1. Introduction
Bacillus thuringiensis (
Bt) is a Gram-positive, spore-forming bacterium which produces crystalline inclusions during sporulation [
1]. These crystalline bodies can house one or more proteins (such as Cry toxins) that possess considerable efficacy and specificity towards invertebrates, which has made these
Bt Cry toxins valuable to agronomic interests, both as insecticidal agents and as genes expressed in transgenic crops [
1]. Cry toxin insect specificity is derived from numerous factors, which include crystal dissolution via compatible pH conditions of the invertebrate midgut, proteolytic cleavage of the protoxin by means of a compatible gut protease, and receptor recognition and binding [
1,
2].
Cry toxins possess a three-domain toxic core structure, which is highly conserved despite significant divergences in their sequence and target specificity [
1]. They all require the same criteria for functionality, namely an environment in which to solubilize the crystal-encased protein, and the proteolytic processing of the N-terminus required for activation [
2]. Further still, many known receptors for these Cry toxins are shared across different orders of insects, such as the glycosyl-phosphatidylinositol (GPI) anchored aminopeptidases and alkaline phosphatases [
2]. Though the Cry toxins are mostly known and commercialized for their efficacy against insects, a small subgroup of Cry toxins known as the parasporins possess activity against human cancer cells whilst having no known invertebrate targets. Parasporin-3, also known as Cry41Aa, is toxic towards some human cancer cells, which include hepatocellular carcinoma and myeloid leukaemia [
3]. We also know that proteolytic activation is required for this activity [
4]. Amongst all the specificity determinants between Cry proteins and their targets, receptor interaction is thought to be the most significant step in determining the cytotoxicity of a Cry protein [
2]. Multiple receptors have been identified and their interactions with Cry toxins characterized, including some for parasporins [
5]. These include Beclin-1 for Parasporin 1 and GPI-anchor protein receptors for Parasporin 2 [
6]. The mode of action of a Cry protein may not be limited to one type of receptor; studies have suggested that a toxin might interact with multiple receptors in each insect species and that the interactions between these receptors might enhance the toxic effect [
7]. In this work, we have explored the binding characteristics of the Cry41Aa protoxin against hepatocellular carcinoma cells and have demonstrated cross-over between Cry41Aa and some insecticidal Cry toxins, speaking to a closer evolutionary relationship.
2. Materials and Methods
2.1. Strains, Plasmids, and Constructs
2.1.1. Bacillus thuringiensis Strain 4D7
An acrystalliferous strain of Bacillus thuringiensis subsp. kurstaki was used for the production of all Cry toxins within this work. This strain came from the Bacillus Genetic Stock Center (Ohio State University, OH, USA).
2.1.2. Toxin Expression Plasmids
pHT315 is an
E. coli/
Bt shuttle vector containing an origin of replication for both
E. coli and
Bt, as well as resistance genes for ampicillin and erythromycin [
8]. We have previously used this plasmid as the basis of an expression vector for Cry2Aa and Cry2Ab [
9]. In this work, pHTCry41Aa was created by replacing the ORF2 and
cry2A genes from these constructs with
cry41Aa and its ORF3 gene. For pHTCry41AaGFP, the non-essential beta-trefoil domain at the C-terminus of Cry41Aa [
3] was deleted and GFP was added after the 713th amino acid. To prevent cleavage of the GFP during activation, a unique PreScission protease site (LEVLFQGP) was added after the 40th amino acid, as previously described [
4]. In pHTCry41AaHA, an HA tag (YPYDVPDYA) was inserted in pHTCry41Aa after the 825th codon. To create the disabled (DIP) version of Cry41Aa, an A126E mutation was introduced into pHTCry41AaHA. Cry1Ac, Cry1Ca, Cry1Da and Cry4Aa expression plasmids were created by replacing the
cry2Aa gene from pHTCry2Aa with the alternative coding region. A DIP form of Cry1Ca was created by introducing the mutations N98C and D143C, as identified by Jerga et al. [
10]. Finally, Cry11Aa was expressed from a construct in which the gene was cloned in pSVP27 [
11].
2.2. Growth, Solubilization and Activation of Cry Toxins
Bacillus thuringiensis bacteria, containing the Cry toxin genes used in these experiments, were inoculated to flasks containing 1 L half concentration LB broth with the addition of 5 µg/mL erythromycin (chloramphenicol for Cry11Aa expression) and incubated for 3 days at 30 °C, 200 rpm. Several rounds of sonication and centrifugation were undertaken to fully lyse the cells before the spores and crystals were resuspended in ddH2O. All Cry toxins were solubilized by pelleting the crude cell lysate, followed by resuspension in 50 mM Na2CO3 pH 10.5 containing 5 mM dithiothreitol (DTT) and incubation at 37 °C for one hour. Samples were centrifuged at 16,873 g for 2 min, and the supernatant was collected. For the activation of Cry41AaGFP, Pierce HRV 3C protease (1 Unit per 100 µg Thermo Fisher Scientific, Waltham, MA, USA) was used in PreScission reaction buffer and left for 16 h at 4 °C. The enzymatic reaction was stopped by the addition of a Complete mini EDTA-free proteinase inhibitor cocktail solution (Sigma-Aldrich, St. Louis, MO, USA). Cry41Aa and Cry41HADIP were activated by treating the solubilized samples with 1 mg/mL chymotrypsin at 37 °C for one hour. For Cry1Ca and Cry11Aa, trypsin was used instead of chymotrypsin.
2.3. Cell Culture
The adherent hepatocellular carcinoma cell line HepG2, purchased from ECACC, was routinely cultured in nunclon surface-treated T-25, T-75, and T-175 flasks (Nunc), using a high glucose DMEM (4.5 g/L, Gibco Life Technologies, Waltham, MA, USA) containing 1% penicillin-streptomycin-L-glutamate (PSG; Gibco Life Technologies) and 10% fetal bovine serum (FCS; Gibco Life Technologies) and incubated at 37 °C with 5% CO2 humidified air.
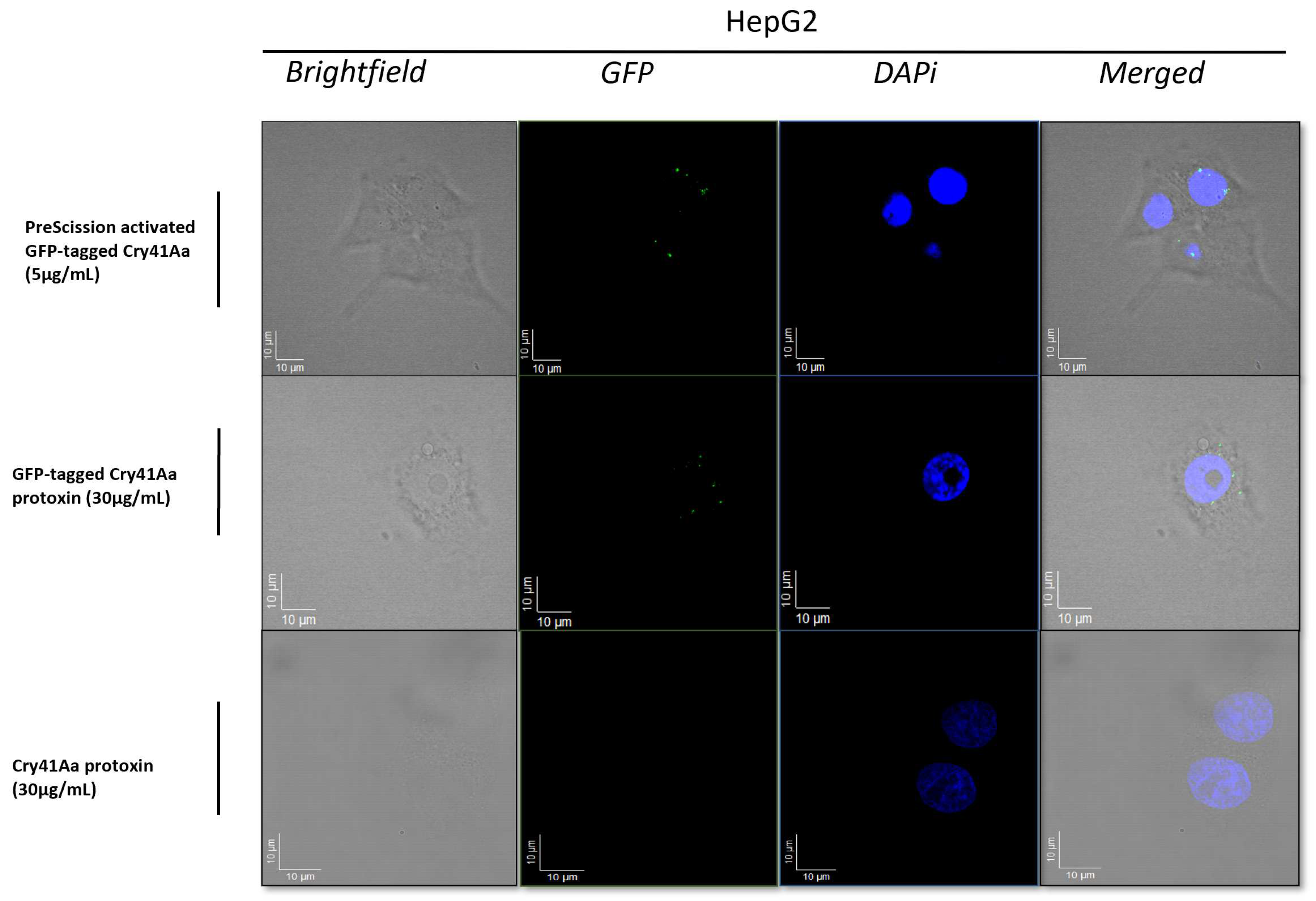
2.4. Fluorescence Microscopy Using GFP-Labelled Cry41Aa
HepG2 cells were placed on a sterile microscope cover glass at a density of 2.5 × 105 cells/mL in serum-free DMEM treated with PSG (1%) for 16 h in a 6-well culture plate. Appropriate treatments were added to each well for one hour, the medium was aspirated, and each well washed with PBS. A 4% formaldehyde solution (Sigma-Aldrich) was used to fix the samples. Coverslips were mounted on microscope glass slides using mounting media (plus DAPI), and the edges were sealed. The following parameters were set on the fluorescence microscope (Zeiss LSM880, Oberkochen, Germany): a 64× objective (with immersion oil) was used to obtain a Z-stack scan of cells from bottom to top (cell surface) to obtain micrographs of the whole cell surface. A GFP filter was used to scan the cells for bound GFP-tagged Cry41Aa on the surface of treated cells, whereas a blue filter was used to visualize DAPI staining.
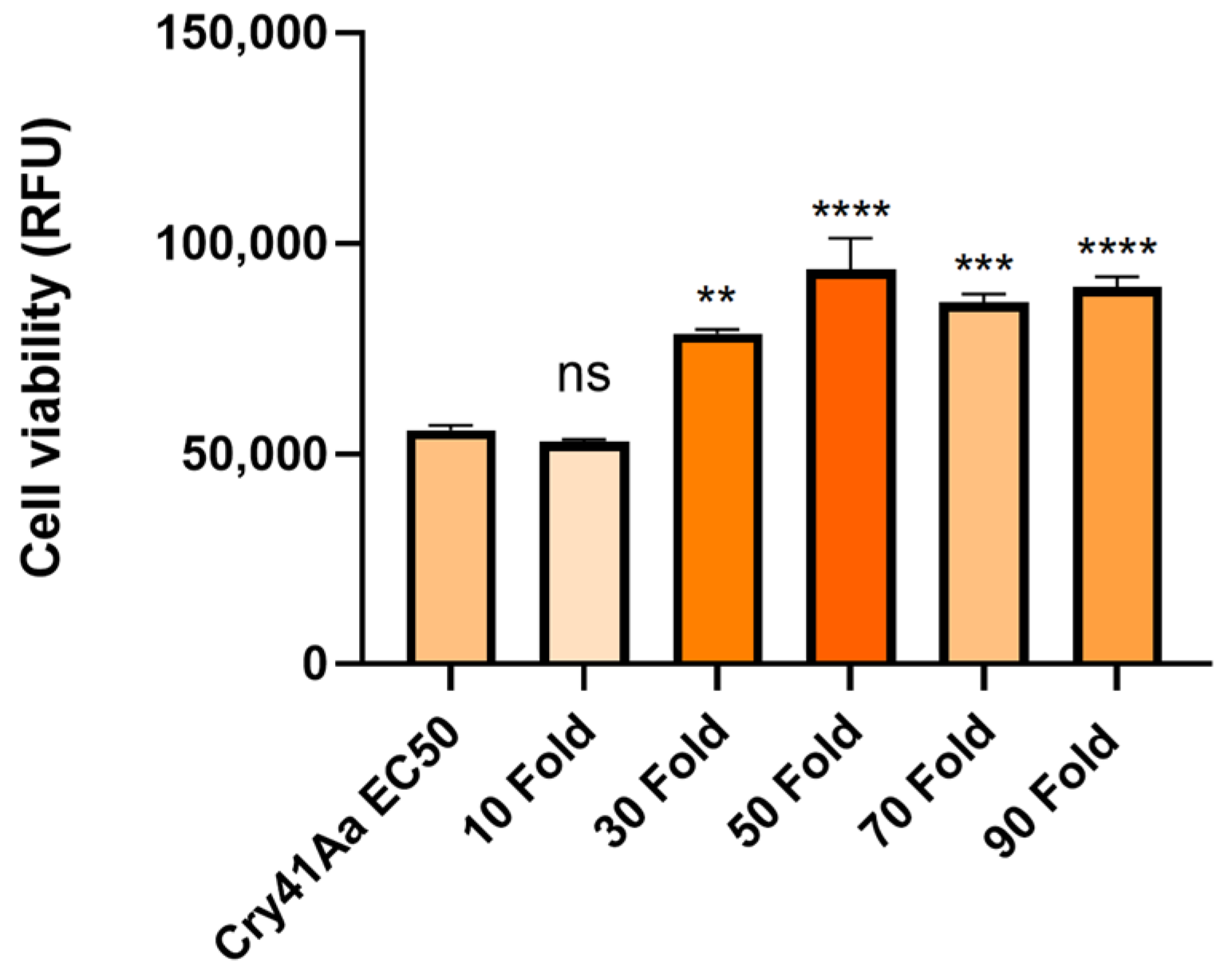
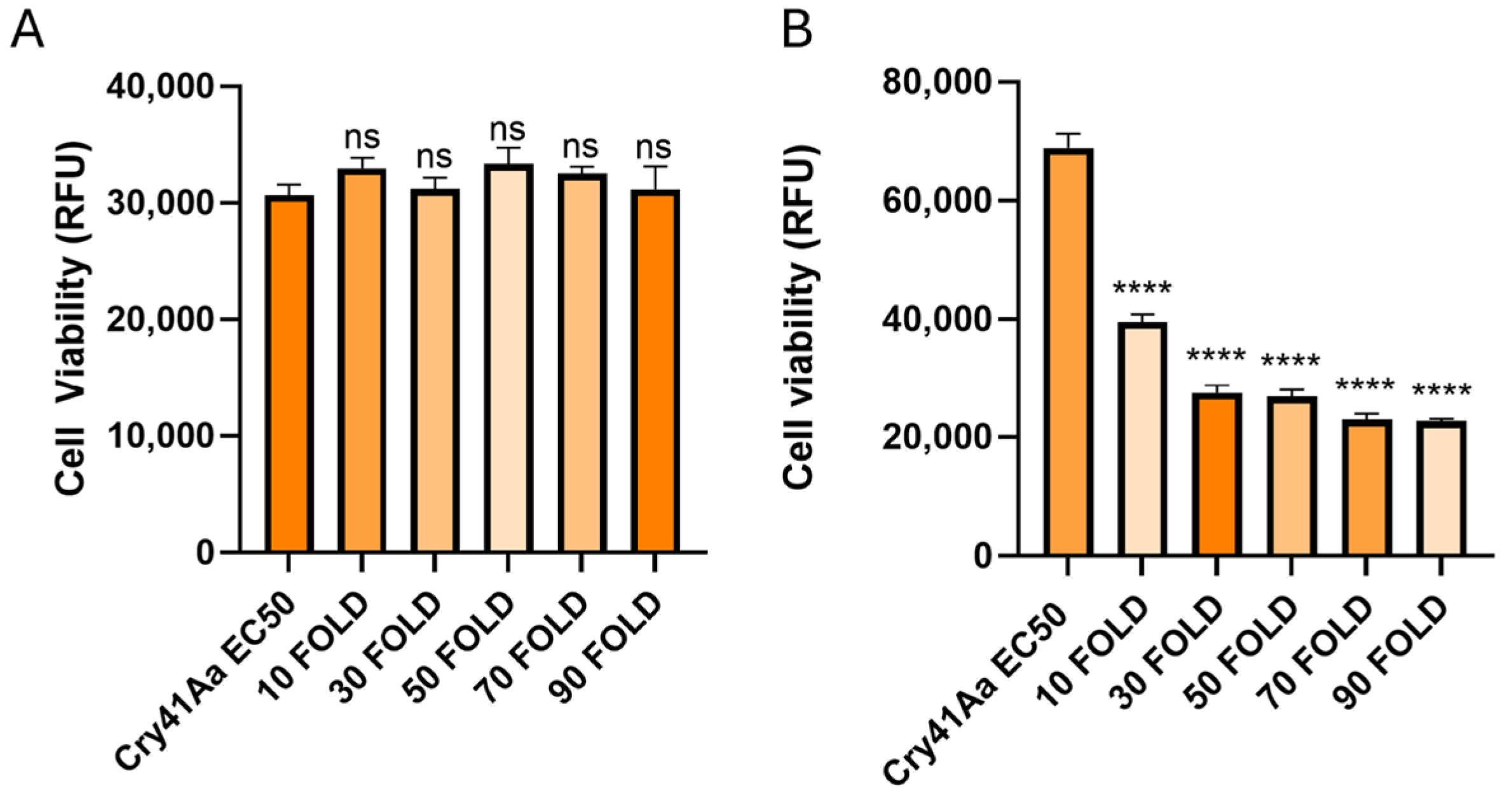
2.5. Cellular Competition Assays
Cellular competition assays were carried out in 96-well culture plates (Nunc). Cells were seeded in a volume of 90 µL at a concentration of 2.5 × 105 cells/mL, and allowed to adhere overnight whilst incubated at 37 °C with 5% CO2 humidified air. A total of 10 µL of either experimental or control samples were added to wells in triplicate for each condition. For the experimental wells, combinations of Cry41Aa at an EC50 dose of 2 µg/mL and 10–180 fold amounts of competitor were added. Control wells included the application of 10 µL buffer only (50 mM Na2CO3 pH 10.5), Cry41Aa alone, the competitor toxin at the highest concentration used, or Triton X-100 (0.1%). DMEM was added to additional wells at a volume of 100 µL in triplicate to act as a background fluorescent control. The cells were then incubated for 24 h.
Cell viability was measured the following day using the CellTiter-Blue cell viability assay kit (Promega, Chuo City, Japan). A total of 20 µL of the CellTiter-Blue reagent alamarBlue was added to each well. The plate was incubated for 2 h, before reading using a GloMax-Multi Detection System from Promega. The fluorescence was measured using a green filter with an excitation wavelength of 525 nm and an emission wavelength range of 580–640 nm.
2.6. Cellular Binding Assays
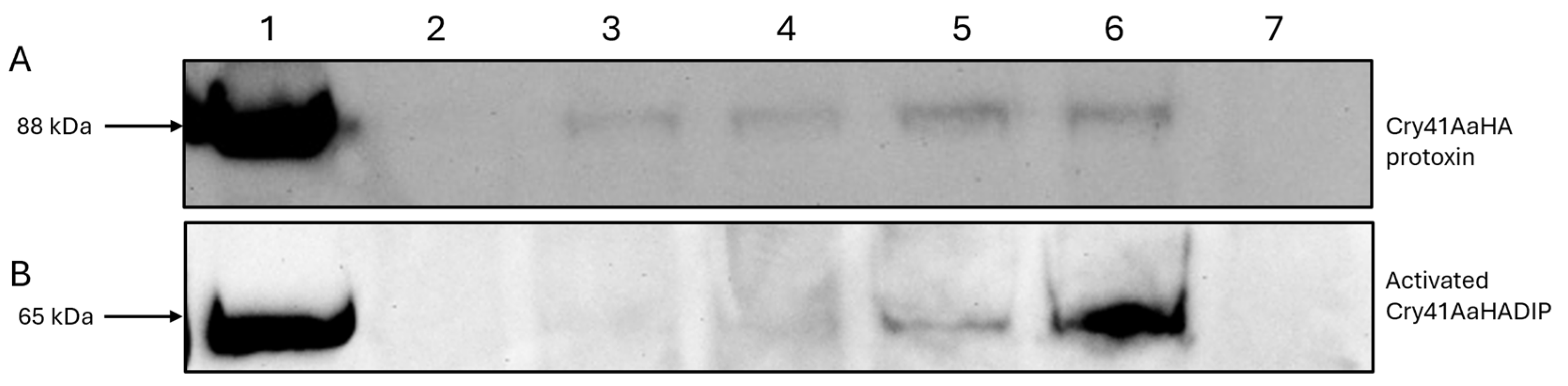
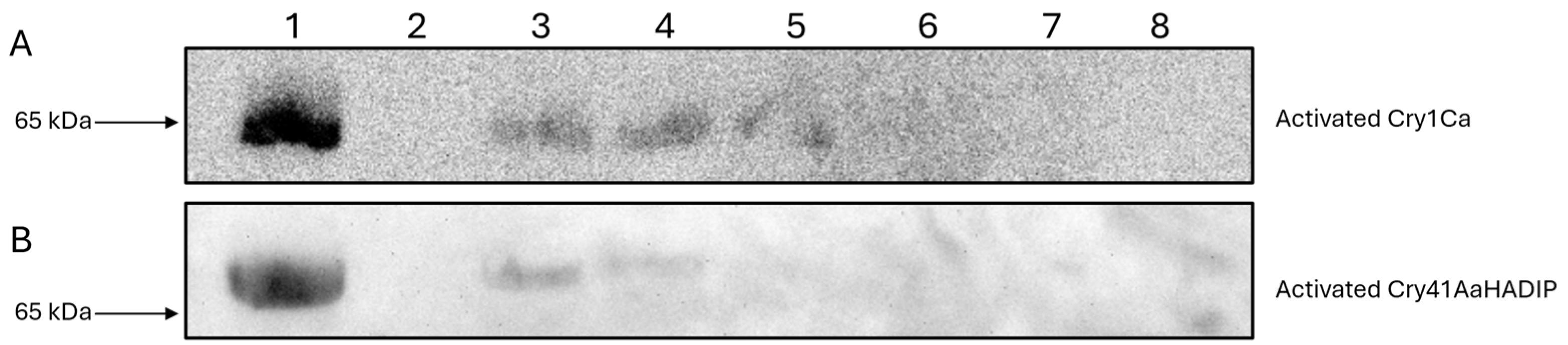
Cells were seeded to 6-well culture plates (Nunc), 1 mL per well at a concentration of 1 × 106 cells/mL, and incubated overnight to allow cells to adhere to the plates. Existing DMEM within the 6-well plates was aspirated off, and the toxin/DMEM solutions were added, with a control well receiving only 50 mM Na2CO3 pH 10.5 buffer. Plates were incubated in the presence of toxin for 1 h. After incubation, DMEM and any unbound toxin were aspirated off, and each well was washed twice in PBS. A total of 70 µL of resuspension buffer (20 mM Tris HCl pH 8 containing 1× phosphatase inhibitors and 1× protease inhibitors) was added to each well, and cells were transferred into the buffer using cell scrapers (Corning, Corning, NY, USA). A total of 70 µL of whole cell lysate was collected from each well and placed in ice-cold Eppendorf tubes to be later analyzed via Western blotting. All binding experiments were repeated at least three times, and the blots contained within this paper are representative of each experimental set.
2.7. Western Blotting
Whole cell lysates (10 µL) were combined with 10 µL of 2× SDS-ME (SDS loading buffer + 5% β-mercaptoethanol) and heated at 98 °C for 10 min. These samples were loaded onto a 7.5% SDS PAGE gel at 15 µL per well and ran for 40 min at 200 V. The gel was removed and incubated in semi dry transfer buffer for 5 min while shaking. The protein samples were then transferred to a nitrocellulose membrane (Bio-Rad, Hercules, CA, USA, 0.45 µm) using a Bio-Rad Trans-Blot Semi-Dry Transfer Cell system at 100 mA for 1 h. The nitrocellulose membrane was then equilibrated in 20 mL of 1× PBS buffer for 5 min before incubation for 1 h with 10 mL blocking buffer (1× PBS containing 0.02% tween-20 and 5% dry milk). The blocking buffer was poured off, and the membrane was incubated for a further hour in 5 mL of blocking buffer containing 5 µL of an HRP conjugated anti-HA antibody (Abcam, Cambridge, UK). The membrane was washed in PBS-T 3 times, with a 5 min incubation with each wash. The last wash was poured off, and the membrane was incubated in an enhanced luminol-based chemiluminescent detection (ECL) buffer for 2 min (10 mL of 100 mM Tris HCl pH 8.5, 3 µL H2O2, 25 µL p-coumaric acid (14.7 mg/mL in DMSO), and 50 µL luminol (88.6 mg/mL in DMSO). The signal was then detected using a UVP ChemStudio imaging system with a 10 min exposure time and displayed within the UVP Life Science VisionWorks software (version 8.20.17096.9551).
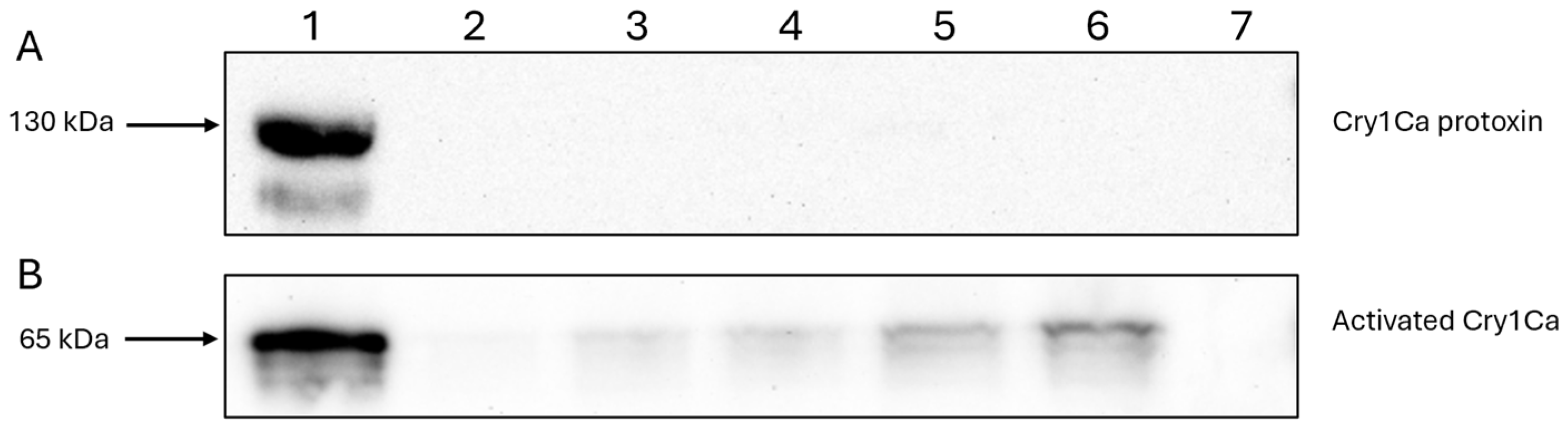
For detecting Cry1Ca, the Western blotting process remained the same, save for the use of an in house Cry1C specific primary antibody and a secondary anti-rabbit IgG HRP conjugated antibody (Cell Signalling Technology, Danvers, MA, USA).
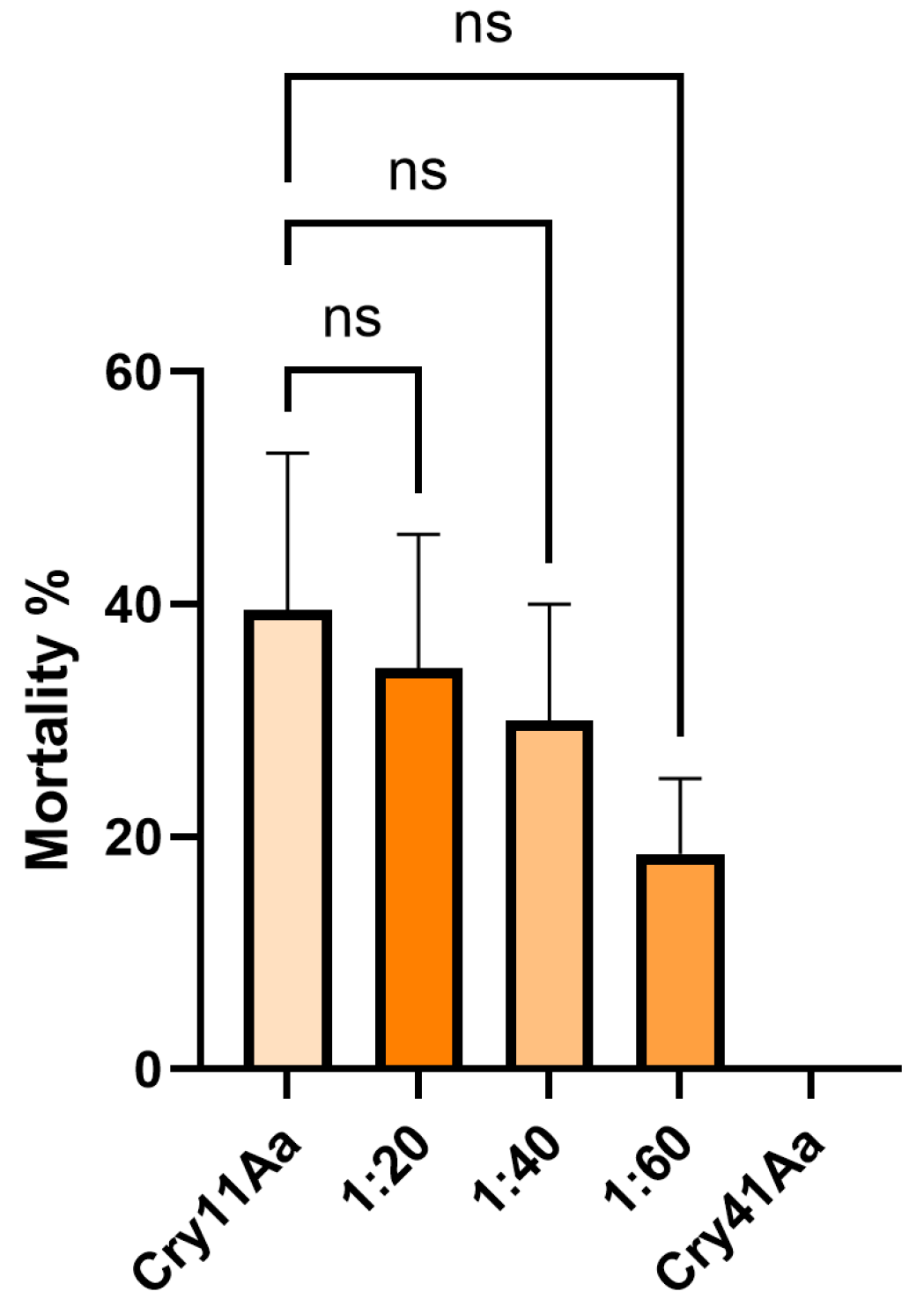
2.8. Aedes aegypti Bioassays
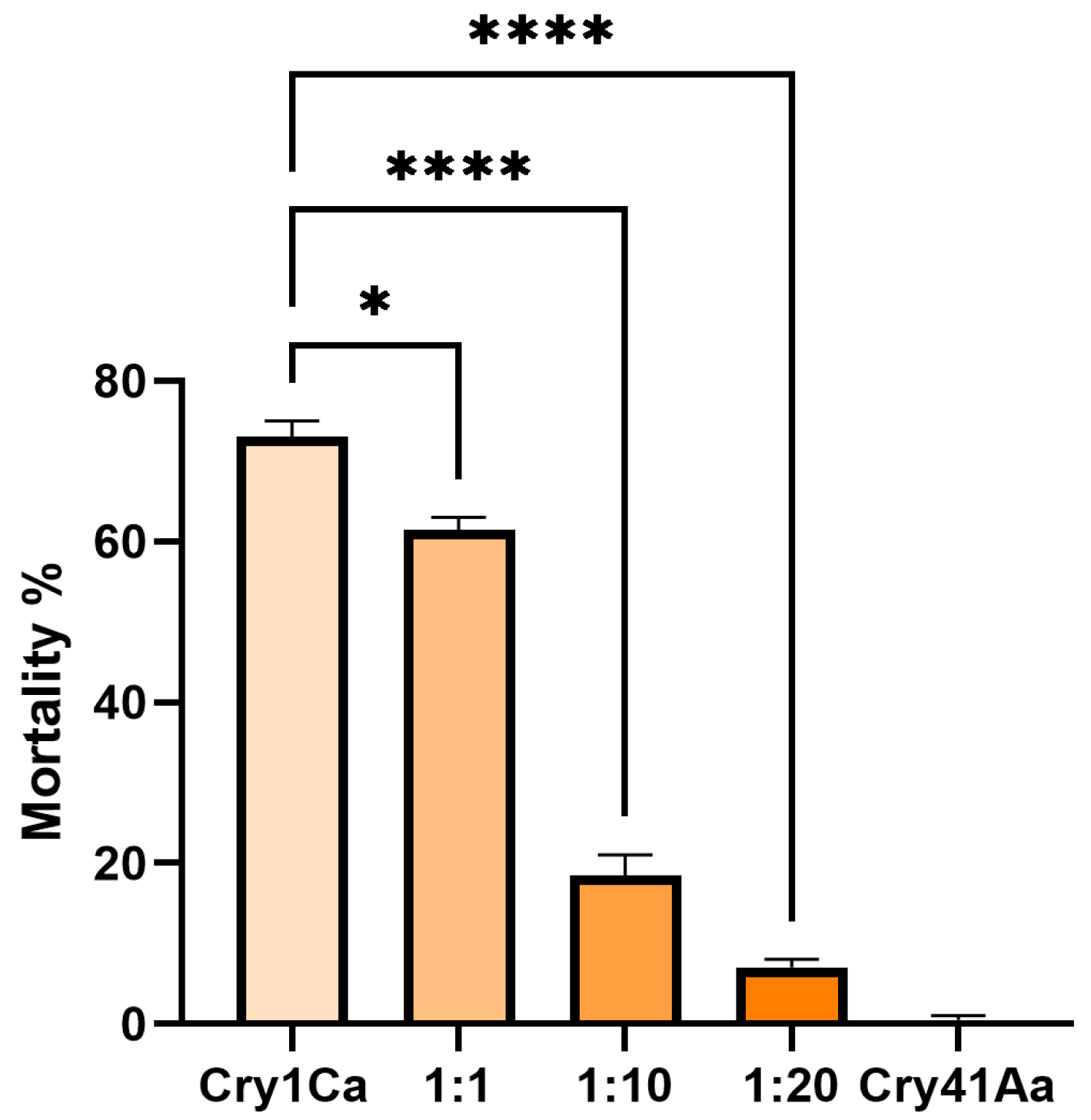
Aedes aegypti eggs were originally provided by InfraVec2, the adults were fed on defibrinated horse blood and a 10% sucrose solution, and the larvae on goldfish flakes. The insects were kept at 27 °C, 65% humidity. Larvae were selected at the 3rd instar stage of development and placed in 24-well plates, 5 larvae per well and 100 larvae total per condition. Cry1Ca was added at 25 µg/mL, and Cry41Aa was added as a competitor at various concentrations. Water-only and Cry41Aa-only conditions were used as controls. Water in the wells was topped up to a total volume of 3 mL. Larvae were incubated at 27 °C for 24 h, after which the total alive and dead larvae for each condition were recorded.
2.9. Data Analysis
All data analysis was carried out using GraphPad Prism version 10.2.2 (Dotmatics, Boston, MA, USA).
4. Discussion
N-terminal activation is believed to be an important step in the mode of action of Cry toxins [
12]. Cross-linking studies, using disulfide bonds, have shown that the activation of Cry toxins through the cleavage of the N-terminal peptide segment results in a conformational change, leading to domain I moving away from domains II and III, the insertion of the helical bundle into the membrane, and pore formation [
14,
15]. Bravo et al., 2007, postulated that this conformational change may expose a region within domain II responsible for toxin–receptor interaction [
16]. Our work shows that N-terminal cleavage of Cry41Aa is not required for the toxin to bind to HepG2 cells. It is thought that Cry41Aa, like other Cry toxins, still requires proteolytic processing to insert and form a pore within the membrane, as evidenced by the fact the protoxin is not toxic towards cells [
3]. The binding of the Cry41Aa protoxin was confirmed through both fluorescent microscopy and immunoblotting, and it was found to compete with the activated toxin, indicating shared binding sites. Though the Cry41Aa protoxin has been proven to be non-toxic towards HepG2 cells even at relatively high concentrations, a lack of toxicity at any concentration cannot be ruled out [
3]. Tabashnik et al., 2015, discussed a dual mode of action hypothesis following findings that Cry1Ab and Cry1Ac protoxins were more potent than the activated toxins in insects with resistance to these toxins, indicating that the mechanisms of resistance can highlight alternative pathways for protoxins to exert toxicity [
17].
This study also found that some insecticidal toxins could exert a synergistic effect with Cry41Aa against HepG2 cells, despite these insecticidal toxins not possessing toxicity in their own right. The synergistic effects of
Bt Cry toxins have been reported in studies since the 1980s [
18], and can be the result of several different mechanisms in which combinations of two or more Cry toxins can exert a greater than expected influence [
19,
20]. The formation of hetero-oligomers between closely related Cry toxins has been shown to confer advantages in receptor binding and subsequent toxicity over monomers and homo-oligomers [
21], and thus result in synergism. Toxins may even serve as membrane binding targets for other toxins, as Cyt1A was shown to serve as a functional, membrane-bound receptor for Cry11Aa in
Aedes aegypti [
22].
Of the toxins tested within this study, several were found capable of exerting a synergistic effect with Cry41Aa. Interestingly, all of those with this activity also have activity against
Aedes aegypti, although it is currently unclear if there is any significance to this observation. Among these, Cry1Ca was shown to bind HepG2 cells in its active, but not protoxin, form. Despite its ability to bind, the exposure of HepG2 cells to activated Cry1Ca, or any of the other insecticidal Cry toxins used, failed to illicit any cell swelling, a characteristic early cellular response of HepG2 cells to Cry41Aa exposure [
3]. This absence of cellular response suggests that, whilst possessing the ability to bind a putative receptor is certainly a prerequisite for Cry toxin specificity, other factors such as the orientation of binding or the ability to insert into the host membrane also affect toxicity. This study found that the synergism between Cry41Aa and Cry1Ca was also observed when utilizing a DIP variant of Cry1Ca, indicating that Cry1Ca’s ability to oligomerize had no impact on the synergism observed, likely ruling out the possibility of hetero-oligomer formation as the reason for the synergism. The various interactions between Cry1Ca and Cry41Aa prompted the development of a two-receptor hypothesis. We hypothesize that two receptors exist for Cry41Aa in HepG2 cells, and that one is less effective than the other. Cry41Aa can bind both targets, whilst Cry1Ca, Cry11Aa, and Cry4Aa Cry toxins can only bind to one, the hypothesized “less effective” target. In doing so, these Cry toxins facilitate a synergistic effect by blocking binding to the less effective site, thereby allowing more of the Cry41Aa to bind to the “more effective” target. The observable result is a decrease in cell viability beyond that observed in controls featuring Cry41Aa alone, where more of the toxin has bound to the less or non-effective receptor.
The synergism between the Cry toxins and Cry41Aa in HepG2 cells differs from the effect observed in
Aedes aegypti larval competition studies. Competition studies have long been a means by which shared receptors and cross-resistance between different Cry toxins have been explored in insect systems [
23]. We observed that Cry41Aa could inhibit the toxicity of Cry1Ca and Cry11Aa in larval assays, which suggests that Cry41Aa can bind the same receptor in the
Aedes midgut. Though the putative receptor for Cry1Ca in
Aedes aegypti is unknown, it has been elucidated that the receptor for Cry11Aa in
Aedes is a glycosylphosphatidylinositol (GPI) anchored alkaline phosphatase [
24]. Therefore, it is possible that an homologous alkaline phosphatase within mammalian cells may represent a shared binding site for Cry1Ca, Cry11Aa, Cry4Aa and Cry41Aa.
Though long considered to be a non-insecticidal and non-hemolytic Cry toxin, there exists the possibility that Cry41Aa does possess some form of insecticidal ability. Still, any insect species in which it may have an effect has yet to be found. Given Cry41Aa’s interesting crossover with the Aedes-active toxins detailed in this work, it could be argued that this parasporin may share a close evolutionary history with Cry toxins, and that perhaps another factor, such as changes to the biochemistry of domain I, have imbued this protein with pore-forming capabilities in mammalian cells rather than in insects.