Comparative Transcriptome Analysis of Pueraria lobata Provides Candidate Genes Involved in Puerarin Biosynthesis and Its Regulation
Abstract
1. Introduction
2. Materials and Methods
2.1. Plant Materials and Chemicals
2.2. Phytochemical Analysis
2.3. RNA Extraction and Transcriptome Sequencing
2.4. Mapping to the P. lobata Genome and Data Analysis
2.5. Phylogenetic Analysis
2.6. Statistical Analysis
3. Results
3.1. Differential Accumulation of Puerarin and the Isoflavone Aglycones in the Pueraria Tissues
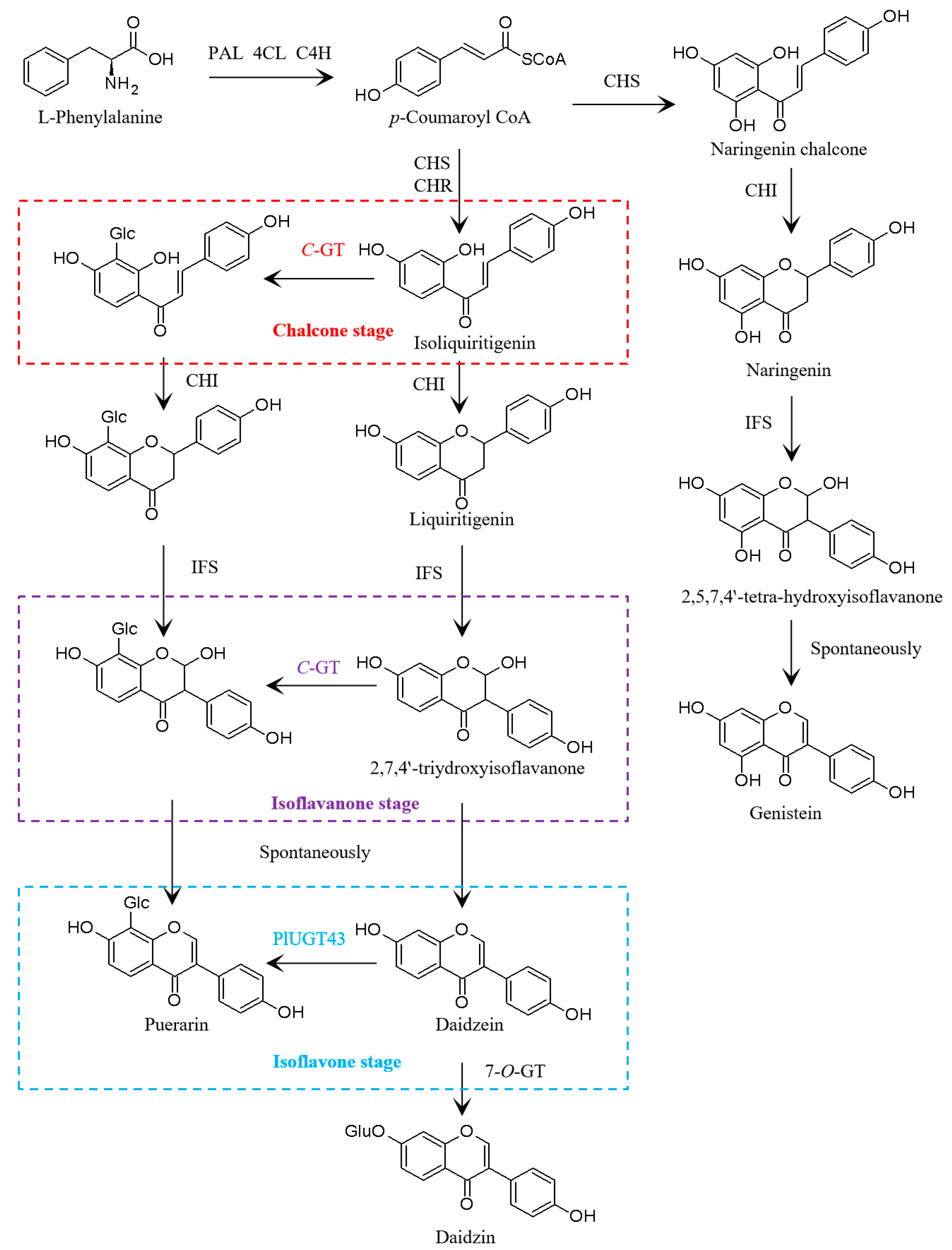
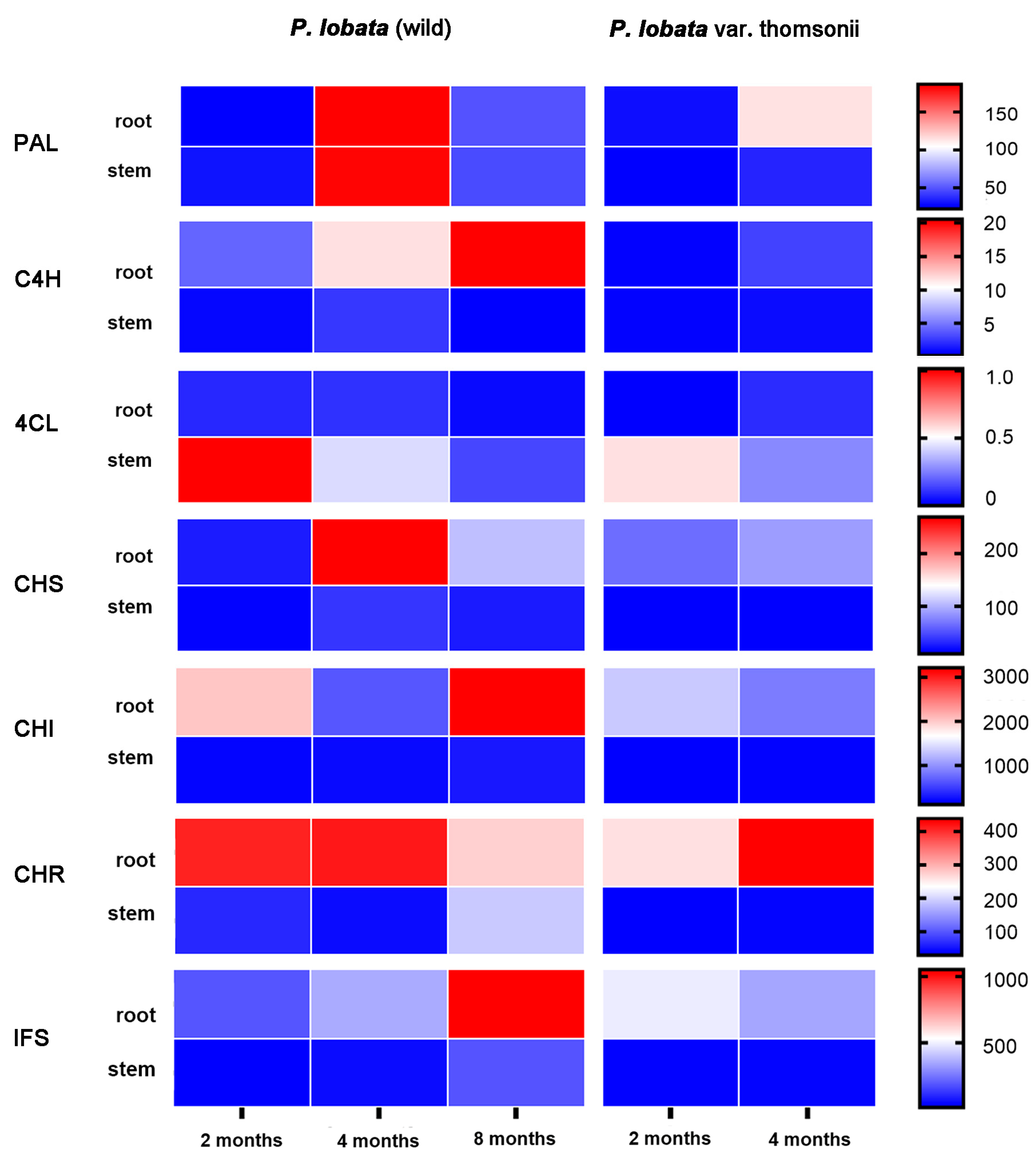
3.2. Transcriptomic Analysis of Isoflavone Biosynthetic Genes in P. lobata
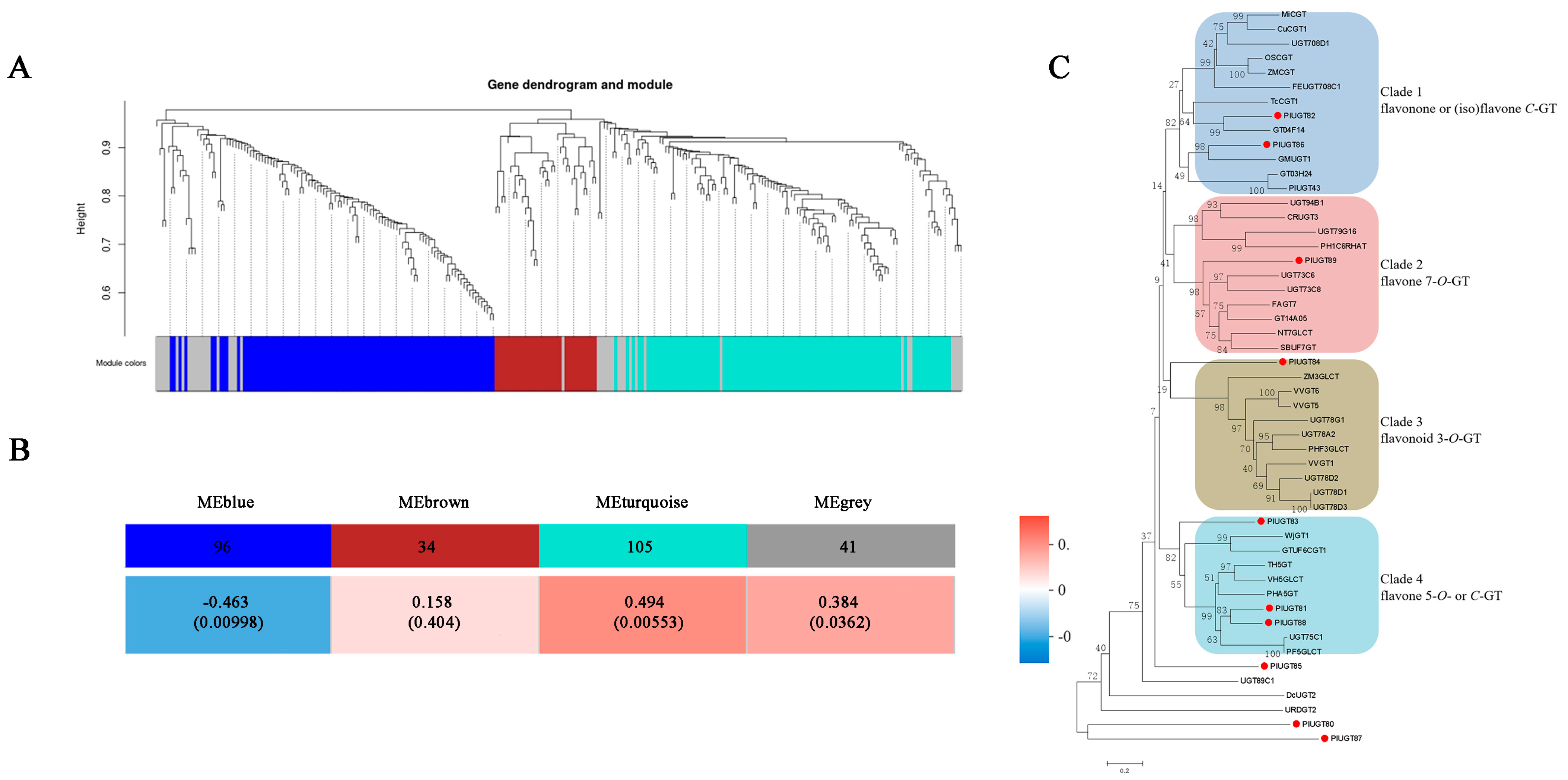
3.3. Identification of Specific UGT Genes Correlated to Puerarin Biosynthesis
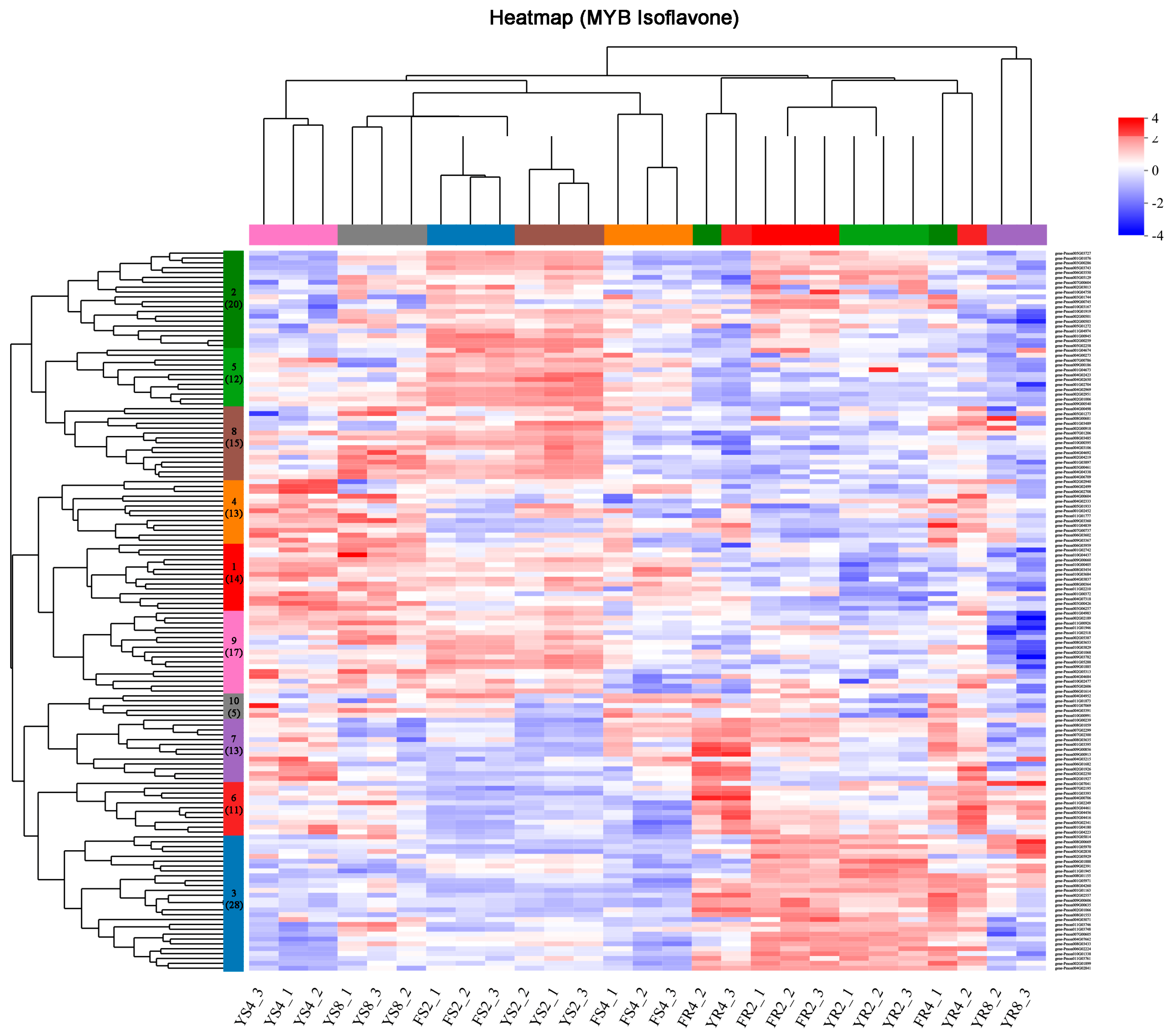
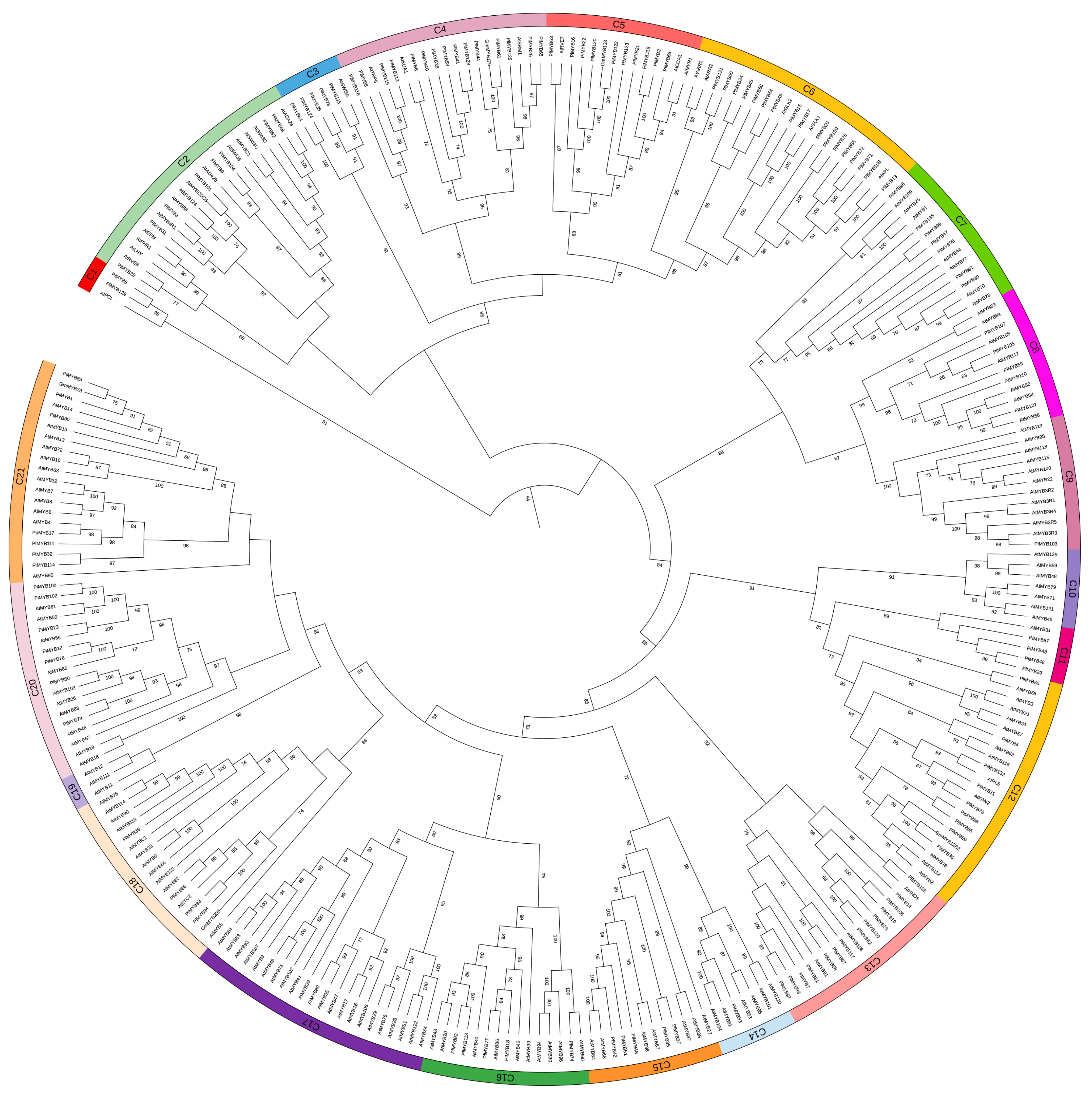
3.4. Identification of Specific Transcription Factors Important for Isoflavone Biosynthesis
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Wang, S.; Zhang, S.; Wang, S.; Gao, P.; Dai, L. A comprehensive review on Pueraria: Insights on its chemistry and medicinal value. Biomed. Pharmacother. 2020, 131, 110734. [Google Scholar] [CrossRef]
- Wong, K.H.; Li, G.Q.; Li, K.M.; Razmovski-Naumovski, V.; Chan, K. Kudzu root: Traditional uses and potential medicinal benefits in diabetes and cardiovascular diseases. J. Ethnopharmacol. 2011, 134, 584–607. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Y.X.; Zhang, H.; Peng, C. Puerarin: A review of pharmacological effects. Phytother. Res. 2014, 28, 961–975. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Y.X.; Zhang, H.; Peng, C. Effects of Puerarin on the Prevention and Treatment of Cardiovascular Diseases. Front. Pharmacol. 2021, 12, 771793. [Google Scholar] [CrossRef] [PubMed]
- Kulczynski, B.; Gramza-Michalowska, A.; Suliburska, J.; Sidor, A. Puerarin-an isoflavone with beneficial effects on bone health. Front. Biosci. 2021, 26, 1653–1667. [Google Scholar] [CrossRef]
- Murahari, M.; Singh, V.; Chaubey, P.; Suvarna, V. A Critical Review on Anticancer Mechanisms of Natural Flavonoid Puerarin. Anti-Cancer Agents Med. Chem. 2020, 20, 678–686. [Google Scholar] [CrossRef]
- Wang, H.X.; Zeng, M.S.; Ye, Y.; Liu, J.Y.; Xu, P.P. Antiviral activity of puerarin as potent inhibitor of influenza virus neuraminidase. Phytother. Res. 2021, 35, 324–336. [Google Scholar] [CrossRef]
- Jung, W.; Yu, O.; Lau, S.M.; O’Keefe, D.P.; Odell, J.; Fader, G.; McGonigle, B. Identification and expression of isoflavone synthase, the key enzyme for biosynthesis of isoflavones in legumes. Nat. Biotechnol. 2000, 18, 208–212. [Google Scholar] [CrossRef]
- Vogt, T. Phenylpropanoid biosynthesis. Mol. Plant 2010, 3, 2–20. [Google Scholar] [CrossRef]
- Wang, X.; Li, C.; Zhou, C.; Li, J.; Zhang, Y. Molecular characterization of the C-glucosylation for puerarin biosynthesis in Pueraria lobata. Plant J. Cell Mol. Biol. 2017, 90, 535–546. [Google Scholar] [CrossRef]
- Inoue, T.; Fujita, M. Biosynthesis of puerarin in Pueraria root. Chem. Pharm. Bull. 1977, 25, 3226–3231. [Google Scholar] [CrossRef]
- He, X.; Blount, J.W.; Ge, S.; Tang, Y.; Dixon, R.A. A genomic approach to isoflavone biosynthesis in kudzu (Pueraria lobata). Planta 2011, 233, 843–855. [Google Scholar] [CrossRef] [PubMed]
- Adolfo, L.M.; Burks, D.; Rao, X.; Alvarez-Hernandez, A.; Dixon, R.A. Evaluation of pathways to the C-glycosyl isoflavone puerarin in roots of kudzu (Pueraria montana lobata). Plant Direct 2022, 6, e442. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Li, Z.; Li, C.; Gou, J.; Zhang, Y. Molecular cloning and characterization of an isoflavone 7-O-glucosyltransferase from Pueraria lobata. Plant Cell Rep. 2014, 33, 1173–1185. [Google Scholar] [CrossRef] [PubMed]
- Prasain, J.K.; Reppert, A.; Jones, K.; Moore, D.R., 2nd; Barnes, S.; Lila, M.A. Identification of isoflavone glycosides in Pueraria lobata cultures by tandem mass spectrometry. Phytochem. Anal. 2007, 18, 50–59. [Google Scholar] [CrossRef]
- Chu, S.; Wang, J.; Zhu, Y.; Liu, S.; Zhou, X.; Zhang, H.; Wang, C.E.; Yang, W.; Tian, Z.; Cheng, H.; et al. An R2R3-type MYB transcription factor, GmMYB29, regulates isoflavone biosynthesis in soybean. PLoS Genet. 2017, 13, 1006770. [Google Scholar] [CrossRef]
- Han, X.; Yin, Q.; Liu, J.; Jiang, W.; Di, S.; Pang, Y. GmMYB58 and GmMYB205 are seed-specific activators for isoflavonoid biosynthesis in Glycine max. Plant Cell Rep. 2017, 36, 1889–1902. [Google Scholar] [CrossRef]
- Shang, X.; Yi, X.; Xiao, L.; Zhang, Y.; Huang, D.; Xia, Z.; Ou, K.; Ming, R.; Zeng, W.; Wu, D.; et al. Chromosomal-level genome and multi-omics dataset of Pueraria lobata var. thomsonii provide new insights into legume family and the isoflavone and puerarin biosynthesis pathways. Hortic. Res. 2022, 9, uhab035. [Google Scholar] [CrossRef]
- Wang, X.; Li, S.; Li, J.; Li, C.; Zhang, Y. De novo transcriptome sequencing in Pueraria lobata to identify putative genes involved in isoflavones biosynthesis. Plant Cell Rep. 2015, 34, 733–743. [Google Scholar] [CrossRef]
- Cerveau, N.; Jackson, D.J. Combining independent de novo assemblies optimizes the coding transcriptome for nonconventional model eukaryotic organisms. BMC Bioinform. 2016, 17, 525. [Google Scholar] [CrossRef]
- Chen, S.; Zhou, Y.; Chen, Y.; Gu, J. Fastp: An ultra-fast all-in-one FASTQ preprocessor. Bioinformatics 2018, 34, i884–i890. [Google Scholar] [CrossRef] [PubMed]
- Kim, D.; Langmead, B.; Salzberg, S.L. HISAT: A fast spliced aligner with low memory requirements. Nat. Methods 2015, 12, 357–360. [Google Scholar] [CrossRef] [PubMed]
- Pertea, M.; Pertea, G.M.; Antonescu, C.M.; Chang, T.C.; Mendell, J.T.; Salzberg, S.L. StringTie enables improved reconstruction of a transcriptome from RNA-seq reads. Nat. Biotechnol. 2015, 33, 290–295. [Google Scholar] [CrossRef]
- Mortazavi, A.; Williams, B.A.; McCue, K.; Schaeffer, L.; Wold, B. Mapping and quantifying mammalian transcriptomes by RNA-Seq. Nat. Methods 2008, 5, 621–628. [Google Scholar] [CrossRef] [PubMed]
- Langfelder, P.; Horvath, S. WGCNA: An R package for weighted correlation network analysis. BMC Bioinform. 2008, 9, 559. [Google Scholar] [CrossRef]
- Tamura, K.; Stecher, G.; Peterson, D.; Filipski, A.; Kumar, S. MEGA6: Molecular Evolutionary Genetics Analysis version 6.0. Mol. Biol. Evol. 2013, 30, 2725–2729. [Google Scholar] [CrossRef]
- Brazier-Hicks, M.; Evans, K.M.; Gershater, M.C.; Puschmann, H.; Steel, P.G.; Edwards, R. The C-glycosylation of flavonoids in cereals. J. Biol. Chem. 2009, 284, 17926–17934. [Google Scholar] [CrossRef]
- Hirade, Y.; Kotoku, N.; Terasaka, K.; Saijo-Hamano, Y.; Fukumoto, A.; Mizukami, H. Identification and functional analysis of 2-hydroxyflavanone C-glucosyltransferase in soybean (Glycine max). FEBS Lett. 2015, 589, 1778–1786. [Google Scholar] [CrossRef]
- Chen, D.; Chen, R.; Wang, R.; Li, J.; Xie, K.; Bian, C.; Sun, L.; Zhang, X.; Liu, J.; Yang, L.; et al. Probing the Catalytic Promiscuity of a Regio- and Stereospecific C-Glycosyltransferase from Mangifera indica. Angew. Chem. 2015, 54, 12678–12682. [Google Scholar] [CrossRef]
- Nagatomo, Y.; Usui, S.; Ito, T.; Kato, A.; Shimosaka, M.; Taguchi, G. Purification, molecular cloning and functional characterization of flavonoid C-glucosyltransferases from Fagopyrum esculentum M. (buckwheat) cotyledon. Plant J. Cell Mol. Biol. 2014, 80, 437–448. [Google Scholar] [CrossRef]
- Ito, T.; Fujimoto, S.; Suito, F.; Shimosaka, M.; Taguchi, G. C-Glycosyltransferases catalyzing the formation of di-C-glucosyl flavonoids in citrus plants. Plant J. Cell Mol. Biol. 2017, 91, 187–198. [Google Scholar] [CrossRef] [PubMed]
- Falcone Ferreyra, M.L.; Rodriguez, E.; Casas, M.I.; Labadie, G.; Grotewold, E.; Casati, P. Identification of a bifunctional maize C- and O-glucosyltransferase. J. Biol. Chem. 2013, 288, 31678–31688. [Google Scholar] [CrossRef] [PubMed]
- He, J.B.; Zhao, P.; Hu, Z.M.; Liu, S.; Kuang, Y.; Zhang, M.; Li, B.; Yun, C.H.; Qiao, X.; Ye, M.; et al. Molecular and Structural Characterization of a Promiscuous C-Glycosyltransferase from Trollius chinensis. Angew. Chem. 2019, 58, 11513–11520. [Google Scholar] [CrossRef] [PubMed]
- Vom Endt, D.; Kijne, J.W.; Memelink, J. Transcription factors controlling plant secondary metabolism: What regulates the regulators? Phytochemistry 2002, 61, 107–114. [Google Scholar] [CrossRef] [PubMed]
- Bian, S.; Li, R.; Xia, S.; Liu, Y.; Jin, D.; Xie, X.; Dhaubhadel, S.; Zhai, L.; Wang, J.; Li, X.; et al. Soybean CCA1-like MYB transcription factor GmMYB133 modulates isoflavonoid biosynthesis. Biochem. Biophys. Res. Commun. 2018, 507, 324–329. [Google Scholar] [CrossRef]
- Anguraj Vadivel, A.K.; McDowell, T.; Renaud, J.B.; Dhaubhadel, S. A combinatorial action of GmMYB176 and GmbZIP5 controls isoflavonoid biosynthesis in soybean (Glycine max). Commun. Biol. 2021, 4, 356. [Google Scholar] [CrossRef]
- Arce-Rodriguez, M.L.; Martinez, O.; Ochoa-Alejo, N. Genome-Wide Identification and Analysis of the MYB Transcription Factor Gene Family in Chili Pepper (Capsicum spp.). Int. J. Mol. Sci. 2021, 22, 2229. [Google Scholar] [CrossRef]
- Wang, X.; Li, C.; Zhou, Z.; Zhang, Y. Identification of Three (Iso)flavonoid Glucosyltransferases From Pueraria lobata. Front. Plant Sci. 2019, 10, 28. [Google Scholar] [CrossRef]
- Li, J.; Li, C.; Gou, J.; Wang, X.; Fan, R.; Zhang, Y. An Alternative Pathway for Formononetin Biosynthesis in Pueraria lobata. Front. Plant Sci. 2016, 7, 861. [Google Scholar] [CrossRef]
- Li, J.; Li, C.; Gou, J.; Zhang, Y. Molecular Cloning and Functional Characterization of a Novel Isoflavone 3′-O-methyltransferase from Pueraria lobata. Front. Plant Sci. 2016, 7, 793. [Google Scholar] [CrossRef]
- Zhang, Y.; Teoh, K.H.; Reed, D.W.; Maes, L.; Goossens, A.; Olson, D.J.; Ross, A.R.; Covello, P.S. The molecular cloning of artemisinic aldehyde Delta11(13) reductase and its role in glandular trichome-dependent biosynthesis of artemisinin in Artemisia annua. J. Biol. Chem. 2008, 283, 21501–21508. [Google Scholar] [CrossRef] [PubMed]
- Funaki, A.; Waki, T.; Noguchi, A.; Kawai, Y.; Yamashita, S.; Takahashi, S.; Nakayama, T. Identification of a Highly Specific Isoflavone 7-O-glucosyltransferase in the soybean (Glycine max (L.) Merr.). Plant Cell Physiol. 2015, 56, 1512–1520. [Google Scholar] [CrossRef] [PubMed]
- Gutmann, A.; Nidetzky, B. Switching between O- and C-Glycosyltransferase through Exchange of Active-Site Motifs. Angew. Chem. 2012, 51, 12879–12883. [Google Scholar] [CrossRef] [PubMed]

Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Xi, H.; Zhu, Y.; Sun, W.; Tang, N.; Xu, Z.; Shang, X.; Zhang, Y.; Yan, H.; Li, C. Comparative Transcriptome Analysis of Pueraria lobata Provides Candidate Genes Involved in Puerarin Biosynthesis and Its Regulation. Biomolecules 2023, 13, 170. https://doi.org/10.3390/biom13010170
Xi H, Zhu Y, Sun W, Tang N, Xu Z, Shang X, Zhang Y, Yan H, Li C. Comparative Transcriptome Analysis of Pueraria lobata Provides Candidate Genes Involved in Puerarin Biosynthesis and Its Regulation. Biomolecules. 2023; 13(1):170. https://doi.org/10.3390/biom13010170
Chicago/Turabian StyleXi, Huiting, Yaru Zhu, Wenwen Sun, Nan Tang, Zhengqin Xu, Xiaohong Shang, Yansheng Zhang, Huabing Yan, and Changfu Li. 2023. "Comparative Transcriptome Analysis of Pueraria lobata Provides Candidate Genes Involved in Puerarin Biosynthesis and Its Regulation" Biomolecules 13, no. 1: 170. https://doi.org/10.3390/biom13010170
APA StyleXi, H., Zhu, Y., Sun, W., Tang, N., Xu, Z., Shang, X., Zhang, Y., Yan, H., & Li, C. (2023). Comparative Transcriptome Analysis of Pueraria lobata Provides Candidate Genes Involved in Puerarin Biosynthesis and Its Regulation. Biomolecules, 13(1), 170. https://doi.org/10.3390/biom13010170