Abstract
Dilated cardiomyopathy (DCM) encompasses various acquired or genetic diseases sharing a common phenotype. The understanding of pathogenetic mechanisms and the determination of the functional effects of each etiology may allow for tailoring different therapeutic strategies. MicroRNAs (miRNAs) have emerged as key regulators in cardiovascular diseases, including DCM. However, their specific roles in different DCM etiologies remain elusive. Here, we applied mRNA-seq and miRNA-seq to identify the gene and miRNA signature from myocardial biopsies from four patients with DCM caused by volume overload (VCM) and four with ischemic DCM (ICM). Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) enrichment analysis were used for differentially expressed genes (DEGs). The miRNA–mRNA interactions were identified by Pearson correlation analysis and miRNA target-prediction programs. mRNA-seq and miRNA-seq were validated by qRT-PCR and miRNA–mRNA interactions were validated by luciferase assays. We found 112 mRNAs and five miRNAs dysregulated in VCM vs. ICM. DEGs were positively enriched for pathways related to the extracellular matrix (ECM), mitochondrial respiration, cardiac muscle contraction, and fatty acid metabolism in VCM vs. ICM and negatively enriched for immune-response-related pathways, JAK-STAT, and NF-kappa B signaling. We identified four pairs of negatively correlated miRNA–mRNA: miR-218-5p-DDX6, miR-218-5p-TTC39C, miR-218-5p-SEMA4A, and miR-494-3p-SGMS2. Our study revealed novel miRNA–mRNA interaction networks and signaling pathways for VCM and ICM, providing novel insights into the development of these DCM etiologies.
1. Introduction
Dilated cardiomyopathy (DCM) is characterized by an enlarged and dysfunctional left ventricle [1] that encompasses various etiological causes [2]. Globally, this condition is the most common cause of heart failure (HF) and heart transplantation [3]. Ischemic cardiomyopathy (ICM) is one of the most common types of DCM. Although ICM and DCM can lead to similar symptoms of HF, evidence suggests that distinct DCM etiologies may produce different structural and/or functional phenotypes and may respond differently to therapy [4,5,6]. Aortic regurgitation is a valve disease characterized by the inadequate closure of the aortic valve during diastole, resulting in reverse blood flow from the aorta to the left ventricle [7]. Chronic valve regurgitation causes left-ventricle volume overload, leading to a left-ventricle dilatation that remains silent for a long period of time [8]. However, AR patients suddenly progress towards symptoms and HF after this long stable period [9]. Despite the worse prognosis associated with HF due to ICM, current therapies are relatively indifferent to the disease’s etiology. This reflects an incomplete understanding of the biological mechanisms contributing to HF. Hence, a better understanding of the basic pathological mechanisms of distinct DCM etiologies could provide new insight into diverse mechanisms and pave the way for new precision medicine strategies.
MicroRNA (miRNAs) are small endogenous non-coding RNAs that negatively regulate gene expression at the posttranscriptional level. Mechanistic studies have indicated that miRNAs regulate mRNA expression by partially or complementary binding to the complementary sites in the 3′-untranslated regions (3′UTR), inducing mRNA degradation and translational repression [10]. A single miRNA can target a cluster of protein-coding genes, and a single gene can be targeted by many miRNAs [11,12]. miRNAs show unique expression patterns in different cells, tissues, developmental stages, and disease stages. Their influence on gene regulation is key to understanding biological processes in complex diseases [13]. In this regard, an increasing number of studies have demonstrated that miRNAs play a crucial role in various DCM etiologies and have been suggested as potential therapeutic targets [14,15,16,17]. In addition, it has been shown that specific miRNA expression profiles exist for cardiomyopathies of diverse etiologies [18,19,20,21,22].
Prior studies have demonstrated distinct transcriptome signatures between ICM and DCM hearts by RNA-seq [23,24]. However, the differential miRNA and mRNA pattern expression between different DCM etiologies to distinguish etiology-specific molecular pathways, particularly in human tissue models, are scarce. Li et al. analyzed the paired miRNA–mRNA expression profiles of ICM and non-ICM from data downloaded from gene-expression and hybridization-array data repositories (GEO ID: GSE46224) [25]. However, some patients in both cohorts also presented with valve diseases such as pulmonic valve regurgitation, mitral valve regurgitation, AR, and tricuspid valve regurgitation.
In the present study, we aim to determine the mechanisms of miRNA regulation in two distinct DCM etiologies. For this purpose, we used RNA sequencing (RNA-seq) and small RNA-seq to identify etiology-specific transcriptome signatures in myocardial biopsies from volume overload DCM (VCM) and ICM patients without any valve disease. Furthermore, we analyzed the miRNA–mRNA interactome in order to reveal a previously unknown pathway regulating VCM and ICM progression.
2. Materials and Methods
2.1. Study Population
The study was carried out on myocardial samples from patients undergoing surgery at the Puerta del Mar, Cádiz University Hospital, Spain, after obtaining ethical authorization (133/2019). The population was distributed into two groups: volume overload cardiomyopathy, VCM (n = 9), and ICM (n = 6). The ICM cohort included patients with a history of acute myocardial infarction or significant coronary-artery stenosis (left main artery greater than 50% or any major coronary artery greater than 75%) [26]. The VCM cohort included patients with severe AR that fulfilled the criteria for aortic valve replacement [9,27]. The etiology was determined by two independent cardiologists. None of the patients had familial DCM criteria, additional cardiovascular disease, or any inflammatory, tumor, or infectious disease that could influence our results. All subjects were older than 18 years of age. Complete clinical information, including family and personal history, symptoms of HF, and pharmacological information, was obtained from each patient. Transthoracic echocardiogram and electrocardiogram were performed on all individuals. All patients underwent coronary-artery catheterization prior to surgery as recommended by the European Society of Cardiology guidelines [9]. Only 4 VCM and 4 ICM samples were used for mRNA-seq and 4 VCM and 3 ICM samples for miRNA-seq due to technical issues. The entire population of samples was used to validate the sequencing results.
2.2. Myocardial Tissue Collection
Human left ventricular transmural core biopsies were collected by experienced cardiothoracic surgeons at the Puerta del Mar Hospital University. Samples were obtained from 15 patients undergoing valve-replacement surgery or coronary-artery bypass grafting. After entering extracorporeal circulation and performing aortic clamping, the anterior descending coronary artery and the interventricular septum were located. A disposable 18 G/15 cm biopsy needle (Quick-core, Cook Medical, Bloomington, IN, USA) was used to extract the ventricular biopsy as described [28]. Samples were biopsied from the left anterolateral free wall between the anterior descending coronary artery and the marginal oblique, close to the apex. No complications were observed during this technique. Cardiac tissue samples were collected, frozen in liquid nitrogen, and stored at −80 °C.
2.3. RNA Isolation and qRT-PCR
All qRT-PCR experiments followed MIQE guidelines [29]. Total RNA was extracted and purified by using TRI Reagent (Sigma-Aldrich, St. Louis, MO, USA) according to the manufacturer’s instructions, followed by DNase treatment and purification using RNA clean and a concentrator-5 kit (Zymo Research, Irvine, CA, USA). RNA was quantified using a Qubit RNA High-Sensitivity Assay kit in the Qubit® 2.0 Fluorometer (Life Technologies, Carlsbad, CA, USA). mRNA was reverse-transcribed employing a PrimeScript RT reagent Kit (Takara, Shiga, Japan) following the manufacturer’s instructions. Relative quantification of mRNA levels was executed using forward and reverse primers at 100 nM each (primer pairs shown in Table 1) and iTaq Universal SYBR Green Supermix (Bio-Rad, Hercules, CA, USA). The mRNA levels were normalized to GAPDH and ß-ACTIN. For miRNA expression analysis, 5 ng of RNA were reverse-transcribed using a miRCURY LNA RT Kit (Qiagen, Hilden, Germany) and amplified with hsa-miR-106b-3p (339306, Qiagen), hsa-miR-193b-5p (339306, Qiagen), hsa-miR-218-5p (339306, Qiagen), hsa-miR-487b-3p (339306, Qiagen), and hsa-miR-494-3p (339306, Qiagen) miRCURY LNA miRNA PCR primer set and a miRCURY LNA SYBR Green PCR Kit (Qiagen, Hilden, Germany) according to the manufacturer’s instruction. All qRT-PCRs were performed on a CFX96 Real-Time PCR system (Bio-Rad, Hercules, CA, USA). The miRNA expression was normalized against U6 snRNA (v2) and a 5S rRNA miRCURY LNA miRNA PCR primer set, and data were analyzed using the 2−ΔΔCt algorithm.

Table 1.
Quantitative real-time polymerase chain reaction primer pair sequence.
2.4. RNA-Seq Analysis and Bioinformatics
The quality and integrity of the total RNA were controlled on the Agilent Technologies 2100 Bioanalyzer. Standard-specific mRNA sequencing (mRNA-seq) libraries were generated using the NEBNext Ultra II Directional RNA Library Prep Kit for Illumina using the NEBNext Poly(A) mRNA Magnetic Isolation Module (New England Biolabs, Ipswich, MA, USA), and single-end libraries were sequenced on an Illumina SE75 Platform with an output of ~70 M reads per sample. Standard miRNA libraries were generated using the NEXTFLEX small RNA-seq kit v3 (Perkin Elmer, Waltham, MA, USA), and single-end libraries were sequenced on an Illumina SE75 Platform with an output of ~20 M reads per sample.
For trimming and aligning raw data, fastq sequence reads were uploaded to the European version of the Galaxy platform [30]. Reads were trimmed with the Trim Galore software (Galaxy Version 0.6.7+galaxy0) and aligned to the built-in human reference genome, Dec. 2013 (GRCh38/hg38), with the RNA STAR Gapped-read mapper (Galaxy Version 2.7.10b+galaxy3) [31]. For gene-expression analyses, bam files were downloaded from the Galaxy server and further analyzed with the different RStudio packages downloaded from the Bioconductor website (http://bioconductor.org, accessed on 2 March 2023). Reads were assigned to genes by means of the “featureCounts” function of the “Rsubread” package, version 2.10.5. [32] and annotation files human release 43 (GRCh38.p13) (https://www.gencodegenes.org/human/release_43.html, accessed on 2 March 2023) and Chromosomal coordinates of Homo sapiens microRNAs (https://www.mirbase.org/ftp/CURRENT/genomes/hsa.gff3, accessed on 2 March 2023) for mRNA and miRNA analysis, respectively. Only the mapped reads were used to calculate gene expressions. The library size of each experimental point ranged from 58,431,140 to 76,443,844 sequences and from 7,530,361 to 27,506,550 sequences for mRNA and miRNA analysis, respectively. The differential gene-expression analyses were performed with package ‘DESeq2’ version 1.36.0 [https://doi.org/10.1186/s13059-014-0550-8, accessed on 2 March 2023]. All the gene comparisons with a p value < 0.05 and an abslog2 fold change (FC) > 0.5 were considered differentially expressed under the experimental conditions. For miRNA–target transcript interaction network analysis, first, we computed Pearson correlation coefficients for all miRNA–mRNA couples available in each group of samples (VCM and ICM). The next step consisted of intersecting the significant negative correlations with the predicted miRNA–mRNA potential interactions from DIANA-microT and ElMMo PITA databases [33,34]. In addition, in order to minimize the false-positive ratio, the information from experimentally validated miRNA–mRNA target pairs extracted from TarBase and miRTarbase databases was also implemented [35,36]. This experimental approach is similar to the one carried out by Vila-Casadesús et al. (2016) [37] previously. Functional enrichment analyses and Gene Set Enrichment Analysis (GSEA)-based [38] Kyoto Encyclopedia of Genes and Genomes (KEGG) and Gene Ontology (GO) analyses were conducted with the “clusterProfiler” package version 3.6.0. [39,40]. The gene sets with a p-value < 0.05 were considered overrepresented under the experimental conditions.
2.5. Luciferase Reporter Assay
Dual-luciferase reporter assay was performed as described previously [41].
DDX6, TTC39C, SEMA4A, and SGMS2 3’UTR fragments were PCR-amplified and cloned into the pMIR-REPORT vector. The 3T3 fibroblasts (ATCC) were cultured with DMEM (Gibco, Waltham, MA, USA) supplemented with 10% fetal bovine serum and 1% penicillin/streptomycin and seeded overnight at 50–60% confluence in 24-well plates. Then, the cells were co-transfected using Lipofectamine 3000 (Thermo Fisher Scientific, Waltham, MA, USA) with 100 ng of the luciferase vector carrying the 3’UTR fragment and 50 nM of the mirVana mimics has-miR-218-5p or has-miR-494-3p (4464066, Thermo Fisher Scientific, Waltham, MA, USA), along with 300 ng of pcLux vector control for internal normalization.
PCR-based site-directed mutagenesis was performed using the Bio-Rad iPROOF PCR kit. Primers, including MD218_DDX6_Fw (5′-GTCCCTCTTAAACCACAGAC-3′), MD218_DDX6_Rv (5′-GTCTGTGGTTTAAGAGGGAC-3′), MD218_SEMA4_Fw (5′-CTCAAGAGCAGAGAGA-3′), MD218_SEMA4_Rv (5′-TCTCTCTGCTCTTGAG-3′), MD494_SMGS2_Fw (5’-ACACTGCAGCTGCCAC-3’), and MD494_SMGS2_Rv (5’-GTGGCAGCTGCAGTGT-3’). The underlined nucleotides are referenced to nucleotides changed in site-directed mutagenesis and were used to introduce mutations into the sequence complementary to seed sequences of miR-218 and miR-494 within the 3′ UTRs of DDX6, SEMA4, and SMGS2, respectively. Each luciferase assay was carried out in triplicate and repeated in at least three distinct biological samples to obtain representative means.
Luciferase activity was measured 18 h after transfection by using the Pierce Gaussia Luciferase Flash Assay Kit (Thermo Fisher Scientific, Waltham, MA, USA) and normalized to pcLux vector control by using the Pierce Cypridina Luciferase Flash Assay Kit (Thermo Fisher Scientific, Waltham, MA, USA). In all cases, transfections were carried out in triplicate.
2.6. Statistical Analysis
Data are expressed as mean ± SEM, and n denotes the number of replicates for each experiment. Outliers were identified through the Rout method, using a Q = 1%. The normal distribution of each variable was verified with the Shapiro–Wilk test. Statistical differences (p < 0.05) between the experimental groups were assessed using a two-tailed, unpaired Student’s t test for Gaussian distributions. For non-Gaussian distributions, a Mann–Whitney non-parametric test was used [42]. All the statistical analyses were performed using GraphPad Prism 9.0 software (San Diego, CA, USA).
3. Results
3.1. Baseline Characteristics
The baseline characteristics of the 15 patients with DCM included in this study are shown in Table 2. The ICM cohort included six patients (mean age, 66.33 ± 7.84 years; all male patients) while the VCM group comprised nine patients (mean age, 63.67 ± 11.78 years; one female). There were no significant differences in the echocardiographic variables, such as left-ventricle ejection fraction, dilated left ventricular end-diastolic diameter, left ventricular end-systolic diameters, and left atrial dimension.

Table 2.
Baseline patient characteristics of the ICM and VCM groups. ICM: ischemic cardiomyopathy; VCM: volume overload cardiomyopathy; LVEF: Left-ventricle ejection fraction; LVEDD: left-ventricle end-diastolic diameter; LVESD: Left-ventricle end-systolic diameter; LA: left-atrium diameter.
3.2. VCM and ICM Have Distinct mRNA and miRNA Expression Profiles
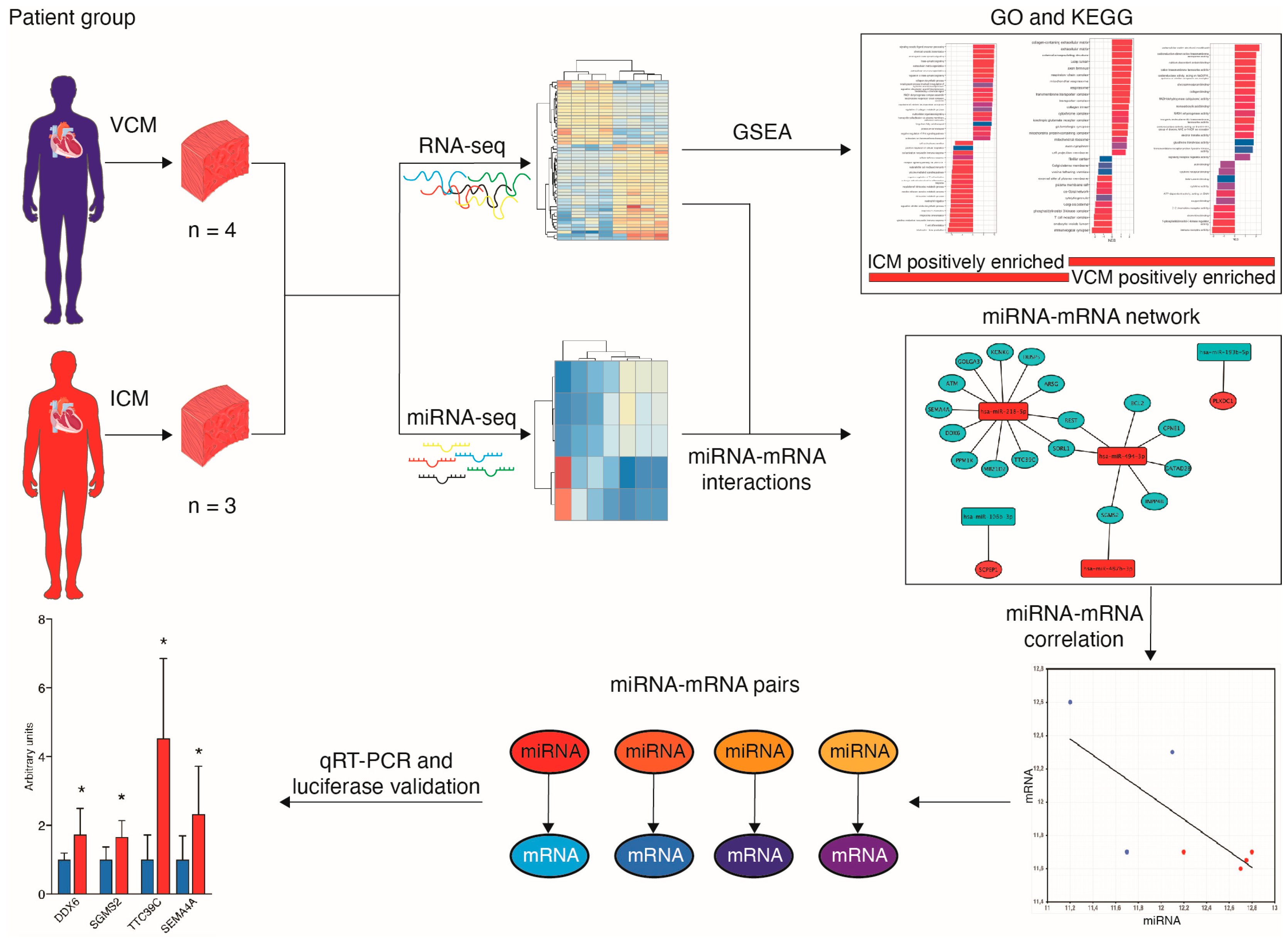
To investigate the mRNA expression differences between DCM etiologies, we performed RNA-seq on left ventricular tissue from four VCM and three ICM patients (Figure 1). We used principal component analysis (PCA) to visualize sample clustering for the most variably expressed genes, and no outliers were observed. PCA indicated that the VCM samples are plainly different from the ICM ones based on mRNA profiling (Figure 2A).
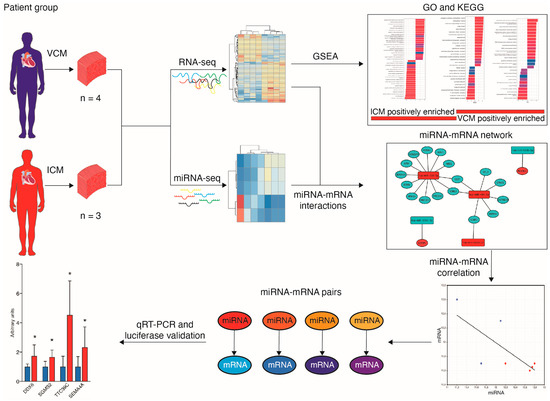
Figure 1.
A flowchart of the study design * p < 0.05.
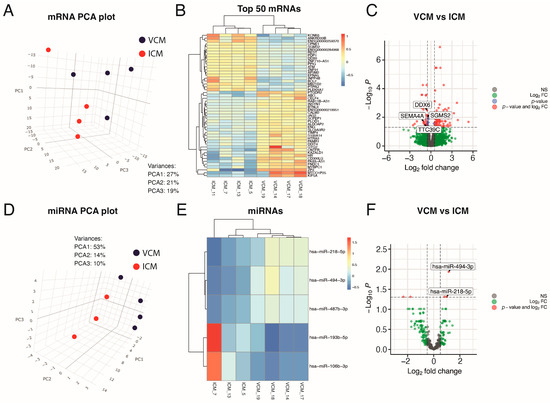
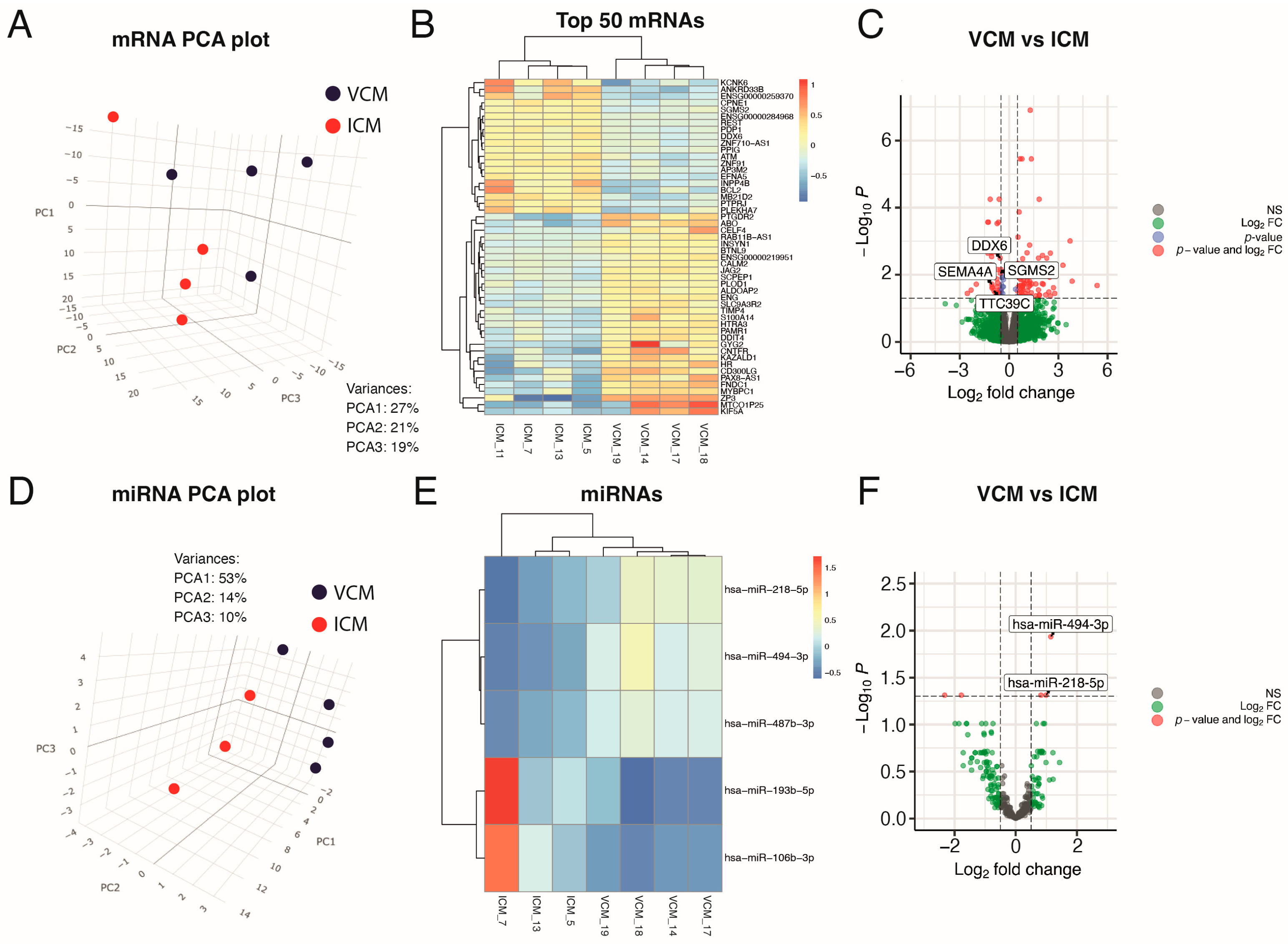
Figure 2.
Exploratory analysis of paired miRNA and mRNA expression in heart samples. (A) 3D principal components analysis plot, based on correlation matrix, for mRNA expression in VCM (n = 4) and ICM (n = 4) tissue samples. (B) Heatmap of the top-50 most differentially expressed mRNAs sorted by absolute FC (all of them having FDR < 0.05). (C) Volcano plot of the mRNAs, highlighting in gray those not statistically significant, with FDR > 0.05 and absolute FC > 1.42 (abslog2FC > 0.5), in blue those with FDR < 0.05 but absolute FC > 1.42 (abslog2FC > 0.5), in green those with absolute FC > 1.42 (abslog2FC > 0.5) but FDR > 0.05, and in red those with FDR < 0.05 and absolute FC > 1.42 (abslog2FC > 0.5). (D) 3D principal components analysis plot, based on correlation matrix, for miRNA expression in VCM (n = 4) and ICM (n = 3) tissue samples. (E) Heatmap of the only 5 differentially expressed miRNAs sorted by absolute FC (all of them having FDR < 0.05). (F) Volcano plot of the miRNAs highlighting in gray those not statistically significant with FDR > 0.05 and absolute FC > 1.42 (abslog2FC > 0.5), in green those with absolute FC > 1.42 (abslog2FC > 0.5) but FDR > 0.05 and in red those with FDR < 0.05 and absolute FC > 1.42 (abslog2FC > 0.5).
The filtered RNA-seq read set identified 18,975 genes and 583 miRNAs. In total, 111 genes were differentially expressed between VCM and ICM, with an abslog2 fold change (FC) > 0.5 using a 5% false discovery rate (FDR) (Supplementary Table S1). Of those genes, 75 were upregulated and 36 were downregulated in VCM relative to ICM (Figure 2B,C).
We also analyzed the miRNA profile by miRNA-seq to investigate the differential expression between these distinct etiologies of DCM. No outliers were observed in the miRNA expression profile in a PCA. According to the PCA, the miRNA expression characterization of the VCM samples differed significantly from that of ICM biopsies (Figure 2D). After sequencing analysis, 634 miRNAs from all myocardial samples were identified. The most highly expressed miRNA in all the myocardial samples was miR-1-3p (Supplementary Table S2). Differential expression analysis identified five miRNAs to be differentially expressed between VCM and ICM (abslog2 FC > 0.5; FDR ≤ 0.05) (Supplementary Table S3). Whereas miR-218-5p, miR-487b-3p, and miR-494-3p were upregulated, miR-106b-3p and miR-193b-5p were downregulated in VCM compared to ICM, respectively (Figure 2E,F).
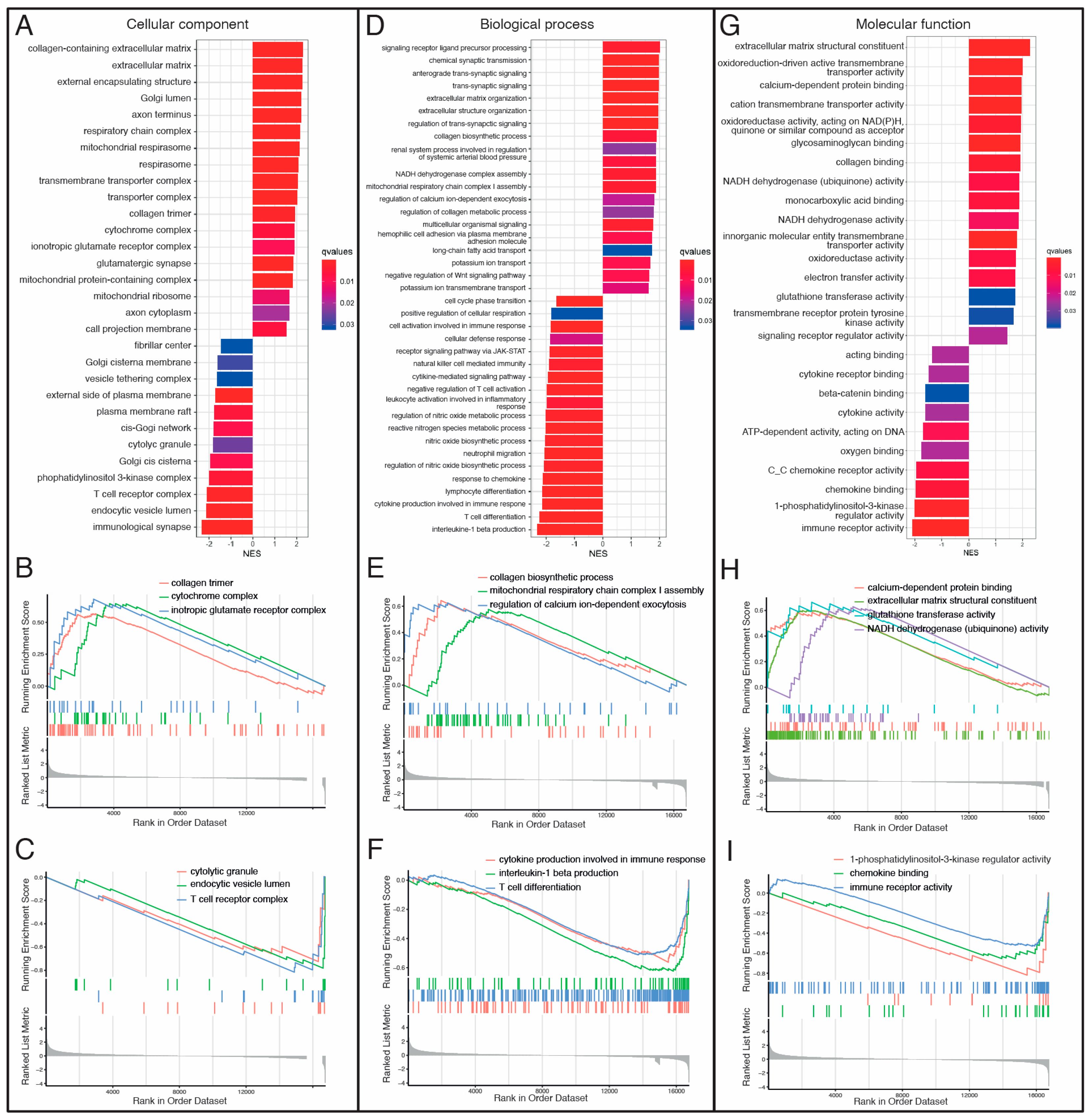
To identify pathways related to each DCM etiology, we performed pathway-enrichment analysis on the differentially expressed genes in our study. We ranked genes in order from increasing to decreasing abslog2-fold expression changes in VCM vs. ICM. Then, GO and KEGG pathway analyses based on GSEA were performed to evaluate key molecules and pathways contributing to each DCM etiology. The most enriched GO terms according to the gene ratio and categorized by the cellular component (CC), biological process (BP), and molecular function (MF) are shown in Figure 3A–I and Supplementary Tables S4–S6. GO analysis revealed extracellular matrix (ECM)-, mitochondria respiration- and immune-response-related pathways as the main terms involved in CC, BP, and MF (Figure 3A–I). In CC, collagen trimmer, cytochrome complex, and ionotropic glutamate receptor complex were among the most upregulated pathways in VCM compared to ICM (Figure 3B), whereas cytolytic granule, endocytic vesicle lumen, and T-cell receptor complex were among the most downregulated pathways (Figure 3C). In BP, collagen biosynthetic process, mitochondrial respiratory chain complex I assembly, and regulation of calcium ion-dependent exocytosis were among the most positively enriched pathways in VCM related to ICM (Figure 3D,E), whereas cytokine production involved in immune response, interleukine-1 (IL-1) beta production, and T-cell differentiation were among the most negatively enriched pathways (Figure 3D,F). Finally, calcium-dependent protein binding, ECM structural constituent, glutathione transferase activity, and NADH dehydrogenase (ubiquinone) activity were among the most positively enriched pathways in VCM related to ICM (Figure 3G,H), whereas 1-phosphatidylinositol-3-kinase regulator activity, chemokine binding, and immune-receptor activity were the most negatively enriched pathways in MF (Figure 3G,I). On the other hand, the results of the KEGG pathway analysis showed that cardiac muscle contraction, fatty acid degradation, and oxidative phosphorylation, among others, were positively enriched in VCM when compared to ICM (Figure 4A,B), whereas B cell receptor, JAK-STAT, and NF-kappa B signaling pathways, among others, were negatively regulated with VCM (Figure 4A,C).
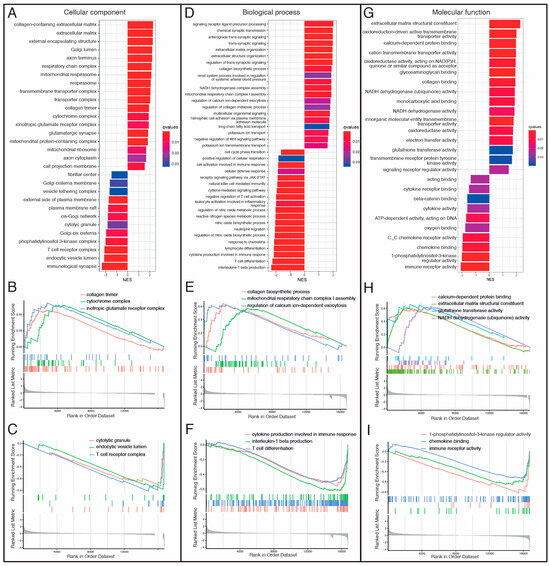
Figure 3.
Gene Ontology (GO) Gene Set Enrichment Analysis (GSEA) between VCM and ICM groups. (A) Bar plot for cellular component (CC) analysis ranked by Normalized Enrichment Score (NES). (B) Gseaplot for “collagen trimer”, “cytochrome complex”, and “ionotropic glutamate receptor” upregulated GO terms showing the running score and preranked list of GSEA and its association of phenotype in CC analysis. (C) Gseaplot for “cytolytic granule”, “endocytic vesicle lumen”, and “T cell receptor complex” downregulated GO terms showing the running score and preranked list of GSEA and its association of phenotype in CC analysis. (D) Bar plot for biological process (BP) analysis ranked by Normalized Enrichment Score (NES). (E) Gseaplot for “collagen biosynthetic process”, “mitochondrial respiratory chain complex I assembly”, and “regulation of calcium ion-dependent exocytosis” upregulated GO terms showing the running score and preranked list of GSEA and its association of phenotype in BP analysis. (F) GSEA plot for “cytokine production involved in immune response”, “interleukin-1 beta production”, and “T cell differentiation” downregulated GO terms showing the running score and preranked list of GSEA and its association of phenotype in BP analysis. (G) Bar plot for molecular function (MF) analysis ranked by Normalized Enrichment Score (NES). (H) GSEA plot for “calcium-dependent protein binding”, “extracellular matrix structural constituent”, “glutathione transferase activity”, and “NADH dehydrogenase (ubiquinone) activity” upregulated GO terms showing the running score and preranked list of GSEA and its association of phenotype in MF analysis. (I) GSEA plot for “1-phosphatidylinositol-3-kinase regulator activity”, “chemokine binding”, and “immune receptor activity” downregulated GO terms showing the running score and preranked list of GSEA and its association of phenotype in MF analysis. NES: normalized enrichment scores.
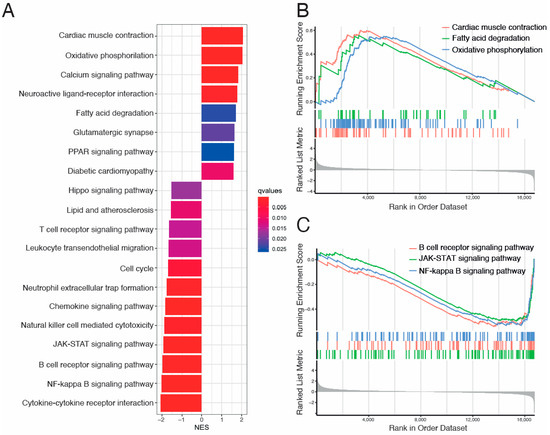
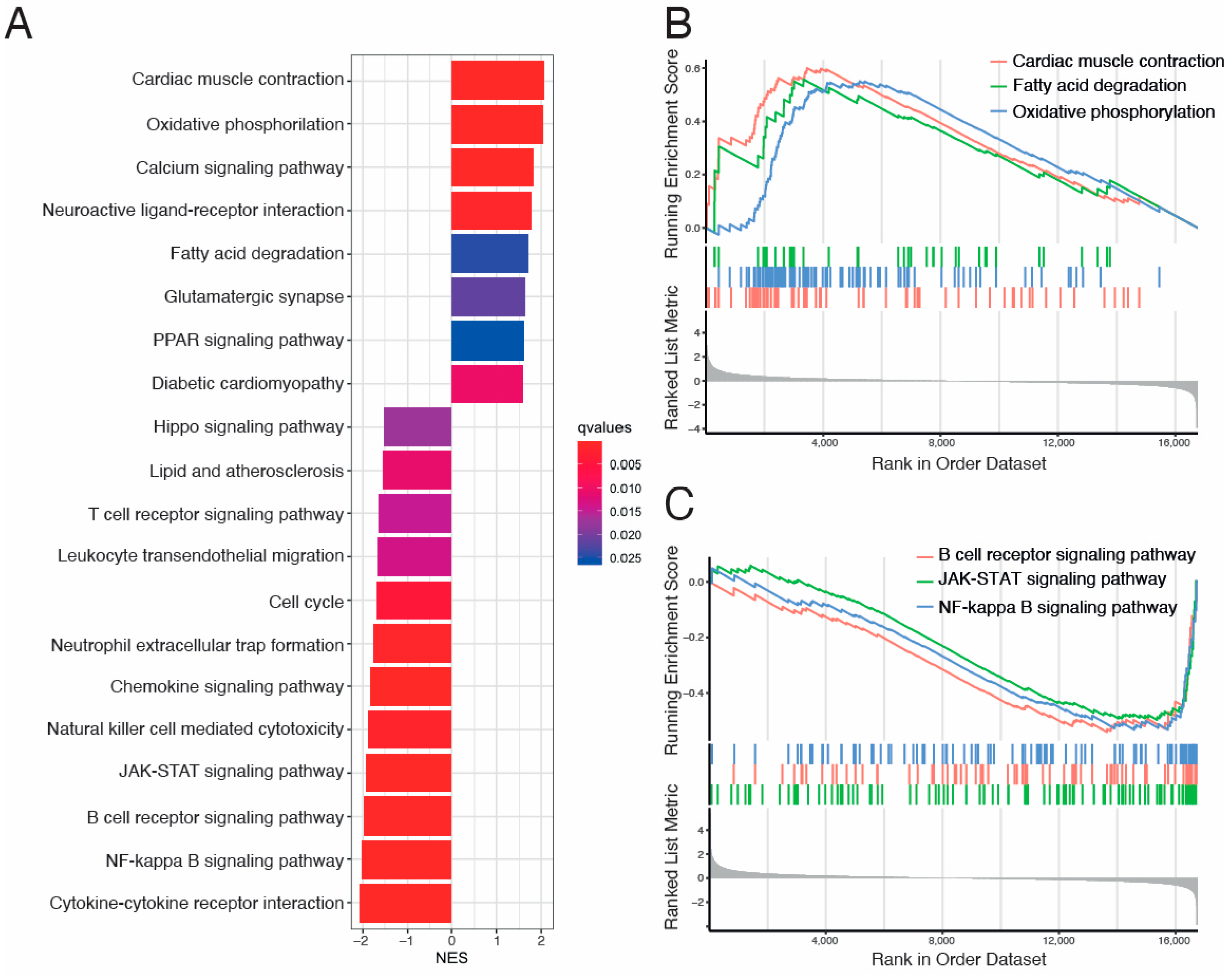
Figure 4.
Kyoto Encyclopedia of Genes and Genomes (KEGG) Gene Set Enrichment Analysis (GSEA) between VCM and ICM groups. (A) Bar plot for KEGG analysis ranked by Normalized Enrichment Score (NES). (B) GSEA plot for “Cardiac muscle contraction”, “Fatty acid degradation”, and “Oxidative phosphorylation” upregulated KEGG terms showing the running score and preranked list of GSEA and its association of phenotype. (C) GSEA plot for “B cell receptor signaling pathway”, “JAK-STAT signaling pathway”, and “NF-kappa B signaling pathway” downregulated KEGG terms showing the running score and preranked list of GSEA and its association of phenotype. NES: normalized enrichment scores.
3.3. miRNA–Target Transcript Interaction Network
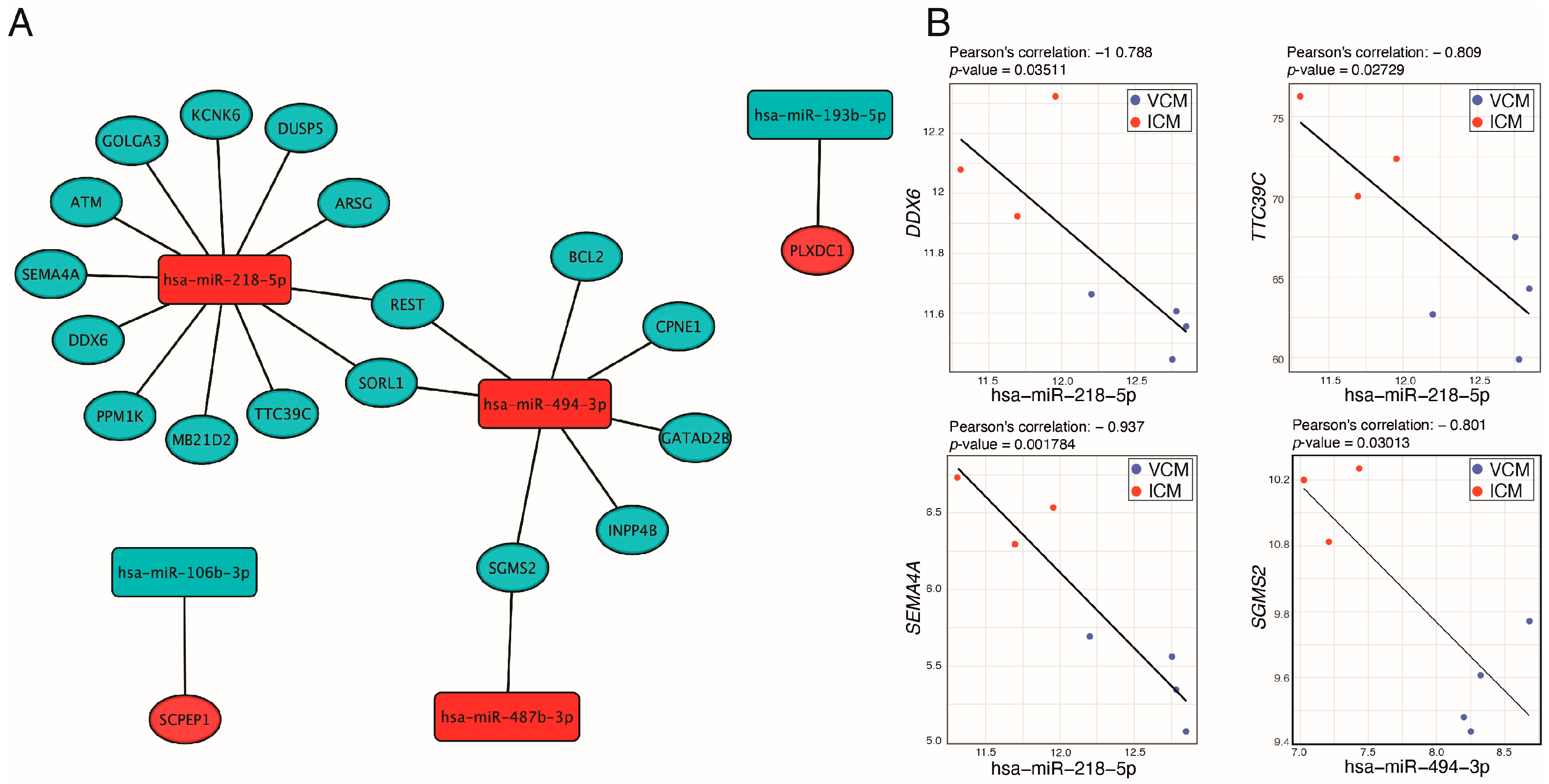
In order to identify miRNA–target pairs that could be playing an active role in the two etiologies of DCM, we focused on those miRNAs that were differentially expressed between VCM and ICM. We selected miRNA–target pairs from three prediction tools (DIANA-microT, ElMMo, and PITA,) and from databases that list experimentally validated miRNA–target pairs (TarBase and miRTarbase) (Table 3). We identified 22 miRNA–target transcript pairs. The resulting miRNA–target transcript interaction network consists of five miRNAs connected to 19 genes (Figure 5A). Of the 22 miRNA–transcript pairs, 13 miRNA–transcript interactions have been previously experimentally validated (Table 3). Afterward, we performed a Pearson correlation between the selected miRNAs and genes. This identified four negatively correlated miRNA–target transcript pairs, miR-218-5p-DDX6, miR-218-5p-TTC39C, miR-218-5p-SEMA4A, and miR-494-3p-SGMS2 (R < −0.7 and p-value < 0.05) (Figure 5B), of which only miR-218-5p-DDX6 has been previously experimentally validated (Table 3).

Table 3.
miRNA–target pairs from three prediction tools, -DIANA-microT, ElMMo and PITA-, and from databases that list experimentally validated miRNA–target pairs (TarBase and miRTarbase).
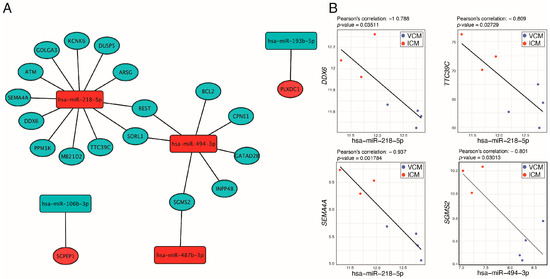
Figure 5.
miRNA–miRNA interaction analysis. (A) Network of selected miRNA–mRNA interactions. (B) Negatively correlated miRNA–mRNA pairs predicted simultaneously, at least, in three used databases in our pipeline. Red and blue nodes mean upregulated and downregulated in VCM vs. ICM, respectively.
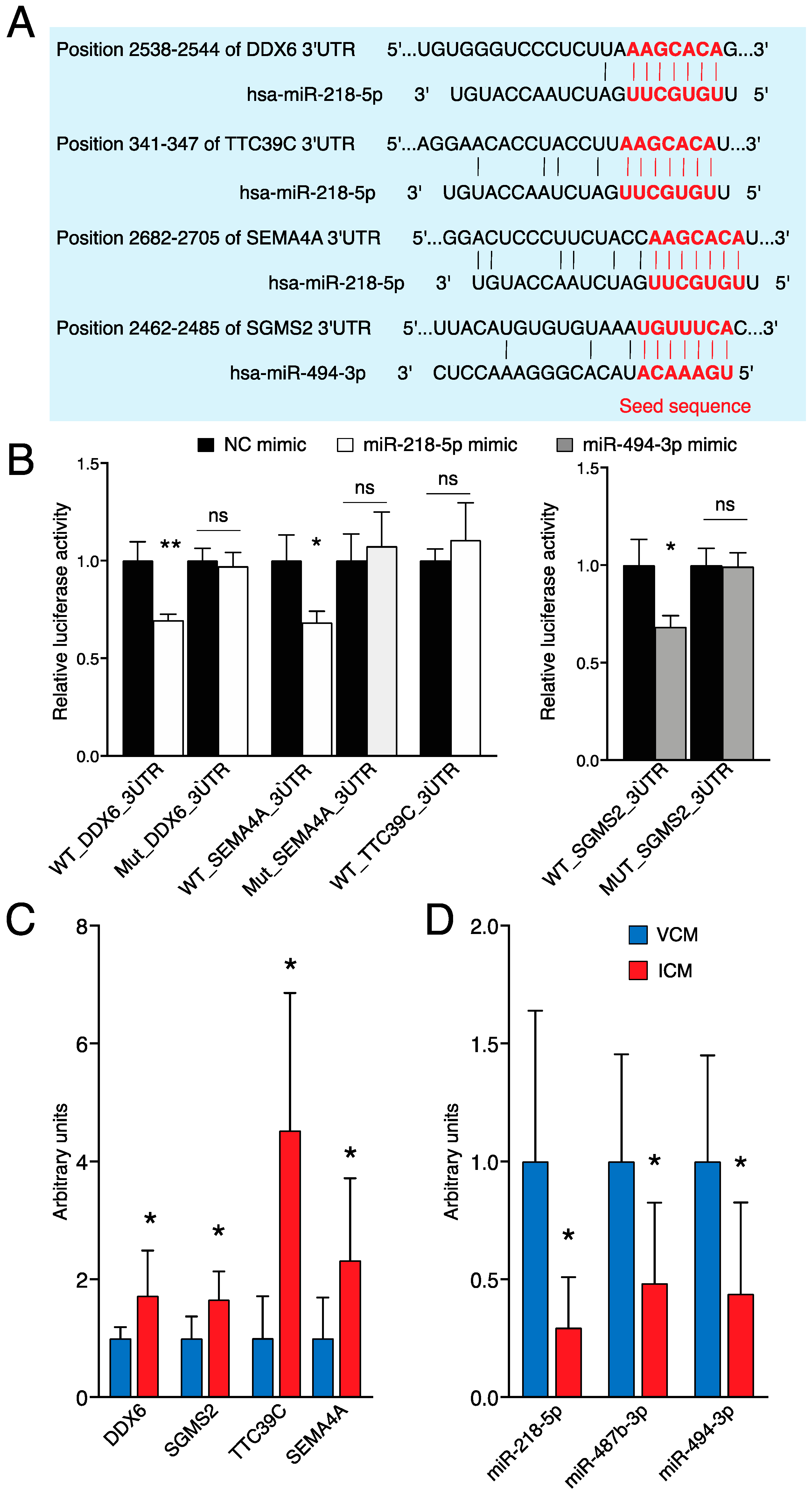
To investigate the physical interaction of the potential miRNA–target transcript pairs, we performed a dual-luciferase reporter assay. The pMIR-REPORT containing the wild-type (WT) or mutant (Mut) DDX6 3’UTR, TTC39C 3’UTR, or SEMA4A 3’UTR was co-transfected with miR-218-5p mimic, and the pMIR-REPORT containing the wild-type (WT) or mutant (Mut) SGMS2 3’UTR was co-transfected with miR-494-3p mimic into 3T3 cells. Figure 6A shows the predicted miRNA binding site in the 3’UTR of its target gene.
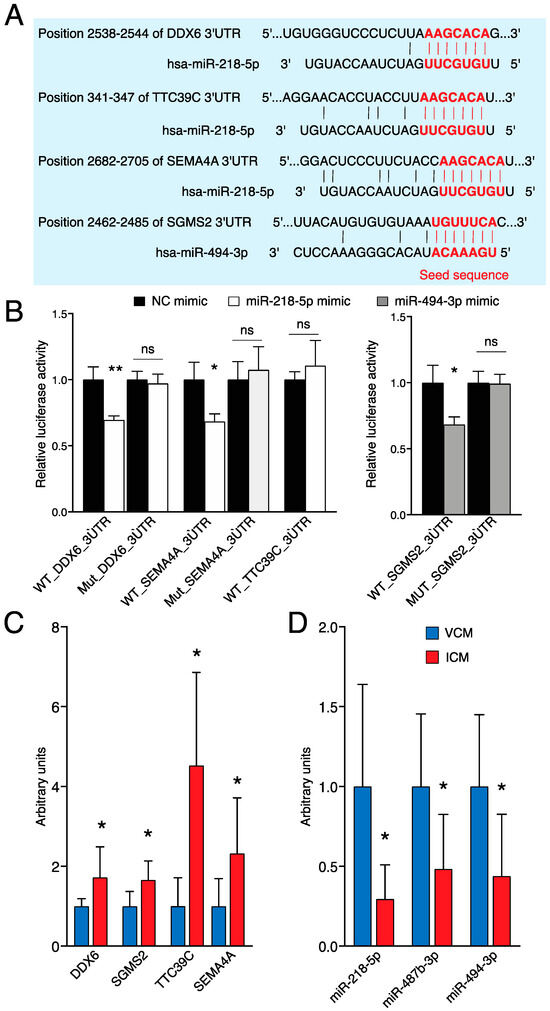
Figure 6.
miR-218-5p targets DDX6 and SEMA4A and miR-494-3p targets SGMS2. (A) Predicted miR-218-5p binding sites in the 3’UTR of DDX6, TTC39C, and SEMA4A, and predicted miR-494-3p binding sites in the 3’UTR of SGMS2. The red nucleotides are referenced to seed sequence and sequence complementary to seed sequence of miRNA and 3’UTR, respectively. (B) Dual-luciferase activity assay in 3T3 cells co-transfected with the pMIR-REPORT miRNA expression reporter vector containing the wild-type (WT) or mutant (Mut) DDX6, TTC39C, or SEMA4A 3’UTR fragment with miR-218-5p mimic, and the pMIR-REPORT miRNA expression reporter vector containing the wild-type (WT) or mutant (Mut) SGMS2 3’UTR fragment with miR-494-3p mimic for 18 h (n = 3). (C) Expression levels of DDX6, TTC39C, SEMA4A, and SGMS2 in VCM (n = 9) vs. ICM (n = 6) analyzed by qRT-PCR. (D) Expression levels of miR-218-5p, miR-487b-3p, and miR-494-3p in VCM (n = 9) vs. ICM (n = 6) analyzed by qRT-PCR. ** p < 0.01; * p < 0.05. ns: nonsignificant; NC: negative control.
Transfection of miR-218-5p mimic significantly reduced the luciferase activity in WT_DDX6_3’UTR and WT_SEMA4A_3’UTR, whereas the luciferase activity in the WT_TTC39C_3’UTR remained unaltered. Additionally, miR-494-3p significantly reduced the luciferase activity of SGMS2 (Figure 6B). Therefore, these results confirmed that DDX6 and SEMA4A are direct targets of miR-218-5p, and SGMS2 is a direct target of miR-494-3p.
3.4. Validation of Differentially Expressed Genes and miRNAs by qRT-PCR
To validate the sequencing data for certain differentially expressed genes, miRNAs were verified in the myocardial tissue of nine VCM and six ICM patients by qRT-PCR. ß-ACTIN and GAPDH were used as an internal control for mRNA qRT-PCRs, whereas U6 and 5s small nuclear RNAs were used for miRNAs. As presented in Figure 6D, significant differences were observed in the expression of the four deregulated genes between the VCM and ICM groups. DDX6, SGMS2, TTC39C, and SEMA4A were upregulated in ICM heart samples compared to VCM (Figure 6D). On the other hand, only the expressions of miR-218-5p, miR-487b-3p, and miR-494-3p, downregulated in ICM samples compared to VCM, were comparable with the sequencing data (Figure 6E).
4. Discussion
DCM is commonly regarded as a final pathway resulting from various underlying causes, which significantly influences both prognosis and outcomes [43]. However, there remains a significant gap in knowledge regarding etiologically based biological pathways associated with DCM. Here, we propose an approach to better understand molecular and cellular processes of distinct DCM etiologies through transcriptome analysis of mRNA and miRNA. Although several studies have analyzed and compared the transcriptome of distinct DCM etiologies against healthy controls [19,23,24], to date, studies comparing the mRNA and miRNA transcriptome of different etiologies with each other remain poor. Using RNA-seq, we identified genes that are differentially expressed between VCM and ICM myocardial biopsies. Pathway analysis with GSEA revealed ECM- and mitochondrial respiration-related pathways as the most common positively enriched biological routes for the GO terms CC, BP, and MF in VCM compared to ICM. In contrast, immune-activation-related pathways were among the most negatively enriched in VCM vs. ICM.
In DCM, the deposition of structural ECM proteins varies based on the mechanical or ischemic trigger. Thus, several studies suggest that, in contrast to the marked deposition of structural ECM due to mechanical alterations (i.e., pressure overload) or injuries (myocardial infarction), volume overload is less profibrotic [44,45]. VCM develops a compensatory response to maintain ventricular shape and size. However, fibrosis in VCM is associated with the loss of interstitial collagen, a prominent ECM degradation [46,47,48,49]. Although the collagen gene expression increases, there is an imbalance in the matrix metalloproteases (MMP) and tissue inhibitor metalloproteases (TIMPs) that correlates with unaltered collagen protein levels [49]. In this sense, Irqsusi et al. demonstrated that MMPs were expressed during mitral valve regurgitation, a model of volume overload, in a dynamic destruction process supported by TIMPs, the expression of which increased with the severity of the mitral regurgitation [50]. Accordingly, our results showed the upregulation of collagen and MMP family genes. The fact that both collagen synthesis and ECM degradation pathways are activated during ventricular remodeling in VCM may explain the enrichment of the ECM-related pathway compared to ICM, even being less fibrotic.
ICM fibrosis is the consequence of a repair process secondary to significant cardiomyocyte death that stimulates the inflammation cascade and the activation of reparative myofibroblasts to protect myocardium integrity. Cardiomyocyte death rapidly activates innate immune pathways that trigger cytokine, chemokine, and adhesion molecule expression, initiating the inflammatory phase and leading to the infiltration of the infarct by leukocytes and T cells [51]. Accordingly, our results showed positive enrichment of immune-response-related pathways in ICM compared to VCM. In addition, biopsy samples collected from the remote zone of ICM hearts are subject to diffuse reactive fibrosis that differs from the reparative fibrosis that occurs in the infarcted zone [52] GSEA analysis also revealed mitochondrial respiration-related pathways as more enriched in VCM compared to ICM, suggesting a more severe mitochondrial dysfunction in the ICM cohort. It is well known that mitochondrial dysfunction contributes to the transition from the normal heart to end-stage-HF, which is the common pathway of DCM, including VCM and ICM [53,54,55]. Therefore, the enrichment of pathways related to mitochondrial respiration in VCM may indicate a more severe mitochondrial dysfunction in ICM than in VCM. It has been suggested that mitochondrial dysfunction may represent only a late phenomenon in the disease process. Therefore, these results may indicate that ICM hearts go into failure faster than VCM. Consistently, rats with infarcted hearts displayed clear impairment and mitochondrial dysfunction at baseline, whereas a model of compensated volume overload showed the absence of major mitochondrial and contractile dysfunction [56,57]. Mitochondrial ion channels allow the fine-tuning of its membrane potential, ROS production, and function of the respiratory chain complexes and are extensively studied in terms of protecting the heart or cardiac cells from ischemia–reperfusion injury [58]. This is consistent with the negative enrichment of mitochondrial respiratory-related pathways, such as respiratory complex I and the cytochrome complex in ICM related to VCM. On the other hand, the ICM cohort presented a downregulation of calcium ion-dependent exocytosis and calcium-dependent protein-binding pathways, which is also related to mitochondrial dysfunction and ischemic heart disease. Ischemia–reperfusion induces a large influx of calcium into the cell and the mitochondrial outer compartment. The subsequent opening of the membrane permeability transition pore in the inner mitochondrial membrane and the resulting calcium overload induces the homeostasis of cardiomyocytes and activates the mitochondrial pathway of apoptosis [59]. Hence, our results support the idea that mitochondria are strongly recommended as a therapeutic target for cardiac ischemia–reperfusion injury [59,60]. It is well-recognized that cardiac ischemia–reperfusion injury following myocardial infarction, is a complex process involving multiple cellular and molecular pathways. Targeting mitochondria offers a promising approach due to their central role in energy production, oxidative stress, and cell-death pathways. Mitochondria-targeted therapies, including antioxidants, mitochondrial uncouplers, and agents that modulate mitochondrial permeability transition, have shown potential in preclinical studies. However, translating these findings into effective clinical therapies remains challenging due to issues such as off-target effects, limited tissue specificity, and inadequate delivery methods [60]. Hence, our findings underscore the importance of mitochondria as therapeutic targets in the management of ICM.
Finally, KEGG enrichment analysis showed cardiac muscle contraction and fatty acid degradation among the most positively enriched pathways in VCM compared to ICM. An enrichment of the cardiac muscle contraction pathway in VCM could be explained by the fact that volume overload is characterized by an eccentric (lengthening) muscle contraction as a consequence of the mechanical stress [61]. Fatty acids are the main energy substrate of the heart via ß-oxidation to produce ATP in mitochondria [61]. Therefore, fatty acid degradation pathway enrichment could be a secondary effect of the less dysfunctional state of mitochondria in VCM. In this sense, we also observed a positive enrichment of the PPAR signaling pathway, involved in fatty acid metabolism and mitochondrial function in the heart [62].
miRNA-seq analysis resulted in five miRNAs deregulated. miR-218-5p, miR-487b-3p, and miR-494-3p were upregulated, whereas miR-106b-3p and miR-193b-5p appeared downregulated in VCM vs. ICM. Among the 22 predicted miRNA–target transcript pairs, only four were negatively correlated, miR-218-5p-DDX6, miR-218-5p-TTC39C, miR-218-5p-SEMA4A, and miR-494-3p-SGMS2, and only three (miR-218-5p-DDX6, miR-218-5p-SEMA4A, and miR-494-3p-SGMS2) displayed physical interaction. DDX6 is an RNA helicase with roles in cellular stress and hypoxia [63], processing (P)-body homeostasis regulating stem-cell plasticity [64], and the miRNA pathway [65,66]. In this regard, our analysis shows that the DDX6 gene is involved in P-body formation in CC, the regulation of cellular macromolecule biosynthetic processes in BP, and is associated with double-stranded RNA binding in MF. Although little is known about its role in the heart, DDX6 has been associated with abnormal heart formation and described as modulating bone morphogenic protein (BMP) signaling [67]. The BMP subfamily belongs to the transforming growth factor beta (TGF-β) superfamily, which is well-documented to be upregulated in animal models of myocardial infarction [68]. In addition, BMP subfamily members have been suggested to exert both pro- and anti-inflammatory actions and may regulate fibrosis in myocardial infarction. Curiously, miR-218-5p, a potential miRNA targeting DDX6, has been also reported to mediate myocardial fibrosis and inflammation after myocardial infarction [69,70,71,72], suggesting therefore that the so far unknown DDX6/miR-218-5p axis could play a regulatory role during fibrosis and inflammation in the infarcted heart. Regarding the SEMA4A, which is involved in cell activation, immune response, T-cell differentiation, and lymphocyte differentiation according to BP, an in vivo cardiac ischemia–reperfusion mouse model showed SEMA4A to be highly expressed in macrophages recruited at the injured area, and it was suggested to exert a role activating angiogenesis and inflammation [73]. Accordingly, our results showed higher expression of SEMA4A in ICM than in VCM. Interestingly, miR-218-5p has been associated with inflammation in a myocardial ischemia–reperfusion injury model [71], suggesting, therefore, the SEMA4A/miR-218-5p axis as a potential target for regulating cardiac inflammation. It is well-known that the inflammatory response following an ischemic event can lead to adverse remodeling, cardiomyocyte injury, and matrix degradation, ultimately contributing to the development of HF. Targeting specific inflammatory pathways can potentially reduce infarct size, promote repair, and prevent adverse remodeling, offering a promising approach to improve outcomes in patients with ICM [74]. This positions inflammation as a therapeutic target for ICM in the context of diversifying therapies against ICM vs. VCM.
So far, nothing is known about the role of TTC39C in the heart. In fact, TTC39C is a little-studied gene. A Pubmed search only resulted in 11 results, and its function is unknown. To date, it has been only related to immunodeficiency [75]. Therefore, further studies are needed to unveil the role of this gene in heart disease. Finally, Sphingomyelin synthase 2 (SGMS2, also SMS2) was upregulated in ICM hearts compared to VCM. Concordantly, SGMS2 was found upregulated in cardiomyocytes in response to hypoxia in vitro [76]. This study concluded that downregulation of SGMS2 may protect cardiomyocytes against hypoxia-induced apoptosis and oxidative stress by enhancing the expression of nuclear factor erythroid 2-related factor 2 (Nrf2), an inhibitor of apoptosis [77]. Interestingly, two studies have reported miR-494-3p, which constitutes a miRNA–target transcript pair together SGMS2, to play a protective role in myocardial ischemia–reperfusion injury, repressing inflammation and apoptosis [78,79]. In addition, SGMS2 has been also described to attenuate inflammatory injury after cerebral ischemia–reperfusion in mice [80]. This indicates that the SGMS2/miR-494-3p axis may be a novel potential target against hypoxia-induced inflammation and/or apoptosis in cardiomyocytes. Treating cardiomyocyte apoptosis in ICM vs. VCM is important because it directly addresses the underlying mechanisms driving HF in each condition. In ICM, cardiomyocyte apoptosis occurs as a result of ischemic injury, leading to a loss of functional heart muscle tissue. This process significantly contributes to the progression of HF by reducing the heart’s ability to pump effectively. In contrast, in VCM, cardiomyocyte apoptosis may be less prominent and may occur due to factors related to valvular dysfunction rather than ischemia [81]. Therefore, targeting cardiomyocyte apoptosis in ICM could be essential to halt the progression of HF and improve outcomes for affected individuals.
To date, most studies have tested differentially expressed genes between ischemic and non-ischemic cardiomyopathy relative to a normal heart [2,23,82,83,84], offering novel insight into unique disease-specific gene expression that exists between different etiologies. However, our approach goes beyond this by directly comparing the transcriptomic profiles of two distinct etiologies of DCM: VCM and ICM. This direct comparison allows us to identify unique gene-expression signatures associated with each etiology, as well as common pathways dysregulated in DCM as a whole. Understanding these differences is crucial for developing personalized treatments tailored to each etiology, potentially improving patient outcomes. Our study offers insights into the molecular heterogeneity of DCM and paves the way for more precise therapeutic interventions.
In conclusion, we found distinct gene-expression profiles in VCM and ICM heart tissues, indicating that distinct mechanisms are involved in the progression of the diseases towards end-stage HF. In addition, we found five differentially expressed miRNAs and identified four previously undescribed miRNA–target transcript pairs that may open new avenues for research into the molecular pathogenesis of DCM.
Despite the valuable insights provided by our study, it is crucial to acknowledge limitations. The small number of differentially expressed miRNAs observed between VCM and ICM is notable. This limitation is largely due to the difficulty in accessing human myocardial tissue samples. Additionally, the inherent variability in human tissue samples complicates analysis and may contribute to the modest number of differentially expressed miRNAs identified. Nevertheless, these findings offer valuable insights into the molecular differences between VCM and ICM, highlighting the potential role of miRNAs in DCM etiologies. Future research with larger sample sizes and comprehensive analyses will be essential to address these limitations and advance our understanding of DCM pathogenesis.
Supplementary Materials
The following supporting information can be downloaded at https://www.mdpi.com/article/10.3390/biom14050524/s1: Table S1: differentially expressed genes; Table S2: miRNA raw counts table; Table S3: differentially expressed miRNAs; Table S4: top 30 enriched GO terms categorized by the CC. Table S5: top 40 enriched GO terms categorized by the BP; Table S6: top 26 enriched GO terms categorized by the MF.
Author Contributions
Conceptualization, R.T., F.B. and F.H.-T.; methodology, F.B., F.H.-T., E.A.-V., M.R.-S., M.Q.-F., A.B.-G., T.D. and C.G.-P.; software, F.H.-T., F.B. and R.T.; validation, F.B., E.A.-V., M.R.-S., M.Q.-F., A.B.-G., T.D. and C.G.-P.; formal analysis, R.T., F.B. and F.H.-T.; investigation, R.T., F.B. and F.H.-T.; resources, R.T. and A.M.; data curation, F.B., R.T. and F.H.-T.; writing—original draft preparation, F.B., F.H.-T. and R.T.; writing—review and editing, F.B., F.H.-T., M.R.-S., M.Q.-F., E.A.-V., C.G.-P., A.M. and R.T.; visualization, F.B., F.H.-T., M.R.-S., M.Q.-F., A.M. and R.T.; supervision, R.T. and A.M.; project administration, R.T. and A.M.; funding acquisition, R.T. All authors have read and agreed to the published version of the manuscript.
Funding
This work was supported by grants in the framework of the European Regional Development Fund (ERDF) Integrated Territorial Initiative (ITI0017_2019) and Fundación de Progreso y Salud PEER (2020-019).
Institutional Review Board Statement
This study was approved by the Andalusian Biomedical Research Ethics Committee (CEICA, reference number PI17/0023). Ethical research principles were followed in accordance with the Helsinki Declaration and the Belmont Report. The study also complied with two legal provisions on research involving human research and the Spanish Organic Law 15/1999 for the Regulation of Automated Processing of Personal Data. All patients gave written informed consent before surgery and prior to their inclusion in the study.
Informed Consent Statement
Informed consent was obtained from all subjects involved in the study.
Data Availability Statement
RNA-seq data generated in this manuscript have been deposited in GEO (Gene Expression Omnibus) of NCBI under accession code GSE243406 study.
Acknowledgments
The authors would like to thank the European Molecular Biology Laboratory GeneCore, Genomics Core Facility (Heidelberg, Germany) staff for processing our samples study.
Conflicts of Interest
The authors declare no conflicts of interest. The funders had no role in the design of the study; in the collection, analyses, or interpretation of data; in the writing of the manuscript; or in the decision to publish the results.
References
- Fatkin, D. Guidelines for the Diagnosis and Management of Familial Dilated Cardiomyopathy. Heart Lung Circ. 2011, 20, 691–693. [Google Scholar] [CrossRef] [PubMed]
- Kittleson, M.M.; Ye, S.Q.; Irizarry, R.A.; Minhas, K.M.; Edness, G.; Conte, J.V.; Parmigiani, G.; Miller, L.W.; Chen, Y.; Hall, J.L.; et al. Identification of a Gene Expression Profile That Differentiates between Ischemic and Nonischemic Cardiomyopathy. Circulation 2004, 110, 3444–3451. [Google Scholar] [CrossRef] [PubMed]
- Merlo, M.; Gentile, P.; Artico, J.; Cannatà, A.; Paldino, A.; De Angelis, G.; Barbati, G.; Alonge, M.; Gigli, M.; Pinamonti, B.; et al. Arrhythmic Risk Stratification in Patients with Dilated Cardiomyopathy and Intermediate Left Ventricular Dysfunction. J. Cardiovasc. Med. 2019, 20, 343–350. [Google Scholar] [CrossRef] [PubMed]
- Merlo, M.; Cannatà, A.; Gobbo, M.; Stolfo, D.; Elliott, P.M.; Sinagra, G. Evolving Concepts in Dilated Cardiomyopathy. Eur. J. Heart Fail. 2018, 20, 228–239. [Google Scholar] [CrossRef] [PubMed]
- Liew, C.-C.; Dzau, V.J. Molecular Genetics and Genomics of Heart Failure. Nat. Rev. Genet. 2004, 5, 811–825. [Google Scholar] [CrossRef] [PubMed]
- Hannenhalli, S.; Putt, M.E.; Gilmore, J.M.; Wang, J.; Parmacek, M.S.; Epstein, J.A.; Morrisey, E.E.; Margulies, K.B.; Cappola, T.P. Transcriptional Genomics Associates FOX Transcription Factors with Human Heart Failure. Circulation 2006, 114, 1269–1276. [Google Scholar] [CrossRef] [PubMed]
- Dewaswala, N.; Chait, R. Aortic Regurgitation. In StatPearls; StatPearls Publishing: Treasure Island, FL, USA, 2023. [Google Scholar]
- Walpot, J.; Vermeiren, G.L.; Al Mafragi, A.; Malbrain, M.L.N.G. Comprehensive Assessment of the Aortic Valve in Critically Ill Patients for the Non-Cardiologist. Part II: Chronic Aortic Regurgitation of the Native Valve. Anaesthesiol. Intensive Ther. 2021, 53, 55–68. [Google Scholar] [CrossRef]
- Otto, C.M.; Nishimura, R.A.; Bonow, R.O.; Carabello, B.A.; Erwin, J.P.; Gentile, F.; Jneid, H.; Krieger, E.V.; Mack, M.; McLeod, C.; et al. 2020 ACC/AHA Guideline for the Management of Patients with Valvular Heart Disease: A Report of the American College of Cardiology/American Heart Association Joint Committee on Clinical Practice Guidelines. Circulation 2021, 143, e72–e227. [Google Scholar] [CrossRef]
- Callis, T.E.; Wang, D.-Z. Taking microRNAs to Heart. Trends Mol. Med. 2008, 14, 254–260. [Google Scholar] [CrossRef]
- Bartel, D.P. MicroRNAs: Target Recognition and Regulatory Functions. Cell 2009, 136, 215–233. [Google Scholar] [CrossRef]
- Friedman, R.C.; Farh, K.K.-H.; Burge, C.B.; Bartel, D.P. Most Mammalian mRNAs Are Conserved Targets of microRNAs. Genome Res. 2009, 19, 92–105. [Google Scholar] [CrossRef] [PubMed]
- Yuan, Z.; Murakoshi, N.; Xu, D.; Tajiri, K.; Okabe, Y.; Aonuma, K.; Murakata, Y.; Li, S.; Song, Z.; Shimoda, Y.; et al. Identification of Potential Dilated Cardiomyopathy-Related Targets by Meta-Analysis and Co-Expression Analysis of Human RNA-Sequencing Datasets. Life Sci. 2022, 306, 120807. [Google Scholar] [CrossRef] [PubMed]
- Calderon-Dominguez, M.; Belmonte, T.; Quezada-Feijoo, M.; Ramos-Sánchez, M.; Fernández-Armenta, J.; Pérez-Navarro, A.; Cesar, S.; Peña-Peña, L.; Vea, À.; Llorente-Cortés, V.; et al. Emerging Role of microRNAs in Dilated Cardiomyopathy: Evidence Regarding Etiology. Transl. Res. 2020, 215, 86–101. [Google Scholar] [CrossRef] [PubMed]
- Alonso-Villa, E.; Bonet, F.; Hernandez-Torres, F.; Campuzano, Ó.; Sarquella-Brugada, G.; Quezada-Feijoo, M.; Ramos, M.; Mangas, A.; Toro, R. The Role of MicroRNAs in Dilated Cardiomyopathy: New Insights for an Old Entity. Int. J. Mol. Sci. 2022, 23, 13573. [Google Scholar] [CrossRef] [PubMed]
- Laggerbauer, B.; Engelhardt, S. MicroRNAs as Therapeutic Targets in Cardiovascular Disease. J. Clin. Investig. 2022, 132, e159179. [Google Scholar] [CrossRef] [PubMed]
- Sessa, F.; Salerno, M.; Esposito, M.; Cocimano, G.; Pomara, C. miRNA Dysregulation in Cardiovascular Diseases: Current Opinion and Future Perspectives. Int. J. Mol. Sci. 2023, 24, 5192. [Google Scholar] [CrossRef] [PubMed]
- Ikeda, S.; Kong, S.W.; Lu, J.; Bisping, E.; Zhang, H.; Allen, P.D.; Golub, T.R.; Pieske, B.; Pu, W.T. Altered microRNA Expression in Human Heart Disease. Physiol. Genom. 2007, 31, 367–373. [Google Scholar] [CrossRef]
- Yang, K.-C.; Yamada, K.A.; Patel, A.Y.; Topkara, V.K.; George, I.; Cheema, F.H.; Ewald, G.A.; Mann, D.L.; Nerbonne, J.M. Deep RNA Sequencing Reveals Dynamic Regulation of Myocardial Noncoding RNAs in Failing Human Heart and Remodeling With Mechanical Circulatory Support. Circulation 2014, 129, 1009–1021. [Google Scholar] [CrossRef] [PubMed]
- Poller, W.; Dimmeler, S.; Heymans, S.; Zeller, T.; Haas, J.; Karakas, M.; Leistner, D.-M.; Jakob, P.; Nakagawa, S.; Blankenberg, S.; et al. Non-Coding RNAs in Cardiovascular Diseases: Diagnostic and Therapeutic Perspectives. Eur. Heart J. 2018, 39, 2704–2716. [Google Scholar] [CrossRef]
- Mably, J.D.; Wang, D.-Z. Long Non-Coding RNAs in Cardiac Hypertrophy and Heart Failure: Functions, Mechanisms and Clinical Prospects. Nat. Rev. Cardiol. 2023, 21, 326–345. [Google Scholar] [CrossRef]
- Lu, P.; Ding, F.; Xiang, Y.K.; Hao, L.; Zhao, M. Noncoding RNAs in Cardiac Hypertrophy and Heart Failure. Cells 2022, 11, 777. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Morley, M.; Brandimarto, J.; Hannenhalli, S.; Hu, Y.; Ashley, E.A.; Tang, W.H.W.; Moravec, C.S.; Margulies, K.B.; Cappola, T.P.; et al. RNA-Seq Identifies Novel Myocardial Gene Expression Signatures of Heart Failure. Genomics 2015, 105, 83–89. [Google Scholar] [CrossRef] [PubMed]
- Sweet, M.E.; Cocciolo, A.; Slavov, D.; Jones, K.L.; Sweet, J.R.; Graw, S.L.; Reece, T.B.; Ambardekar, A.V.; Bristow, M.R.; Mestroni, L.; et al. Transcriptome Analysis of Human Heart Failure Reveals Dysregulated Cell Adhesion in Dilated Cardiomyopathy and Activated Immune Pathways in Ischemic Heart Failure. BMC Genom. 2018, 19, 812. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Liu, C.Y.; Li, Y.S.; Xu, J.; Li, D.G.; Han, D. Deep RNA Sequencing Elucidates microRNA-Regulated Molecular Pathways in Ischemic Cardiomyopathy and Nonischemic Cardiomyopathy. Genet. Mol. Res. 2016, 15, 15027465. [Google Scholar] [CrossRef] [PubMed]
- Levine, G.N.; Bates, E.R.; Bittl, J.A.; Brindis, R.G.; Fihn, S.D.; Fleisher, L.A.; Granger, C.B.; Lange, R.A.; Mack, M.J.; Mauri, L.; et al. 2016 ACC/AHA Guideline Focused Update on Duration of Dual Antiplatelet Therapy in Patients With Coronary Artery Disease: A Report of the American College of Cardiology/American Heart Association Task Force on Clinical Practice Guidelines: An Update of the 2011 ACCF/AHA/SCAI Guideline for Percutaneous Coronary Intervention, 2011 ACCF/AHA Guideline for Coronary Artery Bypass Graft Surgery, 2012 ACC/AHA/ACP/AATS/PCNA/SCAI/STS Guideline for the Diagnosis and Management of Patients With Stable Ischemic Heart Disease, 2013 ACCF/AHA Guideline for the Management of ST-Elevation Myocardial Infarction, 2014 AHA/ACC Guideline for the Management of Patients with Non–ST-Elevation Acute Coronary Syndromes, and 2014 ACC/AHA Guideline on Perioperative Cardiovascular Evaluation and Management of Patients Undergoing Noncardiac Surgery. Circulation 2016, 134, e123–e155. [Google Scholar] [CrossRef]
- Elliott, P.; Andersson, B.; Arbustini, E.; Bilinska, Z.; Cecchi, F.; Charron, P.; Dubourg, O.; Kuhl, U.; Maisch, B.; McKenna, W.J.; et al. Classification of the Cardiomyopathies: A Position Statement from the European Society of Cardiology Working Group on Myocardial and Pericardial Diseases. Eur. Heart J. 2007, 29, 270–276. [Google Scholar] [CrossRef] [PubMed]
- Grankvist, R.; Chireh, A.; Sandell, M.; Mukarram, A.K.; Jaff, N.; Berggren, I.; Persson, H.; Linde, C.; Arnberg, F.; Lundberg, J.; et al. Myocardial Micro-Biopsy Procedure for Molecular Characterization with Increased Precision and Reduced Trauma. Sci. Rep. 2020, 10, 8029. [Google Scholar] [CrossRef] [PubMed]
- Bustin, S.A.; Benes, V.; Garson, J.A.; Hellemans, J.; Huggett, J.; Kubista, M.; Mueller, R.; Nolan, T.; Pfaffl, M.W.; Shipley, G.L.; et al. The MIQE Guidelines: Minimum Information for Publication of Quantitative Real-Time PCR Experiments. Clin. Chem. 2009, 55, 611–622. [Google Scholar] [CrossRef] [PubMed]
- Afgan, E.; Baker, D.; Batut, B.; van den Beek, M.; Bouvier, D.; Čech, M.; Chilton, J.; Clements, D.; Coraor, N.; Grüning, B.A.; et al. The Galaxy Platform for Accessible, Reproducible and Collaborative Biomedical Analyses: 2018 Update. Nucleic Acids Res. 2018, 46, W537–W544. [Google Scholar] [CrossRef]
- Dobin, A.; Davis, C.A.; Schlesinger, F.; Drenkow, J.; Zaleski, C.; Jha, S.; Batut, P.; Chaisson, M.; Gingeras, T.R. STAR: Ultrafast Universal RNA-Seq Aligner. Bioinformatics 2013, 29, 15–21. [Google Scholar] [CrossRef]
- Liao, Y.; Smyth, G.K.; Shi, W. The R Package Rsubread Is Easier, Faster, Cheaper and Better for Alignment and Quantification of RNA Sequencing Reads. Nucleic Acids Res. 2019, 47, e47. [Google Scholar] [CrossRef] [PubMed]
- Paraskevopoulou, M.D.; Georgakilas, G.; Kostoulas, N.; Vlachos, I.S.; Vergoulis, T.; Reczko, M.; Filippidis, C.; Dalamagas, T.; Hatzigeorgiou, A.G. DIANA-microT Web Server v5.0: Service Integration into miRNA Functional Analysis Workflows. Nucleic Acids Res. 2013, 41, W169–W173. [Google Scholar] [CrossRef] [PubMed]
- Zhao, B.; Xue, B. Improving Prediction Accuracy Using Decision-Tree-Based Meta-Strategy and Multi-Threshold Sequential-Voting Exemplified by miRNA Target Prediction. Genomics 2017, 109, 227–232. [Google Scholar] [CrossRef] [PubMed]
- Sethupathy, P.; Corda, B.; Hatzigeorgiou, A.G. TarBase: A Comprehensive Database of Experimentally Supported Animal microRNA Targets. RNA 2006, 12, 192–197. [Google Scholar] [CrossRef] [PubMed]
- Huang, H.-Y.; Lin, Y.-C.-D.; Cui, S.; Huang, Y.; Tang, Y.; Xu, J.; Bao, J.; Li, Y.; Wen, J.; Zuo, H.; et al. miRTarBase Update 2022: An Informative Resource for Experimentally Validated miRNA-Target Interactions. Nucleic Acids Res. 2022, 50, D222–D230. [Google Scholar] [CrossRef] [PubMed]
- Vila-Casadesús, M.; Gironella, M.; Lozano, J.J. MiRComb: An R Package to Analyse miRNA-mRNA Interactions. Examples across Five Digestive Cancers. PLoS ONE 2016, 11, e0151127. [Google Scholar] [CrossRef] [PubMed]
- Väremo, L.; Nielsen, J.; Nookaew, I. Enriching the Gene Set Analysis of Genome-Wide Data by Incorporating Directionality of Gene Expression and Combining Statistical Hypotheses and Methods. Nucleic Acids Res. 2013, 41, 4378–4391. [Google Scholar] [CrossRef] [PubMed]
- Boyle, E.I.; Weng, S.; Gollub, J.; Jin, H.; Botstein, D.; Cherry, J.M.; Sherlock, G. GO::TermFinder—Open Source Software for Accessing Gene Ontology Information and Finding Significantly Enriched Gene Ontology Terms Associated with a List of Genes. Bioinformatics 2004, 20, 3710–3715. [Google Scholar] [CrossRef] [PubMed]
- Yu, G.; Wang, L.-G.; Han, Y.; He, Q.-Y. clusterProfiler: An R Package for Comparing Biological Themes among Gene Clusters. OMICS J. Integr. Biol. 2012, 16, 284–287. [Google Scholar] [CrossRef] [PubMed]
- Toro, R.; Pérez-Serra, A.; Mangas, A.; Campuzano, O.; Sarquella-Brugada, G.; Quezada-Feijoo, M.; Ramos, M.; Alcalá, M.; Carrera, E.; García-Padilla, C.; et al. miR-16-5p Suppression Protects Human Cardiomyocytes against Endoplasmic Reticulum and Oxidative Stress-Induced Injury. Int. J. Mol. Sci. 2022, 23, 1036. [Google Scholar] [CrossRef]
- Ali, Z.; Bhaskar, S.B. Basic Statistical Tools in Research and Data Analysis. Indian J. Anaesth. 2016, 60, 662–669. [Google Scholar] [CrossRef] [PubMed]
- Sinagra, G.; Elliott, P.M.; Merlo, M. Dilated Cardiomyopathy: So Many Cardiomyopathies! Eur. Heart J. 2020, 41, 3784–3786. [Google Scholar] [CrossRef] [PubMed]
- Frangogiannis, N.G. The Extracellular Matrix in Ischemic and Nonischemic Heart Failure. Circ. Res. 2019, 125, 117–146. [Google Scholar] [CrossRef] [PubMed]
- Frangogiannis, N.G. Cardiac Fibrosis. Cardiovasc. Res. 2021, 117, 1450–1488. [Google Scholar] [CrossRef] [PubMed]
- Frangogiannis, N.G. Fibroblasts and the Extracellular Matrix in Right Ventricular Disease. Cardiovasc. Res. 2017, 113, 1453–1464. [Google Scholar] [CrossRef] [PubMed]
- Nagatomo, Y.; Carabello, B.A.; Coker, M.L.; McDermott, P.J.; Nemoto, S.; Hamawaki, M.; Spinale, F.G. Differential Effects of Pressure or Volume Overload on Myocardial MMP Levels and Inhibitory Control. Am. J. Physiol.-Heart Circ. Physiol. 2000, 278, H151–H161. [Google Scholar] [CrossRef]
- Zheng, J.; Chen, Y.; Pat, B.; Dell’Italia, L.A.; Tillson, M.; Dillon, A.R.; Powell, P.C.; Shi, K.; Shah, N.; Denney, T.; et al. Microarray Identifies Extensive Downregulation of Noncollagen Extracellular Matrix and Profibrotic Growth Factor Genes in Chronic Isolated Mitral Regurgitation in the Dog. Circulation 2009, 119, 2086–2095. [Google Scholar] [CrossRef] [PubMed]
- Corporan, D.; Onohara, D.; Hernandez-Merlo, R.; Sielicka, A.; Padala, M. Temporal Changes in Myocardial Collagen, Matrix Metalloproteinases, and Their Tissue Inhibitors in the Left Ventricular Myocardium in Experimental Chronic Mitral Regurgitation in Rodents. Am. J. Physiol.-Heart Circ. Physiol. 2018, 315, H1269–H1278. [Google Scholar] [CrossRef] [PubMed]
- Irqsusi, M.; Mansouri, A.L.; Ramaswamy, A.; Rexin, P.; Salman, M.; Mahmood, S.; Mirow, N.; Ghazi, T.; Ramzan, R.; Rastan, A.J.; et al. Role of Matrix Metalloproteinases in Mitral Valve Regurgitation: Association between the of MMP-1, MMP-9, TIMP-1, and TIMP-2 Expression, Degree of Mitral Valve Insufficiency, and Pathologic Etiology. J. Card. Surg. 2022, 37, 1613–1622. [Google Scholar] [CrossRef]
- Kong, P.; Christia, P.; Frangogiannis, N.G. The Pathogenesis of Cardiac Fibrosis. Cell. Mol. Life Sci. 2014, 71, 549–574. [Google Scholar] [CrossRef]
- Swynghedauw, B. Molecular Mechanisms of Myocardial Remodeling. Physiol. Rev. 1999, 79, 215–262. [Google Scholar] [CrossRef] [PubMed]
- Okamura, K.; Nakagama, Y.; Takeda, N.; Soma, K.; Sato, T.; Isagawa, T.; Kido, Y.; Sakamoto, M.; Manabe, I.; Hirata, Y.; et al. Therapeutic Targeting of Mitochondrial ROS Ameliorates Murine Model of Volume Overload Cardiomyopathy. J. Pharmacol. Sci. 2019, 141, 56–63. [Google Scholar] [CrossRef] [PubMed]
- Ramaccini, D.; Montoya-Uribe, V.; Aan, F.J.; Modesti, L.; Potes, Y.; Wieckowski, M.R.; Krga, I.; Glibetić, M.; Pinton, P.; Giorgi, C.; et al. Mitochondrial Function and Dysfunction in Dilated Cardiomyopathy. Front. Cell Dev. Biol. 2021, 8, 624216. [Google Scholar] [CrossRef] [PubMed]
- Bisaccia, G.; Ricci, F.; Gallina, S.; Di Baldassarre, A.; Ghinassi, B. Mitochondrial Dysfunction and Heart Disease: Critical Appraisal of an Overlooked Association. Int. J. Mol. Sci. 2021, 22, 614. [Google Scholar] [CrossRef] [PubMed]
- Javadov, S.; Huang, C.; Kirshenbaum, L.; Karmazyn, M. NHE-1 Inhibition Improves Impaired Mitochondrial Permeability Transition and Respiratory Function during Postinfarction Remodelling in the Rat. J. Mol. Cell. Cardiol. 2005, 38, 135–143. [Google Scholar] [CrossRef] [PubMed]
- Marcil, M.; Ascah, A.; Matas, J.; Belanger, S.; Deschepper, C.; Burelle, Y. Compensated Volume Overload Increases the Vulnerability of Heart Mitochondria without Affecting Their Functions in the Absence of Stress. J. Mol. Cell. Cardiol. 2006, 41, 998–1009. [Google Scholar] [CrossRef] [PubMed]
- Singh, H. Mitochondrial Ion Channels in Cardiac Function. Am. J. Physiol.-Cell Physiol. 2021, 321, C812–C825. [Google Scholar] [CrossRef]
- Chang, X.; Liu, R.; Li, R.; Peng, Y.; Zhu, P.; Zhou, H. Molecular Mechanisms of Mitochondrial Quality Control in Ischemic Cardiomyopathy. Int. J. Biol. Sci. 2023, 19, 426–448. [Google Scholar] [CrossRef] [PubMed]
- Marin, W.; Marin, D.; Ao, X.; Liu, Y. Mitochondria as a Therapeutic Target for Cardiac Ischemia-reperfusion Injury (Review). Int. J. Mol. Med. 2020, 47, 485–499. [Google Scholar] [CrossRef]
- Lopaschuk, G.D.; Ussher, J.R.; Folmes, C.D.L.; Jaswal, J.S.; Stanley, W.C. Myocardial Fatty Acid Metabolism in Health and Disease. Physiol. Rev. 2010, 90, 207–258. [Google Scholar] [CrossRef]
- Montaigne, D.; Butruille, L.; Staels, B. PPAR Control of Metabolism and Cardiovascular Functions. Nat. Rev. Cardiol. 2021, 18, 809–823. [Google Scholar] [CrossRef] [PubMed]
- De Vries, S.; Naarmann-de Vries, I.S.; Urlaub, H.; Lue, H.; Bernhagen, J.; Ostareck, D.H.; Ostareck-Lederer, A. Identification of DEAD-Box RNA Helicase 6 (DDX6) as a Cellular Modulator of Vascular Endothelial Growth Factor Expression under Hypoxia*. J. Biol. Chem. 2013, 288, 5815–5827. [Google Scholar] [CrossRef] [PubMed]
- Di Stefano, B.; Luo, E.-C.; Haggerty, C.; Aigner, S.; Charlton, J.; Brumbaugh, J.; Ji, F.; Rabano Jiménez, I.; Clowers, K.J.; Huebner, A.J.; et al. The RNA Helicase DDX6 Controls Cellular Plasticity by Modulating P-Body Homeostasis. Cell Stem Cell 2019, 25, 622–638.e13. [Google Scholar] [CrossRef]
- Chu, C.; Rana, T.M. Translation Repression in Human Cells by MicroRNA-Induced Gene Silencing Requires RCK/P54. PLoS Biol. 2006, 4, e210. [Google Scholar] [CrossRef]
- Eulalio, A.; Rehwinkel, J.; Stricker, M.; Huntzinger, E.; Yang, S.-F.; Doerks, T.; Dorner, S.; Bork, P.; Boutros, M.; Izaurralde, E. Target-Specific Requirements for Enhancers of Decapping in miRNA-Mediated Gene Silencing. Genes Dev. 2007, 21, 2558–2570. [Google Scholar] [CrossRef]
- Kim, J.; Muraoka, M.; Okada, H.; Toyoda, A.; Ajima, R.; Saga, Y. The RNA Helicase DDX6 Controls Early Mouse Embryogenesis by Repressing Aberrant Inhibition of BMP Signaling through miRNA-Mediated Gene Silencing. PLoS Genet. 2022, 18, e1009967. [Google Scholar] [CrossRef]
- Hanna, A.; Frangogiannis, N.G. The Role of the TGF-β Superfamily in Myocardial Infarction. Front. Cardiovasc. Med. 2019, 6, 140. [Google Scholar] [CrossRef]
- Ke, X.; Yang, R.; Wu, F.; Wang, X.; Liang, J.; Hu, X.; Hu, C. Exosomal miR-218-5p/miR-363-3p from Endothelial Progenitor Cells Ameliorate Myocardial Infarction by Targeting the P53/JMY Signaling Pathway. Oxid. Med. Cell. Longev. 2021, 2021, 5529430. [Google Scholar] [CrossRef] [PubMed]
- Sun, B.; Zhao, C.; Mao, Y. MiR-218-5p Mediates Myocardial Fibrosis after Myocardial Infarction by Targeting CX43. Curr. Pharm. Des. 2021, 27, 4504–4512. [Google Scholar] [CrossRef]
- Yang, Y.; Zhao, F.; Yuan, Z.; Wang, C.; Chen, K.; Xiao, W. Inhibition of miR-218-5p Reduces Myocardial Ischemia–Reperfusion Injury in a Sprague-Dawley Rat Model by Reducing Oxidative Stress and Inflammation through MEF2C/NF-κB Pathway. Int. Immunopharmacol. 2021, 101, 108299. [Google Scholar] [CrossRef]
- Guo, J.; Chen, L.-W.; Huang, Z.-Q.; Guo, J.-S.; Li, H.; Shan, Y.; Chen, Z.-R.; Yan, Y.-M.; Zhu, J.-N.; Guo, H.-M.; et al. Suppression of the Inhibitory Effect of Circ_0036176-Translated Myo9a-208 on Cardiac Fibroblast Proliferation by miR-218-5p. J. Cardiovasc. Transl. Res. 2022, 15, 548–559. [Google Scholar] [CrossRef] [PubMed]
- Meda, C.; Molla, F.; De Pizzol, M.; Regano, D.; Maione, F.; Capano, S.; Locati, M.; Mantovani, A.; Latini, R.; Bussolino, F.; et al. Semaphorin 4A Exerts a Proangiogenic Effect by Enhancing Vascular Endothelial Growth Factor—A Expression in Macrophages. J. Immunol. 2012, 188, 4081–4092. [Google Scholar] [CrossRef] [PubMed]
- Saxena, A.; Russo, I.; Frangogiannis, N.G. Inflammation as a Therapeutic Target in Myocardial Infarction: Learning from Past Failures to Meet Future Challenges. Transl. Res. 2016, 167, 152–166. [Google Scholar] [CrossRef]
- Grant, F.M.; Yang, J.; Nasrallah, R.; Clarke, J.; Sadiyah, F.; Whiteside, S.K.; Imianowski, C.J.; Kuo, P.; Vardaka, P.; Todorov, T.; et al. BACH2 Drives Quiescence and Maintenance of Resting Treg Cells to Promote Homeostasis and Cancer Immunosuppression. J. Exp. Med. 2020, 217, e20190711. [Google Scholar] [CrossRef] [PubMed]
- Jin, A.; Cheng, H.; Xia, L.; Ye, S.; Yang, C. Inhibition of Sphingomyelin Synthase 2 Relieves Hypoxia-Induced Cardiomyocyte Injury by Reinforcing Nrf2/ARE Activation via Modulation of GSK-3β. Hum. Exp. Toxicol. 2021, 40, 791–800. [Google Scholar] [CrossRef] [PubMed]
- Niture, S.K.; Jaiswal, A.K. Nrf2 Protein Up-Regulates Antiapoptotic Protein Bcl-2 and Prevents Cellular Apoptosis. J. Biol. Chem. 2012, 287, 9873–9886. [Google Scholar] [CrossRef] [PubMed]
- Mu, N.; Zhang, T.; Zhu, Y.; Lu, B.; Zheng, Q.; Duan, J. The Mechanism by Which miR-494-3p Regulates PGC1-α-Mediated Inhibition of Mitophagy in Cardiomyocytes and Alleviation of Myocardial Ischemia—Reperfusion Injury. BMC Cardiovasc. Disord. 2023, 23, 204. [Google Scholar] [CrossRef] [PubMed]
- Zheng, W.; Xie, Q.; Zhang, Z.; Li, J.; Fang, L.; Li, W. Inhibited HDAC3 or Elevated MicroRNA-494-3p Plays a Protective Role in Myocardial Ischemia-Reperfusion Injury via Suppression of BRD4. Mol. Neurobiol. 2021, 58, 4268–4279. [Google Scholar] [CrossRef] [PubMed]
- Xue, J.; Yu, Y.; Zhang, X.; Zhang, C.; Zhao, Y.; Liu, B.; Zhang, L.; Wang, L.; Chen, R.; Gao, X.; et al. Sphingomyelin Synthase 2 Inhibition Ameliorates Cerebral Ischemic Reperfusion Injury Through Reducing the Recruitment of Toll-like Receptor 4 to Lipid Rafts. J. Am. Heart Assoc. 2019, 8, e012885. [Google Scholar] [CrossRef]
- Sabbah, H. Apoptotic Cell Death in Heart Failure. Cardiovasc. Res. 2000, 45, 704–712. [Google Scholar] [CrossRef]
- Kittleson, M.M.; Minhas, K.M.; Irizarry, R.A.; Ye, S.Q.; Edness, G.; Breton, E.; Conte, J.V.; Tomaselli, G.; Garcia, J.G.N.; Hare, J.M. Gene Expression Analysis of Ischemic and Nonischemic Cardiomyopathy: Shared and Distinct Genes in the Development of Heart Failure. Physiol. Genom. 2005, 21, 299–307. [Google Scholar] [CrossRef] [PubMed]
- Kuner, R.; Barth, A.S.; Ruschhaupt, M.; Buness, A.; Zwermann, L.; Kreuzer, E.; Steinbeck, G.; Poustka, A.; Sültmann, H.; Nabauer, M. Genomic Analysis Reveals Poor Separation of Human Cardiomyopathies of Ischemic and Nonischemic Etiologies. Physiol. Genom. 2008, 34, 88–94. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Onrat, S.T.; Onrat, E.; Ercan Onay, E.; Yalım, Z.; Avşar, A. The Genetic Determination of the Differentiation between Ischemic Dilated Cardiomyopathy and Idiopathic Dilated Cardiomyopathy. Genet. Test. Mol. Biomark. 2018, 22, 644–651. [Google Scholar] [CrossRef] [PubMed]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).