Abstract
Within the field of computer vision, image segmentation and classification serve as crucial tasks, involving the automatic categorization of images into predefined groups or classes, respectively. In this work, we propose a framework designed for simultaneously addressing segmentation and classification tasks in image-processing contexts. The proposed framework is composed of three main modules and focuses on providing transparency, interpretability, and explainability in its operations. The first two modules are used to partition the input image into regions of interest, allowing the automatic and interpretable identification of segmentation regions using clustering techniques. These segmentation regions are then analyzed to select those considered valuable by the user for addressing the classification task. The third module focuses on classification, using an explainable classifier, which relies on hand-crafted transparent features extracted from the selected segmentation regions. By leveraging only the selected informative regions, the classification model is made more reliable and less susceptible to misleading information. The proposed framework’s effectiveness was evaluated in a case study on skin-cancer-segmentation and -classification benchmarks. The experimental analysis highlighted that the proposed framework exhibited comparable performance with the state-of-the-art deep-learning approaches, which implies its efficiency, considering the fact that the proposed approach is also interpretable and explainable.
1. Introduction
Image processing and analysis are vital components of computer vision, enabling machines to comprehend and interpret image data. Among the fundamental tasks in computer vision are image classification and image segmentation, both playing crucial roles in extracting meaningful information from digital images.
Image classification [1] involves the automatic categorization of images into predefined classes or categories. Using advanced deep-learning techniques, such as convolutional layers, the process begins with the extraction of meaningful features from the input image, capturing its unique patterns and characteristics. These features are then fed and processed by dense layers, which learn and recognize patterns within the images [2]. The model’s final layer produces a probability distribution over different classes, indicating the likelihood that the image belongs to each category. Through extensive training on large datasets, convolutional-based neural network (CNN) image-classification models can achieve impressive accuracy, enabling a wide range of applications, from object recognition to medical imaging [3,4,5,6]. However, their architecture and their large number of learnable parameters make it difficult for humans to comprehend why particular predictions are made; since they are lacking in transparency and explainability [7,8], these models are termed black box (BB) models [8].
Image segmentation [6] is a vital task in computer vision, which involves partitioning an image into multiple informative and semantically coherent regions [9]. Unlike image classification, in which the entire image is assigned to a single label, image segmentation aims to identify and distinguish regions (segmentation regions) within the image. Deep learning has proven to be remarkably successful in tackling image segmentation challenges. The U-Net [10], and DeepLab [11] approaches are among the most popular CNN-based architectures designed specifically for accurate and efficient image segmentation. The U-Net employs a symmetric encoder–decoder structure with skip connections to preserve spatial information, while DeepLab incorporates dilated convolutions to capture multi-scale contextual information. These advanced techniques have significantly improved the accuracy and speed of image segmentation, contributing to many real-world applications, especially in the domain of medical-image analysis [10,12]. Nevertheless, the utilization of a deep-learning CNN-based model in order to perform the segmentation task sacrifices the interpretation and explainability of the whole segmentation process; therefore, such models are considered black box (BB) models [13,14].
Recently, Pintelas et al. [15] proposed a new image-classification approach. The authors investigated the segmentation of informative regions from images in order to extract features for improving the performance of an image classifier. They applied their proposed approach to a skin-cancer-prediction problem, reporting promising performance while simultaneously providing some understandable and reliable explanations. Nevertheless, a limitation of this approach was that the final classification model is highly dependent on the accurate segmentation of the specific extracted regions, such as the lesion, its boundary, and the skin region. Such regions cannot be efficiently identified by a hard-crafted approach since the segmentation task does not rely to specific rules, while a general approach similar to decisions made by a human would be much more efficient [15]. This means that the segmentation region’s labels are subjective, since one individual’s decision regarding a segmented area might be different from that of another. In other words, there is no clear objective/ground truth. Therefore, an efficient and accurate segmentation algorithm which would segment the regions of an image’s in a manner that is similar to a human decision-making process should be incorporated.
Furthermore, images with large amounts of useless content (such as background) and/or noisy images usually provide unreliable patterns to image-classification models, which may lead to misleading classification-performance scores. A naïve solution would be to remove these images from the whole training dataset. However, the acquisition of labeled images, such as in medical applications, is, in general, a costly and time-consuming task; hence, the exploitation of any possible reliable information on such images would is valuable for final classification models [16].
In this research work, we propose a new eXplainable segmentation and classification framework, named XSC, which is able to address segmentation and classification tasks, providing transparency, interpretability and explainability abilities. The proposed framework is composed by three major modules: clustering, segmentation, and classification. The first two focus on partitioning the image into regions of interest (image segmentation), while the last focuses on addressing the classification task using an explainable classifier via hand-crafted transparent features extracted from every region of interest (image classification). The basic idea behind the proposed framework is to extract features using clustering techniques from various sub-regions of the input image in order to identify the segmentation regions in an automatic and interpretable way. Next, the selected useful segmentation regions are further exploited to provide valuable information to an explainable classification model. With the term useful, we denote the segmentation regions which are selected by the user as trustful to contain valuable information to perform the classification. It should be noted that segmentation regions, such as image background, offer little or no information to the classification model, and can sometimes mislead it.
The proposed framework was applied, evaluated in a skin-cancer case study as segmentation and classification benchmarks, and compared with state-of-the-art deep-learning approaches proposed in the literature. The main contributions of this work are as follows:
- We propose a new explainable segmentation and classification framework, which manages to achieve a similar level of performance to the state-of-the-art CNN black-box segmentation and classification models, as well as surpassing state-of-the-art white box models for image-classification tasks.
- Our methodology can potentially contribute to the explainable machine-learning area by providing a methodology for developing both accurate and interpretable models in both image-classification and image-segmentation applications.
- Lastly, a contribution of this work is the creation of a specialized segmentation database dedicated to the skin-cancer problem. This dataset, which was meticulously curated and designed, serves as an essential resource for training and validating segmentation models for research purposes in the field of dermatology and skin-cancer diagnosis.
The rest of this paper is organized as follows. Section 2 presents the state-of-the-art image-classification and -segmentation approaches proposed in the literature. Section 3 presents a detailed description of the proposed XSC framework, and Section 4 presents the experimental analysis and results. Finally, Section 5 presents a use-case scenario, and Section 6 summarizes the concluding remarks and our proposals for future work.
2. Related Work
Image classification is an area in machine learning and computer vision in which deep CNNs have flourished due to their remarkable performances [3,4]. These models are trained on large or even huge numbers of images and are composed on a large variety of CNN architectures, such as VGG [1], AlexNet [2], ResNet [17], Inception [18], InceptionResNet [19,20], Xception [21], DenseNet [22], MobileNet [23], NASNetMobile [24], and EfficientNet [25]. In fact, these architectures are utilized as feature extractors, and their knowledge is transferred into dense layers in order to specialize in new specific image-classification problems. It should be noted that the development of a classification model based on these architectures is generally considered a state-of-the-art approach for solving image-classification problems. Nevertheless, beyond the computation cost, which in some applications constitutes a considerable drawback of CNN models, another limitation lies in the lack of interpretation and explanation of their predictions. More specifically, these models are considered BB models, since their prediction and feature-extraction mechanisms are not interpretable. Recent approaches have attempted to interpret and explain such BB models based mainly on post hoc interpretability methods [25,26,27,28,29,30]. Nevertheless, such explanations cannot always be considered reliable and trustful [14]. In contrast, in intrinsic prediction models, the decision function of the model is totally transparent and globally interpretable, and its explanations have the potential to be reliable and trustworthy [15].
In two recent works, Pintelas et al. [14,15] proposed two global intrinsic interpretable and eXplainable prediction models, named ExpClassifier1 and ExpClassifier2 for image-classification tasks. The motivation behind both works is based on the philosophy of extracting interpretable and explainable features from images, which are fed to a linear white-box model for obtaining the prediction. ExpClassifier1 is a partially explainable model, since it utilizes features that are only understandable by image analysts. In contrast, ExpClassifier2 is a generally explainable model, since it utilizes simple defined features, which can be easily understandable in human terms. More specifically, ExpClassifier2 is based on a vector-based feature-hierarchy tree, which combines a segmentation approach and a clustering approach, in order to simplify and efficiently extract valuable information and patterns from images. In fact, the model identified the regions of interest along with their boundaries, which were then used for extracting simple texture features. Next, a filtering procedure was applied for removing redundant and less significant information to create a robust, clear, and explainable final feature representation of the initial input image. The ExpClassifier2 approach was proven to be effective for solving plant-disease and skin-cancer-image-classification tasks, achieving performance scores that were 1–2% lower compared to the state-of-the-art black-box CNN models. This performance can be considered very good, considering the fact that ExpClassifier2 is a totally explainable white-box model. However, ExpClassifier2′s main drawback lies in the segmentation algorithm utilized. More specifically, on noisy and flurry images, where the extraction of regions of interest is subjective and not clearly defined, the utilization of a hard coding-segmentation approach is not efficient. If the segmentation algorithm does not manage to efficiently filter out the noisy regions, then the classification model’s predictions and explanations are less accurate and, ultimately, not trustworthy or reliable.
Image segmentation focuses on dividing an image into multiple meaningful and semantically coherent sections [6]. In particular, it aims to identify and differentiate individual objects or regions, called “segmentation regions”, within the image. The U-Net and DeepLab models are two well-known CNN approaches, specifically designed for precise and efficient image segmentation. Both the U-Net and the DeepLab models have made significant contributions to the field of image segmentation, each offering unique architectural designs that cater to different segmentation requirements and challenges [10,11].
Ronneberger et al. [10] introduced the U-Net architecture, which was designed with a symmetric encoder–decoder structure that enables it to capture both low-level and high-level features in and input image. The encoder part consists of convolutional and pooling layers, which progressively reduce the spatial resolution and extract hierarchical features. On the other hand, the decoder part utilizes up-sampling and skip connections to reconstruct the segmentation map with detailed spatial information. The skipped connections assist in retaining fine-grained details from the encoder, improving the segmentation accuracy, particularly for small objects and boundaries.
Chen et al. [11] proposed another architecture for image segmentation, which focuses on capturing contextual information, named DeepLab. It employs dilated convolutions, which allow the network to enlarge the receptive field without losing resolution. By using different dilated rates, DeepLab can capture multi-scale contextual information effectively. One of its notable versions, DeepLab-V3, utilizes a combination of spatial pyramid pooling and a feature-refinement module to improve the segmentation performance. In fact, this pyramid pooling employs parallel dilated convolutions at different rates to capture various scales of context, while the feature-refinement module refines the final prediction using skip connections. DeepLab has achieved state-of-the-art results in various segmentation challenges, making it a popular choice in the field of computer vision.
Akash et al. [12] recently proposed a two-stage segmentation-classification model designed for COVID detection based on lung-CT-scan images, named U-NET-Xception. It combines the U-Net architecture, which is well-known for its effectiveness in image-segmentation tasks, with the Xception architecture, which is used for its efficient depth-wise separable convolutions. Based on this two-stage approach, the model performs segmentation to identify relevant regions of interest in lung-CT-scan images, and then these segmented regions are passed through a classification stage to determine whether COVID is present.
He et al. [31] proposed Mask R-CNN, in a seminal work that extended the Faster R-CNN object-detection framework to include instance segmentation. The two-stage architecture combines region-proposal generation and instance segmentation, enabling accurate object detection and pixel-level segmentation in the same model. This model has been widely used for various real-world tasks, including instance segmentation and object detection.
Chen et al. [32] proposed the dual-path network (DPN), which is a multi-path architecture for image-classification tasks. It introduces dual path networks with shortcut connections to enhance information flow and improve the learning capacity of the model. While DPN is primarily designed for classification, its multi-path nature allows potential extensions with which to handle segmentation tasks.
Qin et al. [33] proposed a two-stage network designed for salient object detection, named BASNet. It first generates saliency maps to identify the most significant object regions and then uses these regions for further processing. This kind of two-stage approach is relevant to segmentation-classification models as it involves initial localization and subsequent refinement.
In this research, we propose a new explainable framework, named XSC, designed for segmentation and classification tasks in image processing. The framework mainly emphasizes transparency and interpretability, and it is composed of three modules: clustering, segmentation, and classification. The clustering and segmentation modules collaborate to partition the input image into regions of interest, while the classification module exploits the information provided by the previous modules and conducts a prediction utilizing an explainable classifier and hand-crafted features. The XSC framework’s effectiveness was assessed in a skin-cancer case study, demonstrating promising results and comparable performance to the presented state-of-the-art DL models, while maintaining interpretability and explainability.
3. Proposed Methodology
In this work, we propose a new eXplainable machine-learning segmentation and classification framework, named XSC. This framework is composed of three modules: the clustering module, as well as the transparent-segmentation and -classification modules. The primary aim of the proposed framework is to achieve similar performance to the state-of-the-art BB image segmentation and classification CNN models, while simultaneously providing explainability and interpretability. The rationale behind this work is the efficient identification of informative and the discarding of “useless” image regions in order to enhance the final prediction classification performance, in an interpretable and explainable way.
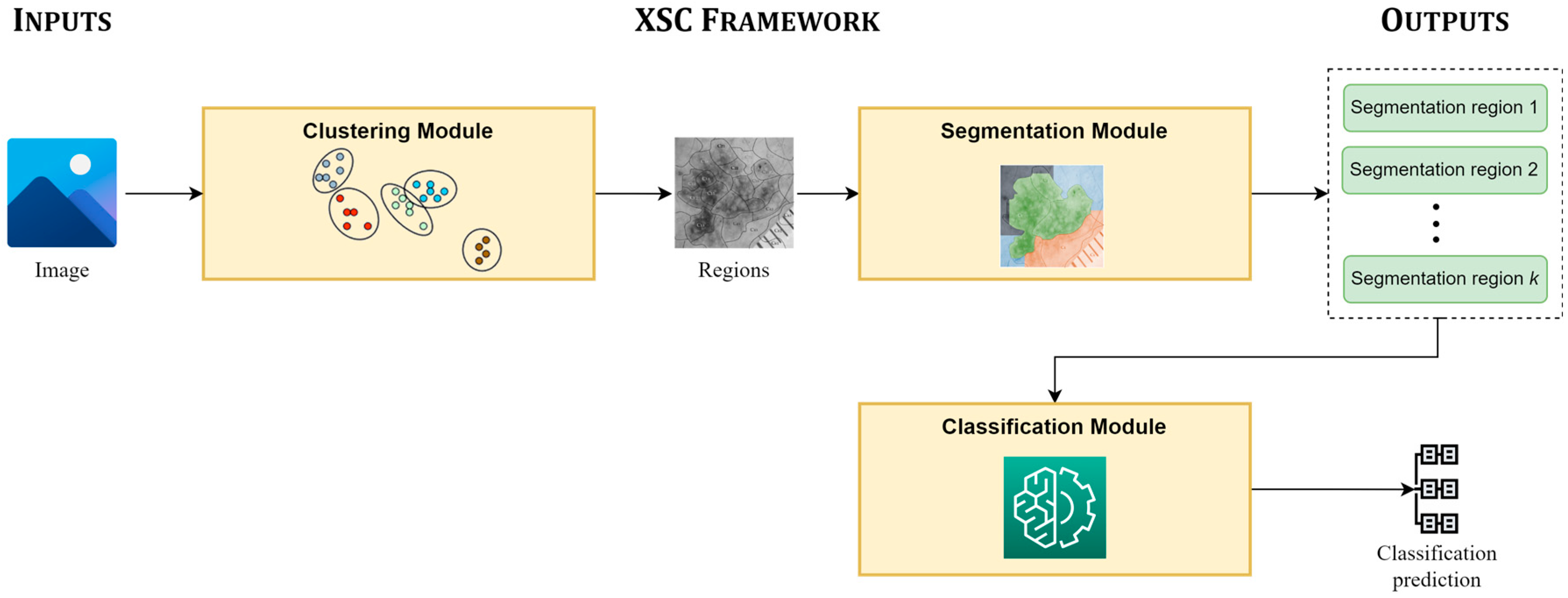
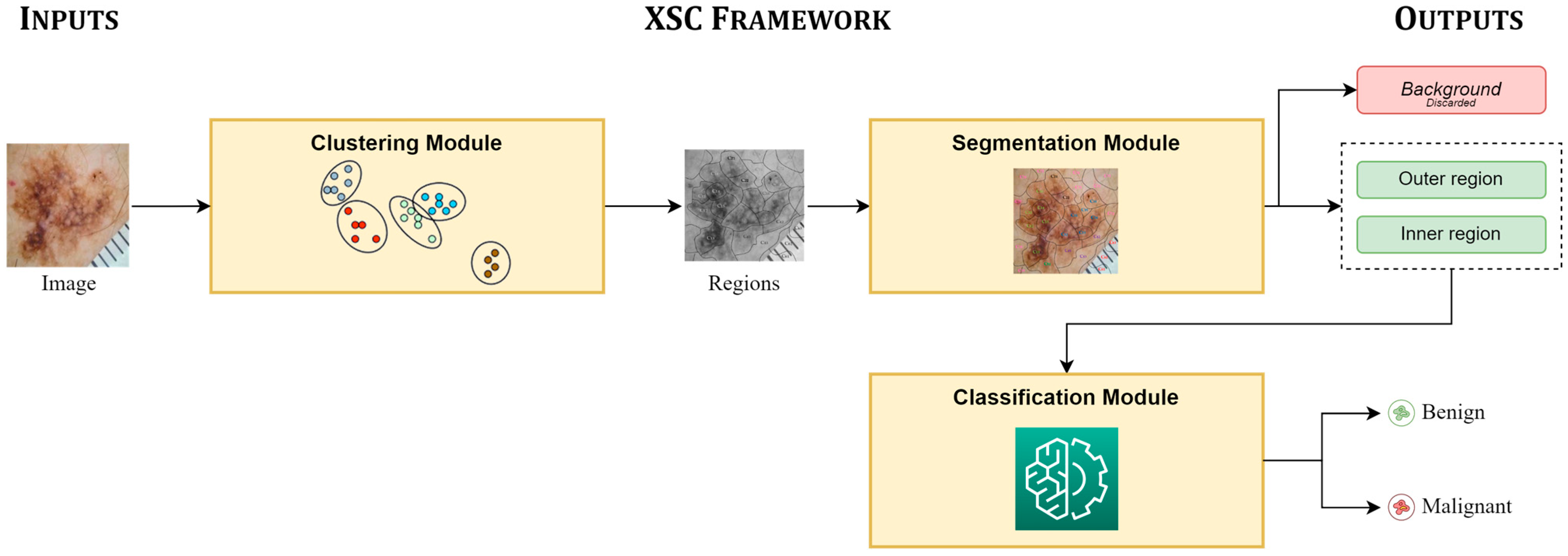
Figure 1 presents the main pipeline, focusing on the inputs of the XSC framework, its modules, and its provided outputs. The proposed framework takes an image as an input, which is initially clustered into unlabeled sub-regions by the clustering module. Next, the segmentation module is applied, in which each sub-region is assigned to a segmentation class; while the regions, which are assigned to the same segmentation class are concatenated and produce the segmentation prediction of XSC. Finally, the classification module mines information from all useful segmented regions for conducting the classification predictions.
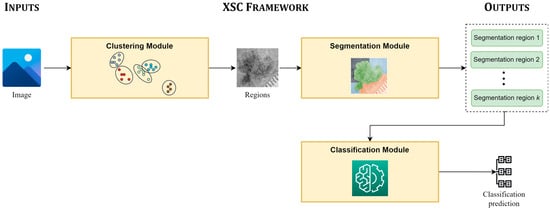
Figure 1.
Main pipeline of the proposed framework (XSC).
3.1. Clustering Module
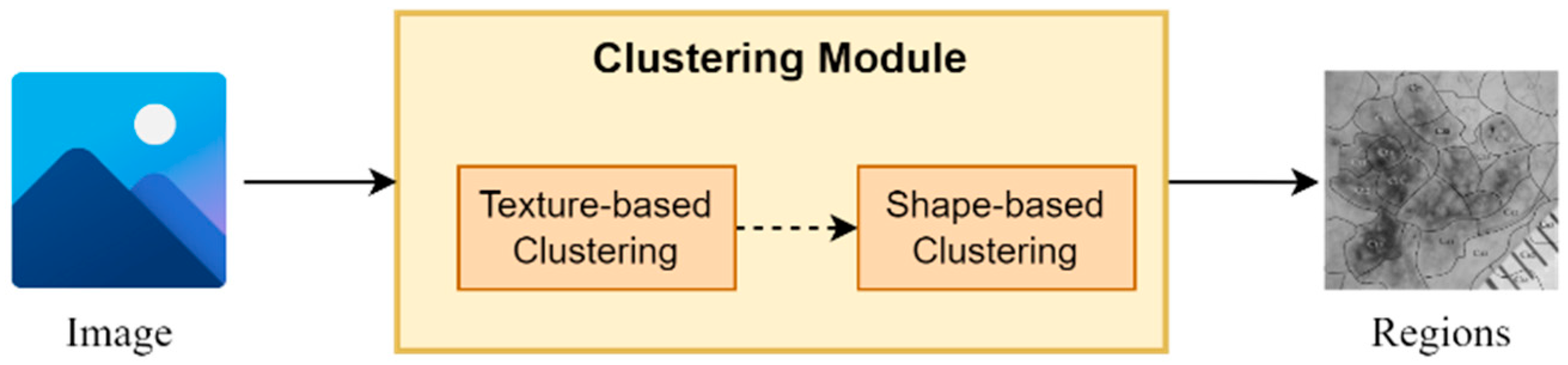
The proposed clustering module is based on a two-stage clustering approach composed of texture-based and a shape-based clustering components (Figure 2). More specifically, for every input image, the texture-based component applies a clustering algorithm, which clusters the image in regions based on its texture information (intensity pixels’ values). Next, the shape-based component for every input region applies a second clustering algorithm, which further splits every region into sub-regions based on their shape information (pixels’ coordinates values). It is worth mentioning that the output of this module is a set of sub-regions, which compose the input image.
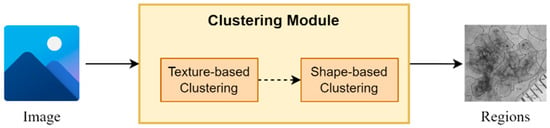
Figure 2.
An abstract overview of the segmentation module.
It should be noted that the texture-based clustering component is responsible for dividing the input image into regions based on their texture information, i.e., the intensity-pixel values. The application of a clustering algorithm results in the development of regions with similar texture patterns and color characteristics. Hence, this component offers a way to capture variations in texture within the image, helping to identify regions with similar visual properties. On the other hand, shape-based clustering component operates in each region obtained from the texture-based clustering component. As a result, every region is further split into sub-regions based on their shape and structure information. This component enables the framework to dissect regions based on their spatial distribution and structural characteristics, thereby assisting in identifying boundaries, edges, and shapes within the regions, leading to a more refined segmentation.
In our experiments, we utilized -means as clustering algorithm [34], which was selected due to simplicity and speed, while the parameter was selected based on silhouette score.
3.2. Segmentation Module
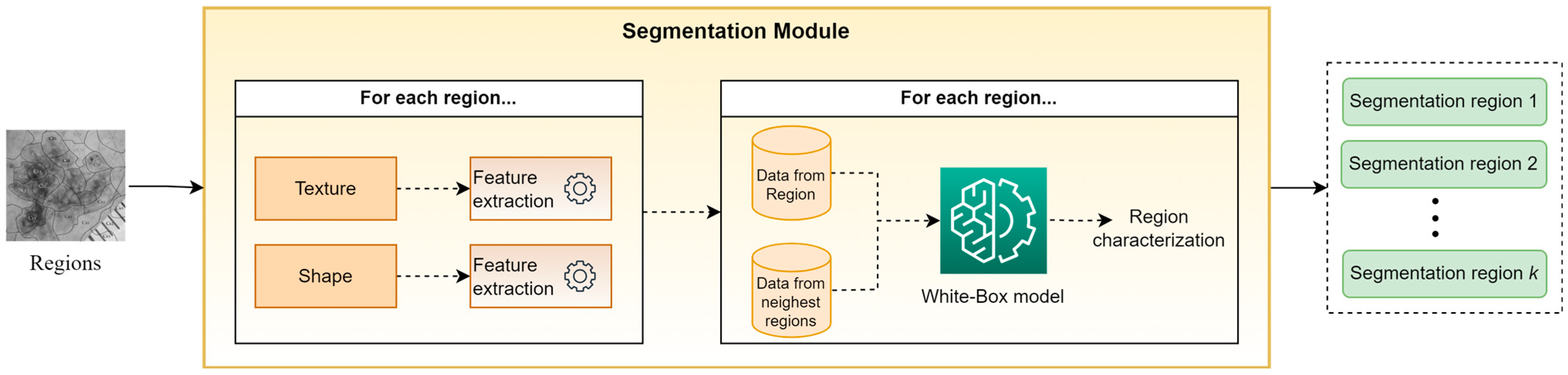
Figure 3 presents an abstract overview of the proposed segmentation module’s workflow. This module takes as input the output of the clustering module, i.e., a set of sub-regions. For every identified sub-region, a trained white box (WB) model is applied in order to classify it in one of the segmentation classes. To perform this task, the WB takes into account hand-crafted features, which are based on descriptive statistics (mean, st.d., median, kurtosis, skewness, etc.) from both the texture and shape of each sub-region, as well as from the sub-regions with the closest centroids. The motivation for this approach is based on the desire to exploit information not only from the current sub-region but also from its neighboring sub-regions. Finally, all sub-regions assigned to the same segmentation class are concatenated, composing the “segmentation regions,” which constitute the outputs of XSC framework relative to the segmentation problem. Note that in our experiments, the selected descriptive statistics were the mean values and the st.d., while parameter was set to 5.
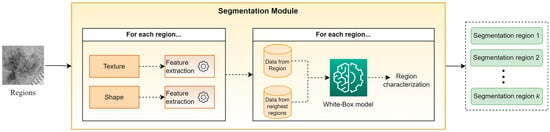
Figure 3.
An abstract overview of the segmentation modules.
Furthermore, in order to create a transparent and interpretable prediction model, we adopted the feature-hierarchy-based-tree philosophy [15] in our segmentation module (Figure 4). This means that our topology is constituted by first-order features, second-order features, and third-order features to explain the segmentation predictions of every input image’s region. The regions constitute the first-order features, while for every region, various feature families (second-order features) are extracted such as “texture features”, and “shape features”. In addition, every feature family is constituted by various specific feature-extraction formulas (third order features) corresponding to its feature-family category. In our case, these formulas are the descriptive statistics described previously. Finally, all the extracted features are fed into a white-box predictor and the explanation of every prediction is interpreted by computing the attributions for every tree branch of the feature-hierarchy tree in a backward way [15].
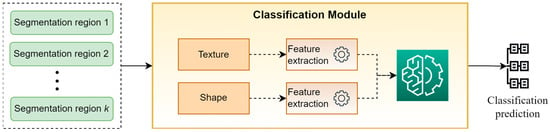
Figure 4.
An abstract overview of the classification modules.
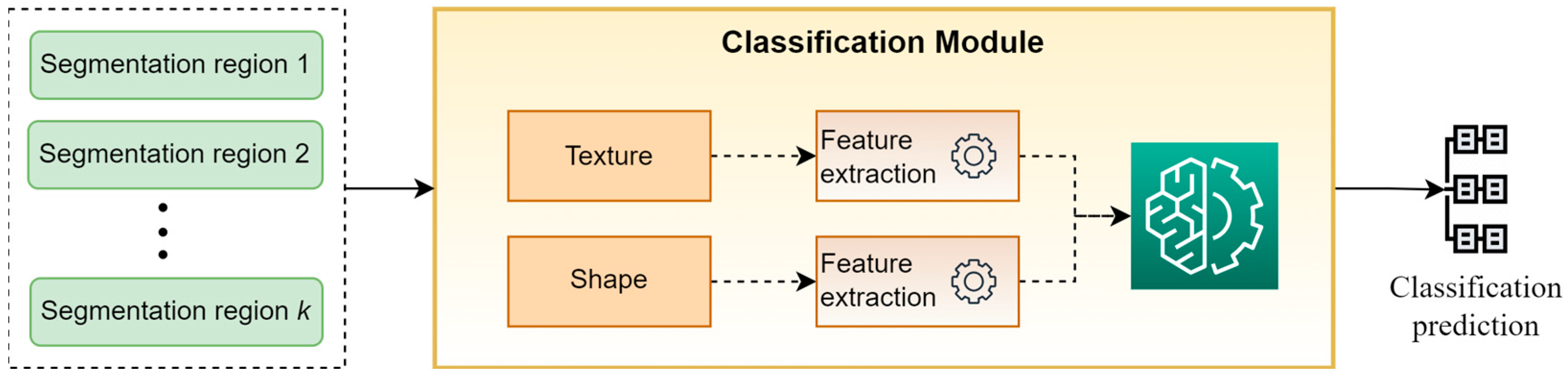
3.3. Classification Module
Figure 4 presents a high-level overview of the classification module. This module takes as input the “segmentation regions” developed by the segmentation module and classifies the original image into a specific category. For performing this task, a set of hand-crafted features based on the descriptive statistics of texture and shape of “segmentation regions” are calculated and are used as input to a WB predictor for providing the final classification prediction. It is worth mentioning that in our experiments, the selected descriptive statistics were the mean values and the st.d., while XGBoost was selected as WB classifier.
3.4. Pseudo-Code of XSC Framework
In this section, we present the pseudo-code of the proposed XSC framework (Algorithm 1) and its main modules i.e., clustering, segmentation, and classification modules, to provide the reader with a high-level description of our proposed approach.
Initially, the input image is imported (Step 1) and a number of pre-processing steps are applied such as resizing, normalization, etc. (Step 2). In Step 3, the clustering module is applied for clustering the image into sub-regions. In Step 4, these sub-regions are exploited by the segmentation module to calculate the “segmentation regions,” which constitutes the XSC’s prediction regarding the segmentation task. Finally, in Step 5, the classification module is applied for classifying the image to a specific category.
| Algorithm 1 XSC Framework. |
| Inputs |
|
| Outputs |
|
| Step 1. Import image |
| Step 2. Preprocess the image (resizing, normalization, etc.) |
| Step 3. Application of clustering module |
| Step 4. Application of segmentation module |
| Step 5. Application of classification module |
Algorithm 2 (clustering module) takes as input the processed image and splits it into sub-regions. Initially, the texture-based component applies a clustering algorithm, which clusters the image into regions based on its texture information (Step 1) while the shape-based component is applied to every region from Step 1 and further splits it into sub-regions. Finally, all created sub-regions, which constitute the output of the module, are gathered.
| Algorithm 2 Clustering module. |
| Inputs |
|
| Outputs |
|
| Step 1. Application of texture-based component for clustering the image into clusters/regions based on texture information. |
| Step 2. Application of shape-based component for splitting every cluster/region in sub-regions |
| Step 3. Create a set containing all generated sub-regions |
Algorithm 3 (segmentation module) takes as input the set of sub-regions developed by the clustering module and produces the “segmentation regions.” In more detail, for each sub-region, hand-crafted features are calculated based on its texture and shape (Steps 2–3). Next, additional hand-crafted features are calculated based on texture and shape of its nearest sub-regions (Steps 4–5). All calculated features are fed to a WB classifier, which assigns the sub-region to one of the segmentation classes (Step 6). Finally, in Step 7, the sub-regions which are assigned to the same segmentation class are merged to create the “segmentation regions.” Note that these regions constitute the prediction of the proposed framework regarding the segmentation task.
| Algorithm 3 Segmentation module. |
| Inputs |
|
| Outputs |
|
| Step 1. For each sub-region |
| Step 2. Compute hand-crafted features based on sub-region’s texture |
| Step 3. Compute hand-crafted features based on sub-region’s shape |
| Step 4. Compute hand-crafted features based on the texture from its nearest sub-regions |
| Step 5. Compute hand-crafted features based on the shape from its nearest sub-regions |
| Step 6. Application of WB classifier for assigning the sub-region to a segmentation class |
| Step 7. Development of segmentation sub-regions: merge sub-regions assigned to the same segmentation class |
Finally, Algorithm 4 (classification module) takes as input the “segmentation regions” and computes for each of them a set of hand-crafted features based on their texture and shape. These features are then fed to a WB classifier for assigning the image to a classification class, which constitutes the output of this module.
| Algorithm 4 Classification module. |
| Inputs |
|
|
| Outputs |
|
| Step 1. For each segmentation sub-region |
| Step 2. Compute hand-crafted features based on segmentation region’s texture |
| Step 3. Compute hand-crafted features based on segmentation region’s shape |
| Step 4. Application of WB classifier for assigning the image to a classification class |
4. Experimental Analysis
In this section, we present our experimental analysis to evaluate the proposed XSC framework on skin-cancer-segmentation and -classification benchmarks.
The dataset utilized in our experiments is a publicly available balanced version of the ISIC dataset [15]; it was selected in order to perform a fair and reliable classification comparison. It is composed of 1400 benign and 1400 malignant image instances (the dataset is available at https://www.kaggle.com/datasets/emmanuelpintelas/segmentation-dataset-for-skin-cancer, accessed on 1 July 2023). Regarding the segmentation problem, all images were manually annotated, while the masks contained three segmentation classes, namely “background,” “outer region,” and “inner region.”
Next, we present a comparison between the performance of the proposed XSC framework against that of state-of-the-art CNN-based approaches for image segmentation and classification of skin-cancer images. More specifically, regarding the segmentation task, XSC was evaluated against U-Net [10], DeepLab [11], U-NET-Xception [12], Mask R-CNN [31], DPN [32], and BASNet [33] using the segmentation metrics [35,36] Dice Coefficient and IoU. For the classification task, XSC was evaluated against InceptionResNet [19], DenseNet [22], EfficientNet [25], U-NET-Xception [12], Mask R-CNN [32], DPN [33] BASNet [34], ExpClassifier1 [14] and ExpClassifier2 [15], using the following classification metrics [31,37]: accuracy (Acc), area under the curve (AUC), sensitivity (Sen), specificity (Spe), positive predictive value (PPV), and negative predictive value (NPV). Note that U-NET-Xception, Mask R-CNN, DPN, and BASNet were implemented with a two-headed architecture, which implies that they are able to segment and classify an image, simultaneously. Finally, the evaluation was performed using 10-fold cross-validation, while in each simulation, 10% of the training data were used as a validation set for optimizing the network, along with the early stopping technique, to avoid overfitting. In addition, it is worth mentioning that all the models exhibited similar training and the testing performances, which implies that no overfitting occurred.
Table 1 summarizes the performances of the models applied on the skin-cancer-segmentation benchmark, while Table 2 presents the experimental results for all the evaluated models applied on the skin-cancer-classification benchmark.

Table 1.
Performance evaluation of the proposed XSC framework against state-of-the-art models on skin-cancer-segmentation benchmark.

Table 2.
Performance evaluation of the proposed XSC framework against state-of-the-art models on skin-cancer-classification benchmark.
Regarding the segmentation problem, the results underline the robustness and competitiveness of the proposed XSC framework compared to the state-of-the-art models such as U-Net, DeepLab, U-NET-Xception, Mask R-CNN, DPN, and BASNet. The proposed model exhibits a Dice Coefficient of 0.838 and an IoU score of 0.725, which implies that it exhibits a better performance than U-Net and Mask R-CNN and a marginally worse performance than U-NET-Xception, DPN, and BASNet.
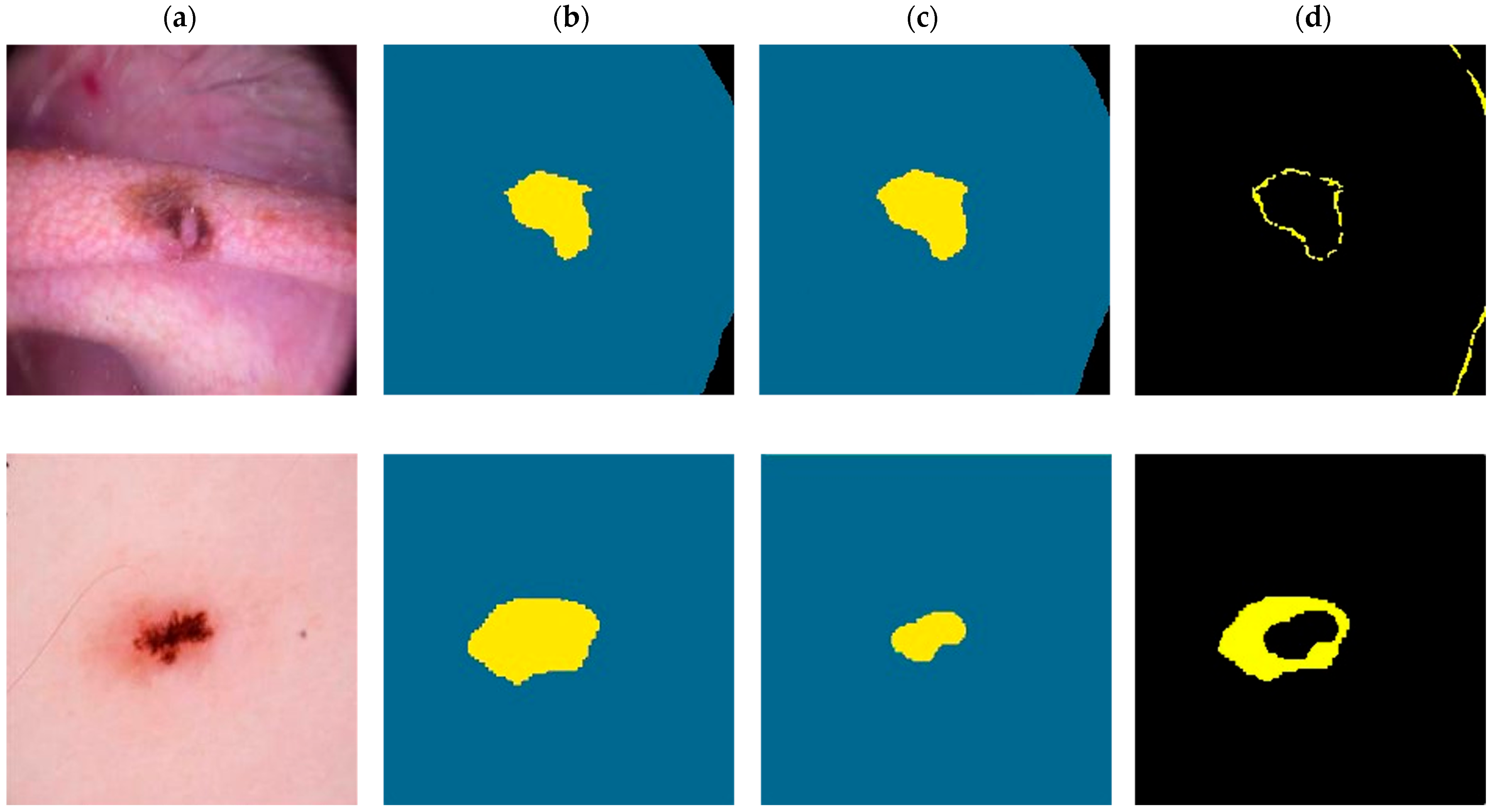
In Figure 5, we present two visual comparisons between the predicted segmentation results provided by the XSC framework and the ground truth. The comparison consists of two use-case examples representing benign and a malignant lesions. Note that in the ground-truth and predicted images, yellow denotes the “inner region,” with blue denoting the “outer region,” and black denoting the “background.”
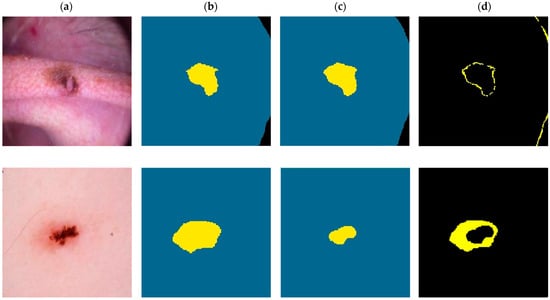

Figure 5.
Visual comparison for the XSC segmentation model for malignant (first-row images) and a benign (second row images) examples. (a) Initial. (b) Ground-truth. (c) Predicted. (d) Difference.
For the malignant case, the XSC framework effectively matched the ground truth with minimal differences, demonstrating the model’s accuracy in capturing the complexity of malignant lesions. This aligns with the general expectations of a robust segmentation model and reinforces the XSC framework’s applicability to real-world clinical scenarios.
However, the benign example provided an interesting insight into the algorithm’s behavior. While the difference between the ground-truth and predicted segmentations was larger in this case, a closer examination revealed that the algorithm prioritized segmenting the main lesion area, ignoring some boundary areas, which were marked in the ground truth. This phenomenon might be interpreted as the model’s intuitive approach to focusing on the essential features of the benign lesion, providing a segmentation which encapsulates the lesion’s core characteristics. This unique behavior could be an advantageous property, reflecting a more clinically relevant segmentation approach, which prioritizes lesion features that are crucial to diagnostic processes.
Regarding the classification problem, the proposed XSC framework achieved a comparable performance to state-of-the-art models. More specifically, the proposed framework achieved an accuracy of 0.881, which was slightly worse than those of DenseNet (0.883), U-NET-Xception (0.897), and BASNet (0.892). Similar conclusions can be made regarding the AUC metrics and the balance between the Sen and the Spe. In addition, the XSC outperformed Mask R-CNN, as well as the interpretable models ExpClassifier1 and ExpClassifier2, relative to all the performance metrics. By taking these into consideration, we were able to conclude that the proposed XSC framework presented a promising performance by achieving this level of accuracy while being able to provide interpretability and explainability. It is worth highlighting that the ability to achieve competitive performance while remaining interpretable is particularly valuable in medical applications, where explainability is essential for gaining insights into the decision-making process.
To summarize, the experimental results demonstrate that the proposed XSC framework successfully combines a very good performance with explainability and interpretability in both segmentation and classification tasks. As a result, the proposed approach may stand out as a promising solution, especially in medical-image analysis, as it provides state-of-the-art performance together with interpretability, allowing researchers and practitioners to gain a better understanding of the model’s predictions.
5. Use-Case Scenario
Next, we present an example from the application of the XSC framework to the skin-cancer segmentation and classification problem. For completeness, Figure 6 presents an abstract overview of the XSC framework applied on the skin-cancer-classification and -segmentation benchmarks. At this point, we recall that each image needed to be segmented into three regions, “background,” “outer region,” and “inner region,” in relation to the segmentation problem, while, it also needed to be classified as “benign” or “malignant” in relation to the classification problem.
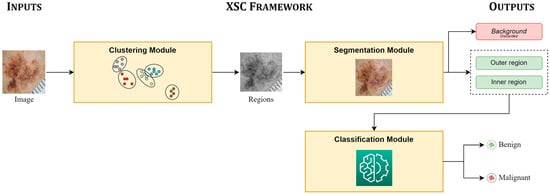
Figure 6.
Overview of XSC framework applied in skin-cancer case study.
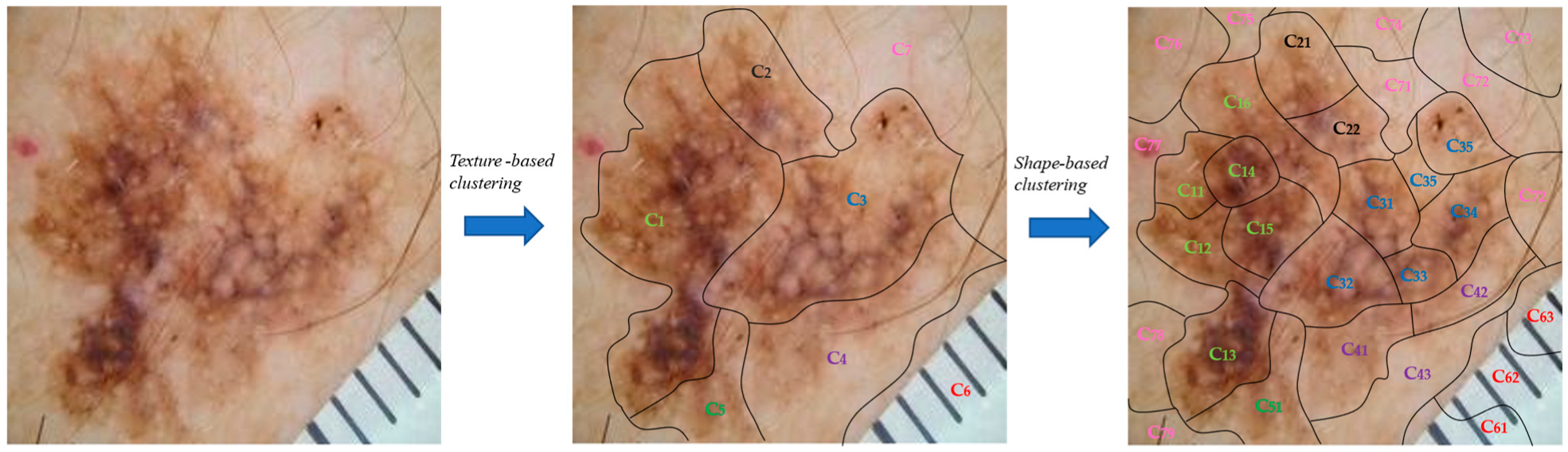
Initially, the clustering module was applied to the input image to split it into sub-regions. More specifically, the texture-based component split the image into the seven (7) clusters/regions: , while the shape-based component was applied, which further split every generated cluster/region into sub-regions (Figure 7).
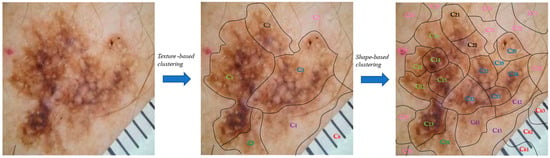
Figure 7.
Presentation of the proposed two-stage clustering algorithm via an image example.
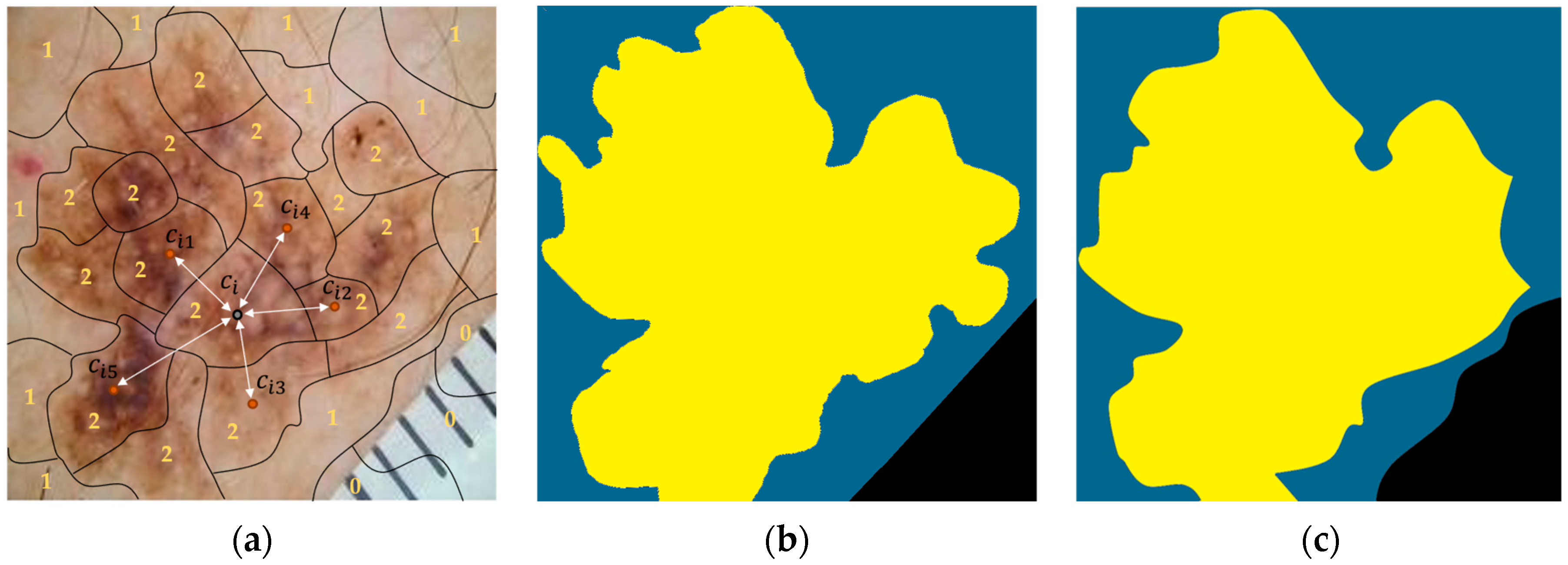
Next, the calculated sub-regions were provided to the segmentation module, which assigned each of them to one of three segmentation classes, i.e., “background,” “outer region,” and “inner region,” using a white-box XGBoost classifier. We recall that the classifier uses information (hand-crafted features) from the regions, as well as their five (5) nearest regions, and it is based on the XGBoost algorithm. Finally, the labeled regions were concatenated, forming the output segmentation regions: “background,” “outer region,” and “inner region.” Figure 8a shows the output from the application of the presented procedure, in which 0 stands for “background,” 1 stands for “outer region,” and 2 stands for “inner region.” In addition, all the sub-regions assigned to the same segmentation class were concatenated, composing the “segmentation regions,” which constituted the output of the XSC framework in relation to the segmentation problem. The ground truth is presented, along with these regions, in Figure 8b and Figure 8c, respectively.
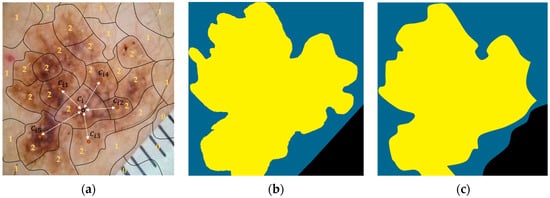

Figure 8.
An image example presenting the proposed feature-extraction procedure for our segmentation model. (a) Initial. (b) Ground-truth. (c) Predicted.
Finally, during the application of the classification module, a set of features based on descriptive statistics from both the outer and the inner region was calculated. These features were fed to a white box (XGBoost) model to provide its predictions for the original image (“benign” or “malignant”). It is worth mentioning that the “background” region had no useful information to provide to the classifier; thus, it was discarded during the classification process.
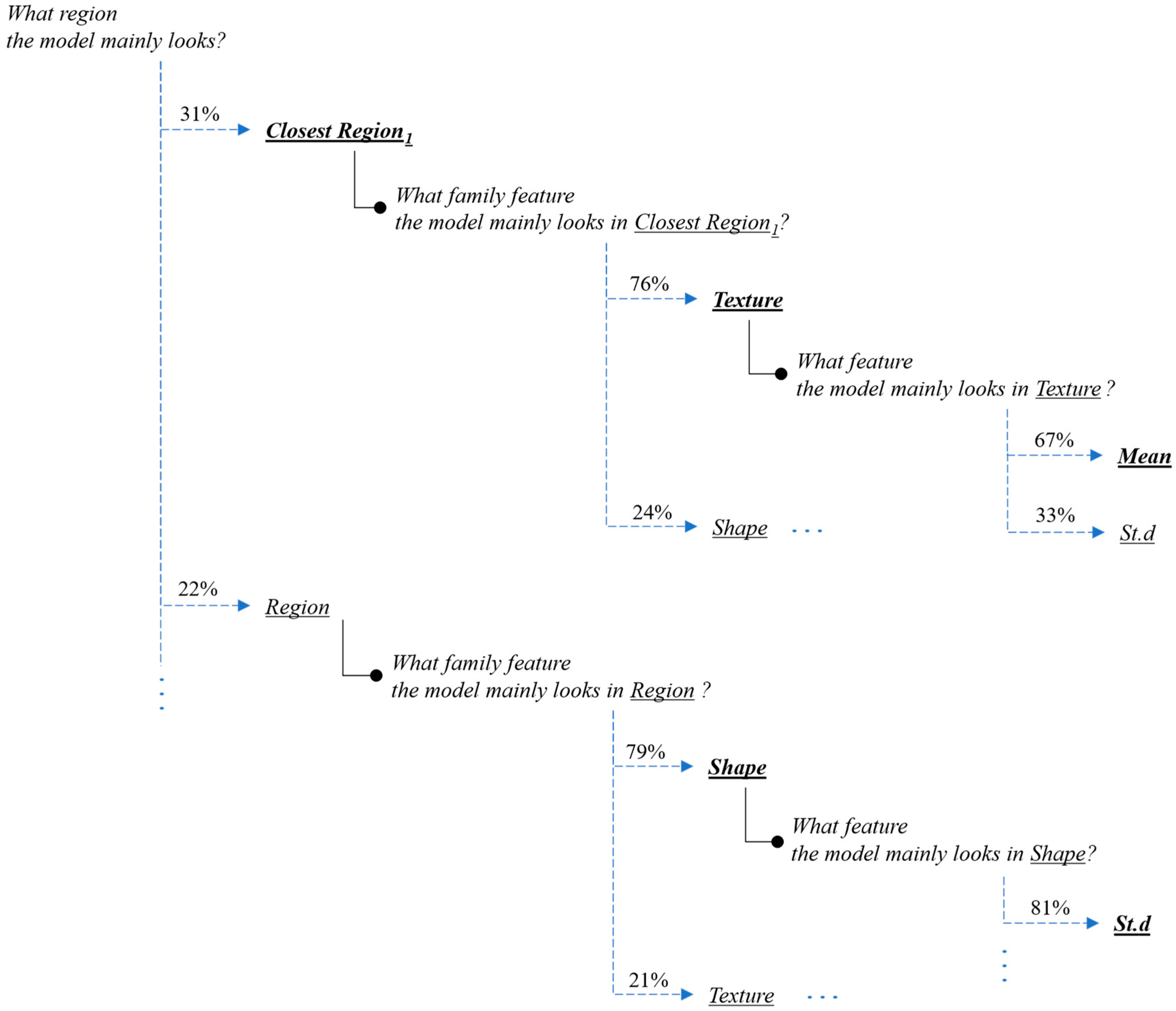
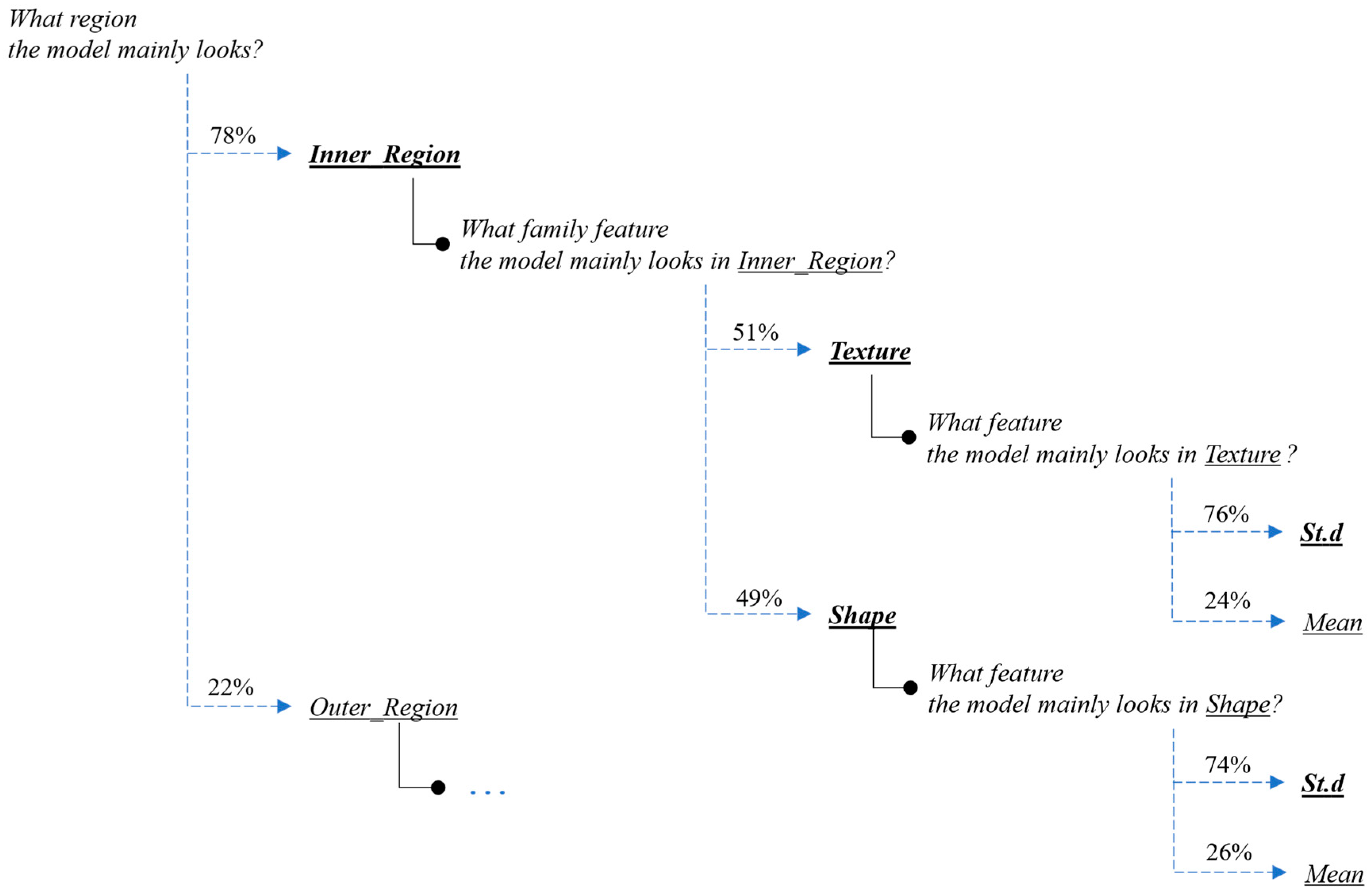
Next, we present the interpretation and the explanations of the proposed framework’s predictions for this case-study example. Figure 9 and Figure 10 present the overall feature attributions of the segmentation and classification modules, respectively, while Figure 11 and Figure 12 present the interpretations of the feature-hierarchy tree by the segmentation and classification modules, respectively.
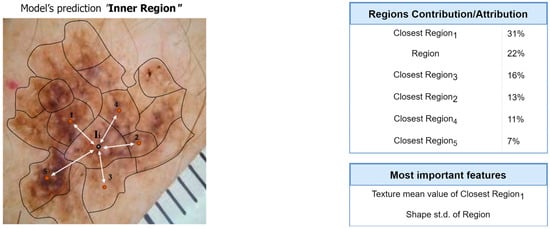
Figure 9.
Computation of overall contributions/attributions for the segmentation model.
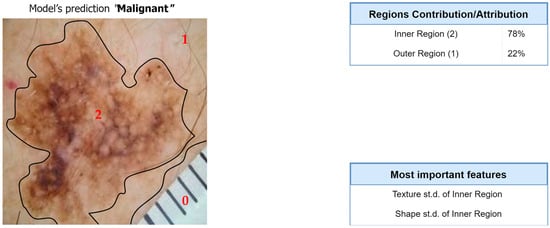
Figure 10.
Computation of overall contributions/attributions for the classification model.
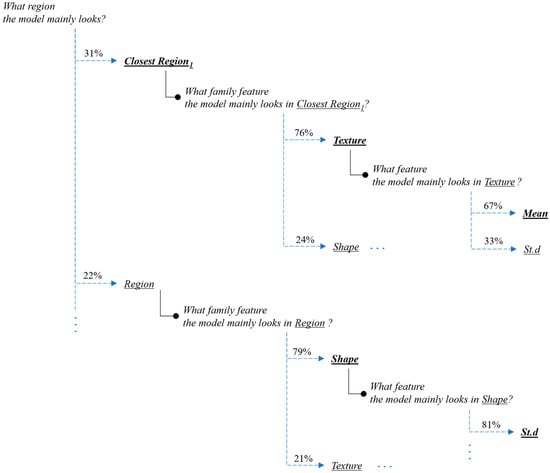
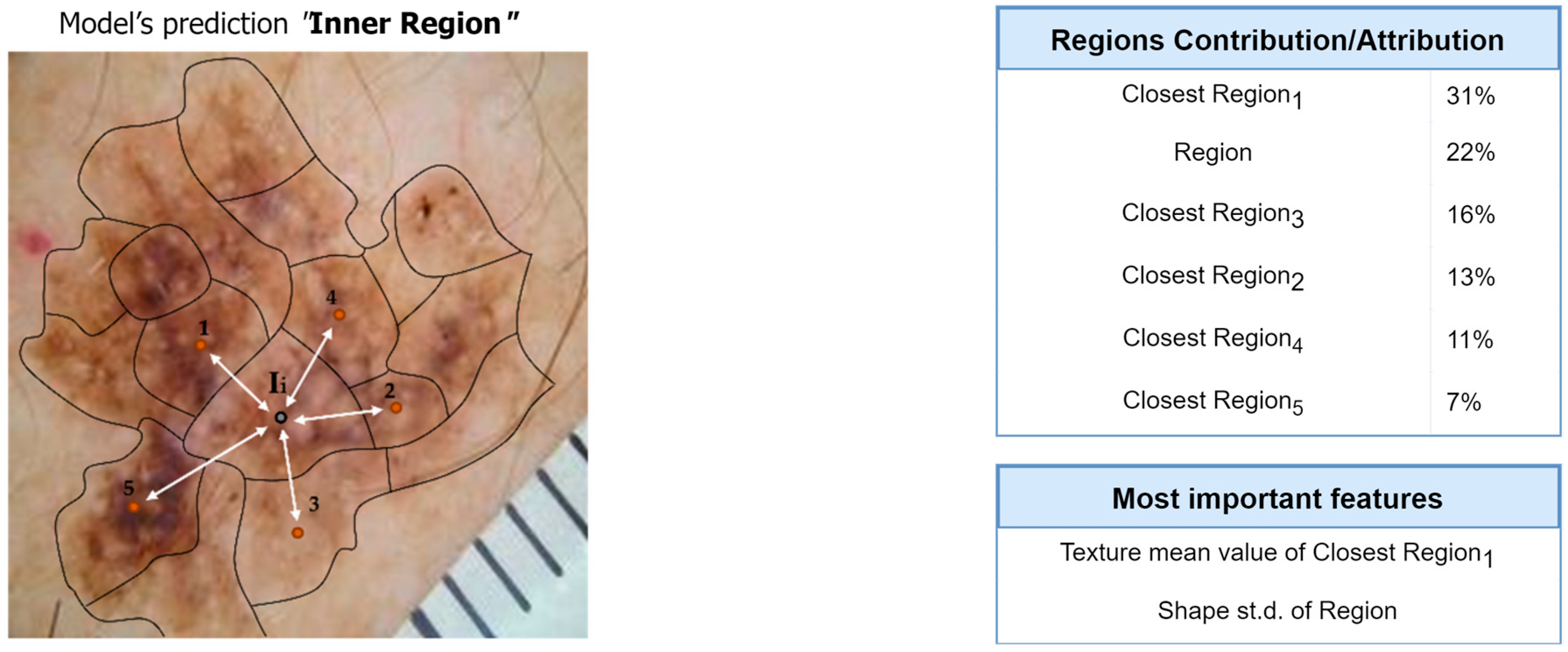
Figure 11.
Interpretation of the feature-hierarchy tree for a case-study instance (segmentation).
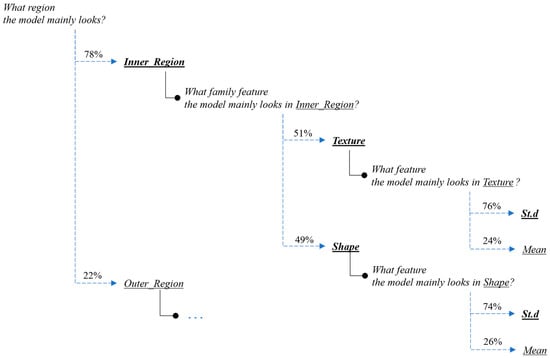
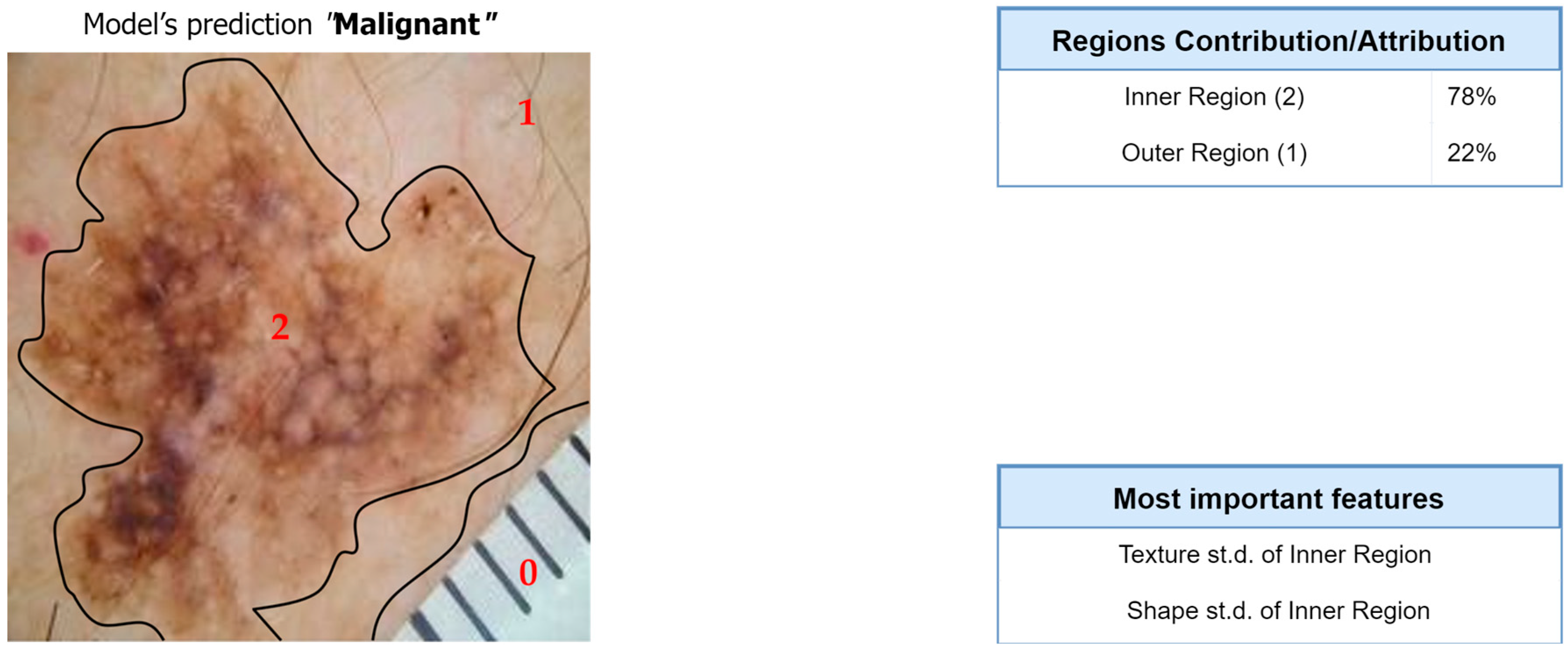
Figure 12.
Interpretation of the feature-hierarchy tree for a case-study instance (classification).
Regarding the segmentation prediction of the XSC framework, it was found that the WB (XGBoost) model in the segmentation module mainly decides based on the texture features of its five nearest regions rather than the features from the region. In addition, this model mainly decides based on the mean texture value of its closest region1 and the shape st.d of its region. Regarding the classification prediction of the proposed framework, it is shown that the WB (XGBoost) model in the classification module mainly decides based on both the texture’s st.d and shape’s st.d features of the inner region.
6. Discussion and Conclusions
In this work, we propose a new framework for simultaneously addressing segmentation and classification tasks in image processing. This framework focuses on transparency, interpretability, and explainability, and it is composed of three main modules: clustering, segmentation, and classification.
The two former modules are used to automatically identify segmentation regions in the input image, while the third module uses an explainable classifier, which is based on hand-crafted transparent features from selected segmentation regions to classify the input image. It is worth highlighting that in our proposed framework, we adopted a two-stage clustering methodology (Section 3.1), which considers both texture and shape information. The motivation behind our approach is two-fold:
- Interpretability: The two-stage clustering approach provides a more interpretable segmentation process. By initially focusing on texture-based clustering and then refining it with shape-based clustering, we can better understand how the image is partitioned into regions. Each stage provided a specific emphasis, allowing us to attribute texture-related characteristics to one stage and shape-related characteristics to the other. This interpretability is important in the context of explainable machine learning, since it allows researchers to gain insights into the contributions of different features during the segmentation process.
- Robustness: The separation of texture-based clustering from shape-based clustering can make the framework more robust to variations in the data. Different types of image may have different dominant features, and by applying separate clustering techniques, we are able to adapt to various image characteristics more effectively. This flexibility allows the framework to handle a wider range of image data and, potentially, improve segmentation performance.
In addition, another advantage of the proposed XSC framework regarding the classification module is that the selection of exploiting information from the selected region only during the classification process focuses on enhancing the reliability of the classification model by reducing susceptibility to misleading information.
The performance and the effectiveness of the proposed XSC framework were demonstrated through a skin-cancer-segmentation and -classification benchmark, in which we presented some promising results compared to state-of-the-art deep learning approaches. Encouragingly, the framework achieved a performance that was comparable with those of black-box models, which is especially noteworthy given its inherent interpretability and explainability.
Finally, another contribution of this work is the creation of a specialized segmentation database dedicated to the skin-cancer problem. This dataset, which was meticulously curated and designed, serves as an essential resource for training and validating segmentation models for research purposes in the field of dermatology and skin-cancer diagnosis, aiming to foster future research and collaboration in this vital medical area.
At this point, it is worth mentioning that the clinical implementation of the XSC framework could be a considerable step towards more transparent and informed decision making in the diagnosis of skin cancer. The explainability aspect of our model can serve as a bridge between complex machine-learning algorithms and medical practitioners, offering insights which are intuitive and relevant to clinical practice. Note that for medical practitioners, the ability to understand the reasoning behind a model’s segmentation and classification predictions can facilitate more confident and informed decision making. The interpretations presented, such as the overall feature attributions and the interpretation of the feature-hierarchy tree, could allow medical staff to understand the specific features and regions on which the model focuses. This transparency not only supports the diagnostic process but also enables medical staff to critically evaluate and corroborate the model’s findings with their expert judgments.
The explainability of the XSC framework can also be useful to patients by demystifying the often complex process of medical diagnoses. More specifically, by presenting the model’s reasoning in an interpretable manner, patients can be offered a clearer understanding of how the diagnosis was reached. This could lead to increased trust in the diagnostic process, enhanced patient–doctor communication, and a more personalized healthcare experience.
Furthermore, the integration of the proposed framework into existing clinical workflows could enhance the efficiency and accuracy of skin-cancer diagnosis. In the modern era, the combination of state-of-the-art DL models with explainable techniques offers a solution, which aligns with the growing demand for transparent and interpretable healthcare AI technologies. As a result, this alignment with both medical and technological needs could pave the way for broader adoption and more meaningful effects in various clinical scenarios.
Although the proposed research has demonstrated promising results and provides considerable progress in the field of explainable machine learning for segmentation and classification tasks, there are some limitations to this study, which require our consideration for future research. Firstly, an interesting idea is to enhance the clustering process by incorporating and exploring the performance of sophisticated clustering algorithms [38,39]. Algorithms, which belong to the class of spectral clustering, hierarchical clustering, and density-based clustering, may offer superior partitioning of image regions, leading to better accuracy and interpretability in the resulting segmentation, and, therefore, improving the overall performance of the proposed framework. Secondly, this work utilizes hand-crafted features based on descriptive statistics. The incorporation of more advanced and comprehensive hand-crafted feature sets [7] for expanding the feature space might provide richer representations of data, potentially leading to higher accuracy and robustness in both segmentation and classification tasks. Another interesting idea is to evaluate the performance of domain-specific features tailored to an application domain. Addressing these limitations may contribute to a more refined and versatile machine-learning framework, ultimately advancing the field and opening up new avenues for research and application.
Since the proposed XSC framework was only applied on the skin-cancer benchmark, evaluating its robustness and generalization across different datasets and medical conditions is critical. Future work should include the evaluation of the XSC framework on diverse datasets from the medical domain to ensure its reliability and applicability in various scenarios. Additionally, investigating methods to mitigate any potential biases or data drift in the model’s predictions may be important for real-world deployment. Finally, as the proposed framework is inherently interpretable, incorporating additional techniques for model explanation and visualization could be valuable. Techniques like saliency maps, attention mechanisms, and gradient-based attribution methods can provide deeper insights into the decision-making process of the model [29,30]. Combining these interpretability techniques with the existing framework may further enhance the trust in and adoption of the proposed solution in clinical settings.
In conclusion, addressing the aforementioned limitations and exploring the suggested future research directions may contribute to the refinement and versatility of the XSC framework for image segmentation and classification. By continuously advancing the field of explainable machine learning, we aim to facilitate more reliable, interpretable, and transparent models, ultimately benefiting the medical community and paving the way for broader applications in various image-based domains.
Author Contributions
Conceptualization, E.P.; methodology, E.P.; software, E.P.; validation, E.P., I.E.L.; formal analysis, E.P. and I.E.L.; investigation, E.P.; resources, E.P.; data curation, E.P.; writing original draft preparation, E.P.; writing review and editing, E.P. and I.E.L.; visualization, I.E.L.; supervision, I.E.L. All authors have read and agreed to the published version of the manuscript.
Funding
This research received no external funding.
Data Availability Statement
The dataset is available at https://www.kaggle.com/datasets/emmanuelpintelas/segmentation-dataset-for-skin-cancer (accessed on 1 July 2023).
Conflicts of Interest
The authors declared no potential conflict of interest with respect to the research, authorship, and/or publication of this article.
References
- Simonyan, K.; Zisserman, A. Very deep convolutional networks for large-scale image recognition. arXiv 2014, arXiv:1409.1556. [Google Scholar]
- Krizhevsky, A.; Sutskever, I.; Hinton, G.E. ImageNet classification with deep convolutional neural networks. Commun. ACM 2017, 60, 84–90. [Google Scholar] [CrossRef]
- Lu, L.; Wang, X.; Carneiro, G.; Yang, L. (Eds.) Deep Learning and Convolutional Neural Networks for Medical Imaging and Clinical Informatics; Springer International Publishing: Berlin/Heidelberg, Germany, 2019. [Google Scholar]
- Hemanth, D.J.; Estrela, V.V. (Eds.) Deep Learning for Image Processing Applications; IOS Press: Amsterdam, The Netherlands, 2017; Volume 31. [Google Scholar]
- Rajpurkar, P.; Irvin, J.; Ball, R.L.; Zhu, K.; Yang, B.; Mehta, H.; Lungren, M.P. Deep learning for chest radiograph diagnosis: A retrospective comparison of the CheXNeXt algorithm to practicing radiologists. PLoS Med. 2018, 15, e1002686. [Google Scholar] [CrossRef] [PubMed]
- Long, J.; Shelhamer, E.; Darrell, T. Fully convolutional networks for semantic segmentation. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Boston, MA, USA, 7–12 June 2015; pp. 3431–3440. [Google Scholar]
- Pintelas, E.; Livieris, I.E.; Pintelas, P. Explainable Feature Extraction and Prediction Framework for 3D Image Recognition Applied to Pneumonia Detection. Electronics 2023, 12, 2663. [Google Scholar] [CrossRef]
- Pintelas, E.; Livieris, I.E.; Pintelas, P. A grey-box ensemble model exploiting black-box accuracy and white-box intrinsic interpretability. Algorithms 2020, 13, 17. [Google Scholar] [CrossRef]
- Zhang, Z.; Liu, B.; Li, Y. FURSformer: Semantic Segmentation Network for Remote Sensing Images with Fused Heterogeneous Features. Electronics 2023, 12, 3113. [Google Scholar] [CrossRef]
- Ronneberger, O.; Fischer, P.; Brox, T. U-net: Convolutional networks for biomedical image segmentation. In Proceedings of the Medical Image Computing and Computer-Assisted Intervention–MICCAI 2015: 18th International Conference, Munich, Germany, 5–9 October 2015; Proceedings Part III 18. Springer International Publishing: Berlin/Heidelberg, Germany, 2015; pp. 234–241. [Google Scholar]
- Chen, L.C.; Papandreou, G.; Kokkinos, I.; Murphy, K.; Yuille, A.L. Deeplab: Semantic image segmentation with deep convolutional nets, atrous convolution, and fully connected CRFs. IEEE Trans. Pattern Anal. Mach. Intell. 2017, 40, 834–848. [Google Scholar] [CrossRef] [PubMed]
- Akash Guna, R.T.; Rahul, K.; Sikha, O.K. U-NET Xception: A Two-Stage Segmentation-Classification Model for COVID Detection from Lung CT Scan Images. In Proceedings of the International Conference on Innovative Computing and Communications: Proceedings of ICICC 2022, Delhi, India, 19–20 February 2022; Springer: Singapore, 2022; Volume 1, pp. 335–343. [Google Scholar]
- Molnar, C. Interpretable Machine Learning: A Guide for Making Black Box Models Explainable. 2018. Available online: https://originalstatic.aminer.cn/misc/pdf/Molnar-interpretable-machine-learning_compressed.pdf (accessed on 6 June 2018).
- Pintelas, E.; Liaskos, M.; Livieris, I.E.; Kotsiantis, S.; Pintelas, P. Explainable machine learning framework for image classification problems: Case study on glioma cancer prediction. J. Imaging 2020, 6, 37. [Google Scholar] [CrossRef] [PubMed]
- Pintelas, E.; Liaskos, M.; Livieris, I.E.; Kotsiantis, S.; Pintelas, P. A novel explainable image classification framework: Case study on skin cancer and plant disease prediction. Neural Comput. Appl. 2021, 33, 15171–15189. [Google Scholar] [CrossRef]
- Pintelas, E.; Pintelas, P. A 3D-CAE-CNN model for Deep Representation Learning of 3D images. Eng. Appl. Artif. Intell. 2022, 113, 104978. [Google Scholar] [CrossRef]
- He, K.; Zhang, X.; Ren, S.; Sun, J. Deep residual learning for image recognition. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Las Vegas, NV, USA, 27–30 June 2016; pp. 770–778. [Google Scholar]
- Szegedy, C.; Liu, W.; Jia, Y.; Sermanet, P.; Reed, S.; Anguelov, D.; Rabinovich, A. GoogLeNet/Inception Going Deeper with Convolutions. Available online: https://www.cs.unc.edu/~wliu/papers/GoogLeNet.pdf (accessed on 6 June 2018).
- Szegedy, C.; Ioffe, S.; Vanhoucke, V.; Alemi, A. Inception v4, inception resnet and the impact of residual connections on learning. In Proceedings of the AAAI Conference on Artificial Intelligence, San Francisco, CA, USA, 4–9 February 2017; pp. 4278–4284. [Google Scholar]
- Liu, L.; Ouyang, W.; Wang, X.; Fieguth, P.; Chen, J.; Liu, X.; Pietikäinen, M. Deep learning for generic object detection: A survey. Int. J. Comput. Vis. 2020, 128, 261–318. [Google Scholar] [CrossRef]
- Fran, C. Deep learning with depth wise separable convolutions. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition (CVPR), Honolulu, HI, USA, 21–26 July 2017; Huang, G., Liu, Z., Weinberger, K.Q., van der Maaten, L., Eds.; Densely Connected Convolutional Networks. [Google Scholar]
- Sandler, M.; Howard, A.; Zhu, M.; Zhmoginov, A.; Chen, L.C. Mobilenetv2: Inverted residuals and linear bottlenecks. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Salt Lake City, UT, USA, 18–23 June 2018; pp. 4510–4520. [Google Scholar]
- Zoph, B.; Vasudevan, V.; Shlens, J.; Le, Q.V. Learning transferable architectures for scalable image recognition. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Salt Lake City, UT, USA, 18–23 June 2018; pp. 8697–8710. [Google Scholar]
- Tan, M.; Le, Q. Rethinking model scaling for Convolutional Neural Networks. arXiv 2019, arXiv:1905.11946. [Google Scholar]
- Ribeiro, M.T.; Singh, S.; Guestrin, C. “Why should Ι trust you?” Explaining the predictions of any classifier. In Proceedings of the 22nd ACM SIGKDD International Conference on Knowledge Discovery and Data Mining, San Francisco, CA, USA, 13–17 August 2016; pp. 1135–1144. [Google Scholar]
- Robnik-Šikonja, M.; Bohanec, M. Perturbation-based explanations of prediction models. In Human and Machine Learning: Visible, Explainable, Trustworthy and Transparent; Springer International Publishing: Berlin/Heidelberg, Germany, 2018; pp. 159–175. [Google Scholar]
- Wachter, S.; Mittelstadt, B.; Russell, C. Counterfactual explanations without opening the black box: Automated decisions and the GDPR. Harv. J. Law Technol. 2017, 31, 841. [Google Scholar] [CrossRef]
- Selvaraju, R.R.; Cogswell, M.; Das, A.; Vedantam, R.; Parikh, D.; Batra, D. Grad-cam: Visual explanations from deep networks via gradient-based localization. In Proceedings of the IEEE International Conference on Computer Vision, Venice, Italy, 22–29 October 2017; pp. 618–626. [Google Scholar]
- Lundberg, S.M.; Lee, S.I. A unified approach to interpreting model predictions. Adv. Neural Inf. Process. Syst. 2017, 30, 4768–4777. [Google Scholar]
- Livieris, I.E. Improving the classification efficiency of an ANN utilizing a new training methodology. Informatics 2018, 6, 1. [Google Scholar] [CrossRef]
- He, K.; Gkioxari, G.; Dollár, P.; Girshick, R. Mask R-CNN. In Proceedings of the IEEE International Conference on Computer Vision, Venice, Italy, 22–29 October 2017; pp. 2961–2969. [Google Scholar]
- Chen, Y.; Li, J.; Xiao, H.; Jin, X.; Yan, S.; Feng, J. Dual path networks. Adv. Neural Inf. Process. Syst. 2017, 30, 4470–4478. [Google Scholar] [CrossRef]
- Qin, X.; Zhang, Z.; Huang, C.; Gao, C.; Dehghan, M.; Jagersand, M. Basnet: Boundary-aware salient object detection. In Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition, Long Beach, CA, USA, 16–17 June 2019; pp. 7479–7489. [Google Scholar]
- Shukla, S.; Naganna, S. A review on -means data clustering approach. Int. J. Inf. Comput. Technol. 2014, 4, 1847–1860. [Google Scholar]
- Rahman, M.A.; Wang, Y. Optimizing intersection-over-union in deep neural networks for image segmentation. In Proceedings of the International Symposium on Visual Computing, Las Vegas, NV, USA, 12–14 December 2016; Springer: Cham, Switzerland, 2016; pp. 234–244. [Google Scholar]
- Jha, S.; Kumar, R.; Priyadarshini, I.; Smarandache, F.; Long, H.V. Neutrosophic image segmentation with dice coefficients. Measurement 2019, 134, 762–772. [Google Scholar]
- Livieris, I.E.; Pintelas, E.; Kiriakidou, N.; Stavroyiannis, S. An advanced deep learning model for short-term forecasting US natural gas price and movement. In Proceedings of the Artificial Intelligence Applications and Innovations. AIAI 2020 IFIP WG 12.5 International Workshops: MHDW 2020 and 5G-PINE 2020, Neos Marmaras, Greece, 5–7 June 2020; pp. 165–176. [Google Scholar]
- Liu, C.; Xie, S.; Ma, X.; Huang, Y.; Sui, X.; Guo, N.; Yang, X. A Hierarchical Clustering Obstacle Detection Method Applied to RGB-D Cameras. Electronics 2023, 12, 2316. [Google Scholar]
- Zhang, J.; Li, Z. A Clustered Federated Learning Method of User Behavior Analysis Based on Non-IID Data. Electronics 2023, 12, 1660. [Google Scholar] [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).