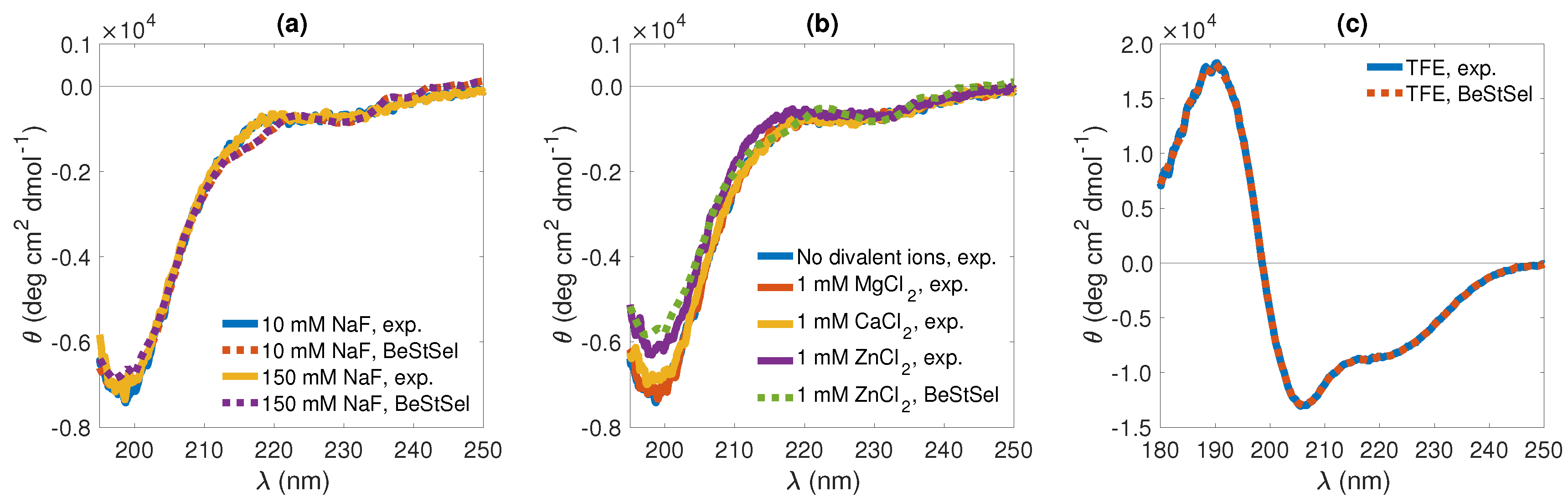
3.2.1. CD Spectroscopy
CD spectroscopy, a technique widely used to study the conformation of proteins in solution [
46,
47,
48], was used with KEIF in order to obtain information about the peptide’s secondary structure. CD spectra were recorded at 10 and 150 mM 1:1 salt (NaF), on the addition of Mg
2+, Ca
2+ and Zn
2+ cations in the form of chloride salts, as well as in organic solvent TFE (
Figure 3a,
Figure 3b and
Figure 3c, respectively). In aqueous solution (TRIS buffer) and irrespective of salt concentration, the obtained CD spectra were characteristic of a disordered structure [
46,
47], and appeared to be completely insensitive to a 15-fold change in salt concentration (
Figure 3a). The disordered structure is likely promoted by intrachain electrostatic repulsion caused by the relatively high density of positively charged amino acid residues. As expected, on the basis of the high similarity of the two spectra, the BeStSel [
5,
6] fitting of the two datasets returned highly similar secondary-structure elements where irregular (other) structures constituted the largest portion (see
Table 2). The fits also pointed to a considerable fraction of
-strands, whereas helical structure elements were absent.
Whereas KEIF secondary structure appeared to be essentially insensitive to the presence of divalent Ca
2+ and Mg
2+ cations, again deduced from recorded CD spectra, the addition of Zn
2+ ions served to make the minimum at around 200 nm somewhat less pronounced (
Figure 3b); however, the effect on the corresponding structural elements returned from BeStSel [
5,
6] fitting is almost negligible. KEIF’s apparent insensitivity to the presence of divalent cations was not surprising, considering that amino acids typically involved in metal ion co-ordination via their polar side-chain atoms—thiolate-carrying Cys (C), imidazole-carrying His (H), and carboxylate-carrying Glu (E) and Asp (D), collectively known as CHED [
49]—are scarce. Moreover, the rather high density of cationic amino acid residues, as opposed to anionic ones, likely makes KEIF–cation interactions electrostatically unfavourable.
The situation was very different when KEIF is suspended in TFE (
Figure 3c). In this organic solvent, as indicated by the development of a double minimum at 208 and 220 nm and a maximum at 192 nm [
46,
47], helical content considerably increases, mainly at the expense of the portion of
-strands (
Table 2). Similar observations were made for the human-saliva protein histatin 5, which has disordered conformation in aqueous solution [
50], but adopts a more helical conformation in TFE [
51,
52].
3.2.2. SAXS Measurements
Conformational information about the single chain of KEIF was obtained by performing SAXS experiments. The resulting form factor, Kratky plot, and distance-distribution function are depicted in
Figure 4, in comparison to the EOM fit and obtained results from MD simulations.
Figure 4a shows the obtained form factor, whose shape indicated natively unfolded behaviour. Further investigation of the data, in the form of the Kratky plot (
Figure 4b), revealed the typical curve shape of a fully flexible and extended protein/peptide. The EOM fit conformed well with the experiment data (
).
Estimations of the radius of gyration were obtained using Guinier approximation (up to
), the
and the EOM. As shown in
Table 3, Guinier approximation provides the smallest estimation, and
the largest, although the difference between the two was only 0.1 nm (5.5%). The estimation from the EOM was close to an average of the two values, and corresponded to deviations of only 2.2–3.3%. Estimations of the maximal dimension were also obtained from the
and the EOM, which are also shown in
Table 3. A larger discrepancy of approximately 2 nm (33.3%) was found between the two estimated values.
3.2.3. Atomistic Simulations
Atomistic MD simulations were performed to complement the experiment studies, and to obtain additional insight about the conformational properties of KEIF in bulk solution. Simulation convergence was assessed considering probability-distribution functions, autocorrelation functions, and block-average-error estimates of the radius of gyration and end-to-end distance (see
Figures S3–S6). PCA was also utilised for this assessment (
Figure S7). Discussion of the convergence is referred to the Supplementary Materials. To assess the validity of the simulations, simulation results were compared to the experiment results. Scattering curves were procured from the concatenated simulation trajectory by the use of CRYSOL (version 2.8.2) [
32] and compared to the experiment SAXS curves and the curves from the EOM (see
Figure 4). The curves were found to be very similar. The radius of gyration from the simulation was, however, found to be smaller than what was obtained from analysis of the experiment data (see
Table 3), although the percentage difference was only 7.1–12.6%. Because of the good correspondence with the experiment SAXS results, the simulated data were considered to be sufficiently valid to be used as accurate single-chain representation.
Cluster analysis was performed on the concatenated MD simulation trajectory to obtain representative structures. Eight clusters were found with an RMSD cutoff of 0.99 Å, and the top six (99.75%) were compared to the six structures that were obtained from EOM analysis in
Table 4. A large majority of the MD structures were found in the first two clusters at this cutoff. However, if using an RMSD cutoff of 0.70 Å or 0.50 Å, cluster sizes became of more equal size, and the top eight clusters summed up to 58.72% and 16.74%, respectively. For more thorough analysis of the structures, distance maps showing the distance between amino acid residues in the representative structures were created (see
Figure 5). By studying these maps, details otherwise unnoticed were found. For example, evidence of cation–
interactions was observed between Phe-6 and (i) Arg-20 in the MD 3 structure (see
Figure 6), (ii) Gln-27 in the MD 4 structure, and (iii) Lys-3 in the MD 5 structure. The remaining close distances seemed to arise due to hydrogen bonds and dispersion interactions, although a few electrostatic interactions were also observed. A contact map, instead showing the probability of contacts within a cutoff of 4.0 Å throughout the entire concatenated simulation, is presented in
Figure 7. Here, the most probable contact was found between Leu-23 and Gln-27. Other notable contacts were found between residues Leu-13 and Arg-16, Arg-16 and Val-30, as well as between Leu-17 and Arg-20.
The secondary structure per amino acid of KEIF from the MD simulation was analysed using the DSSP algorithm, and is visualised in
Figure 8a. Although most of the structure was dominated by coils and bends, a few residues also showed propensity for turns and
-structures. The helical content was found to be negligible. Unfortunately, this analysis did not include the PPII helix. To account for PPII helices, DSSPPII analysis was utilised on the representative structures from the top six clusters of the MD simulations (see
Table 5). While the N-terminal half of the first structure was dominated by random coil conformation, a more local order was found towards the C-terminus as distinguished turns around a small PPII helix at Asp-21–Pro-22–Leu-23, followed by an isolated
-bridge between Thr-29 and Thr-32. The small PPII helix around residues 21–24 was conserved in the top three cluster structures, although PPII helices were present in all structures. Particularly, the fourth structure seemed to have strong PPII propensity. A small
-helix was found at residues 4–6 in the second structure, whereas the sixth structure contained evidence of
-sheet formation. These results did not contradict what was observed by CD spectroscopy (
Figure 3). The presence of PPII in the conformational ensemble of KEIF was also in line with what was seen in the Kratky plot (
Figure 4), that is, mainly flexible but extended conformations. A Ramachandran plot (
Figure 8b) was also produced from the simulated results that showed a high count in the region of
, which also supported a significant PPII content. The plot also shows a fairly high count of
-structures, but only little
-helical content, which corroborated the CD spectroscopy results (
Table 2).