaliana PLD gene, it was found that the gene members of the same subfamily had similar exon and intron structures (
Figure 4B), but they were not exactly the same, indicating that functional differentiation had taken place in potato PLD gene in the long process of evolution [
37]. Conservative motif analysis showed that the number and sequence of motifs in the same subfamily were similar; only the motifs in the PLD ζ subfamily were different, which may be due to the acquisition or loss of conserved motifs in the PLD gene family in the process of evolution [
38]. The C2-PLD subfamily had five motifs, and the PXPH-PLD subfamily had three motifs. Motif 2 contained a regular-expression sequence “IYIENQ[F/Y]F”. The seventh amino acid of this sequence, Phenylalanine (F), appeared in all PX/PH-PLDs, but was often substituted by Tyrosine (Y) in the C2-PLDs [
39]. Motif 4 contained a highly conserved core triad “ERF” in the C2-PLDs [
40].
The promoter area containing the cis-action element is considered to be related to gene expression of plant growth and adaptation [
41]. It is predicted that its cis-acting element is displayed, and the potato PLD promoter region contains many acts associated with non-biological stress and hormone response, such as drought (MBS), low temperature (LTR), light response (G-Box), abscisic acid (ABRE), Salicylic acid (TCA-element) (
Figure 5), etc. It showed that these genes are regulated by low temperature and drought and also respond to the influence of exogenous hormones on plants, validated by qRT-PCR. qRT-PCR gene expression was basically in line with the prediction of cis-acting elements. StPLD gene expression changed under drought and ABA treatment. Among all the cis-acting elements, ABRE had the largest number of elements, indicating that most of the potato PLD members may be related to the stress regulated by abscisic acid. Abscisic acid responds to abiotic stress and also regulates stomata closure and root growth [
42]. The subsequent real-time quantitative results showed that compared with the control, the gene expression of the tissue-cultured plantlets treated with abscisic acid was upregulated. It is inferred that ABA plays an important role in the growth and development of potatoes.
Detecting the expression of StPLD gene by real-time quantitative qRT-PCR found that when plants were exposed to drought, high temperature, salt stress, and other adversities, they used a variety of methods to deal with adversities. At this time, the mechanism of plant genes shows similarities [
43]. In this experiment, the expression patterns of 16 StPLD genes under three abiotic stress and one hormone treatment were analyzed. The expression levels of all genes under the four treatments showed different degrees of increase and decrease (
Figure 9), which were consistent with the function of the cis-acting element predicted above. The quantitative results showed that the expression of 11 genes increased the most under high temperature treatment, which was similar to the previous study that BrPLD of cabbage [
44] had upregulated expression of nine genes under high temperature treatment at 37 °C. One CbPLD was cloned in Alpine ion mustard. A CbPLD gene was cloned in Chorispora bungeana, and it was found that CbPLD was induced at 40 °C, indicating that the gene may be regulated by high temperature [
45]. In Jatropha curcas, the enzyme activity of JcPLDα increases with temperature and reaches a peak at 60 °C, which also indicates that the JcPLDα gene is affected by high temperature [
46]. These results all indicate that the PLD gene family can regulate plant growth and development when plants are under high temperature stress. In addition, under low temperature stress, PLD may be involved in the disintegration of plasma membrane and the change in membrane fluidity. AtPLDδ has been proven to be a positive regulator of freezing stress tolerance [
47].
Studies have shown that under 20% polyethylene glycol (PEG6000) simulated drought stress, the expression of four genes in the aboveground part is significantly upregulated, and the expression of nine genes in the underground part is significantly upregulated. Under ABA treatment, the expression levels of seven genes were significantly upregulated in the aboveground part, and the expression levels of six genes in the underground part were significantly upregulated [
3]. The quantitative results of StPLDs showed that the expression levels of 11 genes were significantly upregulated under drought stress (
Figure 9C), and the expression levels of seven genes were significantly upregulated under ABA treatment (
Figure 9D), which was similar to the quantitative results of maize. Under drought stress, plants can adapt by regulating stomata. GPA1 protein controls the activity of PLDα, and GPA1 protein and PLDα1 influence the process of ABA-induced stomata opening and closing through interaction [
39]; it shows that PLDα1 plays an important role in regulating the opening and closing of stomata. The quantitative results under drought stress showed that not only the genes of the PLDα subfamily were significantly upregulated, but the genes of other sub-families were also significantly upregulated, indicating that other sub-families are also involved in the process of stomata opening and closing [
47]. Studies have shown that chickpea PLD [
14] genes are exclusively upregulated in the upper part of the ground under drought stress, indicating that they may play a role in the closure of stomata, thereby reducing transpiration and water loss. Prior to this, PLD has been thought to regulate stomata closure under drought and hypertonic stress [
48,
49]. These are enough to show that the PLD gene family plays an important role in regulating the opening and closing of plant stomata in response to drought stress [
50]. In
Arabidopsis thaliana under salt stress, PLDα1 and its derivative PA regulate the mitogen-activated protein kinase (MAPK) signal components MAPK6 and MAPK3 [
51]. MAPK6 is activated by PLDα1-derived PA and interacts with SOS1(Na
+/H
+ reverse transport protein) and phosphorylates it. Phosphorylation leads to the activation of SOS1. The activated SOS1 effectively eliminates excess Na+ from the cell, thereby preventing the cell’s Na
+ toxicity [
52]. In addition, PLDα-1 derived PA also binds to the microtubule-associated protein MAP65-1, which helps stabilize the microtubules, thereby improving salt tolerance [
53].
The tissue expression pattern of genes is closely related to functional characteristics [
31]. This study used potato tissue expression transcript data to analyze the expression of the StPLD family in 12 tissues. Potato PLD genes were abundantly expressed in petals, stamens, carpels, and sepals (
Figure 8A), indicating that the expression level of PLD genes in the flowering period is higher than other periods, in particular, the three genes StPLDα5, StPLDβ1, and StPLDδ4 were significantly expressed in the stamens. It can be speculated that these three genes may prolong or advance the flowering period of potato. Using qRT-PCR technology to detect the gene expression changes of potato PLD gene under drought, high temperature, salt stress, and ABA treatment, expression profile analysis showed that the expression of StPLDβ2 and StPLDβ3 decreased under salt stress, and the expression of StPLDα5 decreased under drought stress, which was consistent with the subsequent quantitative results.