1. Introduction
Male infertility is a global health problem that affects approximately 7% of the male population [
1]. It is a complex pathological condition that involves environmental and genetic factors [
2]. More importantly, however, over the past 40 years, sperm counts have decreased by half, and this decline is characterized as alarming [
3]. Overall, studies indicate that infertility is on the rise, and one in six couples trying to conceive worldwide is diagnosed as infertile [
4]. The male factor also contributes to 40–50% of all cases of infertility [
5]. Therefore, there is a growing need for a better understanding of the molecular mechanisms involved in male infertility, leading to new approaches for its prevention, diagnosis, and treatment [
6].
In recent years, with the development of high-throughput sequencing technologies, non-coding RNAs (ncRNAs), including long non-coding RNAs (lncRNAs), circular (circRNAs), and microRNAs (miRNAs), have emerged as important regulators in the pathogenesis of male infertility, and it is also suggested that many of these molecules have the potential to be used as biomarkers [
7,
8]. Previous reviews have extensively discussed the expression profiles and regulatory functions in male infertility [
9,
10]; therefore, these topics are not discussed here.
Circular RNAs (circRNAs) are single-stranded, continuous-loop RNA transcripts that are covalently closed. In general, precursor mRNAs (pre-mRNAs) are transcribed by RNA polymerase II (Pol II), and then they undergo a process called “canonical splicing” that is catalyzed by the spliceosomal machinery and involves the splicing of introns and ligation of exons for the formation of a linear RNA transcript (i.e., mature messenger RNA (mRNA)) with 5′–3′ polarity. However, these canonical spliceosome-mediated pre-mRNAs can also undergo a process called “back splicing” and form circular RNAs in this way. “Back-splicing” is a type of alternative splicing in which the 3′ end of an exon binds to the 5′ end of its own or an upstream exon via a 3′,5′-phosphodiester link, generating a closed structure with a back-splicing junction site [
11,
12,
13]. Due to this structure, they are relatively stable and less susceptible to degradation by Ribonuclease R (RNase R) [
14]. RNase R is an exoribonuclease that has the ability to degrade structured RNA molecules such as mRNAs or ribosomal RNA (rRNA) [
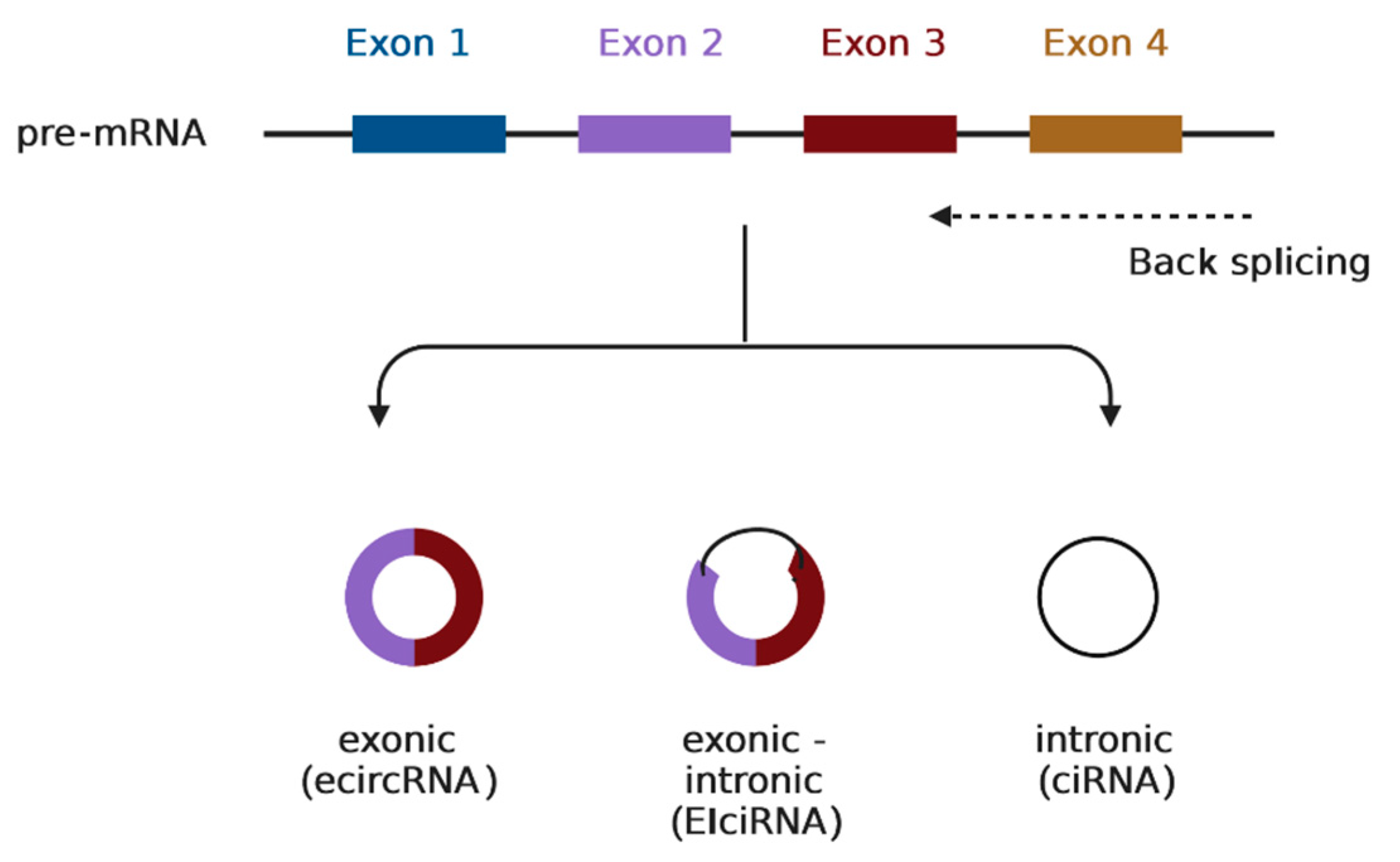
15]. As circRNAs may arise from exons or introns, they are classified into three main subtypes of circRNAs: exonic (ecircRNAs), intronic (ciRNAs), and exonic-intronic (EIciRNAs) [
16] (
Figure 1). Furthermore, circRNAs exert their function through various mechanisms [
13]. Among others, some studies demonstrate that they can act as miRNA sponges or competing endogenous RNAs (ceRNAs) and upregulate the miRNA target genes, contributing to gene regulation at the post-transcriptional level [
17]. In addition, they have complex interactions with several proteins [
13]. They can serve as protein sponges or decoys and affect their function [
18]. They can also regulate parental gene expression through diverse mechanisms, such as translational modulation and post-translational modification [
19]. Finally, their high abundance, stability, and evolutionary conservation between species are some of their intriguing characteristics [
14,
20], and growing evidence suggests their involvement in many diseases, such as cardiovascular disease, diabetes, cancer, etc. [
11].
Regarding reproduction, these molecules are gradually gaining the attention of researchers, but despite several sporadic reports about their expression patterns and their potential role in processes associated with fertility, the research on circRNAs in this field is still in its infancy. Many studies also suggest that circRNAs may function as potential biomarkers or therapeutic and pharmaceutical targets in reproduction research [
21]. However, it is not clear exactly how circRNAs contribute to male infertility. Most of the data have been obtained mainly from experiments in cell lines and animal models, and limited information is available on whether these results are validated in humans. There are many heterogeneous studies reporting findings on circRNAs and male infertility, but to our knowledge, there are no systematic reviews summarizing the circRNAs that have been associated with male infertility and its specific subtypes or that could serve as biomarkers. A comparison of the studies focusing on humans could be important to identify similar patterns of expression of circRNAs and circRNAs with a regulatory role in male infertility.
Therefore, to address this gap and determine how far science has advanced regarding circRNAs and male reproduction, the present systematic review aimed to summarize existing studies on humans about the role of circRNAs in male infertility. With this approach, we aim to provide a comprehensive summary of the circRNAs involved in male infertility, which will be of particular interest to the life science community and could serve as candidate biomarkers or therapeutic targets in the future.
4. Discussion
Recent advances in molecular technologies, as well as the wide use of genome sequencing and transcriptomic analyses, have enhanced our understanding of male infertility. CircRNAs are a new class of RNA molecules with cell- and tissue-specific expression and are conserved among eukaryotes [
56]. Furthermore, they contribute to the regulation of many cellular processes and are released in body fluids, where they are highly stable. Therefore, the use of circRNAs as novel biomarkers shows great promise and may eventually have a significant impact on clinical practice for male infertility diagnosis and prognosis. Their study could also provide a new perspective on our understanding of the molecular mechanisms and pathogenesis of male infertility. Thus, we reviewed studies investigating the role of circRNAs in male infertility and summarized DE circRNAs from all published studies in humans, evaluating the processes and pathways that are affected, mainly through circRNA-miRNA interactions, as well as the diagnostic performance of circRNAs.
In the present systematic review, 13 articles were included in the final sample analyzed. Azoospermia is the most well-studied subtype of male infertility regarding circRNAs, probably because it is the most well-defined. Therefore, more than half of the studies were about azoospermia. Most studies focus on the role of circRNAs in NOA. The circRNAs reported in the studies included in the present systematic review to be involved in NOA are presented in
Figure 6. Furthermore, the above studies confirm the continuous crosstalk of circRNAs and microRNAs. Usually, circRNAs act as miRNA inhibitors or sponges, and with this mechanism, they participate in regulating mRNA expression in various human diseases, including male infertility. As observed above, many cellular processes and pathways are deregulated by circRNA-miRNA interactions, leading to NOA pathogenesis. The development of biomarkers for screening patients undergoing micro-TESE was also investigated in NOA patients in the publication of Ji et al. (2021) [
46].
Finally, particular subtypes of azoospermia have also received attention, and specific circRNAs have been associated with them. For example, the main circRNAs associated with SCOS and the pathways deregulated by them are presented in
Figure 7.
It should be noted that
Figure 6 and
Figure 7 present circRNAs that were further investigated with extra experiments in the articles included in the present systematic review. These circRNAs were specifically selected based on the availability of more information provided by the above articles regarding their potential role, interactions, or mechanism of action related to male infertility. As a result, these figures serve as a summary of the main findings derived from these experiments described in the articles included in the present systematic review, focusing on these selected circRNAs.
Of the above circRNAs, hsa_circRNA_101373 has been reported in a study performed in rats and is associated with light-induced ovarian dysfunction [
57]. Some circRNAs have also been implicated to have a role in cancer, e.g., hsa_circ_0000277 is involved in esophageal cancer [
58,
59], whereas hsa_circ_0007773 has been indicated to inhibit the progression of hepatocellular carcinoma [
60]. Interestingly, the other circRNAs have not been associated with specific diseases or with the regulation of specific cellular processes.
Furthermore, some studies included in the present systematic review focused on the role of circRNAs in asthenozoospermia. Two studies [
36,
37] report that some of the DE circRNAs identified (circUSP54) are mainly associated with mitochondrial physiology and energy production. These results are consistent with previous studies that associate decreased sperm motility with defects in mitochondria, the powerhouse of the cell [
61,
62,
63]. Furthermore, the most important finding of these studies is that the circRNA expression pattern is characterized by high plasticity. circBoule RNA levels also exert lower levels in asthenozoospermic men, and these circRNAs play a role in heat stress-induced fertility decline [
40].
Regarding other subtypes of male infertility, only a preliminary study about circRNAs in OAZ was found [
39]. This was particularly focused on the circRNA profile of exosomes derived from seminal plasma. In this study, the important role of circRNA-microRNA-mRNA networks in male infertility is again highlighted, as in previous cases, but notably, the authors suggest that exosome-derived circRNAs could be used as potential noninvasive biomarkers. Of the identified DE circRNAs, hsa_circ_0003823 is associated with esophageal cancer, according to previous research [
64].
One more study included in the present systematic review was performed in a mixed sample of infertile men and supported the use of circRNAs found in seminal plasma extracellular vesicles as biomarkers of male infertility [
41]. In general, the diagnostic potential of circRNAs is supported by many studies regarding various diseases, as they exert high tissue specificity, are stable in body fluids, and are easily detected [
65].
Finally, one study [
53] was conducted on normozoospermic men, differentiating spermatozoa into two populations based on the Percoll gradient. They identified DE circRNAs between spermatozoa of high and poor quality, and furthermore, they indicate that circRNAs may be transmitted to the oocyte during fertilization. Such studies may shed light on cases of normozoospermic men that are unable to achieve pregnancy and require further research, highlighting that complex mechanisms are involved in male infertility.
It should also be noted that no common DE circRNAs were observed between the different studies or the different subtypes of male infertility. This may be due to the variability of the methods used in the above studies and draws attention to issues of comparability between sequencing and array-based technologies. In general, array-based methods are usually considered more prone to false positives, especially when combined with limited sample sizes [
41], and thus, the results of these studies should be interpreted with caution.
Based on the above and taking into consideration the methodology we followed, the present systematic review has some significant limitations. First, studies on animals (both experimental animals and livestock) were excluded because our main goal was to report DE circRNAs that have been identified and validated in humans and thus have a greater potential for clinical applications. However, with this approach, some information may have been overlooked. Another limitation of the present systematic review is that we were unable to retrieve every article. Specifically, despite our best efforts, we were unable to retrieve one article [
66]. This source may refer to an abstract of a conference presentation, which would be excluded from our systematic review according to our pre-defined exclusion criteria. However, since we were unable to find this article, there is a possibility that relevant information may have been overlooked. Furthermore, because we removed studies that were not published in English, we may have overlooked significant papers that were published in other languages. Another limitation of the present systematic review is that it focused solely on articles reporting DE circRNAs between men with varying fertility statuses (fertile or infertile). However, there are other four studies that, although not reporting DE circRNAs, offer valuable insights. More specifically, Dong et al. (2016) [
67] conducted a comprehensive profiling of circRNAs in the testicular tissues of healthy men and identified a large number of novel circRNAs. They further investigated the expression patterns and characteristics of these circRNAs. The study found that circRNAs derived from the human testis exhibited tissue-specific expression and were enriched in specific biological processes and pathways. Additionally, the researchers identified certain circRNAs that were detectable in the seminal plasma, indicating their potential as noninvasive biomarkers for male reproductive health. Similarly, Ragusa et al. (2019) [
68] investigated the presence and interaction of a specific circRNA, circNAPEPLD, in both human and murine spermatozoa. The study revealed the expression of circNAPEPLD in both human and murine spermatozoa through various experimental techniques, and the researchers found that circNAPEPLD physically interacts with miRNAs present in oocytes, suggesting a potential regulatory role during fertilization. These findings may shed light on the molecular mechanisms underlying reproductive processes and provide a basis for further investigation into the functional significance of circRNAs in fertility and fertilization. Tang et al. (2020) [
69] performed another type of study as they aimed to understand the regulatory mechanisms involved in circRNA biogenesis and the impact of m
6A modification on this process. More specifically, they identified specific m
6A-modified regions within precursor messenger RNAs (pre-mRNAs) that contribute to circRNA formation. By manipulating the m
6A machinery, they were able to modulate circRNA abundance, proving that these modifications play a crucial role in circRNA biogenesis in male germ cells. Lastly, Liu et al. (2020) [
70] investigated the effects of l-carnitine on the expression of miRNAs in the spermatozoa of individuals with asthenozoospermia. The study demonstrated that l-carnitine treatment influences the expression levels of specific miRNAs. By employing high-throughput sequencing and bioinformatics analysis, they identified the miRNAs that are differentially expressed upon l-carnitine treatment, and additionally, they explored the regulatory network involving these miRNAs and their associated molecules. Among other molecules, they report interactions with specific circRNAs. Thus, all the above may have led to the small number of studies included in the present systematic review jeopardizing solid conclusions, but in general, it is notable that this review has identified surprisingly little research conducted to date about circRNAs and male infertility.
Several limitations also arose from the existing studies included in this systematic review. One is that some studies do not present the full list of differentially expressed circRNAs, preventing the identification of common circRNAs between studies or circRNAs that are deregulated in many subtypes of male infertility. Second, the studies developed very different experimental designs and used different technologies, such as the sequencing- and array-based approaches mentioned above, preventing any comparative or cross-analysis. Third, one limitation is the variability in the definitions of differential expression used among the included studies. The studies included employed diverse fold change thresholds to determine differential expression of circRNAs. This variability in defining differential expression may introduce discrepancies and hinder direct comparison and interpretation of the results. Therefore, future research may benefit from establishing standardized guidelines for defining differential expression of circRNAs to enhance comparability and reproducibility in the field of male infertility. Fourth, some of the included studies used normozoospermic males as controls based on seminogram results rather than those confirmed fertile by previous pregnancy outcomes. For example, Oluwayiose et al. (2022) [
41] used as a control group normozoospermic men recruited from couples undergoing IVF. This can result in inconsistencies and limit the generalizability of findings. Furthermore, another limitation of this systematic review is the incomplete availability of information regarding both the mapping and circRNA calling processes in most of the included studies. Mapping involves aligning sequencing reads to a reference genome, while circRNA calling focuses on specifically identifying circular RNAs from the mapped reads. The mapping method and circRNA calling algorithms employed can significantly influence the identification and quantification of circRNAs. However, due to the lack of detailed information in some studies, it is challenging to assess the specific mapping methods and circRNA calling algorithms utilized, potentially introducing variability in circRNA detection and expression quantification. The variation in these methods across studies could impact the comparability and interpretation of the findings. Finally, the main limitation of most studies included is their small sample sizes. This can lead to unreliable results, and thus, validation in larger cohorts is required.
Despite the limitations described above, our study has key strengths. To our knowledge, this is the first systematic review investigating published research linking circRNAs to male fertility and summarizing the main findings of the existing literature. Furthermore, the use of PRISMA guidelines and the developed search strategy were intended to be as inclusive as possible by expanding the search to two renowned databases, such as PubMed and Web of Science. Therefore, we believe that this systematic review will be of particular interest to the life science community and clinical doctors engaged in the field of infertility. The list of DE circRNAs reported here will also help biologists unravel the role of circRNAs in male infertility and pave the way for future studies.
Regarding future research, compared with other noncoding RNAs, such as lncRNAs and miRNAs, studies of circRNAs in male infertility are just at the beginning. This systematic review points out a huge knowledge gap on the role of circRNAs in subtypes of male infertility other than azoospermia. There are no studies about circRNAs and specific sperm defects, such as teratozoospermia, oligozoospermia, or their combination; therefore, future studies should focus on these directions. In general, most of the circRNAs identified as deregulated have not been associated in previous studies with other diseases or molecular mechanisms, which implies that more research is needed to explore the exact biological functions of circRNAs and their role. Third, most of the interactions between circRNAs and miRNAs reported in the identified studies were predicted using bioinformatics models; therefore, more experimental studies are needed to investigate specific interactions and the networks of circRNAs. For diagnostic applications, more studies should also focus on the sensitivity and specificity of circRNAs applied to male infertility subtypes and evaluate their diagnostic value. Furthermore, existing studies in humans about circRNAs are mainly single experiments with a small number of subjects, which undermines the robustness of the results. Thus, it is essential to perform studies that use a large number of individuals or samples to validate previous findings or conduct new experiments focusing on circRNAs. Collaboration between institutions or research centers would be beneficial in this approach, but unfortunately, such collaborations have not been common in the studies carried out to date. Finally, some of the studies identified in the present systematic review suggest that spermatozoa are cells with unique epigenetic signatures characterized by unique and differential quantitative and qualitative fingerprints of circRNAs [
36]. This result is intriguing and raises questions about the real effectiveness of IVF techniques, such as the intracytoplasmic sperm injection (ICSI) procedure, which has been considered a useful approach to selecting good-quality spermatozoa for fertilization. It also raises the question of whether this unique epigenetic signature has an impact on the offspring, as the use of IVF is on the rise. Thus, further research is also required in this direction.