Extracellular Matrix Proteomics: The mdx-4cv Mouse Diaphragm as a Surrogate for Studying Myofibrosis in Dystrophinopathy
Abstract
:1. Introduction
2. Proteomic Strategies to Study the Extracellular Matrix
2.1. Bottom-Up Proteomics versus Top-Down Proteomics
2.2. Sample Handling and Protein Extraction for Gel-Free and Bottom-Up Proteomics
2.3. Sample Handling and Protein Extraction for Gel-Based and Top-Down Proteomics
2.4. Extracellular-Matrix-Enrichment-Based Proteomic-Profiling Approaches
2.5. Mass Spectrometry and Data Acquisition
3. The Extracellular Matrix of Skeletal Muscles
- The stabilization of skeletal muscle fibers, and the provision of tissue strength and elasticity;
- Structural support via physical scaffolding, and the provision of an embedding medium for motor units, capillaries, and pools of satellite cells;
- The provision of neuromuscular homeostasis;
- Mechanical force transduction from muscle fibers to the surrounding tissues;
- The provision of cell–matrix interactions for the support of cell adhesion and cell migration during embryonic myogenesis, adult myogenesis, and tissue repair;
- Cell–matrix support during cell differentiation, maturation, remodeling, fiber transitions, and muscle aging.
4. The mdx-4cv Mouse Model and Dystrophinopathy-Associated Myofibrosis
4.1. Duchenne Muscular Dystrophy and Fibrosis
4.2. Characterization of the mdx-4cv Mouse within the Context of Other Dystrophic Models
4.3. Histological and Biochemical Characterization of Fibrosis in the mdx-4cv Mouse Diaphragm
4.4. Proteomic Profiling of Fibrosis in the mdx-4cv Mouse Diaphragm
- (i)
- Basal-lamina-enriched extracellular matrix proteins:
- Collagen IV (COL4A1, COL4A2, COL4A3);
- Collagen XV (COL15A1);
- Collagen XVIII (COL18A1);
- Nidogen-2/Osteonidogen (NID2).
- (ii)
- Endomysium-enriched extracellular matrix proteins:
- Collagen V (COL5A1, COL5A2);
- Collagen VI (COL6A1, COL6A2, COL6A5, COL6A6).
- (iii)
- Matrisomal proteins of the endomysium, perimysium, and epimysium:
- Fibronectin (FN1);
- Vitronectin (VTN);
- Dermatopontin (DPT).
- (iv)
- Small leucine-rich proteoglycans:
- Asporin (ASPN);
- Biglycan (BGN);
- Decorin (DCN);
- Lumican (LUM);
- Mimecan/Osteoglycin (OGN).
- (v)
- Non-structural matricellular proteins:
- Periostin (POSTN).
- (vi)
- Extracellular-matrix-associated repair proteins:
- Annexin-2 (ANXA2);
- Annexin-6 (ANXA6).
- (vii)
- Adhesion receptors:
- Integrin, alpha-7 (ITGA7);
- Integrin, beta-1 (ITGB1);
- Dystroglycan, alpha/beta (DAG1).
- (viii)
- Myotendinous-junction-enriched proteins:
- Collagen XII (COL12A1).
- (ix)
- Tendon-enriched proteins:
- Collagen I (COL1A1, COL1A2).
5. Conclusions
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Mukherjee, P.; Roy, S.; Ghosh, D.; Nandi, S.K. Role of animal models in biomedical research: A review. Lab. Anim. Res. 2022, 38, 18. [Google Scholar] [CrossRef]
- Duan, D.; Goemans, N.; Takeda, S.; Mercuri, E.; Aartsma-Rus, A. Duchenne muscular dystrophy. Nat. Rev. Dis. Primers 2021, 7, 13. [Google Scholar] [CrossRef]
- McGreevy, J.W.; Hakim, C.H.; McIntosh, M.A.; Duan, D. Animal models of Duchenne muscular dystrophy: From basic mechanisms to gene therapy. Dis. Model. Mech. 2015, 8, 195–213. [Google Scholar] [CrossRef] [Green Version]
- Swiderski, K.; Lynch, G.S. Murine models of Duchenne muscular dystrophy: Is there a best model? Am. J. Physiol.-Cell Physiol. 2021, 321, C409–C412. [Google Scholar] [CrossRef]
- Stirm, M.; Fonteyne, L.M.; Shashikadze, B.; Stöckl, J.B.; Kurome, M.; Keßler, B.; Zakhartchenko, V.; Kemter, E.; Blum, H.; Arnold, G.J.; et al. Pig models for Duchenne muscular dystrophy—From disease mechanisms to validation of new diagnostic and therapeutic concepts. Neuromuscul. Disord. 2022, 32, 543–556. [Google Scholar] [CrossRef] [PubMed]
- Henrot, P.; Blervaque, L.; Dupin, I.; Zysman, M.; Esteves, P.; Gouzi, F.; Hayot, M.; Pomiès, P.; Berger, P. Cellular interplay in skeletal muscle regeneration and wasting: Insights from animal models. J. Cachexia Sarcopenia Muscle 2023, 14, 745–757. [Google Scholar] [CrossRef] [PubMed]
- Cohen, S.; Nathan, J.A.; Goldberg, A.L. Muscle wasting in disease: Molecular mechanisms and promising therapies. Nat. Rev. Drug Discov. 2015, 14, 58–74. [Google Scholar] [CrossRef]
- Henderson, N.C.; Rieder, F.; Wynn, T.A. Fibrosis: From mechanisms to medicines. Nature 2020, 587, 555–566. [Google Scholar] [CrossRef] [PubMed]
- Antar, S.A.; Ashour, N.A.; Marawan, M.E.; Al-Karmalawy, A.A. Fibrosis: Types, Effects, Markers, Mechanisms for Disease Progression, and Its Relation with Oxidative Stress, Immunity, and Inflammation. Int. J. Mol. Sci. 2023, 24, 4004. [Google Scholar] [CrossRef]
- Mahdy, M.A.A. Skeletal muscle fibrosis: An overview. Cell Tissue Res. 2019, 375, 575–588. [Google Scholar] [CrossRef]
- Smith, L.R.; Barton, E.R. Regulation of fibrosis in muscular dystrophy. Matrix Biol. 2018, 68–69, 602–615. [Google Scholar] [CrossRef] [PubMed]
- Lamandé, S.R.; Bateman, J.F. Collagen VI disorders: Insights on form and function in the extracellular matrix and beyond. Matrix Biol. 2018, 71–72, 348–367. [Google Scholar] [CrossRef] [PubMed]
- Zanotti, S.; Magri, F.; Salani, S.; Napoli, L.; Ripolone, M.; Ronchi, D.; Fortunato, F.; Ciscato, P.; Velardo, D.; D’Angelo, M.G.; et al. Extracellular Matrix Disorganization and Sarcolemmal Alterations in COL6-Related Myopathy Patients with New Variants of COL6 Genes. Int. J. Mol. Sci. 2023, 24, 5551. [Google Scholar] [CrossRef] [PubMed]
- Villar-Quiles, R.N.; Donkervoort, S.; de Becdelièvre, A.; Gartioux, C.; Jobic, V.; Foley, A.R.; McCarty, R.M.; Hu, Y.; Menassa, R.; Michel, L.; et al. Clinical and Molecular Spectrum Associated with COL6A3 c.7447A>G p.(Lys2483Glu) Variant: Elucidating its Role in Collagen VI-related Myopathies. J. Neuromuscul. Dis. 2021, 8, 633–645. [Google Scholar] [CrossRef]
- Klingler, W.; Jurkat-Rott, K.; Lehmann-Horn, F.; Schleip, R. The role of fibrosis in Duchenne muscular dystrophy. Acta Myol. 2012, 31, 184–195. [Google Scholar]
- Kharraz, Y.; Guerra, J.; Pessina, P.; Serrano, A.L.; Muñoz-Cánoves, P. Understanding the process of fibrosis in Duchenne muscular dystrophy. Biomed. Res. Int. 2014, 2014, 965631. [Google Scholar] [CrossRef] [Green Version]
- Ohlendieck, K.; Swandulla, D. Molecular pathogenesis of Duchenne muscular dystrophy-related fibrosis. Pathologe 2017, 38, 21–29. [Google Scholar] [CrossRef] [Green Version]
- Hoffman, E.P. The discovery of dystrophin, the protein product of the Duchenne muscular dystrophy gene. FEBS J. 2020, 287, 3879–3887. [Google Scholar] [CrossRef]
- Ohlendieck, K.; Swandulla, D. Complexity of skeletal muscle degeneration: Multi-systems pathophysiology and organ crosstalk in dystrophinopathy. Pflügers Arch. 2021, 473, 1813–1839. [Google Scholar] [CrossRef]
- Dowling, P.; Gargan, S.; Murphy, S.; Zweyer, M.; Sabir, H.; Swandulla, D.; Ohlendieck, K. The Dystrophin Node as Integrator of Cytoskeletal Organization, Lateral Force Transmission, Fiber Stability and Cellular Signaling in Skeletal Muscle. Proteomes 2021, 9, 9. [Google Scholar] [CrossRef]
- Wilson, D.G.S.; Tinker, A.; Iskratsch, T. The role of the dystrophin glycoprotein complex in muscle cell mechanotransduction. Commun. Biol. 2022, 5, 1022. [Google Scholar] [CrossRef] [PubMed]
- Zabłocka, B.; Górecki, D.C.; Zabłocki, K. Disrupted Calcium Homeostasis in Duchenne Muscular Dystrophy: A Common Mechanism behind Diverse Consequences. Int. J. Mol. Sci. 2021, 22, 11040. [Google Scholar] [CrossRef] [PubMed]
- Dowling, P.; Gargan, S.; Swandulla, D.; Ohlendieck, K. Proteomic profiling of impaired excitation-contraction coupling and abnormal calcium handling in muscular dystrophy. Proteomics 2022, 22, e2200003. [Google Scholar] [CrossRef]
- Dubinin, M.V.; Belosludtsev, K.N. Ion Channels of the Sarcolemma and Intracellular Organelles in Duchenne Muscular Dystrophy: A Role in the Dysregulation of Ion Homeostasis and a Possible Target for Therapy. Int. J. Mol. Sci. 2023, 24, 2229. [Google Scholar] [CrossRef]
- Rosenberg, A.S.; Puig, M.; Nagaraju, K.; Hoffman, E.P.; Villalta, S.A.; Rao, V.A.; Wakefield, L.M.; Woodcock, J. Immune-mediated pathology in Duchenne muscular dystrophy. Sci. Transl. Med. 2015, 7, 299rv4. [Google Scholar] [CrossRef] [Green Version]
- Serrano, A.L.; Muñoz-Cánoves, P. Fibrosis development in early-onset muscular dystrophies: Mechanisms and translational implications. Semin. Cell Dev. Biol. 2017, 64, 181–190. [Google Scholar] [CrossRef]
- Tidball, J.G.; Welc, S.S.; Wehling-Henricks, M. Immunobiology of Inherited Muscular Dystrophies. Compr. Physiol. 2018, 8, 1313–1356. [Google Scholar]
- Muraine, L.; Bensalah, M.; Butler-Browne, G.; Bigot, A.; Trollet, C.; Mouly, V.; Negroni, E. Update on anti-fibrotic pharmacotherapies in skeletal muscle disease. Curr. Opin. Pharmacol. 2023, 68, 102332. [Google Scholar] [CrossRef]
- Serrano, A.L.; Muñoz-Cánoves, P. Mouse Models of Muscle Fibrosis. Methods Mol. Biol. 2021, 2299, 357–370. [Google Scholar] [PubMed]
- Sleboda, D.A.; Stover, K.K.; Roberts, T.J. Diversity of extracellular matrix morphology in vertebrate skeletal muscle. J. Morphol. 2020, 281, 160–169. [Google Scholar] [CrossRef] [PubMed]
- Purslow, P.P. The Structure and Role of Intramuscular Connective Tissue in Muscle Function. Front. Physiol. 2020, 11, 495. [Google Scholar] [CrossRef] [PubMed]
- Schüler, S.C.; Liu, Y.; Dumontier, S.; Grandbois, M.; Le Moal, E.; Cornelison, D.; Bentzinger, C.F. Extracellular matrix: Brick and mortar in the skeletal muscle stem cell niche. Front. Cell Dev. Biol. 2022, 10, 1056523. [Google Scholar] [CrossRef]
- Moratal, C.; Arrighi, N.; Dechesne, C.A.; Dani, C. Control of Muscle Fibro-Adipogenic Progenitors by Myogenic Lineage is Altered in Aging and Duchenne Muscular Dystrophy. Cell. Physiol. Biochem. 2019, 53, 1029–1045. [Google Scholar]
- Molina, T.; Fabre, P.; Dumont, N.A. Fibro-adipogenic progenitors in skeletal muscle homeostasis, regeneration and diseases. Open Biol. 2021, 11, 210110. [Google Scholar] [CrossRef]
- Giuliani, G.; Rosina, M.; Reggio, A. Signaling pathways regulating the fate of fibro/adipogenic progenitors (FAPs) in skeletal muscle regeneration and disease. FEBS J. 2022, 289, 6484–6517. [Google Scholar] [CrossRef]
- Im, W.B.; Phelps, S.F.; Copen, E.H.; Adams, E.G.; Slightom, J.L.; Chamberlain, J.S. Differential expression of dystrophin isoforms in strains of mdx mice with different mutations. Hum. Mol. Genet. 1996, 5, 1149–1153. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Holland, A.; Murphy, S.; Dowling, P.; Ohlendieck, K. Pathoproteomic profiling of the skeletal muscle matrisome in dystrophinopathy associated myofibrosis. Proteomics 2016, 16, 345–366. [Google Scholar] [CrossRef] [PubMed]
- Bulfield, G.; Siller, W.G.; Wight, P.A.; Moore, K.J. X chromosome-linked muscular dystrophy (mdx) in the mouse. Proc. Natl. Acad. Sci. USA 1984, 81, 1189–1192. [Google Scholar] [CrossRef]
- Tanabe, Y.; Esaki, K.; Nomura, T. Skeletal muscle pathology in X chromosome-linked muscular dystrophy (mdx) mouse. Acta Neuropathol. 1986, 69, 91–95. [Google Scholar] [CrossRef]
- Partridge, T.A. The mdx mouse model as a surrogate for Duchenne muscular dystrophy. FEBS J. 2013, 280, 4177–4186. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chapman, V.M.; Miller, D.R.; Armstrong, D.; Caskey, C.T. Recovery of induced mutations for X chromosome-linked muscular dystrophy in mice. Proc. Natl. Acad. Sci. USA 1989, 86, 1292–1296. [Google Scholar] [CrossRef] [PubMed]
- Pearson-White, S.H. DMD(mdx3Cv) and DMD(mdx4Cv) dystrophin mutations in mice: Rapid polymerase chain reaction genotyping. Neuromuscul. Disord. 2002, 12, 366–370. [Google Scholar] [CrossRef] [PubMed]
- Banks, G.B.; Combs, A.C.; Chamberlain, J.S. Sequencing protocols to genotype mdx, mdx(4cv), and mdx(5cv) mice. Muscle Nerve 2010, 42, 268–270. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Shin, J.H.; Hakim, C.H.; Zhang, K.; Duan, D. Genotyping mdx, mdx3cv, and mdx4cv mice by primer competition polymerase chain reaction. Muscle Nerve 2011, 43, 283–286. [Google Scholar] [CrossRef] [Green Version]
- Danko, I.; Chapman, V.; Wolff, J.A. The frequency of revertants in mdx mouse genetic models for Duchenne muscular dystrophy. Pediatr. Res. 1992, 32, 128–131. [Google Scholar] [CrossRef] [Green Version]
- Gutpell, K.M.; Hrinivich, W.T.; Hoffman, L.M. Skeletal muscle fibrosis in the mdx/utrn+/- mouse validates its suitability as a murine model of Duchenne muscular dystrophy. PLoS ONE 2015, 10, e0117306. [Google Scholar] [CrossRef] [Green Version]
- Giovarelli, M.; Arnaboldi, F.; Zecchini, S.; Cornaghi, L.B.; Nava, A.; Sommariva, M.; Clementi, E.G.I.; Gagliano, N. Characterisation of Progressive Skeletal Muscle Fibrosis in the Mdx Mouse Model of Duchenne Muscular Dystrophy: An In Vivo and In Vitro Study. Int. J. Mol. Sci. 2022, 23, 8735. [Google Scholar] [CrossRef]
- Lazzarin, M.C.; Dos Santos, J.F.; Quintana, H.T.; Pidone, F.A.M.; de Oliveira, F. Duchenne muscular dystrophy progression induced by downhill running is accompanied by increased endomysial fibrosis and oxidative damage DNA in muscle of mdx mice. J. Mol. Histol. 2023, 54, 41–54. [Google Scholar] [CrossRef]
- Stedman, H.H.; Sweeney, H.L.; Shrager, J.B.; Maguire, H.C.; Panettieri, R.A.; Petrof, B.; Narusawa, M.; Leferovich, J.M.; Sladky, J.T.; Kelly, A.M. The mdx mouse diaphragm reproduces the degenerative changes of Duchenne muscular dystrophy. Nature 1991, 352, 536–539. [Google Scholar] [CrossRef]
- Sahani, R.; Wallace, C.H.; Jones, B.K.; Blemker, S.S. Diaphragm muscle fibrosis involves changes in collagen organization with mechanical implications in Duchenne muscular dystrophy. J. Appl. Physiol. 2022, 132, 653–672. [Google Scholar] [CrossRef]
- Wohlgemuth, R.P.; Feitzinger, R.M.; Henricson, K.E.; Dinh, D.T.; Brashear, S.E.; Smith, L.R. The extracellular matrix of dystrophic mouse diaphragm accounts for the majority of its passive stiffness and is resistant to collagenase digestion. Matrix Biol. Plus. 2023, 18, 100131. [Google Scholar] [CrossRef] [PubMed]
- Zweyer, M.; Sabir, H.; Dowling, P.; Gargan, S.; Murphy, S.; Swandulla, D.; Ohlendieck, K. Histopathology of Duchenne muscular dystrophy in correlation with changes in proteomic biomarkers. Histol. Histopathol. 2022, 37, 101–116. [Google Scholar]
- Dubuisson, N.; Versele, R.; Planchon, C.; Selvais, C.M.; Noel, L.; Abou-Samra, M.; Davis-López de Carrizosa, M.A. Histological Methods to Assess Skeletal Muscle Degeneration and Regeneration in Duchenne Muscular Dystrophy. Int. J. Mol. Sci. 2022, 23, 16080. [Google Scholar] [CrossRef]
- Desguerre, I.; Mayer, M.; Leturcq, F.; Barbet, J.P.; Gherardi, R.K.; Christov, C. Endomysial fibrosis in Duchenne muscular dystrophy: A marker of poor outcome associated with macrophage alternative activation. J. Neuropathol. Exp. Neurol. 2009, 68, 762–773. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gargan, S.; Dowling, P.; Zweyer, M.; Henry, M.; Meleady, P.; Swandulla, D.; Ohlendieck, K. Proteomic Identification of Markers of Membrane Repair, Regeneration and Fibrosis in the Aged and Dystrophic Diaphragm. Life 2022, 12, 1679. [Google Scholar] [CrossRef] [PubMed]
- Omenn, G.S. Reflections on the HUPO Human Proteome Project, the Flagship Project of the Human Proteome Organization, at 10 Years. Mol. Cell. Proteom. 2021, 20, 100062. [Google Scholar] [CrossRef] [PubMed]
- Omenn, G.S.; Lane, L.; Overall, C.M.; Pineau, C.; Packer, N.H.; Cristea, I.M.; Lindskog, C.; Weintraub, S.T.; Orchard, S.; Roehrl, M.H.A.; et al. The 2022 Report on the Human Proteome from the HUPO Human Proteome Project. J. Proteome Res. 2022, 22, 1024–1042. [Google Scholar] [CrossRef]
- Wilhelm, M.; Schlegl, J.; Hahne, H.; Gholami, A.M.; Lieberenz, M.; Savitski, M.M.; Ziegler, E.; Butzmann, L.; Gessulat, S.; Marx, H.; et al. Mass-spectrometry-based draft of the human proteome. Nature 2014, 509, 582–587. [Google Scholar] [CrossRef] [PubMed]
- Kim, M.S.; Pinto, S.M.; Getnet, D.; Nirujogi, R.S.; Manda, S.S.; Chaerkady, R.; Madugundu, A.K.; Kelkar, D.S.; Isserlin, R.; Jain, S.; et al. A draft map of the human proteome. Nature 2014, 509, 575–581. [Google Scholar] [CrossRef] [Green Version]
- Adhikari, S.; Nice, E.C.; Deutsch, E.W.; Lane, L.; Omenn, G.S.; Pennington, S.R.; Paik, Y.K.; Overall, C.M.; Corrales, F.J.; Cristea, I.M.; et al. A high-stringency blueprint of the human proteome. Nat. Commun. 2020, 11, 5301. [Google Scholar] [CrossRef]
- Højlund, K.; Yi, Z.; Hwang, H.; Bowen, B.; Lefort, N.; Flynn, C.R.; Langlais, P.; Weintraub, S.T.; Mandarino, L.J. Characterization of the human skeletal muscle proteome by one-dimensional gel electrophoresis and HPLC-ESI-MS/MS. Mol. Cell. Proteom. 2008, 7, 257–267. [Google Scholar] [CrossRef] [Green Version]
- Parker, K.C.; Walsh, R.J.; Salajegheh, M.; Amato, A.A.; Krastins, B.; Sarracino, D.A.; Greenberg, S.A. Characterization of human skeletal muscle biopsy samples using shotgun proteomics. J. Proteome Res. 2009, 8, 3265–3277. [Google Scholar] [CrossRef] [PubMed]
- Capitanio, D.; Moriggi, M.; Gelfi, C. Mapping the human skeletal muscle proteome: Progress and potential. Expert Rev. Proteom. 2017, 14, 825–839. [Google Scholar] [CrossRef]
- Schiaffino, S.; Reggiani, C.; Murgia, M. Fiber type diversity in skeletal muscle explored by mass spectrometry-based single fiber proteomics. Histol. Histopathol. 2020, 35, 239–246. [Google Scholar]
- Murgia, M.; Nogara, L.; Baraldo, M.; Reggiani, C.; Mann, M.; Schiaffino, S. Protein profile of fiber types in human skeletal muscle: A single-fiber proteomics study. Skelet. Muscle 2021, 11, 24. [Google Scholar] [CrossRef]
- Dowling, P.; Gargan, S.; Swandulla, D.; Ohlendieck, K. Identification of Subproteomic Markers for Skeletal Muscle Profiling. Methods Mol. Biol. 2023, 2596, 291–302. [Google Scholar]
- Randles, M.; Lennon, R. Applying Proteomics to Investigate Extracellular Matrix in Health and Disease. Curr. Top. Membr. 2015, 76, 171–196. [Google Scholar]
- Taha, I.N.; Naba, A. Exploring the extracellular matrix in health and disease using proteomics. Essays Biochem. 2019, 63, 417–432. [Google Scholar]
- Naba, A. Ten Years of Extracellular Matrix Proteomics: Accomplishments, Challenges, and Future Perspectives. Mol. Cell. Proteom. 2023, 22, 100528. [Google Scholar] [CrossRef] [PubMed]
- Hynes, R.O.; Naba, A. Overview of the matrisome—An inventory of extracellular matrix constituents and functions. Cold Spring Harb. Perspect. Biol. 2012, 4, a004903. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Naba, A.; Clauser, K.R.; Ding, H.; Whittaker, C.A.; Carr, S.A.; Hynes, R.O. The extracellular matrix: Tools and insights for the “omics” era. Matrix Biol. 2016, 49, 10–24. [Google Scholar] [CrossRef] [PubMed]
- Shao, X.; Taha, I.N.; Clauser, K.R.; Gao, Y.T.; Naba, A. MatrisomeDB: The ECM-protein knowledge database. Nucleic Acids Res. 2020, 48, D1136–D1144. [Google Scholar] [CrossRef] [PubMed]
- Shao, X.; Gomez, C.D.; Kapoor, N.; Considine, J.M.; Grams, C.; Gao, Y.T.; Naba, A. MatrisomeDB 2.0: 2023 updates to the ECM-protein knowledge database. Nucleic Acids Res. 2023, 51, D1519–D1530. [Google Scholar] [CrossRef] [PubMed]
- Krasny, L.; Paul, A.; Wai, P.; Howard, B.A.; Natrajan, R.C.; Huang, P.H. Comparative proteomic assessment of matrisome enrichment methodologies. Biochem. J. 2016, 473, 3979–3995. [Google Scholar] [CrossRef] [Green Version]
- Krasny, L.; Huang, P.H. Advances in the proteomic profiling of the matrisome and adhesome. Expert Rev. Proteom. 2021, 18, 781–794. [Google Scholar] [CrossRef]
- Dussoyer, M.; Page, A.; Delolme, F.; Rousselle, P.; Nyström, A.; Moali, C. Comparison of extracellular matrix enrichment protocols for the improved characterization of the skin matrisome by mass spectrometry. J. Proteom. 2022, 251, 104397. [Google Scholar] [CrossRef]
- Ercan, H.; Resch, U.; Hsu, F.; Mitulovic, G.; Bileck, A.; Gerner, C.; Yang, J.W.; Geiger, M.; Miller, I.; Zellner, M. A Practical and Analytical Comparative Study of Gel-Based Top-Down and Gel-Free Bottom-Up Proteomics Including Unbiased Proteoform Detection. Cells 2023, 12, 747. [Google Scholar] [CrossRef]
- Manes, N.P.; Nita-Lazar, A. Application of targeted mass spectrometry in bottom-up proteomics for systems biology research. J. Proteom. 2018, 189, 75–90. [Google Scholar] [CrossRef]
- Ohlendieck, K. Top-Down Proteomics and Comparative 2D-DIGE Analysis. Methods Mol. Biol. 2023, 2596, 19–38. [Google Scholar] [PubMed]
- Dowling, P.; Gargan, S.; Swandulla, D.; Ohlendieck, K. Fiber-Type Shifting in Sarcopenia of Old Age: Proteomic Profiling of the Contractile Apparatus of Skeletal Muscles. Int. J. Mol. Sci. 2023, 24, 2415. [Google Scholar] [CrossRef]
- Mansuri, M.S.; Williams, K.; Nairn, A.C. Uncovering biology by single-cell proteomics. Commun. Biol. 2023, 6, 381. [Google Scholar] [CrossRef] [PubMed]
- Dowling, P.; Murphy, S.; Ohlendieck, K. Proteomic profiling of muscle fibre type shifting in neuromuscular diseases. Expert Rev. Proteom. 2016, 13, 783–799. [Google Scholar] [CrossRef] [PubMed]
- Sielaff, M.; Kuharev, J.; Bohn, T.; Hahlbrock, J.; Bopp, T.; Tenzer, S.; Distler, U. Evaluation of FASP, SP3, and iST Protocols for Proteomic Sample Preparation in the Low Microgram Range. J. Proteome Res. 2017, 16, 4060–4072. [Google Scholar] [CrossRef]
- Wiśniewski, J.R.; Zougman, A.; Mann, M. Combination of FASP and StageTip-based fractionation allows in-depth analysis of the hippocampal membrane proteome. J. Proteome Res. 2009, 8, 5674–5678. [Google Scholar] [CrossRef]
- Xie, Z.; Feng, Q.; Zhang, S.; Yan, Y.; Deng, C.; Ding, C.F. Advances in proteomics sample preparation and enrichment for phosphorylation and glycosylation analysis. Proteomics 2022, 22, e2200070. [Google Scholar] [CrossRef]
- Wiśniewski, J.R.; Zougman, A.; Nagaraj, N.; Mann, M. Universal sample preparation method for proteome analysis. Nat. Methods 2009, 6, 359–362. [Google Scholar] [CrossRef]
- Wiśniewski, J.R. Filter Aided Sample Preparation—A tutorial. Anal. Chim. Acta 2019, 1090, 23–30. [Google Scholar] [CrossRef]
- Erde, J.; Loo, R.R.; Loo, J.A. Improving Proteome Coverage and Sample Recovery with Enhanced FASP (eFASP) for Quantitative Proteomic Experiments. Methods Mol. Biol. 2017, 1550, 11–18. [Google Scholar]
- Kulak, N.A.; Pichler, G.; Paron, I.; Nagaraj, N.; Mann, M. Minimal, encapsulated proteomic-sample processing applied to copy-number estimation in eukaryotic cells. Nat. Methods 2014, 11, 319–324. [Google Scholar] [CrossRef] [PubMed]
- Kostas, J.C.; Greguš, M.; Schejbal, J.; Ray, S.; Ivanov, A.R. Simple and Efficient Microsolid-Phase Extraction Tip-Based Sample Preparation Workflow to Enable Sensitive Proteomic Profiling of Limited Samples (200 to 10,000 Cells). J. Proteome Res. 2021, 20, 1676–1688. [Google Scholar] [CrossRef] [PubMed]
- Hughes, C.S.; Moggridge, S.; Müller, T.; Sorensen, P.H.; Morin, G.B.; Krijgsveld, J. Single-pot, solid-phase-enhanced sample preparation for proteomics experiments. Nat. Protoc. 2019, 14, 68–85. [Google Scholar] [CrossRef]
- Dagley, L.F.; Infusini, G.; Larsen, R.H.; Sandow, J.J.; Webb, A.I. Universal Solid-Phase Protein Preparation (USP3) for Bottom-up and Top-down Proteomics. J. Proteome Res. 2019, 18, 2915–2924. [Google Scholar] [CrossRef] [PubMed]
- Cai, X.; Xue, Z.; Wu, C.; Sun, R.; Qian, L.; Yue, L.; Ge, W.; Yi, X.; Liu, W.; Chen, C.; et al. High-throughput proteomic sample preparation using pressure cycling technology. Nat. Protoc. 2022, 17, 2307–2325. [Google Scholar] [CrossRef] [PubMed]
- Ma, F.; Liu, F.; Xu, W.; Li, L. Surfactant and Chaotropic Agent Assisted Sequential Extraction/On-Pellet Digestion (SCAD) for Enhanced Proteomics. J. Proteome Res. 2018, 17, 2744–2754. [Google Scholar] [CrossRef] [PubMed]
- Doellinger, J.; Schneider, A.; Hoeller, M.; Lasch, P. Sample Preparation by Easy Extraction and Digestion (SPEED)—A Universal, Rapid, and Detergent-free Protocol for Proteomics Based on Acid Extraction. Mol. Cell. Proteom. 2020, 19, 209–222. [Google Scholar] [CrossRef] [PubMed]
- Dowling, P.; Gargan, S.; Zweyer, M.; Henry, M.; Meleady, P.; Swandulla, D.; Ohlendieck, K. Protocol for the Bottom-Up Proteomic Analysis of Mouse Spleen. STAR Protoc. 2020, 1, 100196. [Google Scholar] [CrossRef]
- Murphy, S.; Ohlendieck, K. Proteomic profiling of large myofibrillar proteins from dried and long-term stored polyacrylamide gels. Anal. Biochem. 2018, 543, 8–11. [Google Scholar] [CrossRef] [PubMed]
- Jin, Y.; Wen, M.; Yuan, Q.; Zhang, J.; Tan, W. Beneficial effects of Coomassie staining on proteomic analysis employing PAGE separation followed with whole-gel slicing, in-gel digestion and quantitative LC-MS/MS. J. Chromatogr. B Analyt. Technol. Biomed. Life Sci. 2019, 1110–1111, 25–35. [Google Scholar] [CrossRef] [PubMed]
- Chen, C.; Wen, M.; Jin, Y. 1DE-MS Profiling for Proteoform-Correlated Proteomic Analysis, by Combining SDS-PAGE, Whole-Gel Slicing, Quantitative LC-MS/MS, and Reconstruction of Gel Distributions of Several Thousands of Proteins. J. Proteome Res. 2022, 21, 2311–2330. [Google Scholar] [CrossRef] [PubMed]
- Dowling, P.; Zweyer, M.; Swandulla, D.; Ohlendieck, K. Characterization of Contractile Proteins from Skeletal Muscle Using Gel-Based Top-Down Proteomics. Proteomes 2019, 7, 25. [Google Scholar] [CrossRef] [Green Version]
- Zhan, X.; Li, B.; Zhan, X.; Schlüter, H.; Jungblut, P.R.; Coorssen, J.R. Innovating the Concept and Practice of Two-Dimensional Gel Electrophoresis in the Analysis of Proteomes at the Proteoform Level. Proteomes 2019, 7, 36. [Google Scholar] [CrossRef] [Green Version]
- Carbonara, K.; Padula, M.P.; Coorssen, J.R. Quantitative assessment confirms deep proteome analysis by integrative top-down proteomics. Electrophoresis 2023, 44, 472–480. [Google Scholar] [CrossRef]
- Arentz, G.; Weiland, F.; Oehler, M.K.; Hoffmann, P. State of the art of 2D DIGE. Proteom. Clin. Appl. 2015, 9, 277–288. [Google Scholar] [CrossRef] [PubMed]
- Blundon, M.; Ganesan, V.; Redler, B.; Van, P.T.; Minden, J.S. Two-Dimensional Difference Gel Electrophoresis. Methods Mol. Biol. 2019, 1855, 229–247. [Google Scholar]
- Ohlendieck, K. Comparative 3-Sample 2D-DIGE Analysis of Skeletal Muscles. Methods Mol. Biol. 2023, 2596, 127–146. [Google Scholar] [PubMed]
- Naba, A.; Clauser, K.R.; Hoersch, S.; Liu, H.; Carr, S.A.; Hynes, R.O. The matrisome: In silico definition and in vivo characterization by proteomics of normal and tumor extracellular matrices. Mol. Cell. Proteom. 2012, 11, M111.014647. [Google Scholar] [CrossRef] [Green Version]
- Hill, R.C.; Calle, E.A.; Dzieciatkowska, M.; Niklason, L.E.; Hansen, K.C. Quantification of extracellular matrix proteins from a rat lung scaffold to provide a molecular readout for tissue engineering. Mol. Cell. Proteom. 2015, 14, 961–973. [Google Scholar] [CrossRef] [Green Version]
- Schiller, H.B.; Fernandez, I.E.; Burgstaller, G.; Schaab, C.; Scheltema, R.A.; Schwarzmayr, T.; Strom, T.M.; Eickelberg, O.; Mann, M. Time- and compartment-resolved proteome profiling of the extracellular niche in lung injury and repair. Mol. Syst. Biol. 2015, 11, 819. [Google Scholar] [CrossRef] [PubMed]
- Brown, K.A.; Chen, B.; Guardado-Alvarez, T.M.; Lin, Z.; Hwang, L.; Ayaz-Guner, S.; Jin, S.; Ge, Y. A photocleavable surfactant for top-down proteomics. Nat. Methods 2019, 16, 417–420. [Google Scholar] [CrossRef]
- Knott, S.J.; Brown, K.A.; Josyer, H.; Carr, A.; Inman, D.; Jin, S.; Friedl, A.; Ponik, S.M.; Ge, Y. Photocleavable Surfactant-Enabled Extracellular Matrix Proteomics. Anal. Chem. 2020, 92, 15693–15698. [Google Scholar] [CrossRef] [PubMed]
- Gilbert, T.W.; Sellaro, T.L.; Badylak, S.F. Decellularization of tissues and organs. Biomaterials 2006, 27, 3675–3683. [Google Scholar] [CrossRef]
- Fernández-Pérez, J.; Ahearne, M. The impact of decellularization methods on extracellular matrix derived hydrogels. Sci. Rep. 2019, 9, 14933. [Google Scholar] [CrossRef] [Green Version]
- Mendibil, U.; Ruiz-Hernandez, R.; Retegi-Carrion, S.; Garcia-Urquia, N.; Olalde-Graells, B.; Abarrategi, A. Tissue-Specific Decellularization Methods: Rationale and Strategies to Achieve Regenerative Compounds. Int. J. Mol. Sci. 2020, 21, 5447. [Google Scholar] [CrossRef] [PubMed]
- Domon, B.; Aebersold, R. Mass spectrometry and protein analysis. Science 2006, 312, 212–217. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhang, Y.; Fonslow, B.R.; Shan, B.; Baek, M.C.; Yates, J.R., 3rd. Protein analysis by shotgun/bottom-up proteomics. Chem. Rev. 2013, 113, 2343–2394. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yates, J.R.; Ruse, C.I.; Nakorchevsky, A. Proteomics by mass spectrometry: Approaches, advances, and applications. Annu. Rev. Biomed. Eng. 2009, 11, 49–79. [Google Scholar] [CrossRef] [Green Version]
- Yates, J.R., 3rd. Recent technical advances in proteomics. F1000Research 2019, 8, 351. [Google Scholar] [CrossRef] [Green Version]
- Nesvizhskii, A.I.; Vitek, O.; Aebersold, R. Analysis and validation of proteomic data generated by tandem mass spectrometry. Nat. Methods 2007, 4, 787–797. [Google Scholar] [CrossRef] [PubMed]
- Evangelista, A.J.; Ferreira, T.L. Matrix-assisted laser desorption/ionization time-of-flight mass spectrometry in the diagnosis of microorganisms. Future Microbiol. 2022, 17, 1409–1419. [Google Scholar] [CrossRef]
- Sabidó, E.; Selevsek, N.; Aebersold, R. Mass spectrometry-based proteomics for systems biology. Curr. Opin. Biotechnol. 2012, 23, 591–597. [Google Scholar] [CrossRef]
- Rauniyar, N.; Yates, J.R., 3rd. Isobaric labeling-based relative quantification in shotgun proteomics. J. Proteome Res. 2014, 13, 5293–5309. [Google Scholar] [CrossRef] [Green Version]
- Beller, N.C.; Hummon, A.B. Advances in stable isotope labeling: Dynamic labeling for spatial and temporal proteomic analysis. Mol. Omics 2022, 18, 579–590. [Google Scholar] [CrossRef] [PubMed]
- Chen, X.; Sun, Y.; Zhang, T.; Shu, L.; Roepstorff, P.; Yang, F. Quantitative Proteomics Using Isobaric Labeling: A Practical Guide. Genom. Proteom. Bioinform. 2021, 19, 689–706. [Google Scholar] [CrossRef]
- Chahrour, O.; Cobice, D.; Malone, J. Stable isotope labelling methods in mass spectrometry-based quantitative proteomics. J. Pharm. Biomed. Anal. 2015, 113, 2–20. [Google Scholar] [CrossRef] [PubMed]
- Xing, T.; Wang, C.; Zhao, X.; Dai, C.; Zhou, G.; Xu, X. Proteome Analysis Using Isobaric Tags for Relative and Absolute Analysis Quantitation (iTRAQ) Reveals Alterations in Stress-Induced Dysfunctional Chicken Muscle. J. Agric. Food Chem. 2017, 65, 2913–2922. [Google Scholar] [CrossRef]
- Aebersold, R.; Mann, M. Mass-spectrometric exploration of proteome structure and function. Nature 2016, 537, 347–355. [Google Scholar] [CrossRef] [PubMed]
- Sobsey, C.A.; Ibrahim, S.; Richard, V.R.; Gaspar, V.; Mitsa, G.; Lacasse, V.; Zahedi, R.P.; Batist, G.; Borchers, C.H. Targeted and Untargeted Proteomics Approaches in Biomarker Development. Proteomics 2020, 20, e1900029. [Google Scholar] [CrossRef] [PubMed]
- Mann, M.; Kumar, C.; Zeng, W.F.; Strauss, M.T. Artificial intelligence for proteomics and biomarker discovery. Cell Syst. 2021, 12, 759–770. [Google Scholar] [CrossRef] [PubMed]
- Fenaille, F.; Barbier Saint-Hilaire, P.; Rousseau, K.; Junot, C. Data acquisition workflows in liquid chromatography coupled to high resolution mass spectrometry-based metabolomics: Where do we stand? J. Chromatogr. A 2017, 1526, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Kitata, R.B.; Yang, J.C.; Chen, Y.J. Advances in data-independent acquisition mass spectrometry towards comprehensive digital proteome landscape. Mass Spectrom. Rev. 2022, e21781. [Google Scholar] [CrossRef]
- Abdollahi, M.; Segura, P.A.; Beaudry, F. Is nontargeted data acquisition for target analysis (nDATA) in mass spectrometry a forward-thinking analytical approach? Biomed. Chromatogr. 2023, 37, e5531. [Google Scholar] [CrossRef] [PubMed]
- Krasny, L.; Bland, P.; Kogata, N.; Wai, P.; Howard, B.A.; Natrajan, R.C.; Huang, P.H. SWATH mass spectrometry as a tool for quantitative profiling of the matrisome. J. Proteom. 2018, 189, 11–22. [Google Scholar] [CrossRef]
- Wood, A.J.; Lin, C.H.; Li, M.; Nishtala, K.; Alaei, S.; Rossello, F.; Sonntag, C.; Hersey, L.; Miles, L.B.; Krisp, C.; et al. FKRP-dependent glycosylation of fibronectin regulates muscle pathology in muscular dystrophy. Nat. Commun. 2021, 12, 2951. [Google Scholar] [CrossRef] [PubMed]
- Kontostathi, G.; Makridakis, M.; Bitsika, V.; Tsolakos, N.; Vlahou, A.; Zoidakis, J. Development and Validation of Multiple Reaction Monitoring (MRM) Assays for Clinical Applications. Methods Mol. Biol. 2019, 1959, 205–223. [Google Scholar] [PubMed]
- Cho, B.G.; Gutierrez Reyes, C.D.; Goli, M.; Gautam, S.; Banazadeh, A.; Mechref, Y. Targeted N-Glycan Analysis with Parallel Reaction Monitoring Using a Quadrupole-Orbitrap Hybrid Mass Spectrometer. Anal. Chem. 2022, 94, 15215–15222. [Google Scholar] [CrossRef]
- Thorsen, A.F.; Riber, L.P.S.; Rasmussen, L.M.; Overgaard, M. A targeted multiplex mass spectrometry method for quantitation of abundant matrix and cellular proteins in formalin-fixed paraffin embedded arterial tissue. J. Proteom. 2023, 272, 104775. [Google Scholar] [CrossRef]
- Maaß, S. Absolute Protein Quantification Using AQUA-Calibrated 2D-PAGE. Methods Mol. Biol. 2018, 1841, 141–162. [Google Scholar]
- Gillies, A.R.; Lieber, R.L. Structure and function of the skeletal muscle extracellular matrix. Muscle Nerve 2011, 44, 318–331. [Google Scholar] [CrossRef] [Green Version]
- Lieber, R.L.; Meyer, G. Structure-Function relationships in the skeletal muscle extracellular matrix. J. Biomech. 2023, 152, 111593. [Google Scholar] [CrossRef]
- Carbonara, K.; Andonovski, M.; Coorssen, J.R. Proteomes Are of Proteoforms: Embracing the Complexity. Proteomes 2021, 9, 38. [Google Scholar] [CrossRef] [PubMed]
- Schaffer, L.V.; Millikin, R.J.; Miller, R.M.; Anderson, L.C.; Fellers, R.T.; Ge, Y.; Kelleher, N.L.; LeDuc, R.D.; Liu, X.; Payne, S.H.; et al. Identification and Quantification of Proteoforms by Mass Spectrometry. Proteomics 2019, 19, e1800361. [Google Scholar] [CrossRef] [PubMed]
- Schneider, M.; Al-Shareffi, E.; Haltiwanger, R.S. Biological functions of fucose in mammals. Glycobiology 2017, 27, 601–618. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Basak, T.; Vega-Montoto, L.; Zimmerman, L.J.; Tabb, D.L.; Hudson, B.G.; Vanacore, R.M. Comprehensive Characterization of Glycosylation and Hydroxylation of Basement Membrane Collagen IV by High-Resolution Mass Spectrometry. J. Proteome Res. 2016, 15, 245–258. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Galligan, J.J.; Petersen, D.R. The human protein disulfide isomerase gene family. Hum. Genom. 2012, 6, 6. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Stefanelli, V.L.; Choudhury, S.; Hu, P.; Liu, Y.; Schwenzer, A.; Yeh, C.R.; Chambers, D.M.; von Beck, K.; Li, W.; Segura, T.; et al. Citrullination of fibronectin alters integrin clustering and focal adhesion stability promoting stromal cell invasion. Matrix Biol. 2019, 82, 86–104. [Google Scholar] [CrossRef] [PubMed]
- Murphy, S.; Zweyerm, M.; Raucamp, M.; Henry, M.; Meleady, P.; Swandulla, D.; Ohlendieck, K. Proteomic profiling of the mouse diaphragm and refined mass spectrometric analysis of the dystrophic phenotype. J. Muscle Res. Cell Motil. 2019, 40, 9–28. [Google Scholar] [CrossRef] [PubMed]
- Halper, J. Basic Components of Connective Tissues and Extracellular Matrix: Fibronectin, Fibrinogen, Laminin, Elastin, Fibrillins, Fibulins, Matrilins, Tenascins and Thrombospondins. Adv. Exp. Med. Biol. 2021, 1348, 105–126. [Google Scholar] [PubMed]
- Holmberg, J.; Durbeej, M. Laminin-211 in skeletal muscle function. Cell Adhes. Migr. 2013, 7, 111–121. [Google Scholar] [CrossRef] [Green Version]
- Goddi, A.; Schroedl, L.; Brey, E.M.; Cohen, R.N. Laminins in metabolic tissues. Metabolism 2021, 120, 154775. [Google Scholar] [CrossRef]
- Rogers, R.S.; Nishimune, H. The role of laminins in the organization and function of neuromuscular junctions. Matrix Biol. 2017, 57–58, 86–105. [Google Scholar] [CrossRef] [Green Version]
- Singhal, N.; Martin, P.T. Role of extracellular matrix proteins and their receptors in the development of the vertebrate neuromuscular junction. Dev. Neurobiol. 2011, 71, 982–1005. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kadler, K.E.; Baldock, C.; Bella, J.; Boot-Handford, R.P. Collagens at a glance. J. Cell Sci. 2007, 120, 1955–1958. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Goody, M.F.; Sher, R.B.; Henry, C.A. Hanging on for the ride: Adhesion to the extracellular matrix mediates cellular responses in skeletal muscle morphogenesis and disease. Dev. Biol. 2015, 401, 75–91. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Grefte, S.; Adjobo-Hermans, M.J.W.; Versteeg, E.M.M.; Koopman, W.J.H.; Daamen, W.F. Impaired primary mouse myotube formation on crosslinked type I collagen films is enhanced by laminin and entactin. Acta Biomater. 2016, 30, 265–276. [Google Scholar] [CrossRef] [PubMed]
- Fox, M.A.; Ho, M.S.; Smyth, N.; Sanes, J.R. A synaptic nidogen: Developmental regulation and role of nidogen-2 at the neuromuscular junction. Neural Dev. 2008, 3, 24. [Google Scholar] [CrossRef] [Green Version]
- Ricard-Blum, S. The collagen family. Cold Spring Harb. Perspect. Biol. 2011, 3, a004978. [Google Scholar] [CrossRef] [Green Version]
- Mienaltowski, M.J.; Gonzales, N.L.; Beall, J.M.; Pechanec, M.Y. Basic Structure, Physiology, and Biochemistry of Connective Tissues and Extracellular Matrix Collagens. Adv. Exp. Med. Biol. 2021, 1348, 5–43. [Google Scholar]
- Cescon, M.; Gattazzo, F.; Chen, P.; Bonaldo, P. Collagen VI at a glance. J. Cell Sci. 2015, 128, 3525–3531. [Google Scholar] [CrossRef] [Green Version]
- Cescon, M.; Gregorio, I.; Eiber, N.; Borgia, D.; Fusto, A.; Sabatelli, P.; Scorzeto, M.; Megighian, A.; Pegoraro, E.; Hashemolhosseini, S.; et al. Collagen VI is required for the structural and functional integrity of the neuromuscular junction. Acta Neuropathol. 2018, 136, 483–499. [Google Scholar] [CrossRef] [PubMed]
- Hu, M.; Ling, Z.; Ren, X. Extracellular matrix dynamics: Tracking in biological systems and their implications. J. Biol. Eng. 2022, 16, 13. [Google Scholar] [CrossRef]
- Zheng, Z.; Granado, H.S.; Li, C. Fibromodulin, a Multifunctional Matricellular Modulator. J. Dent. Res. 2023, 102, 125–134. [Google Scholar] [CrossRef] [PubMed]
- Vanhoutte, D.; Schips, T.G.; Kwong, J.Q.; Davis, J.; Tjondrokoesoemo, A.; Brody, M.J.; Sargent, M.A.; Kanisicak, O.; Yi, H.; Gao, Q.Q.; et al. Thrombospondin expression in myofibers stabilizes muscle membranes. Elife 2016, 5, e17589. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.; An, J.; Zhu, D.; Chen, H.; Lin, A.; Kang, J.; Liu, W.; Kang, X. Periostin: An emerging activator of multiple signaling pathways. J. Cell. Commun. Signal. 2022, 16, 515–530. [Google Scholar] [CrossRef]
- Boppart, M.D.; Mahmassani, Z.S. Integrin signaling: Linking mechanical stimulation to skeletal muscle hypertrophy. Am. J. Physiol.-Cell Physiol. 2019, 317, C629–C641. [Google Scholar] [CrossRef]
- Ibraghimov-Beskrovnaya, O.; Ervasti, J.M.; Leveille, C.J.; Slaughter, C.A.; Sernett, S.W.; Campbell, K.P. Primary structure of dystrophin-associated glycoproteins linking dystrophin to the extracellular matrix. Nature 1992, 355, 696–702. [Google Scholar] [CrossRef]
- Ohlendieck, K. Towards an understanding of the dystrophin-glycoprotein complex: Linkage between the extracellular matrix and the membrane cytoskeleton in muscle fibers. Eur. J. Cell Biol. 1996, 69, 1–10. [Google Scholar]
- Murphy, S.; Ohlendieck, K. The biochemical and mass spectrometric profiling of the dystrophin complexome from skeletal muscle. Comput. Struct. Biotechnol. J. 2015, 14, 20–27. [Google Scholar] [CrossRef] [Green Version]
- Sciandra, F.; Bozzi, M.; Bigotti, M.G.; Brancaccio, A. The multiple affinities of α-dystroglycan. Curr. Protein Pept. Sci. 2013, 14, 626–634. [Google Scholar] [CrossRef] [PubMed]
- Sciandra, F.; Gawlik, K.I.; Brancaccio, A.; Durbeej, M. Dystroglycan: A possible mediator for reducing congenital muscular dystrophy? Trends Biotechnol. 2007, 25, 262–268. [Google Scholar] [CrossRef] [PubMed]
- Jacobson, K.R.; Lipp, S.; Acuna, A.; Leng, Y.; Bu, Y.; Calve, S. Comparative Analysis of the Extracellular Matrix Proteome across the Myotendinous Junction. J. Proteome Res. 2020, 19, 3955–3967. [Google Scholar] [CrossRef] [PubMed]
- Izu, Y.; Adams, S.M.; Connizzo, B.K.; Beason, D.P.; Soslowsky, L.J.; Koch, M.; Birk, D.E. Collagen XII mediated cellular and extracellular mechanisms regulate establishment of tendon structure and function. Matrix Biol. 2021, 95, 52–67. [Google Scholar] [CrossRef]
- Karlsen, A.; Gonzalez-Franquesa, A.; Jakobsen, J.R.; Krogsgaard, M.R.; Koch, M.; Kjaer, M.; Schiaffino, S.; Mackey, A.L.; Deshmukh, A.S. The proteomic profile of the human myotendinous junction. iScience 2022, 25, 103836. [Google Scholar] [CrossRef]
- Franchi, M.; Trirè, A.; Quaranta, M.; Orsini, E.; Ottani, V. Collagen structure of tendon relates to function. Sci. World J. 2007, 7, 404–420. [Google Scholar] [CrossRef] [Green Version]
- Croissant, C.; Gounou, C.; Bouvet, F.; Tan, S.; Bouter, A. Annexin-A6 in Membrane Repair of Human Skeletal Muscle Cell: A Role in the Cap Subdomain. Cells 2020, 9, 1742. [Google Scholar] [CrossRef]
- Vidak, E.; Javoršek, U.; Vizovišek, M.; Turk, B. Cysteine Cathepsins and their Extracellular Roles: Shaping the Microenvironment. Cells 2019, 8, 264. [Google Scholar] [CrossRef] [Green Version]
- Spence, M.A.; Mortimer, M.D.; Buckle, A.M.; Minh, B.Q.; Jackson, C.J. A Comprehensive Phylogenetic Analysis of the Serpin Superfamily. Mol. Biol. Evol. 2021, 38, 2915–2929. [Google Scholar] [CrossRef]
- Zhang, W.; Liu, Y.; Zhang, H. Extracellular matrix: An important regulator of cell functions and skeletal muscle development. Cell Biosci. 2021, 11, 65. [Google Scholar] [CrossRef]
- Szklarczyk, D.; Gable, A.L.; Nastou, K.C.; Lyon, D.; Kirsch, R.; Pyysalo, S.; Doncheva, N.T.; Legeay, M.; Fang, T.; Bork, P.; et al. The STRING database in 2021: Customizable protein-protein networks, and functional characterization of user-uploaded gene/measurement sets. Nucleic Acids Res. 2021, 49, D605–D612. [Google Scholar] [CrossRef]
- Guiraud, S.; Aartsma-Rus, A.; Vieira, N.M.; Davies, K.E.; van Ommen, G.J.; Kunkel, L.M. The Pathogenesis and Therapy of Muscular Dystrophies. Annu. Rev. Genom. Hum. Genet. 2015, 16, 281–308. [Google Scholar] [CrossRef] [Green Version]
- Crisafulli, S.; Sultana, J.; Fontana, A.; Salvo, F.; Messina, S.; Trifirò, G. Global epidemiology of Duchenne muscular dystrophy: An updated systematic review and meta-analysis. Orphanet J. Rare Dis. 2020, 15, 141. [Google Scholar] [CrossRef]
- Bladen, C.L.; Salgado, D.; Monges, S.; Foncuberta, M.E.; Kekou, K.; Kosma, K.; Dawkins, H.; Lamont, L.; Roy, A.J.; Chamova, T.; et al. The TREAT-NMD DMD Global Database: Analysis of more than 7,000 Duchenne muscular dystrophy mutations. Hum. Mutat. 2015, 36, 395–402. [Google Scholar] [CrossRef] [Green Version]
- Doglio, L.; Pavan, E.; Pernigotti, I.; Petralia, P.; Frigo, C.; Minetti, C. Early signs of gait deviation in Duchenne muscular dystrophy. Eur. J. Phys. Rehabil. Med. 2011, 47, 587–594. [Google Scholar] [PubMed]
- Migliorelli, C.; Gómez-Martinez, M.; Subías-Beltrán, P.; Claramunt-Molet, M.; Idelsohn-Zielonka, S.; Mas-Hurtado, E.; Miralles, F.; Montolio, M.; Roselló-Ruano, M.; Medina-Cantillo, J. Multidimensional Biomechanics-Based Score to Assess Disease Progression in Duchenne Muscular Dystrophy. Sensors 2023, 23, 831. [Google Scholar] [CrossRef]
- Sienko, S.; Buckon, C.; Bagley, A.; Fowler, E.; Heberer, K.; Staudt, L.; Sison-Williamson, M.; McDonald, C.; Sussman, M. Kinematic changes in gait in boys with Duchenne Muscular Dystrophy: Utility of the Gait Deviation Index, the Gait Profile Score and the Gait Variable Scores. Gait Posture 2023, 100, 157–164. [Google Scholar] [CrossRef]
- Meyers, T.A.; Townsend, D. Cardiac Pathophysiology and the Future of Cardiac Therapies in Duchenne Muscular Dystrophy. Int. J. Mol. Sci. 2019, 20, 4098. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Power, L.C.; O’Grady, G.L.; Hornung, T.S.; Jefferies, C.; Gusso, S.; Hofman, P.L. Imaging the heart to detect cardiomyopathy in Duchenne muscular dystrophy: A review. Neuromuscul. Disord. 2018, 28, 717–730. [Google Scholar] [CrossRef]
- Vianello, A.; Arcaro, G.; Guarnieri, G.; Sukthi, A.; Molena, B.; Turato, C.; Braccioni, F.; Gallan, F.; Lugato, F.; Turrin, M.; et al. Non-Invasive Ventilation for Acute Respiratory Failure in Duchenne Muscular Dystrophy Patients. Arch. Bronconeumol. 2021, 57, 666–668. [Google Scholar] [CrossRef] [PubMed]
- Birnkrant, D.J.; Bello, L.; Butterfield, R.J.; Carter, J.C.; Cripe, L.H.; Cripe, T.P.; McKim, D.A.; Nandi, D.; Pegoraro, E. Cardiorespiratory management of Duchenne muscular dystrophy: Emerging therapies, neuromuscular genetics, and new clinical challenges. Lancet Respir. Med. 2022, 10, 403–420. [Google Scholar] [CrossRef]
- Tsaknakis, K.; Jäckle, K.; Lüders, K.A.; Lorenz, H.M.; Braunschweig, L.; Hell, A.K. Reduced bone mineral density in adolescents with Duchenne Muscular Dystrophy (DMD) and scoliosis. Osteoporos. Int. 2022, 33, 2011–2018. [Google Scholar] [CrossRef]
- Conway, K.M.; Gedlinske, A.; Mathews, K.D.; Perlman, S.; Johnson, N.; Butterfield, R.; Hung, M.; Bounsanga, J.; Matthews, D.; Oleszek, J.; et al. A population-based study of scoliosis among males diagnosed with a dystrophinopathy identified by the Muscular Dystrophy Surveillance, Tracking, and Research Network (MD STARnet). Muscle Nerve 2022, 65, 193–202. [Google Scholar] [CrossRef]
- Messina, S.; Vita, G.L. Clinical management of Duchenne muscular dystrophy: The state of the art. Neurol. Sci. 2018, 39, 1837–1845. [Google Scholar] [CrossRef]
- Snow, W.M.; Anderson, J.E.; Jakobson, L.S. Neuropsychological and neurobehavioral functioning in Duchenne muscular dystrophy: A review. Neurosci. Biobehav. Rev. 2013, 37, 743–752. [Google Scholar] [CrossRef]
- Chieffo, D.P.R.; Moriconi, F.; Pane, M.; Lucibello, S.; Ferraroli, E.; Norcia, G.; Ricci, M.; Capasso, A.; Cicala, G.; Buchignani, B.; et al. A Longitudinal Follow-Up Study of Intellectual Function in Duchenne Muscular Dystrophy over Age: Is It Really Stable? J. Clin. Med. 2023, 12, 403. [Google Scholar] [CrossRef]
- Pascual-Morena, C.; Cavero-Redondo, I.; Álvarez-Bueno, C.; Jiménez-López, E.; Saz-Lara, A.; Martínez-García, I.; Martínez-Vizcaíno, V. Global prevalence of intellectual developmental disorder in dystrophinopathies: A systematic review and meta-analysis. Dev. Med. Child Neurol. 2023, 65, 734–744. [Google Scholar] [CrossRef] [PubMed]
- Birnkrant, D.J.; Bushby, K.; Bann, C.M.; Apkon, S.D.; Blackwell, A.; Brumbaugh, D.; Case, L.E.; Clemens, P.R.; Hadjiyannakis, S.; Pandya, S.; et al. Diagnosis and management of Duchenne muscular dystrophy, part 1: Diagnosis, and neuromuscular, rehabilitation, endocrine, and gastrointestinal and nutritional management. Lancet Neurol. 2018, 17, 251–267. [Google Scholar] [CrossRef] [Green Version]
- Birnkrant, D.J.; Bushby, K.; Bann, C.M.; Alman, B.A.; Apkon, S.D.; Blackwell, A.; Case, L.E.; Cripe, L.; Hadjiyannakis, S.; Olson, A.K.; et al. Diagnosis and management of Duchenne muscular dystrophy, part 2: Respiratory, cardiac, bone health, and orthopaedic management. Lancet Neurol. 2018, 17, 347–361. [Google Scholar] [CrossRef] [Green Version]
- Birnkrant, D.J.; Bushby, K.; Bann, C.M.; Apkon, S.D.; Blackwell, A.; Colvin, M.K.; Cripe, L.; Herron, A.R.; Kennedy, A.; Kinnett, K.; et al. Diagnosis and management of Duchenne muscular dystrophy, part 3: Primary care, emergency management, psychosocial care, and transitions of care across the lifespan. Lancet Neurol. 2018, 17, 445–455. [Google Scholar] [CrossRef] [Green Version]
- Koenig, M.; Hoffman, E.P.; Bertelson, C.J.; Monaco, A.P.; Feener, C.; Kunkel, L.M. Complete cloning of the Duchenne muscular dystrophy (DMD) cDNA and preliminary genomic organization of the DMD gene in normal and affected individuals. Cell 1987, 50, 509–517. [Google Scholar] [CrossRef] [PubMed]
- Forrest, S.M.; Cross, G.S.; Flint, T.; Speer, A.; Robson, K.J.; Davies, K.E. Further studies of gene deletions that cause Duchenne and Becker muscular dystrophies. Genomics 1988, 2, 109–114. [Google Scholar] [CrossRef] [PubMed]
- Kunkel, L.M.; Hoffman, E.P. Duchenne/Becker muscular dystrophy: A short overview of the gene, the protein, and current diagnostics. Br. Med. Bull. 1989, 45, 630–643. [Google Scholar] [CrossRef] [PubMed]
- Hoffman, E.P.; Brown, R.H., Jr.; Kunkel, L.M. Dystrophin: The protein product of the Duchenne muscular dystrophy locus. Cell 1987, 51, 919–928. [Google Scholar] [CrossRef]
- Hoffman, E.P.; Hudecki, M.S.; Rosenberg, P.A.; Pollina, C.M.; Kunkel, L.M. Cell and fiber-type distribution of dystrophin. Neuron 1988, 1, 411–420. [Google Scholar] [CrossRef]
- Bonilla, E.; Samitt, C.E.; Miranda, A.F.; Hays, A.P.; Salviati, G.; DiMauro, S.; Kunkel, L.M.; Hoffman, E.P.; Rowland, L.P. Duchenne muscular dystrophy: Deficiency of dystrophin at the muscle cell surface. Cell 1988, 54, 447–452. [Google Scholar] [CrossRef]
- Ohlendieck, K.; Matsumura, K.; Ionasescu, V.V.; Towbin, J.A.; Bosch, E.P.; Weinstein, S.L.; Sernett, S.W.; Campbell, K.P. Duchenne muscular dystrophy: Deficiency of dystrophin-associated proteins in the sarcolemma. Neurology 1993, 43, 795–800. [Google Scholar] [CrossRef] [PubMed]
- Lieber, R.L.; Ward, S.R. Cellular mechanisms of tissue fibrosis. 4. Structural and functional consequences of skeletal muscle fibrosis. Am. J. Physiol.-Cell Physiol. 2013, 305, C241–C252. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Levi, O.; Genin, O.; Angelini, C.; Halevy, O.; Pines, M. Inhibition of muscle fibrosis results in increases in both utrophin levels and the number of revertant myofibers in Duchenne muscular dystrophy. Oncotarget 2015, 6, 23249–23260. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zanotti, S.; Bragato, C.; Zucchella, A.; Maggi, L.; Mantegazza, R.; Morandi, L.; Mora, M. Anti-fibrotic effect of pirfenidone in muscle derived-fibroblasts from Duchenne muscular dystrophy patients. Life Sci. 2016, 145, 127–136. [Google Scholar] [CrossRef] [PubMed]
- Miyatake, S.; Shimizu-Motohashi, Y.; Takeda, S.; Aoki, Y. Anti-inflammatory drugs for Duchenne muscular dystrophy: Focus on skeletal muscle-releasing factors. Drug Des. Devel. Ther. 2016, 10, 2745–2758. [Google Scholar] [PubMed] [Green Version]
- March, J.T.; Golshirazi, G.; Cernisova, V.; Carr, H.; Leong, Y.; Lu-Nguyen, N.; Popplewell, L.J. Targeting TGFβ Signaling to Address Fibrosis Using Antisense Oligonucleotides. Biomedicines 2018, 6, 74. [Google Scholar] [CrossRef] [Green Version]
- Durbeej, M.; Campbell, K.P. Muscular dystrophies involving the dystrophin-glycoprotein complex: An overview of current mouse models. Curr. Opin. Genet. Dev. 2002, 12, 349–361. [Google Scholar] [CrossRef]
- Sciandra, F.; Bigotti, M.G.; Giardina, B.; Bozzi, M.; Brancaccio, A. Genetic Engineering of Dystroglycan in Animal Models of Muscular Dystrophy. Biomed. Res. Int. 2015, 2015, 635792. [Google Scholar] [CrossRef] [Green Version]
- Wilson, K.; Faelan, C.; Patterson-Kane, J.C.; Rudmann, D.G.; Moore, S.A.; Frank, D.; Charleston, J.; Tinsley, J.; Young, G.D.; Milici, A.J. Duchenne and Becker Muscular Dystrophies: A Review of Animal Models, Clinical End Points, and Biomarker Quantification. Toxicol. Pathol. 2017, 45, 961–976. [Google Scholar] [CrossRef] [Green Version]
- van Putten, M.; Lloyd, E.M.; de Greef, J.C.; Raz, V.; Willmann, R.; Grounds, M.D. Mouse models for muscular dystrophies: An overview. Dis. Models Mech. 2020, 13, dmm043562. [Google Scholar] [CrossRef] [Green Version]
- Sztretye, M.; Szabó, L.; Dobrosi, N.; Fodor, J.; Szentesi, P.; Almássy, J.; Magyar, Z.É.; Dienes, B.; Csernoch, L. From Mice to Humans: An Overview of the Potentials and Limitations of Current Transgenic Mouse Models of Major Muscular Dystrophies and Congenital Myopathies. Int. J. Mol. Sci. 2020, 21, 8935. [Google Scholar] [CrossRef]
- Zaynitdinova, M.I.; Lavrov, A.V.; Smirnikhina, S.A. Animal models for researching approaches to therapy of Duchenne muscular dystrophy. Transgenic Res. 2021, 30, 709–725. [Google Scholar] [CrossRef]
- Yucel, N.; Chang, A.C.; Day, J.W.; Rosenthal, N.; Blau, H.M. Humanizing the mdx mouse model of DMD: The long and the short of it. NPJ Regen. Med. 2018, 3, 4. [Google Scholar] [CrossRef] [Green Version]
- Holland, A.; Ohlendieck, K. Proteomic profiling of the dystrophin-deficient mdx phenocopy of dystrophinopathy-associated cardiomyopathy. Biomed. Res. Int. 2014, 2014, 246195. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Holland, A.; Dowling, P.; Zweyer, M.; Swandulla, D.; Henry, M.; Clynes, M.; Ohlendieck, K. Proteomic profiling of cardiomyopathic tissue from the aged mdx model of Duchenne muscular dystrophy reveals a drastic decrease in laminin, nidogen and annexin. Proteomics 2013, 13, 2312–2323. [Google Scholar] [CrossRef]
- Yu, X.; Bao, B.; Echigoya, Y.; Yokota, T. Dystrophin-deficient large animal models: Translational research and exon skipping. Am. J. Transl. Res. 2015, 7, 1314–1331. [Google Scholar] [PubMed]
- Kornegay, J.N. The golden retriever model of Duchenne muscular dystrophy. Skelet. Muscle 2017, 7, 9. [Google Scholar] [CrossRef]
- Duan, D. Duchenne muscular dystrophy gene therapy in the canine model. Hum. Gene Ther. Clin. Dev. 2015, 26, 57–69. [Google Scholar] [CrossRef] [PubMed]
- Cassano, M.; Berardi, E.; Crippa, S.; Toelen, J.; Barthelemy, I.; Micheletti, R.; Chuah, M.; Vandendriessche, T.; Debyser, Z.; Blot, S.; et al. Alteration of cardiac progenitor cell potency in GRMD dogs. Cell Transpl. 2012, 21, 1945–1967. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lardenois, A.; Jagot, S.; Lagarrigue, M.; Guével, B.; Ledevin, M.; Larcher, T.; Dubreil, L.; Pineau, C.; Rouger, K.; Guével, L. Quantitative proteome profiling of dystrophic dog skeletal muscle reveals a stabilized muscular architecture and protection against oxidative stress after systemic delivery of MuStem cells. Proteomics 2016, 16, 2028–2042. [Google Scholar] [CrossRef]
- Tamiyakul, H.; Kemter, E.; Kösters, M.; Ebner, S.; Blutke, A.; Klymiuk, N.; Flenkenthaler, F.; Wolf, E.; Arnold, G.J.; Fröhlich, T. Progressive Proteome Changes in the Myocardium of a Pig Model for Duchenne Muscular Dystrophy. iScience 2020, 23, 101516. [Google Scholar] [CrossRef] [PubMed]
- Regensburger, A.P.; Fonteyne, L.M.; Jüngert, J.; Wagner, A.L.; Gerhalter, T.; Nagel, A.M.; Heiss, R.; Flenkenthaler, F.; Qurashi, M.; Neurath, M.F.; et al. Detection of collagens by multispectral optoacoustic tomography as an imaging biomarker for Duchenne muscular dystrophy. Nat. Med. 2019, 25, 1905–1915. [Google Scholar] [CrossRef]
- Smith, L.R.; Hammers, D.W.; Sweeney, H.L.; Barton, E. Increased collagen cross-linking is a signature of dystrophin-deficient muscle. Muscle Nerve 2016, 54, 71–78. [Google Scholar] [CrossRef] [Green Version]
- Dupont-Versteegden, E.E.; McCarter, R.J. Differential expression of muscular dystrophy in diaphragm versus hindlimb muscles of mdx mice. Muscle Nerve 1992, 15, 1105–1110. [Google Scholar] [CrossRef]
- Ritter, P.; Nübler, S.; Buttgereit, A.; Smith, L.R.; Mühlberg, A.; Bauer, J.; Michael, M.; Kreiß, L.; Haug, M.; Barton, E.; et al. Myofibrillar Lattice Remodeling Is a Structural Cytoskeletal Predictor of Diaphragm Muscle Weakness in a Fibrotic mdx (mdx Cmah-/-) Model. Int. J. Mol. Sci. 2022, 23, 10841. [Google Scholar] [CrossRef]
- Murphy, S.; Zweyer, M.; Mundegar, R.R.; Henry, M.; Meleady, P.; Swandulla, D.; Ohlendieck, K. Concurrent Label-Free Mass Spectrometric Analysis of Dystrophin Isoform Dp427 and the Myofibrosis Marker Collagen in Crude Extracts from mdx-4cv Skeletal Muscles. Proteomes 2015, 3, 298–327. [Google Scholar] [CrossRef] [Green Version]
- Murphy, S.; Zweyer, M.; Henry, M.; Meleady, P.; Mundegar, R.R.; Swandulla, D.; Ohlendieck, K. Proteomic analysis of the sarcolemma-enriched fraction from dystrophic mdx-4cv skeletal muscle. J. Proteom. 2019, 191, 212–227. [Google Scholar] [CrossRef]
- Judge, L.M.; Haraguchiln, M.; Chamberlain, J.S. Dissecting the signaling and mechanical functions of the dystrophin-glycoprotein complex. J. Cell Sci. 2006, 119, 1537–1546. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Arecco, N.; Clarke, C.J.; Jones, F.K.; Simpson, D.M.; Mason, D.; Beynon, R.J.; Pisconti, A. Elastase levels and activity are increased in dystrophic muscle and impair myoblast cell survival, proliferation and differentiation. Sci. Rep. 2016, 6, 24708. [Google Scholar] [CrossRef] [Green Version]
- Abdulhady, H.; Sakr, H.M.; Elsayed, N.S.; El-Sobky, T.A.; Fahmy, N.; Saadawy, A.M.; Elsedfy, H. Ambulatory Duchenne muscular dystrophy children: Cross-sectional correlation between function, quantitative muscle ultrasound and MRI. Acta Myol. 2022, 41, 1–14. [Google Scholar] [PubMed]
- Peng, F.; Xu, H.; Song, Y.; Xu, K.; Li, S.; Cai, X.; Guo, Y.; Gong, L. Utilization of T1-Mapping for the pelvic and thigh muscles in Duchenne Muscular Dystrophy: A quantitative biomarker for disease involvement and correlation with clinical assessments. BMC Musculoskelet. Disord. 2022, 23, 681. [Google Scholar] [CrossRef] [PubMed]
- Veeger, T.T.J.; van de Velde, N.M.; Keene, K.R.; Niks, E.H.; Hooijmans, M.T.; Webb, A.G.; de Groot, J.H.; Kan, H.E. Baseline fat fraction is a strong predictor of disease progression in Becker muscular dystrophy. NMR Biomed. 2022, 35, e4691. [Google Scholar] [CrossRef]
- Barnard, A.M.; Hammers, D.W.; Triplett, W.T.; Kim, S.; Forbes, S.C.; Willcocks, R.J.; Daniels, M.J.; Senesac, C.R.; Lott, D.J.; Arpan, I.; et al. Evaluating Genetic Modifiers of Duchenne Muscular Dystrophy Disease Progression Using Modeling and MRI. Neurology 2022, 99, e2406–e2416. [Google Scholar] [CrossRef]
- Tichy, E.D.; Mourkioti, F. A new method of genotyping MDX4CV mice by PCR-RFLP analysis. Muscle Nerve 2017, 56, 522–524. [Google Scholar] [CrossRef]
- Mitrpant, C.; Fletcher, S.; Iversen, P.L.; Wilton, S.D. By-passing the nonsense mutation in the 4 CV mouse model of muscular dystrophy by induced exon skipping. J. Gene Med. 2009, 11, 46–56. [Google Scholar] [CrossRef]
- Li, S.; Kimura, E.; Ng, R.; Fall, B.M.; Meuse, L.; Reyes, M.; Faulkner, J.A.; Chamberlain, J.S. A highly functional mini-dystrophin/GFP fusion gene for cell and gene therapy studies of Duchenne muscular dystrophy. Hum. Mol. Genet. 2006, 15, 1610–1622. [Google Scholar] [CrossRef] [PubMed]
- Kimura, E.; Li, S.; Gregorevic, P.; Fall, B.M.; Chamberlain, J.S. Dystrophin delivery to muscles of mdx mice using lentiviral vectors leads to myogenic progenitor targeting and stable gene expression. Mol. Ther. 2010, 18, 206–213. [Google Scholar] [CrossRef]
- Ramos, J.N.; Hollinger, K.; Bengtsson, N.E.; Allen, J.M.; Hauschka, S.D.; Chamberlain, J.S. Development of Novel Micro-dystrophins with Enhanced Functionality. Mol. Ther. 2019, 27, 623–635. [Google Scholar] [CrossRef] [Green Version]
- Hersh, J.; Condor Capcha, J.M.; Iansen Irion, C.; Lambert, G.; Noguera, M.; Singh, M.; Kaur, A.; Dikici, E.; Jiménez, J.J.; Shehadeh, L.A.; et al. Peptide-Functionalized Dendrimer Nanocarriers for Targeted Microdystrophin Gene Delivery. Pharmaceutics 2021, 13, 2159. [Google Scholar] [CrossRef]
- Eren, S.A.; Tastan, C.; Karadeniz, K.B.; Turan, R.D.; Cakirsoy, D.; Kancagi, D.D.; Yilmaz, S.U.; Oztatlici, M.; Oztatlici, H.; Ozer, S.; et al. Lentiviral micro-dystrophin gene treatment into late-stage mdx mice for Duchene Muscular Dystrophy disease. Curr. Gene Ther. 2023, 23, 304–315. [Google Scholar] [CrossRef] [PubMed]
- Kimura, E.; Han, J.J.; Li, S.; Fall, B.; Ra, J.; Haraguchi, M.; Tapscott, S.J.; Chamberlain, J.S. Cell-lineage regulated myogenesis for dystrophin replacement: A novel therapeutic approach for treatment of muscular dystrophy. Hum. Mol. Genet. 2008, 17, 2507–2517. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Muir, L.A.; Nguyen, Q.G.; Hauschka, S.D.; Chamberlain, J.S. Engraftment potential of dermal fibroblasts following in vivo myogenic conversion in immunocompetent dystrophic skeletal muscle. Mol. Ther. Methods Clin. Dev. 2014, 1, 14025. [Google Scholar] [CrossRef]
- Stuelsatz, P.; Shearer, A.; Li, Y.; Muir, L.A.; Ieronimakis, N.; Shen, Q.W.; Kirillova, I.; Yablonka-Reuveni, Z. Extraocular muscle satellite cells are high performance myo-engines retaining efficient regenerative capacity in dystrophin deficiency. Dev. Biol. 2015, 397, 31–44. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bengtsson, N.E.; Hall, J.K.; Odom, G.L.; Phelps, M.P.; Andrus, C.R.; Hawkins, R.D.; Hauschka, S.D.; Chamberlain, J.R.; Chamberlain, J.S. Muscle-specific CRISPR/Cas9 dystrophin gene editing ameliorates pathophysiology in a mouse model for Duchenne muscular dystrophy. Nat. Commun. 2017, 8, 14454. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Xu, L.; Zhang, C.; Li, H.; Wang, P.; Gao, Y.; Mokadam, N.A.; Ma, J.; Arnold, W.D.; Han, R. Efficient precise in vivo base editing in adult dystrophic mice. Nat. Commun. 2021, 12, 3719. [Google Scholar] [CrossRef]
- Murphy, S.; Zweyer, M.; Mundegar, R.R.; Swandulla, D.; Ohlendieck, K. Comparative gel-based proteomic analysis of chemically crosslinked complexes in dystrophic skeletal muscle. Electrophoresis 2018, 39, 1735–1744. [Google Scholar] [CrossRef] [Green Version]
- Murphy, S.; Zweyer, M.; Mundegar, R.R.; Swandulla, D.; Ohlendieck, K. Chemical crosslinking analysis of β-dystroglycan in dystrophin-deficient skeletal muscle. HRB Open Res. 2018, 1, 17. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Murphy, S.; Zweyer, M.; Henry, M.; Meleady, P.; Mundegar, R.R.; Swandulla, D.; Ohlendieck, K. Subproteomic profiling of sarcolemma from dystrophic mdx-4cv skeletal muscle. Data Brief. 2018, 17, 980–993. [Google Scholar] [CrossRef] [PubMed]
- Gargan, S.; Dowling, P.; Zweyer, M.; Reimann, J.; Henry, M.; Meleady, P.; Swandulla, D.; Ohlendieck, K. Mass Spectrometric Profiling of Extraocular Muscle and Proteomic Adaptations in the mdx-4cv Model of Duchenne Muscular Dystrophy. Life 2021, 11, 595. [Google Scholar] [CrossRef] [PubMed]
- Murphy, S.; Dowling, P.; Zweyer, M.; Mundegar, R.R.; Henry, M.; Meleady, P.; Swandulla, D.; Ohlendieck, K. Proteomic analysis of dystrophin deficiency and associated changes in the aged mdx-4cv heart model of dystrophinopathy-related cardiomyopathy. J. Proteom. 2016, 145, 24–36. [Google Scholar] [CrossRef]
- Murphy, S.; Zweyer, M.; Henry, M.; Meleady, P.; Mundegar, R.R.; Swandulla, D.; Ohlendieck, K. Proteomic profiling of liver tissue from the mdx-4cv mouse model of Duchenne muscular dystrophy. Clin. Proteom. 2018, 15, 34. [Google Scholar] [CrossRef] [PubMed]
- Dowling, P.; Zweyer, M.; Raucamp, M.; Henry, M.; Meleady, P.; Swandulla, D.; Ohlendieck, K. Proteomic and cell biological profiling of the renal phenotype of the mdx-4cv mouse model of Duchenne muscular dystrophy. Eur. J. Cell Biol. 2020, 99, 151059. [Google Scholar] [CrossRef]
- Dowling, P.; Zweyer, M.; Raucamp, M.; Henry, M.; Meleady, P.; Swandulla, D.; Ohlendieck, K. Dataset on the mass spectrometry-based proteomic profiling of the kidney from wild type and the dystrophic mdx-4cv mouse model of X-linked muscular dystrophy. Data Brief. 2020, 28, 105067. [Google Scholar] [CrossRef] [PubMed]
- Dowling, P.; Gargan, S.; Zweyer, M.; Henry, M.; Meleady, P.; Swandulla, D.; Ohlendieck, K. Proteome-wide Changes in the mdx-4cv Spleen due to Pathophysiological Cross Talk with Dystrophin-Deficient Skeletal Muscle. iScience 2020, 23, 101500. [Google Scholar] [CrossRef]
- Dowling, P.; Gargan, S.; Zweyer, M.; Sabir, H.; Henry, M.; Meleady, P.; Swandulla, D.; Ohlendieck, K. Proteomic profiling of the interface between the stomach wall and the pancreas in dystrophinopathy. Eur. J. Transl. Myol. 2021, 31, 9627. [Google Scholar]
- Murphy, S.; Zweyer, M.; Henry, M.; Meleady, P.; Mundegar, R.R.; Swandulla, D.; Ohlendieck, K. Label-free mass spectrometric analysis reveals complex changes in the brain proteome from the mdx-4cv mouse model of Duchenne muscular dystrophy. Clin. Proteom. 2015, 12, 27. [Google Scholar] [CrossRef] [Green Version]
- Murphy, S.; Dowling, P.; Zweyer, M.; Henry, M.; Meleady, P.; Mundegar, R.R.; Swandulla, D.; Ohlendieck, K. Proteomic profiling of mdx-4cv serum reveals highly elevated levels of the inflammation-induced plasma marker haptoglobin in muscular dystrophy. Int. J. Mol. Med. 2017, 39, 1357–1370. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Murphy, S.; Zweyer, M.; Mundegar, R.R.; Swandulla, D.; Ohlendieck, K. Dataset on the comparative proteomic profiling of mouse saliva and serum from wild type versus the dystrophic mdx-4cv mouse model of dystrophinopathy. Data Brief. 2018, 21, 1236–1245. [Google Scholar] [CrossRef] [PubMed]
- Gargan, S.; Dowling, P.; Zweyer, M.; Swandulla, D.; Ohlendieck, K. Identification of marker proteins of muscular dystrophy in the urine proteome from the mdx-4cv model of dystrophinopathy. Mol. Omics 2020, 16, 268–278. [Google Scholar] [CrossRef]
- Murphy, S.; Zweyer, M.; Mundegar, R.R.; Swandulla, D.; Ohlendieck, K. Proteomic identification of elevated saliva kallikrein levels in the mdx-4cv mouse model of Duchenne muscular dystrophy. Biochem. Biophys. Rep. 2018, 18, 100541. [Google Scholar] [CrossRef] [PubMed]
- Wu, J.; Matthias, N.; Lo, J.; Ortiz-Vitali, J.L.; Shieh, A.W.; Wang, S.H.; Darabi, R. A Myogenic Double-Reporter Human Pluripotent Stem Cell Line Allows Prospective Isolation of Skeletal Muscle Progenitors. Cell Rep. 2018, 25, 1966–1981.e4. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sakai-Takemura, F.; Narita, A.; Masuda, S.; Wakamatsu, T.; Watanabe, N.; Nishiyama, T.; Nogami, K.; Blanc, M.; Takeda, S.; Miyagoe-Suzuki, Y. Premyogenic progenitors derived from human pluripotent stem cells expand in floating culture and differentiate into transplantable myogenic progenitors. Sci. Rep. 2018, 8, 6555. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Arpke, R.W.; Darabi, R.; Mader, T.L.; Zhang, Y.; Toyama, A.; Lonetree, C.L.; Nash, N.; Lowe, D.A.; Perlingeiro, R.C.; Kyba, M. A new immuno-, dystrophin-deficient model, the NSG-mdx(4Cv) mouse, provides evidence for functional improvement following allogeneic satellite cell transplantation. Stem Cells 2013, 31, 1611–1620. [Google Scholar] [CrossRef] [Green Version]
- Elhussieny, A.; Nogami, K.; Sakai-Takemura, F.; Maruyama, Y.; Takemura, N.; Soliman, W.T.; Takeda, S.; Miyagoe-Suzuki, Y. Mesenchymal stem cells derived from human induced pluripotent stem cells improve the engraftment of myogenic cells by secreting urokinase-type plasminogen activator receptor (uPAR). Stem Cell Res. Ther. 2021, 12, 532. [Google Scholar] [CrossRef]
- Chang, A.C.; Ong, S.G.; LaGory, E.L.; Kraft, P.E.; Giaccia, A.J.; Wu, J.C.; Blau, H.M. Telomere shortening and metabolic compromise underlie dystrophic cardiomyopathy. Proc. Natl. Acad. Sci. USA 2016, 113, 13120–13125. [Google Scholar] [CrossRef]
- Ziemba, M.; Barkhouse, M.; Uaesoontrachoon, K.; Giri, M.; Hathout, Y.; Dang, U.J.; Gordish-Dressman, H.; Nagaraju, K.; Hoffman, E.P. Biomarker-focused multi-drug combination therapy and repurposing trial in mdx mice. PLoS ONE 2021, 16, e0246507. [Google Scholar] [CrossRef]
- Deng, J.; Zhang, J.; Shi, K.; Liu, Z. Drug development progress in Duchenne muscular dystrophy. Front. Pharmacol. 2022, 13, 950651. [Google Scholar] [CrossRef]
- Eser, G.; Topaloğlu, H. Current Outline of Exon Skipping Trials in Duchenne Muscular Dystrophy. Genes 2022, 13, 1241. [Google Scholar] [CrossRef] [PubMed]
- Chung Liang, L.; Sulaiman, N.; Yazid, M.D. A Decade of Progress in Gene Targeted Therapeutic Strategies in Duchenne Muscular Dystrophy: A Systematic Review. Front. Bioeng. Biotechnol. 2022, 10, 833833. [Google Scholar] [CrossRef]
- Chemello, F.; Olson, E.N.; Bassel-Duby, R. Crispr-Editing Therapy For Duchenne Muscular Dystrophy. Hum. Gene Ther. 2023, 34, 379–387. [Google Scholar] [CrossRef] [PubMed]
- Patterson, G.; Conner, H.; Groneman, M.; Blavo, C.; Parmar, M.S. Duchenne muscular dystrophy: Current treatment and emerging exon skipping and gene therapy approach. Eur. J. Pharmacol. 2023, 947, 175675. [Google Scholar] [CrossRef]
- Heydemann, A.; Bieganski, G.; Wachowiak, J.; Czarnota, J.; Niezgoda, A.; Siemionow, K.; Ziemiecka, A.; Sikorska, M.H.; Bozyk, K.; Tullius, S.G.; et al. Dystrophin Expressing Chimeric (DEC) Cell Therapy for Duchenne Muscular Dystrophy: A First-in-Human Study with Minimum 6 Months Follow-up. Stem Cell Rev. Rep. 2023. [Google Scholar] [CrossRef] [PubMed]
- Barnard, A.M.; Lott, D.J.; Batra, A.; Triplett, W.T.; Forbes, S.C.; Riehl, S.L.; Willcocks, R.J.; Smith, B.K.; Vandenborne, K.; Walter, G.A. Imaging respiratory muscle quality and function in Duchenne muscular dystrophy. J. Neurol. 2019, 266, 2752–2763. [Google Scholar] [CrossRef] [PubMed]
- Boussaïd, G.; Stalens, C.; Devaux, C.; Segovia-Kueny, S.; Lofaso, F.; Reveillere, C. Impact of Mechanical Ventilation Methods on the Life Perception of Subjects With Duchenne Muscular Dystrophy: French Cross-Sectional Survey. Respir. Care 2020, 65, 1712–1720. [Google Scholar] [CrossRef]
- Kipke, J.; Birnkrant, D.J.; Jin, J.B.; Aneja, A.; Bahler, R.C. A systematic review of pharmacologic therapies for the cardiomyopathy of Duchenne muscular dystrophy. Pediatr. Pulmonol. 2021, 56, 782–795. [Google Scholar] [CrossRef]
- Ricotti, V.; Selby, V.; Ridout, D.; Domingos, J.; Decostre, V.; Mayhew, A.; Eagle, M.; Butler, J.; Guglieri, M.; Van der Holst, M.; et al. Respiratory and upper limb function as outcome measures in ambulant and non-ambulant subjects with Duchenne muscular dystrophy: A prospective multicentre study. Neuromuscul. Disord. 2019, 29, 261–268. [Google Scholar] [CrossRef]
- Fayssoil, A.; Chaffaut, C.; Ogna, A.; Stojkovic, T.; Lamothe, L.; Mompoint, D.; Meng, P.; Prigent, H.; Clair, B.; Behin, A.; et al. Echographic Assessment of Diaphragmatic Function in Duchenne Muscular Dystrophy from Childhood to Adulthood. J. Neuromuscul. Dis. 2019, 6, 55–64. [Google Scholar] [CrossRef] [PubMed]
- Beastrom, N.; Lu, H.; Macke, A.; Canan, B.D.; Johnson, E.K.; Penton, C.M.; Kaspar, B.K.; Rodino-Klapac, L.R.; Zhou, L.; Janssen, P.M.; et al. mdx(⁵cv) mice manifest more severe muscle dysfunction and diaphragm force deficits than do mdx Mice. Am. J. Pathol. 2011, 179, 2464–2474. [Google Scholar] [CrossRef] [PubMed]
- Holland, A.; Henry, M.; Meleady, P.; Winkler, C.K.; Krautwald, M.; Brinkmeier, H.; Ohlendieck, K. Comparative Label-Free Mass Spectrometric Analysis of Mildly versus Severely Affected mdx Mouse Skeletal Muscles Identifies Annexin, Lamin, and Vimentin as Universal Dystrophic Markers. Molecules 2015, 20, 11317–11344. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Murphy, S.; Brinkmeier, H.; Krautwald, M.; Henry, M.; Meleady, P.; Ohlendieck, K. Proteomic profiling of the dystrophin complex and membrane fraction from dystrophic mdx muscle reveals decreases in the cytolinker desmoglein and increases in the extracellular matrix stabilizers biglycan and fibronectin. J. Muscle Res. Cell Motil. 2017, 38, 251–268. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Holland, A.; Dowling, P.; Meleady, P.; Henry, M.; Zweyer, M.; Mundegar, R.R.; Swandulla, D.; Ohlendieck, K. Label-free mass spectrometric analysis of the mdx-4cv diaphragm identifies the matricellular protein periostin as a potential factor involved in dystrophinopathy-related fibrosis. Proteomics 2015, 15, 2318–2331. [Google Scholar] [CrossRef] [PubMed]
- Dalton, C.J.; Lemmon, C.A. Fibronectin: Molecular Structure, Fibrillar Structure and Mechanochemical Signaling. Cells 2021, 10, 2443. [Google Scholar] [CrossRef] [PubMed]
- Cynthia Martin, F.; Hiller, M.; Spitali, P.; Oonk, S.; Dalebout, H.; Palmblad, M.; Chaouch, A.; Guglieri, M.; Straub, V.; Lochmüller, H.; et al. Fibronectin is a serum biomarker for Duchenne muscular dystrophy. Proteom. Clin. Appl. 2014, 8, 269–278. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Carberry, S.; Zweyer, M.; Swandulla, D.; Ohlendieck, K. Proteomics reveals drastic increase of extracellular matrix proteins collagen and dermatopontin in the aged mdx diaphragm model of Duchenne muscular dystrophy. Int. J. Mol. Med. 2012, 30, 229–234. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Okamoto, O.; Fujiwara, S. Dermatopontin, a novel player in the biology of the extracellular matrix. Connect. Tissue Res. 2006, 47, 177–189. [Google Scholar] [CrossRef]
- Kato, A.; Okamoto, O.; Ishikawa, K.; Sumiyoshi, H.; Matsuo, N.; Yoshioka, H.; Nomizu, M.; Shimada, T.; Fujiwara, S. Dermatopontin interacts with fibronectin, promotes fibronectin fibril formation, and enhances cell adhesion. J. Biol. Chem. 2011, 286, 14861–14869. [Google Scholar] [CrossRef] [Green Version]
- Sugihara, H.; Teramoto, N.; Nakamura, K.; Shiga, T.; Shirakawa, T.; Matsuo, M.; Ogasawara, M.; Nishino, I.; Matsuwaki, T.; Nishihara, M.; et al. Cellular senescence-mediated exacerbation of Duchenne muscular dystrophy. Sci. Rep. 2020, 10, 16385. [Google Scholar] [CrossRef]
- Signorelli, M.; Tsonaka, R.; Aartsma-Rus, A.; Spitali, P. Multiomic characterization of disease progression in mice lacking dystrophin. PLoS ONE 2023, 18, e0283869. [Google Scholar] [CrossRef]
- Capitanio, D.; Moriggi, M.; Torretta, E.; Barbacini, P.; De Palma, S.; Viganò, A.; Capitanio, D.; Moriggi, M.; Torretta, E.; Barbacini, P.; et al. Comparative proteomic analyses of Duchenne muscular dystrophy and Becker muscular dystrophy muscles: Changes contributing to preserve muscle function in Becker muscular dystrophy patients. J. Cachexia Sarcopenia Muscle 2020, 11, 547–563. [Google Scholar] [CrossRef] [Green Version]
- Capitanio, D.; Moriggi, M.; Barbacini, P.; Torretta, E.; Moroni, I.; Blasevich, F.; Morandi, L.; Mora, M.; Gelfi, C. Molecular fingerprint of BMD patients lacking a portion in the rod domain of dystrophin. Int. J. Mol. Sci. 2020, 23, 2624. [Google Scholar] [CrossRef]
- Doran, P.; Martin, G.; Dowling, P.; Jockusch, H.; Ohlendieck, K. Proteome analysis of the dystrophin-deficient MDX diaphragm reveals a drastic increase in the heat shock protein cvHSP. Proteomics 2006, 6, 4610–4621. [Google Scholar] [CrossRef]
- Guevel, L.; Lavoie, J.R.; Perez-Iratxeta, C.; Rouger, K.; Dubreil, L.; Feron, M.; Talon, S.; Brand, M.; Megeney, L.A. Quantitative proteomic analysis of dystrophic dog muscle. J. Proteome Res. 2011, 10, 2465–2478. [Google Scholar] [CrossRef]
- Rayavarapu, S.; Coley, W.; Cakir, E.; Jahnke, V.; Takeda, S.; Aoki, Y.; Grodish-Dressman, H.; Jaiswal, J.K.; Hoffman, E.P.; Brown, K.J.; et al. Identification of disease specific pathways using in vivo SILAC proteomics in dystrophin deficient mdx mouse. Mol. Cell. Proteom. 2013, 12, 1061–1073. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Fröhlich, T.; Kemter, E.; Flenkenthaler, F.; Klymiuk, N.; Otte, K.A.; Blutke, A.; Krause, S.; Walter, M.C.; Wanke, R.; Wolf, E.; et al. Progressive muscle proteome changes in a clinically relevant pig model of Duchenne muscular dystrophy. Sci. Rep. 2016, 6, 33362. [Google Scholar] [CrossRef] [Green Version]
- van Westering, T.L.E.; Johansson, H.J.; Hanson, B.; Coenen-Stass, A.M.L.; Lomonosova, Y.; Tanihata, J.; Motohashi, N.; Yokota, T.; Takeda, S.; Lehtiö, J.; et al. Mutation-independent proteomic signatures of pathological progression in murine models of Duchenne muscular dystrophy. Mol. Cell. Proteom. 2020, 19, 2047–2068. [Google Scholar] [CrossRef] [PubMed]
- Holland, A.; Carberry, S.; Ohlendieck, K. Proteomics of the dystrophin-glycoprotein complex and dystrophinopathy. Curr. Protein Pept. Sci. 2013, 14, 680–697. [Google Scholar] [CrossRef] [PubMed]
- Carr, S.J.; Zahedi, R.P.; Lochmüller, H.; Roos, A. Mass spectrometry-based protein analysis to unravel the tissue pathophysiology in Duchenne muscular dystrophy. Proteom. Clin. Appl. 2018, 12, 1700071. [Google Scholar] [CrossRef]
- Al-Khalili Szigyarto, C. Duchenne Muscular Dystrophy: Recent advances in protein biomarkers and the clinical application. Expert Rev. Proteom. 2020, 17, 365–375. [Google Scholar] [CrossRef] [PubMed]
- Gawlik, K.I. At the Crossroads of Clinical and Preclinical Research for Muscular Dystrophy—Are We Closer to Effective Treatment for Patients? Int. J. Mol. Sci. 2018, 19, 1490. [Google Scholar] [CrossRef] [Green Version]
- Dowling, P.; Murphy, S.; Zweyer, M.; Raucamp, M.; Swandulla, D.; Ohlendieck, K. Emerging proteomic biomarkers of X-linked muscular dystrophy. Expert Rev. Mol. Diagn. 2019, 19, 739–755. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Grounds, M.D.; Terrill, J.R.; Al-Mshhdani, B.A.; Duong, M.N.; Radley-Crabb, H.G.; Arthur, P.G. Biomarkers for Duchenne muscular dystrophy: Myonecrosis, inflammation and oxidative stress. Dis. Model Mech. 2020, 13, dmm043638. [Google Scholar] [CrossRef] [Green Version]
- Norris, R.A.; Damon, B.; Mironov, V.; Kasyanov, V.; Ramamurthi, A.; Moreno-Rodriguez, R.; Trusk, T.; Potts, J.D.; Goodwin, R.L.; Davis, J.; et al. Periostin regulates collagen fibrillogenesis and the biomechanical properties of connective tissues. J. Cell. Biochem. 2007, 101, 695–711. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ozdemir, C.; Akpulat, U.; Sharafi, P.; Yıldız, Y.; Onbaşılar, I.; Kocaefe, C. Periostin is temporally expressed as an extracellular matrix component in skeletal muscle regeneration and differentiation. Gene 2014, 553, 130–139. [Google Scholar] [CrossRef] [PubMed]
- Ito, N.; Miyagoe-Suzuki, Y.; Takeda, S.; Kudo, A. Periostin Is Required for the Maintenance of Muscle Fibers during Muscle Regeneration. Int. J. Mol. Sci. 2021, 22, 3627. [Google Scholar] [CrossRef]
- Lorts, A.; Schwanekamp, J.A.; Baudino, T.A.; McNally, E.M.; Molkentin, J.D. Deletion of periostin reduces muscular dystrophy and fibrosis in mice by modulating the transforming growth factor-β pathway. Proc. Natl. Acad. Sci. USA 2012, 109, 10978–10983. [Google Scholar] [CrossRef]
- Demonbreun, A.R.; Quattrocelli, M.; Barefield, D.Y.; Allen, M.V.; Swanson, K.E.; McNally, E.M. An actin-dependent annexin complex mediates plasma membrane repair in muscle. J. Cell Biol. 2016, 213, 705–718. [Google Scholar] [CrossRef] [Green Version]
- Defour, A.; Medikayala, S.; Van der Meulen, J.H.; Hogarth, M.W.; Holdreith, N.; Malatras, A.; Duddy, W.; Boehler, J.; Nagaraju, K.; Jaiswal, J.K. Annexin A2 links poor myofiber repair with inflammation and adipogenic replacement of the injured muscle. Hum. Mol. Genet. 2017, 26, 1979–1991. [Google Scholar] [CrossRef] [Green Version]
- Bittel, D.C.; Chandra, G.; Tirunagri, L.M.S.; Deora, A.B.; Medikayala, S.; Scheffer, L.; Defour, A.; Jaiswal, J.K. Annexin A2 Mediates Dysferlin Accumulation and Muscle Cell Membrane Repair. Cells 2020, 9, 1919. [Google Scholar] [CrossRef] [PubMed]
- Lennon, N.J.; Kho, A.; Bacskai, B.J.; Perlmutter, S.L.; Hyman, B.T.; Brown, R.H., Jr. Dysferlin interacts with annexins A1 and A2 and mediates sarcolemmal wound-healing. J. Biol. Chem. 2003, 278, 50466–50473. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Croissant, C.; Carmeille, R.; Brévart, C.; Bouter, A. Annexins and Membrane Repair Dysfunctions in Muscular Dystrophies. Int. J. Mol. Sci. 2021, 22, 5276. [Google Scholar] [CrossRef]
- Croissant, C.; Gounou, C.; Bouvet, F.; Tan, S.; Bouter, A. Trafficking of Annexins during Membrane Repair in Human Skeletal Muscle Cells. Membranes 2022, 12, 153. [Google Scholar] [CrossRef] [PubMed]
- Mouw, J.K.; Ou, G.; Weaver, V.M. Extracellular matrix assembly: A multiscale deconstruction. Nat. Rev. Mol. Cell Biol. 2014, 15, 771–785. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Csapo, R.; Gumpenberger, M.; Wessner, B. Skeletal Muscle Extracellular Matrix—What Do We Know About Its Composition, Regulation, and Physiological Roles? A Narrative Review. Front. Physiol. 2020, 11, 253. [Google Scholar] [CrossRef] [Green Version]
- Brightwell, C.R.; Latham, C.M.; Thomas, N.T.; Keeble, A.R.; Murach, K.A.; Fry, C.S. A glitch in the matrix: The pivotal role for extracellular matrix remodeling during muscle hypertrophy. Am. J. Physiol.-Cell Physiol. 2022, 323, C763–C771. [Google Scholar] [CrossRef]
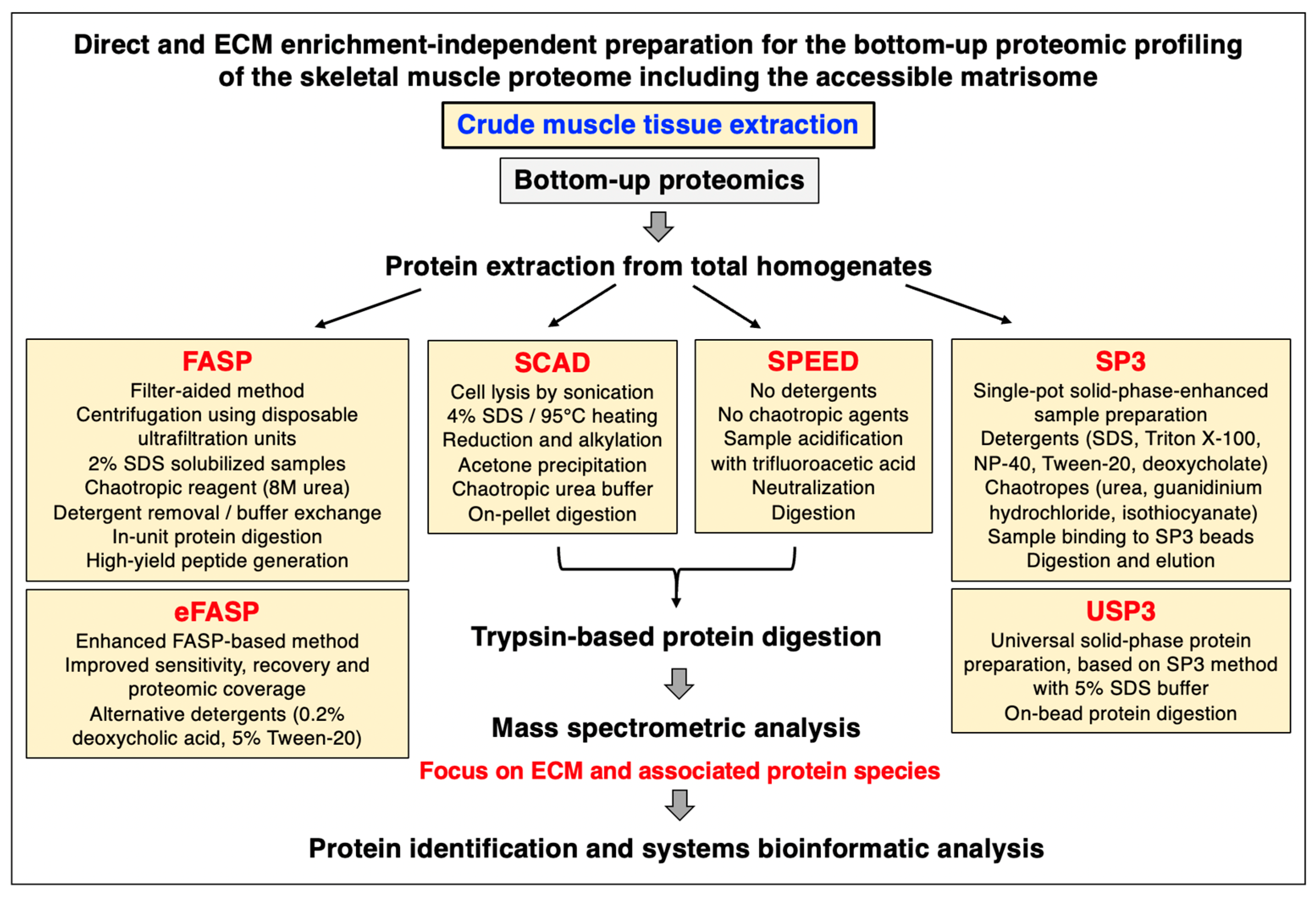

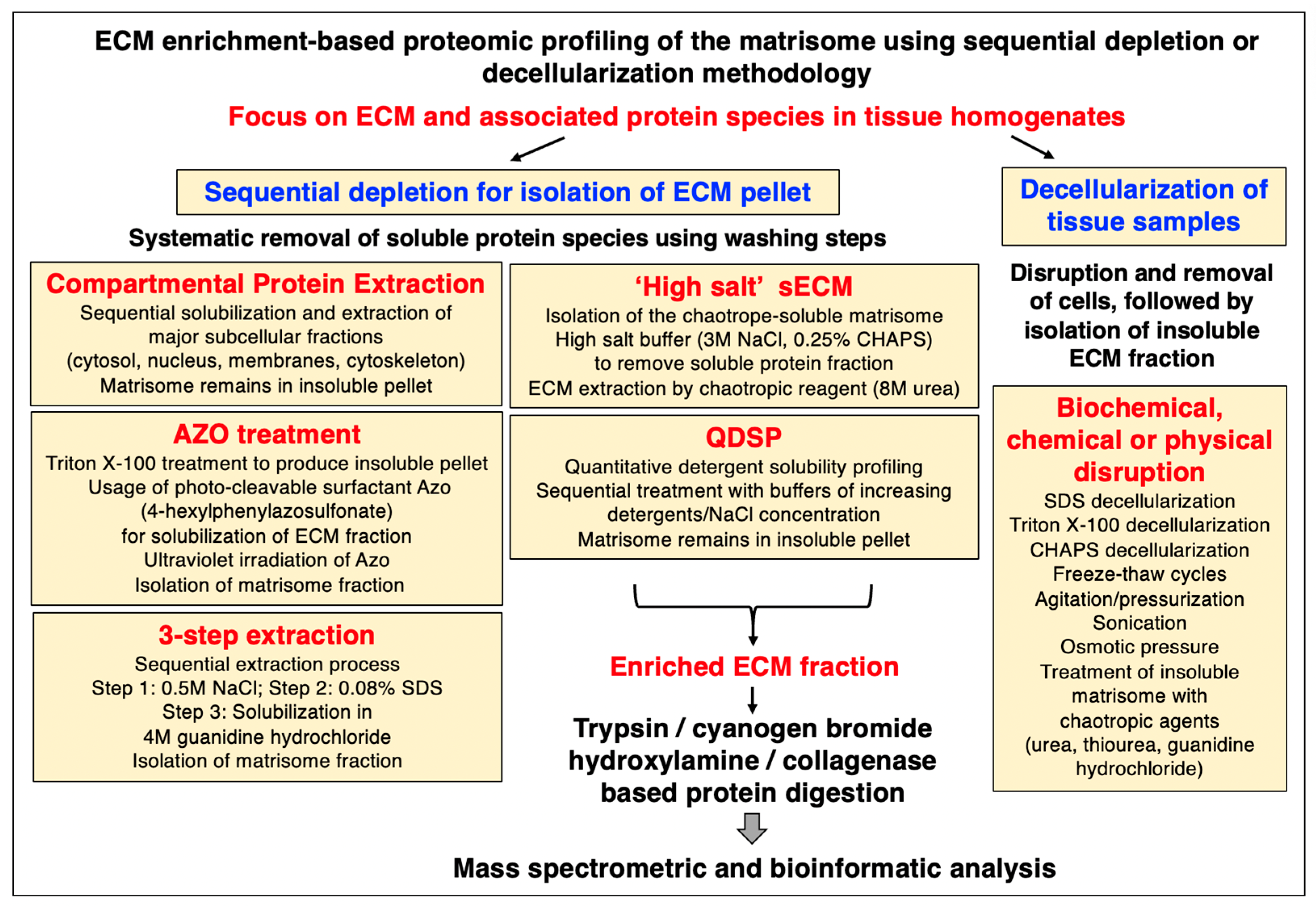
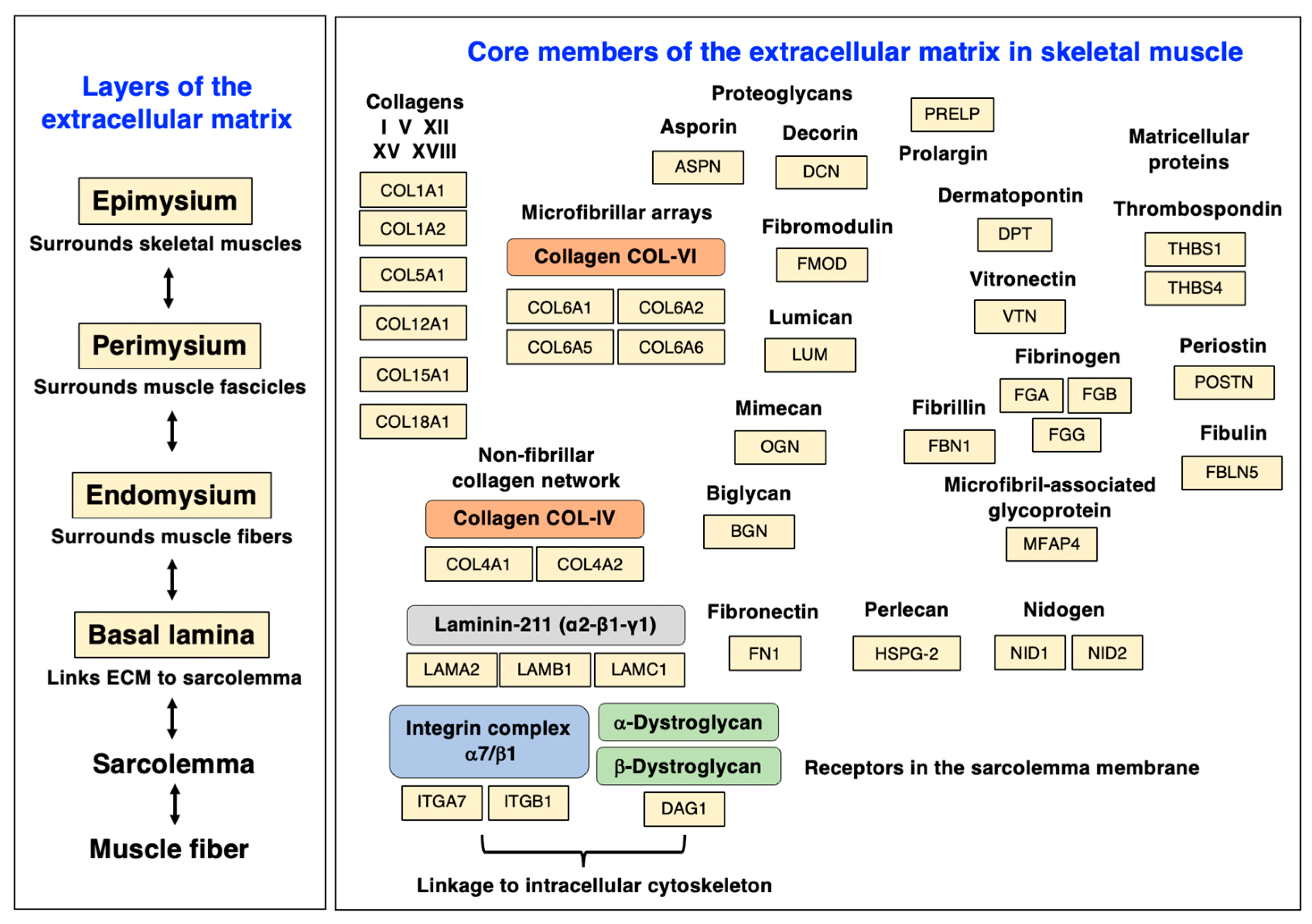





| Accession Number | Extracellular Matrix Protein | Gene Name | Coverage (%) | Peptides | kDa |
|---|---|---|---|---|---|
| (i) Basal-lamina-enriched extracellular matrix proteins | |||||
| Q60675 | Laminin-211, subunit alpha-2 | Lama2 | 21.8 | 47 | 343.6 |
| P02469 | Laminin-211, subunit beta-1 | Lamb1 | 14.9 | 16 | 197.0 |
| P02468 | Laminin-211, subunit gamma-1 | Lamc1 | 30.2 | 31 | 172.2 |
| P02463 | Collagen IV, alpha-1 | Col4a1 | 4.4 | 5 | 160.6 |
| P08122 | Collagen IV, alpha-2 | Col4a2 | 2.5 | 3 | 167.2 |
| O35206 | Collagen XV, alpha-1 | Col15a1 | 10.0 | 9 | 140.4 |
| P39061-2 | Collagen XVIII, alpha-1 | Col18a1 | 2.8 | 3 | 134.1 |
| Q05793 | Perlecan (HSPG-2) | Hspg2 | 20.7 | 43 | 398.0 |
| P10493 | Nidogen-1 (entactin) | Nid1 | 30.1 | 18 | 136.5 |
| O88322 | Nidogen-2 (osteonidogen) | Nid2 | 5.9 | 6 | 153.8 |
| (ii) Endomysium-enriched extracellular matrix proteins | |||||
| O88207 | Collagen V, alpha-1 | Col5a1 | 1.6 | 2 | 183.6 |
| Q04857 | Collagen VI, alpha-1 | Col6a1 | 29.2 | 20 | 108.4 |
| Q02788 | Collagen VI, alpha-2 | Col6a2 | 21.2 | 14 | 110.3 |
| A6H584 | Collagen VI, alpha-5 | Col6a5 | 9.2 | 17 | 289.4 |
| Q8C6K9 | Collagen VI, alpha-6 | Col6a6 | 12.5 | 16 | 246.2 |
| (iii) Extracellular matrix proteins throughout the endomysium, perimysium, and epimysium | |||||
| Q9JK53 | Prolargin | Prelp | 25.7 | 7 | 43.3 |
| Q9QZZ6 | Dermatopontin (tyrosine-rich acidic matrix protein) | Dpt | 19.4 | 2 | 24.0 |
| P29788 | Vitronectin | Vtn | 6.7 | 2 | 54.8 |
| P11276 | Fibronectin | Fn1 | 15.8 | 22 | 272.4 |
| E9PV24 | Fibrinogen, alpha | Fga | 24.2 | 12 | 87.4 |
| Q8K0E8 | Fibrinogen, beta | Fgb | 63.6 | 24 | 54.7 |
| Q8VCM7 | Fibrinogen, gamma | Fgg | 64.5 | 18 | 49.4 |
| Q9D1H9 | Microfibril-associated glycoprotein MFAP4 | Mfap4 | 17.1 | 3 | 28.9 |
| Q61554 | Fibrillin-1 | Fbn1 | 21.6 | 39 | 312.1 |
| (iv) Major muscle-associated small leucine-rich proteoglycans (SLRP type) | |||||
| Q99MQ4 | Asporin | Aspn | 24.1 | 7 | 42.5 |
| P28653 | Biglycan | Bgn | 26.6 | 7 | 41.6 |
| P28654 | Decorin | Dcn | 34.5 | 11 | 39.8 |
| P50608 | Fibromodulin | Fmod | 26.1 | 6 | 43.0 |
| P51885 | Lumican | Lum | 27.2 | 6 | 38.2 |
| Q62000 | Mimecan (Osteoglycin) | Ogn | 32.2 | 7 | 34.0 |
| (v) Matricellular proteins | |||||
| Q62009-5 | Periostin | Postn | 9.6 | 4 | 87.0 |
| P35441 | Thrombospondin-1 | Thbs1 | 15.4 | 12 | 129.6 |
| Q9Z1T2 | Thrombospondin-4 | Thbs4 | 23.7 | 2 | 106.3 |
| Q9WVH9 | Fibulin-5 | Fbln5 | 8.3 | 3 | 50.2 |
| (vi) Major sarcolemmal adhesion proteins/linkers to the extracellular matrix | |||||
| Q61738-4 | Integrin, alpha-7 | Itga7 | 8.5 | 6 | 122.1 |
| P09055 | Integrin, beta-1 | Itgb1 | 19.3 | 11 | 88.2 |
| Q62165 | Dystroglycan, alpha/beta | Dag1 | 4.1 | 3 | 96.8 |
| (vii) Myotendinous-junction-enriched proteins | |||||
| Q60847-2 | Collagen XII, alpha-1 | Col12a1 | 18.5 | 36 | 333.5 |
| Q8CIB5 | FERM-domain containing kindlin 2 (FERMT-2) | Fermt2 | 6.5 | 3 | 77.8 |
| (viii) Tendon-enriched proteins | |||||
| P11087 | Collagen I, alpha-1 | Col1a1 | 5.3 | 5 | 137.9 |
| Q01149 | Collagen I, alpha-2 | Col1a2 | 5.0 | 5 | 129.5 |
| (ix) Cartilage-enriched proteins | |||||
| Q9R0G6 | Cartilage oligomeric matrix protein | Comp | 3.7 | 2 | 82.3 |
| Q66K08 | Cartilage intermediate layer protein 1 | Cilp | 3.9 | 3 | 132.2 |
| D3Z7H8 | Cartilage intermediate layer protein 2 | Cilp2 | 3.6 | 3 | 125.9 |
| (x) Extracellular matrix proteins involved in cellular signaling | |||||
| P82198 | Transforming growth factor-beta-induced protein ig-h3 | Tgfbi | 28.0 | 12 | 74.5 |
| P21981 | Protein-glutamine gamma-glutamyltransferase 2 | Tgm2 | 36.0 | 15 | 77.0 |
| (xi) Extracellular-matrix-associated proteins belonging to the annexin family | |||||
| P07356 | Annexin A2 | Anxa2 | 56.3 | 16 | 38.7 |
| P14824 | Annexin A6 | Anxa6 | 51.0 | 23 | 75.8 |
| (xii) Extracellular matrix regulators | |||||
| Q00897 | Alpha-1-antitrypsin 1-4 (A1AT4) | Serpina1d | 44.6 | 11 | 46.0 |
| P22599 | Alpha-1-antitrypsin 1-2 (A1AT2) | Serpina1b | 38.0 | 9 | 45.2 |
| Q00896 | Alpha-1-antitrypsin 1-3 (A1AT3) | Serpina1c | 35.9 | 9 | 45.8 |
| Q00898 | Alpha-1-antitrypsin 1-5 (A1AT5) | Serpina1e | 26.2 | 8 | 45.9 |
| P19324 | Serpin H1 (SERPH) | Serpinh1 | 9.6 | 3 | 46.5 |
| P07759 | Serine protease inhibitor A3K | Serpina3k | 26.8 | 8 | 46.8 |
| P10605 | Cathepsin B | Ctsb | 22.4 | 5 | 37.3 |
| Mouse Model | Genetic Alteration | Affected Dystrophin Isoforms | Dystrophic Phenotype |
|---|---|---|---|
| mdx (mdx-23) | Exon-23 of Dmd gene; spontaneous premature stop codon | Dp427 | Moderate, but severely affected diaphragm; degeneration–regeneration cycles; considerable number of revertant muscle fibers; mildly affected heart muscle |
| mdx/Dtna | Exon-23 of Dmd gene; plus dKO of Dtna gene | Dp427 | Severely affected general musculature; moderately affected heart muscle |
| mdx/Cmah | Exon-23 of Dmd gene; plus Exon-6 deletion of Cmah gene | Dp427 | Severely affected general musculature; moderately affected diaphragm and heart |
| mdx/Utr | Exon-23 of Dmd gene; plus targeted disruption of Utrn gene | Dp427; plus lacking autosomal dystrophin homologue utrophin (Up395) | Severely affected general musculature including diaphragm; moderately affected heart muscle |
| mdx/α7 | Exon-23 of Dmd gene; plus dKO of Itga7 gene | Dp427 | Severely affected general musculature including diaphragm; mildly affected heart muscle |
| mdx/Myod1 | Exon-23 of Dmd gene; plus dKO of Myod1 gene | Dp427 | Severely affected general musculature including diaphragm, and heart muscle |
| mdx-2cv | Intron-42 of Dmd gene; ENU-mutagenesis-induced shift in reading frame | Dp427, Dp260 | Moderate with large variation in muscle fiber size; severely affected diaphragm |
| mdx-3cv | Intron-65 of Dmd gene; ENU-mutagenesis-induced shift in reading frame | Dp427, Dp260, Dp140, Dp116, Dp71 | Moderate, but severely affected diaphragm; no revertant fibers |
| mdx-4cv | Exon-53 of Dmd gene; ENU-mutagenesis-induced premature stop codon | Dp427, Dp260, Dp140 | Moderate with large variation in muscle fiber size; severely affected diaphragm; fewer revertant fibers than mdx-23 model |
| NSG-mdx-4cv | Exon-54 of Dmd gene; immunodeficient muscular dystrophy model | Dp427, Dp260, Dp140 | Moderate, but severely affected diaphragm; fewer revertant fibers than mdx-23 model |
| mdx-4cv/mTR-G2 | Exon-53 of Dmd gene; KO of mTR; model with humanized telomere lengths | Dp427, Dp260, Dp140 | Severely affected general musculature including diaphragm, and heart muscle |
| mdx-5cv | Exon-10 of Dmd gene; ENU-mutagenesis-induced frame shift deletion | Dp427 | Moderate, but severely affected diaphragm; fewer revertant fibers than mdx-23 model |
| mdx-52 | Exon-52 of Dmd gene; targeted disruption induced point mutation | Dp427, Dp260, Dp140 | Moderate, but severely affected diaphragm; fewer revertant fibers than mdx-23 model |
| mdx-βgeo | Intron-63 of Dmd gene; insertion of β-geo gene trap cassette | Dp427, Dp260, Dp140, Dp116, Dp71 | Moderately affected general musculature |
| Dmd-null | Dmd gene deletion; Cre-loxP system of entire Dmd gene | Dp427, Dp260, Dp140, Dp116, Dp71 | Severely affected general musculature |
| hDMD/mdx-45 | Spontaneous Exon-23 mutation of Dmd gene; plus CRISPR/Cas-mediated Exon-45 deletion in hDMD gene | Dp427, Dp260, Dp140 (murine and human) | Moderate, but severely affected diaphragm; revertant muscle fibers |
| hDMD/mdx-52 | Spontaneous Exon-23 mutation of Dmd gene; plus TALEN-based partial deletion of Exon-52 in hDMD gene | Dp427, Dp260, Dp140 (murine and human) | Moderate, but severely affected diaphragm; revertant muscle fibers |
| Experimental Focus | Bioanalytical Approach | Major Findings | References |
|---|---|---|---|
| (i) Generation, genotyping, and phenotyping of the mdx-4cv model | |||
| Generation of mdx-4cv model | Chemical mutagenesis with N-ethyl-N-nitrosourea (ENU) | Induction of a C-to-T transition at position 7916 in exon-53 of the Dmd gene leading to premature translation termination | [41] |
| Genotyping of mdx-4cv model | DNA sequencing, polymerase chain reaction analysis | Protein-truncating nonsense mutation that introduces premature stop codon in exon-53 of Dmd gene | [42,43,44,237] |
| Characterization of mdx-4cv model | Cell biological and biochemical analyses | Less revertant fibers compared to spontaneous mdx-23 mouse; reduced level of dystrophin–glycoprotein complex | [231] |
| (ii) Evaluation of experimental therapies using the mdx-4cv model | |||
| Exon-skipping therapy | Antisense molecule-based skipping of defective exon | Removal of exon-53 by-passed the protein-truncating mutation and restored the synthesis of semi-functional Dp427-M protein | [238] |
| Micro-dystrophin gene therapy | Virus- or nano-carrier mediated delivery of micro-dystrophin gene | Increased numbers of micro-dystrophin positive fibers using a variety of delivery mechanisms | [239,240,241,242,243] |
| Cell-based therapy | Cell transplantation to introduce mini-dystrophin protein in dystrophic fibers | High levels of mini-dystrophin expression, but no physiological improvements | [244,245,246] |
| Genome editing | CRISPR/Cas9 (clustered regularly interspaced short palindromic repeats)/CRISPR-associated protein 9) | Increased dystrophin expression in CRISPR/Cas9-treated muscles and increased force generation. | [247,248] |
| (iii) Proteomic profiling of skeletal muscle tissue specimens from the mdx-4cv model | |||
| Skeletal muscle, hindlimb | Liquid chromatography– tandem mass spectrometry (LC-MS/MS) | Increased collagen VI, fibronectin, fibrinogen, asporin, annexin-2; reduced dystrophin complex, carbonic anhydrase CA3, parvalbumin, myozenin-2 | [229] |
| Diaphragm muscle | LC-MS/MS | Increased extracellular matrix proteins (collagens, annexins, proteoglycans) and molecular chaperones; decreased dystrophin complex, parvalbumin, carbonic anhydrase CA3, excitation–contraction coupling proteins | [55,146] |
| Extraocular muscle | LC-MS/MS | Mild phenotype lacking drastic changes in protein abundance | [252] |
| Subcellular skeletal muscle fractions | Subcellular fractionation, affinity purification, LC-MS/MS | Increased membrane repair proteins (myoferlin, dysferlin, annexins) and extracellular matrix proteins (collagens); decrease in dystrophin complex | [230,251] |
| Skeletal muscle protein fractions | Chemical crosslinking analysis | Altered patterns of protein interactions in dystrophin-deficient fibers | [249,250] |
| (iv) Proteomic profiling of non-skeletal muscle tissue specimens from the mdx-4cv model | |||
| Heart | LC-MS/MS | Decreased dystrophin complex; identified changes in laminin, periostin, asporin, and lumican, heat shock proteins, mitochondrial and glycolytic enzymes | [253] |
| Liver | LC-MS/MS | Elevated levels of fatty-acid-binding protein FABP5; changes in proteins involved in fatty acid, carbohydrate, and amino acid metabolism | [254] |
| Kidney | LC-MS/MS | Elevated levels of fatty-acid-binding protein FABP1; complex changes in metabolic and bioenergetic enzymes | [255,256] |
| Stomach/pancreas interface | LC-MS/MS | Identification of dystrophin complex in normal stomach muscles; reduced dystrophin complex in mdx-4cv stomach/pancreas-interface | [258] |
| Spleen | LC-MS/MS | Identification of short dystrophin isoform in spleen; altered proteins involved in metabolism, signaling, and cellular architecture; crosstalk between lymphoid system and muscle | [96,257] |
| Brain | LC-MS/MS | Increased levels of gliosis marker GFAP (glial fibrillary acidic protein); altered abundance of a variety of neuronal proteins | [259] |
| (v) Proteomic profiling of biofluids from the mdx-4cv model | |||
| Serum | LC-MS/MS | Increased levels of various muscle damage markers in serum; high levels of the inflammation-induced plasma marker haptoglobin | [260,261] |
| Saliva | LC-MS/MS | Increased levels of kallikrein Kkl-1 and the Klk1-related peptidases Klk1-b1, Klk1-b5 and Klk-b22 | [261,263] |
| Urine | LC-MS/MS | Increased levels of various muscle damage markers in urine; high levels of titin fragments | [262] |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Dowling, P.; Gargan, S.; Zweyer, M.; Swandulla, D.; Ohlendieck, K. Extracellular Matrix Proteomics: The mdx-4cv Mouse Diaphragm as a Surrogate for Studying Myofibrosis in Dystrophinopathy. Biomolecules 2023, 13, 1108. https://doi.org/10.3390/biom13071108
Dowling P, Gargan S, Zweyer M, Swandulla D, Ohlendieck K. Extracellular Matrix Proteomics: The mdx-4cv Mouse Diaphragm as a Surrogate for Studying Myofibrosis in Dystrophinopathy. Biomolecules. 2023; 13(7):1108. https://doi.org/10.3390/biom13071108
Chicago/Turabian StyleDowling, Paul, Stephen Gargan, Margit Zweyer, Dieter Swandulla, and Kay Ohlendieck. 2023. "Extracellular Matrix Proteomics: The mdx-4cv Mouse Diaphragm as a Surrogate for Studying Myofibrosis in Dystrophinopathy" Biomolecules 13, no. 7: 1108. https://doi.org/10.3390/biom13071108





