A Genetic and Environmental Analysis of Inflammatory Factors in Chronic Widespread Pain Using the TwinsUK Cohort
Abstract
1. Introduction
2. Materials and Methods
2.1. Sample
2.2. Inflammatory Factors
2.3. Chronic Widespread Pain
2.4. Design of the Study
2.5. Statistical Analysis
3. Results
3.1. Descriptive Statistics
3.2. Mixed-Effects Logistic Regression of Individual Inflammatory Factors on CWP
3.3. Factor Analysis
3.4. Twin Modelling
3.5. Causal Modelling
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Abbreviations
| CWP | Chronic widespread musculoskeletal pain |
| BMI | Body mass index |
| CRP | C-reactive protein |
| ApoA1 | Apolipoprotein A1 |
| ApoB | Apolipoprotein B |
| PEA | Proximity extension assay |
| QC | Quality control |
| LFESSQ | London Fibromyalgia Epidemiology Study Screening Questionnaire |
References
- Sluka, K.A.; Clauw, D.J. Neurobiology of fibromyalgia and chronic widespread pain. Neuroscience 2016, 338, 114–129. [Google Scholar] [CrossRef] [PubMed]
- Mansfield, K.E.; Sim, J.; Jordan, J.L.; Jordan, K.P. A systematic review and meta-analysis of the prevalence of chronic widespread pain in the general population. Pain 2016, 157, 55–64. [Google Scholar] [CrossRef]
- Andrews, P.; Steultjens, M.; Riskowski, J. Chronic widespread pain prevalence in the general population: A systematic review. Eur. J. Pain 2018, 22, 5–18. [Google Scholar] [CrossRef] [PubMed]
- Gaskin, D.J.; Richard, P. The economic costs of pain in the United States. J. Pain 2012, 13, 715–724. [Google Scholar] [CrossRef] [PubMed]
- Nicholl, B.I.; Macfarlane, G.J.; Davies, K.A.; Morriss, R.; Dickens, C.; McBeth, J. Premorbid psychosocial factors are associated with poor health-related quality of life in subjects with new onset of chronic widespread pain- results from the EPIFUND study. PAIN 2009, 141, 119. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Livshits, G.; Macgregor, A.J.; Gieger, C.; Malkin, I.; Moayyeri, A.; Grallert, H.; Emeny, R.T.; Spector, T.; Kastenmüller, G.; Williams, F.M.K. An omics investigation into chronic widespread musculoskeletal pain reveals epiandrosterone sulfate as a potential biomarker. PAIN 2015, 156, 1845–1851. [Google Scholar] [CrossRef]
- Livshits, G.; Malkin, I.; Freidin, M.B.; Xia, Y.; Gao, F.; Wang, J.; Spector, T.D.; MacGregor, A.; Bell, J.T.; Williams, F.M.K. Genome-wide methylation analysis of a large population sample shows neurological pathways involvement in chronic widespread musculoskeletal pain. PAIN 2017, 158, 1053. [Google Scholar] [CrossRef] [PubMed]
- Ikuta, K.; Ejima, A.; Abe, S.; Shimba, A. Control of immunity and allergy by steroid hormones. Allergol. Int. 2022, 71, 432–436. [Google Scholar] [CrossRef] [PubMed]
- Moulton, V.R. Sex hormones in acquired immunity and autoimmune disease. Front. Immunol. 2018, 9, 2279. [Google Scholar] [CrossRef] [PubMed]
- Mundal, I.; Gråwe, R.W.; Bjørngaard, J.H.; Linaker, O.M.; Fors, E.A. Prevalence and long-term predictors of persistent chronic widespread pain in the general population in an 11-year prospective study: The HUNT study. BMC Musculoskelet. Disord. 2014, 15, 213. [Google Scholar] [CrossRef] [PubMed]
- Matsuda, M.; Huh, Y.; Ji, R.-R. Roles of inflammation, neurogenic inflammation, and neuroinflammation in pain. J. Anesth. 2019, 33, 131–139. [Google Scholar] [CrossRef]
- Zhang, J.-M.; An, J. Cytokines, inflammation, and pain. Int. Anesthesiol. Clin. 2007, 45, 27. [Google Scholar] [CrossRef] [PubMed]
- Rodriguez-Pintó, I.; Agmon-Levin, N.; Howard, A.; Shoenfeld, Y. Fibromyalgia and Cytokines. Immunol. Lett. 2014, 161, 200–203. [Google Scholar] [CrossRef]
- Üçeyler, N.; Häuser, W.; Sommer, C. Systematic review with meta-analysis: Cytokines in fibromyalgia syndrome. BMC Musculoskelet. Disord. 2011, 12, 245. [Google Scholar] [CrossRef]
- Kumbhare, D.; Hassan, S.; Diep, D.; Duarte, F.C.K.; Hung, J.; Damodara, S.; West, D.W.D.; Selvaganapathy, P.R. Potential role of blood biomarkers in patients with Fibromyalgia: A systematic review with meta-analysis. PAIN 2022, 163, 1232. [Google Scholar] [CrossRef] [PubMed]
- Verdi, S.; Abbasian, G.; Bowyer, R.C.E.; Lachance, G.; Yarand, D.; Christofidou, P.; Mangino, M.; Menni, C.; Bell, J.T.; Falchi, M.; et al. TwinsUK: The UK adult twin registry update. Twin Res. Hum. Genet. 2019, 22, 523–529. [Google Scholar] [CrossRef] [PubMed]
- Cederroth, C.R.; Hong, M.-G.; Freydin, M.B.; Edvall, N.K.; Trpchevska, N.; Jarach, C.; Schlee, W.; Schwenk, J.M.; Lopez-Escamez, J.-A.; Gallus, S.; et al. Screening for circulating inflammatory proteins does not reveal plasma biomarkers of constant tinnitus. J. Assoc. Res. Otolaryngol. 2023, 24, 593–606. [Google Scholar] [CrossRef] [PubMed]
- Watanabe, K.; Wilmanski, T.; Diener, C.; Earls, J.C.; Zimmer, A.; Lincoln, B.; Hadlock, J.J.; Lovejoy, J.C.; Gibbons, S.M.; Magis, A.T.; et al. Multiomic body mass index signatures in blood reveal clinically relevant population heterogeneity and variable responses to a healthy lifestyle intervention. Nat. Med. 2023, 29, 996–1008. [Google Scholar] [CrossRef]
- Andrew, T.; Hart, D.J.; Snieder, H.; de Lange, M.; Spector, T.D.; MacGregor, A.J. Are twins and singletons comparable? A study of disease-related and lifestyle characteristics in adult women. Twin Res. Hum. Genet. 2001, 4, 464–477. [Google Scholar] [CrossRef]
- Assarsson, E.; Lundberg, M.; Holmquist, G.; Björkesten, J.; Bucht Thorsen, S.; Ekman, D.; Eriksson, A.; Rennel Dickens, E.; Ohlsson, S.; Edfeldt, G.; et al. Homogenous 96-Plex PEA immunoassay exhibiting high sensitivity, specificity, and excellent scalability. PLoS ONE 2014, 9, e95192. [Google Scholar] [CrossRef] [PubMed]
- Lundberg, M.; Eriksson, A.; Tran, B.; Assarsson, E.; Fredriksson, S. Homogeneous antibody-based proximity extension assays provide sensitive and specific detection of low-abundant proteins in human blood. Nucleic Acids Res. 2011, 39, e102. [Google Scholar] [CrossRef] [PubMed]
- Abe, K.; Beer, J.C.; Nguyen, T.; Ariyapala, I.S.; Holmes, T.H.; Feng, W.; Zhang, B.; Kuo, D.; Luo, Y.; Ma, X.-J.; et al. Cross-platform comparison of highly sensitive immunoassays for inflammatory markers in a COVID-19 cohort. J. Immunol. 2024, 212, 1244–1253. [Google Scholar] [CrossRef] [PubMed]
- Kany, S.; Vollrath, J.T.; Relja, B. Cytokines in inflammatory disease. Int. J. Mol. Sci. 2019, 20, 6008. [Google Scholar] [CrossRef] [PubMed]
- Soares, C.L.R.; Wilairatana, P.; Silva, L.R.; Moreira, P.S.; Vilar Barbosa, N.M.M.; da Silva, P.R.; Coutinho, H.D.M.; de Menezes, I.R.A.; Felipe, C.F.B. Biochemical: Aspects of the inflammatory process a narrative review. Biomed. Pharmacother. 2023, 168, 115764. [Google Scholar] [CrossRef]
- Wautier, J.-L.; Wautier, M.-P. Pro- and Anti-Inflammatory Prostaglandins and Cytokines in Humans: A Mini Review. Int. J. Mol. Sci. 2023, 24, 9647. [Google Scholar] [CrossRef] [PubMed]
- Tao, X.; Tao, R.; Wang, K.; Wu, L. Anti-inflammatory mechanism of apolipoprotein A-I. Front. Immunol. 2024, 15, 1417270. [Google Scholar] [CrossRef] [PubMed]
- Sarmadi, N.; Poustchi, H.; Yari, F.A.; Radmard, A.R.; Karami, S.; Pakdel, A.; Shabani, P.; Khaleghian, A. Anti-inflammatory function of apolipoprotein B-depleted plasma is impaired in non-alcoholic fatty liver disease. PLoS ONE 2022, 17, e0266227. [Google Scholar] [CrossRef] [PubMed]
- White, K.P.; Speechley, M.; Harth, M.; Ostbye, T. The London fibromyalgia epidemiology study: The prevalence of fibromyalgia syndrome in London, Ontario. J. Rheumatol. 1999, 26, 1570–1576. [Google Scholar]
- Livshits, G.; Ni Lochlainn, M.; Malkin, I.; Bowyer, R.; Verdi, S.; Steves, C.J.; Williams, F.M.K. Shared Genetic influence on frailty and chronic widespread pain: A study from TwinsUK. Age Ageing 2018, 47, 119–125. [Google Scholar] [CrossRef] [PubMed]
- Brooks, M.E.; Kristensen, K.; van Benthem, K.J.; Magnusson, A.; Berg, C.W.; Nielsen, A.; Skaug, H.J.; Maechler, M.; Bolker, B.M. glmmTMB Balances speed and flexibility among packages for zero-inflated generalized linear mixed modeling. R J. 2017, 9, 378–400. [Google Scholar] [CrossRef]
- R Core Team. R: A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2023. [Google Scholar]
- Stekhoven, D.J.; Buehlmann, P. MissForest—non-parametric missing value imputation for mixed-type data. Bioinformatics 2012, 28, 112–118. [Google Scholar] [CrossRef] [PubMed]
- Revelle, W. Psych: Procedures for Psychological, Psychometric, and Personality Research; Northwestern University: Evanston, IL, USA, 2024. [Google Scholar]
- Neale, M.C.; Hunter, M.D.; Pritikin, J.N.; Zahery, M.; Brick, T.R.; Kirkpatrick, R.M.; Estabrook, R.; Bates, T.C.; Maes, H.H.; Boker, S.M. OpenMx 2.0: Extended structural equation and statistical modeling. Psychometrika 2016, 81, 535–549. [Google Scholar] [CrossRef]
- Boker, S.M.; Neale, M.C.; Maes, H.H.; Spiegel, M.; Brick, T.R.; Estabrook, R.; Bates, T.C.; Gore, R.J.; Hunter, M.D.; Pritikin, J.N.; et al. OpenMx: Extended Structural Equation Modelling. Psychoetrika 2023, 76, 306–317. [Google Scholar] [CrossRef] [PubMed]
- Bates, T.C.; Maes, H.; Neale, M.C. UMX: Twin and path-based structural equation modeling in R. Twin Res. Hum. Genet. 2019, 22, 27–41. [Google Scholar] [CrossRef] [PubMed]
- Kratzer, G.; Lewis, F.; Comin, A.; Pittavino, M.; Furrer, R. Additive Bayesian network modeling with the R package Abn. J. Stat. Softw. 2023, 105, 1–41. [Google Scholar] [CrossRef]
- Furrer, R.; Kratzer, G.; Lewis, F.I. Abn: Modelling Multivariate Data with Additive Bayesian Networks. J. Open Source Softw. 2023, 9, 6822. [Google Scholar]
- Plummer, M. Rjags: Bayesian Graphical Models Using MCMC. 2024. Available online: https://CRAN.R-project.org/package=rjags (accessed on 24 December 2024).
- Cherny, S.S.; Chowers, M.; Obolski, U. Bayesian network modeling of patterns of antibiotic cross-resistance by bacterial sample source. Commun. Med. 2023, 3, 61. [Google Scholar] [CrossRef] [PubMed]
- Choi, J.; Joseph, L.; Pilote, L. Obesity and C-reactive protein in various populations: A systematic review and meta-analysis. Obes. Rev. 2013, 14, 232–244. [Google Scholar] [CrossRef] [PubMed]
- Li, Q.; Wang, Q.; Xu, W.; Ma, Y.; Wang, Q.; Eatman, D.; You, S.; Zou, J.; Champion, J.; Zhao, L.; et al. C-reactive protein causes adult-onset obesity through chronic inflammatory mechanism. Front. Cell Dev. Biol. 2020, 8, 18. [Google Scholar] [CrossRef]

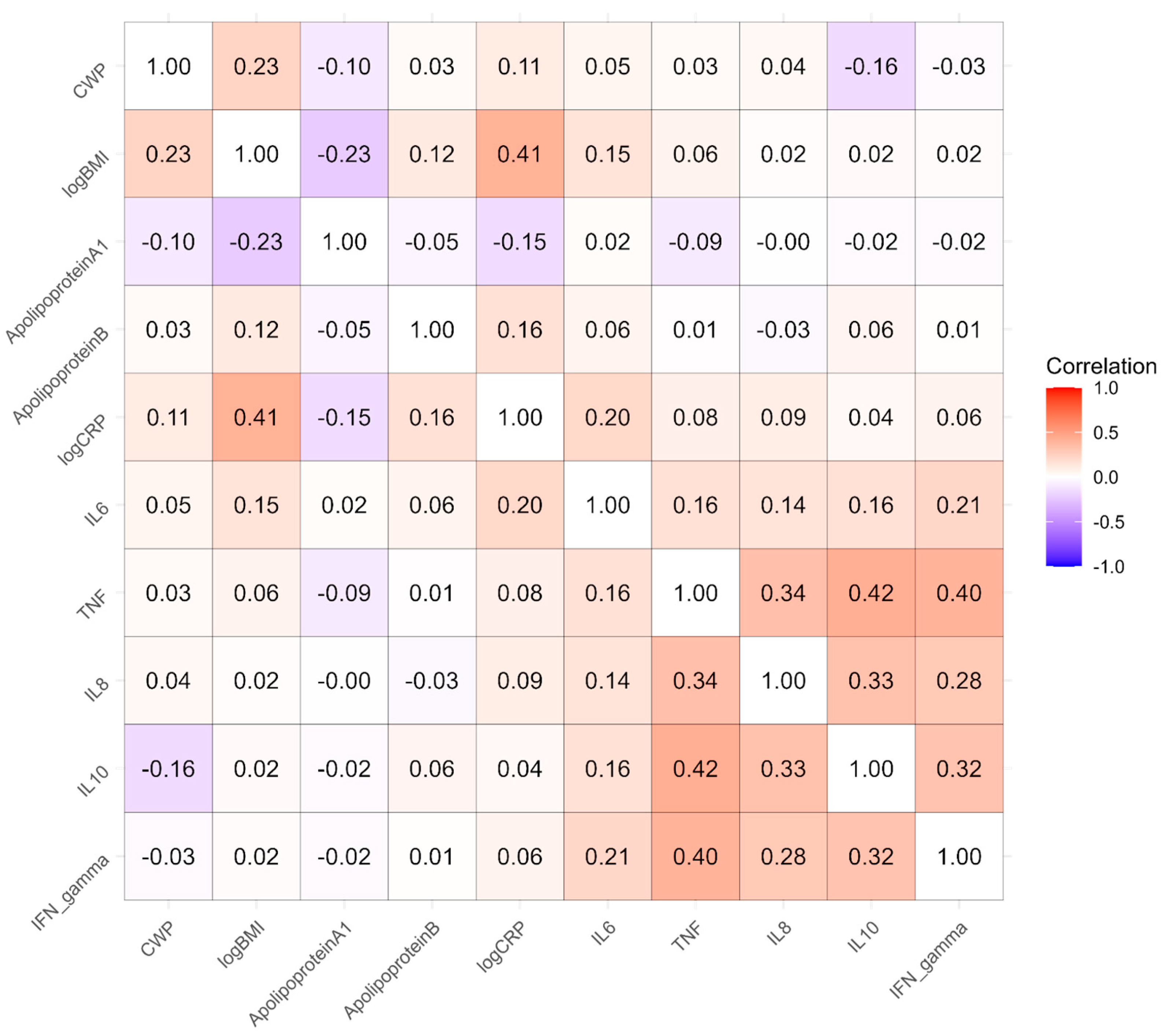
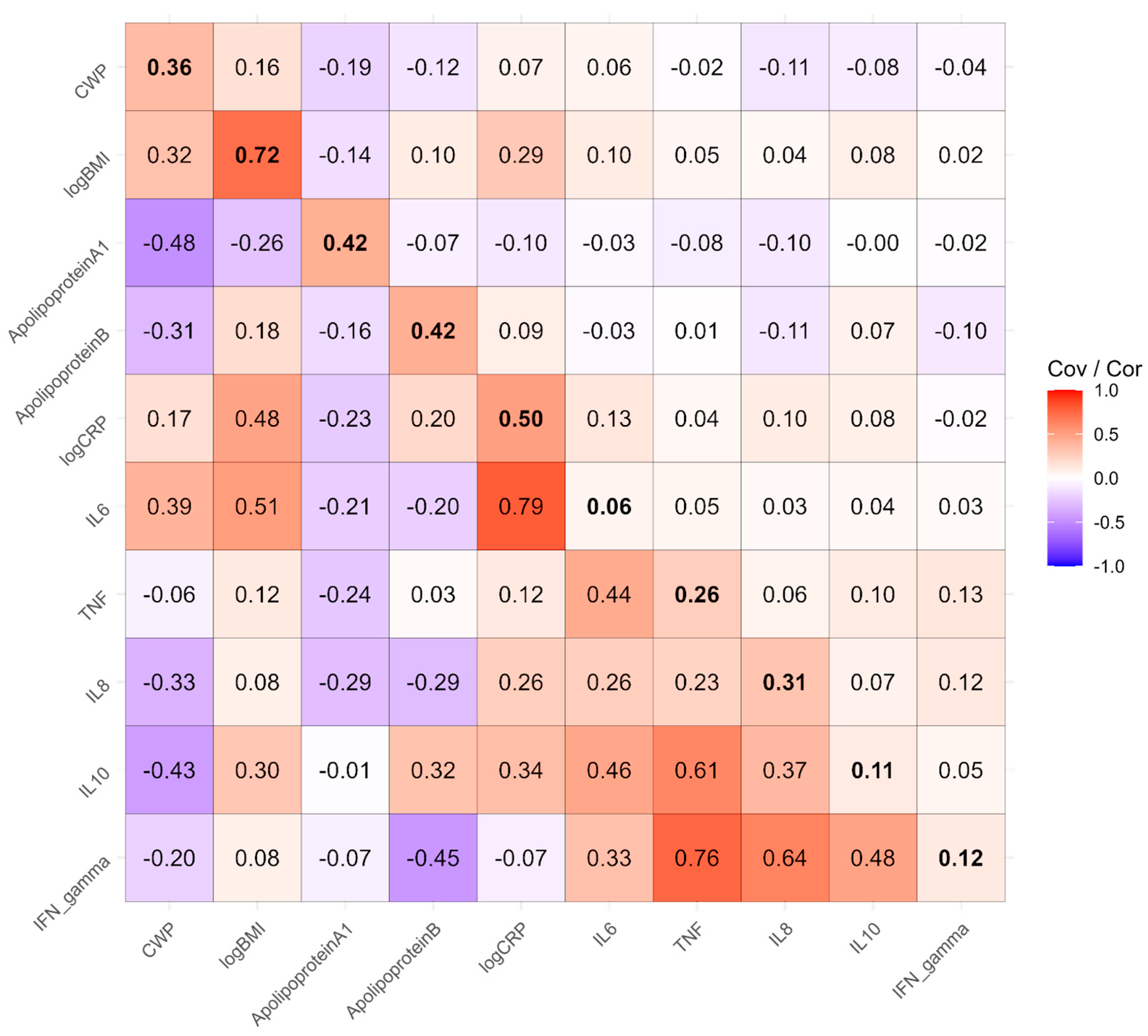


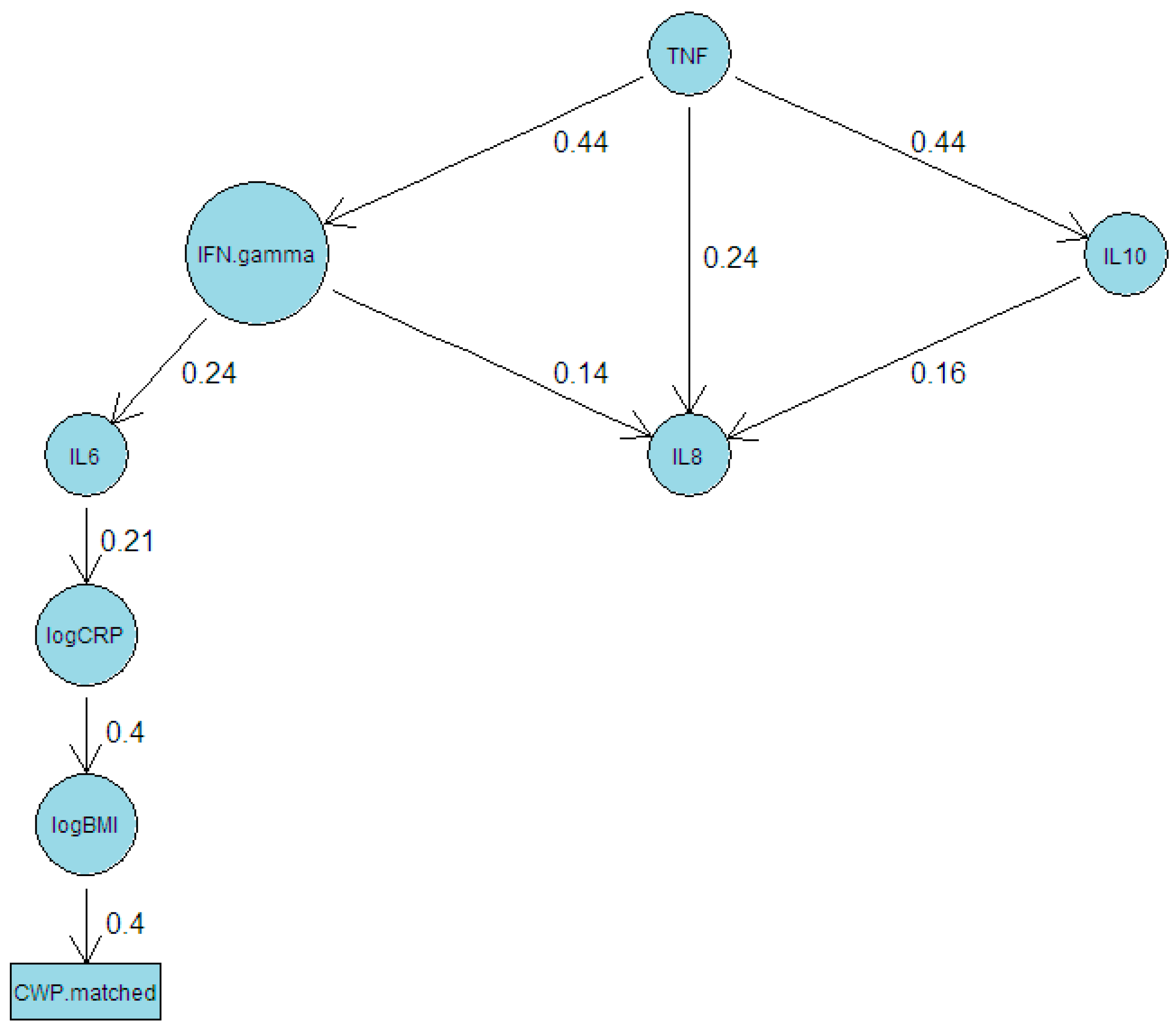

| Factor | N | Estimate | SE | z | p | Adj p |
|---|---|---|---|---|---|---|
| Pro-inflammatory factors | ||||||
| log(CRP) | 870 | 0.432 | 0.242 | 1.785 | 0.074 | 0.535 |
| ApolipoproteinB | 716 | −0.221 | 0.278 | −0.794 | 0.427 | 0.757 |
| CCL11 | 904 | 0.061 | 0.251 | 0.244 | 0.807 | 0.930 |
| CCL23 | 904 | −0.365 | 0.249 | −1.467 | 0.142 | 0.535 |
| CCL3 | 896 | 0.400 | 0.246 | 1.623 | 0.105 | 0.535 |
| CCL4 | 895 | 0.276 | 0.238 | 1.160 | 0.246 | 0.739 |
| CD40 | 904 | 0.127 | 0.253 | 0.502 | 0.616 | 0.862 |
| CXCL1 | 904 | −0.067 | 0.245 | −0.274 | 0.784 | 0.930 |
| CXCL10 | 904 | 0.159 | 0.246 | 0.646 | 0.518 | 0.824 |
| CXCL11 | 902 | −0.139 | 0.247 | −0.563 | 0.573 | 0.862 |
| CXCL5 | 904 | −0.371 | 0.241 | −1.540 | 0.124 | 0.535 |
| CXCL6 | 904 | 0.059 | 0.249 | 0.236 | 0.813 | 0.930 |
| CXCL9 | 903 | 0.382 | 0.267 | 1.430 | 0.153 | 0.535 |
| DNER | 902 | −0.048 | 0.248 | −0.195 | 0.846 | 0.930 |
| EN.RAGE | 904 | −0.359 | 0.239 | −1.498 | 0.134 | 0.535 |
| IFN.gamma | 898 | 0.183 | 0.223 | 0.821 | 0.412 | 0.757 |
| IL.12B | 904 | −0.287 | 0.253 | −1.137 | 0.255 | 0.739 |
| IL.15RA | 782 | 0.431 | 0.242 | 1.784 | 0.074 | 0.535 |
| IL.17C | 795 | 0.238 | 0.236 | 1.010 | 0.312 | 0.739 |
| IL.18R1 | 904 | 0.758 | 0.270 | 2.808 | 0.005 | 0.175 |
| IL18 | 903 | −0.225 | 0.245 | −0.917 | 0.359 | 0.739 |
| IL7 | 879 | −0.018 | 0.254 | −0.070 | 0.944 | 0.956 |
| IL8 | 901 | 0.126 | 0.238 | 0.528 | 0.597 | 0.862 |
| MCP.1 | 901 | 0.223 | 0.240 | 0.930 | 0.353 | 0.739 |
| MCP.2 | 902 | 0.236 | 0.252 | 0.940 | 0.347 | 0.739 |
| MCP.4 | 903 | −0.078 | 0.246 | −0.316 | 0.752 | 0.930 |
| MMP.1 | 904 | −0.266 | 0.250 | −1.067 | 0.286 | 0.739 |
| SLAMF1 | 830 | 0.388 | 0.252 | 1.542 | 0.123 | 0.535 |
| TNF | 901 | 0.012 | 0.216 | 0.055 | 0.956 | 0.956 |
| TNFB | 900 | −0.028 | 0.252 | −0.112 | 0.911 | 0.956 |
| TNFSF14 | 903 | 0.187 | 0.238 | 0.784 | 0.433 | 0.757 |
| TRAIL | 904 | 0.405 | 0.251 | 1.613 | 0.107 | 0.535 |
| TWEAK | 904 | −0.046 | 0.243 | −0.188 | 0.851 | 0.930 |
| uPA | 904 | −0.083 | 0.249 | −0.332 | 0.740 | 0.930 |
| VEGFA | 903 | 0.177 | 0.243 | 0.728 | 0.466 | 0.777 |
| Anti-inflammatory factors | ||||||
| ApolipoproteinA1 | 716 | −0.612 | 0.300 | −2.041 | 0.041 | 0.454 |
| FGF.21 | 897 | −0.052 | 0.243 | −0.213 | 0.831 | 0.914 |
| HGF | 904 | 0.286 | 0.249 | 1.150 | 0.250 | 0.551 |
| IL.10RB | 902 | 0.412 | 0.254 | 1.622 | 0.105 | 0.530 |
| IL10 | 896 | −0.284 | 0.233 | −1.219 | 0.223 | 0.551 |
| LAP.TGF.beta.1 | 902 | −0.055 | 0.241 | −0.227 | 0.821 | 0.914 |
| LIF.R | 902 | 0.361 | 0.248 | 1.459 | 0.145 | 0.530 |
| MMP.10 | 904 | −0.232 | 0.247 | −0.941 | 0.347 | 0.636 |
| SIRT2 | 821 | −0.199 | 0.287 | −0.693 | 0.489 | 0.768 |
| STAMBP | 904 | −0.137 | 0.263 | −0.519 | 0.604 | 0.830 |
| TRANCE | 904 | −0.006 | 0.244 | −0.026 | 0.979 | 0.979 |
| Immunoregulatory factors | ||||||
| ADA | 904 | −0.007 | 0.253 | −0.027 | 0.978 | 0.978 |
| CCL19 | 904 | 0.374 | 0.214 | 1.748 | 0.080 | 0.563 |
| CCL20 | 899 | −0.237 | 0.242 | −0.977 | 0.329 | 0.764 |
| CCL25 | 904 | 0.214 | 0.265 | 0.807 | 0.419 | 0.764 |
| CCL28 | 900 | −0.075 | 0.249 | −0.300 | 0.764 | 0.925 |
| CD244 | 904 | 0.233 | 0.252 | 0.926 | 0.354 | 0.764 |
| CD5 | 904 | 0.057 | 0.250 | 0.226 | 0.821 | 0.925 |
| CD6 | 904 | 0.236 | 0.243 | 0.973 | 0.330 | 0.764 |
| CD8A | 904 | −0.097 | 0.236 | −0.409 | 0.683 | 0.925 |
| CX3CL1 | 903 | 0.446 | 0.249 | 1.789 | 0.074 | 0.563 |
| Flt3L | 903 | 0.128 | 0.249 | 0.515 | 0.607 | 0.925 |
| IL6 | 580 | −0.283 | 0.364 | −0.778 | 0.436 | 0.764 |
| PD.L1 | 903 | −0.187 | 0.240 | −0.780 | 0.435 | 0.764 |
| TNFRSF9 | 902 | −0.045 | 0.255 | −0.177 | 0.859 | 0.925 |
| Multifunctional factors | ||||||
| AXIN1 | 904 | −0.010 | 0.262 | −0.037 | 0.971 | 0.999 |
| CASP.8 | 901 | 0.140 | 0.237 | 0.594 | 0.553 | 0.999 |
| CDCP1 | 901 | 0.683 | 0.289 | 2.364 | 0.018 | 0.235 |
| CSF.1 | 903 | 0.282 | 0.248 | 1.138 | 0.255 | 0.829 |
| CST5 | 901 | −0.264 | 0.265 | −0.995 | 0.319 | 0.831 |
| FGF.19 | 904 | 0.024 | 0.237 | 0.102 | 0.919 | 0.999 |
| NT.3 | 898 | 0.090 | 0.221 | 0.409 | 0.683 | 0.999 |
| OPG | 904 | −0.034 | 0.262 | −0.128 | 0.898 | 0.999 |
| OSM | 904 | 0.000 | 0.234 | 0.002 | 0.999 | 0.999 |
| SCF | 901 | −0.271 | 0.212 | −1.278 | 0.201 | 0.829 |
| ST1A1 | 856 | 0.135 | 0.261 | 0.517 | 0.605 | 0.999 |
| TGF.alpha | 902 | −0.094 | 0.238 | −0.397 | 0.692 | 0.999 |
| X4E.BP1 | 904 | −0.350 | 0.234 | −1.495 | 0.135 | 0.829 |
| Factor | Covariates | N | Estimate | SE | z | p | AUC | adj R2 |
|---|---|---|---|---|---|---|---|---|
| IL6 | No covariates | 580 | −0.282 | 0.362 | −0.779 | 0.436 | 0.542 | 0.002 |
| Age | −0.285 | 0.364 | −0.783 | 0.433 | 0.548 | 0.002 | ||
| Age + Age2 | −0.283 | 0.364 | −0.778 | 0.436 | 0.531 | 0.002 | ||
| Age + Age2 + log(BMI) | −0.419 | 0.393 | −1.065 | 0.287 | 0.591 | 0.012 | ||
| TNF | No covariates | 901 | 0.034 | 0.210 | 0.162 | 0.871 | 0.500 | 0.000 |
| Age | 0.013 | 0.217 | 0.060 | 0.952 | 0.526 | 0.000 | ||
| Age + Age2 | 0.012 | 0.216 | 0.055 | 0.956 | 0.610 | 0.002 | ||
| Age + Age2 + log(BMI) | −0.057 | 0.224 | −0.254 | 0.799 | 0.647 | 0.020 | ||
| IL8 | No covariates | 901 | 0.158 | 0.233 | 0.679 | 0.497 | 0.518 | 0.001 |
| Age | 0.146 | 0.237 | 0.617 | 0.537 | 0.531 | 0.001 | ||
| Age + Age2 | 0.126 | 0.238 | 0.528 | 0.597 | 0.610 | 0.003 | ||
| Age + Age2 + log(BMI) | 0.152 | 0.249 | 0.607 | 0.544 | 0.645 | 0.020 | ||
| log(CRP) | No covariates | 870 | 0.435 | 0.242 | 1.795 | 0.073 | 0.563 | 0.005 |
| Age | 0.432 | 0.243 | 1.777 | 0.076 | 0.564 | 0.005 | ||
| Age + Age2 | 0.432 | 0.242 | 1.785 | 0.074 | 0.613 | 0.007 | ||
| Age + Age2 + log(BMI) | 0.084 | 0.278 | 0.302 | 0.763 | 0.650 | 0.021 | ||
| IL10 | No covariates | 896 | −0.274 | 0.233 | −1.175 | 0.240 | 0.537 | 0.002 |
| Age | −0.287 | 0.235 | −1.223 | 0.221 | 0.550 | 0.002 | ||
| Age + Age2 | −0.284 | 0.233 | −1.219 | 0.223 | 0.599 | 0.004 | ||
| Age + Age2 + log(BMI) | −0.312 | 0.242 | −1.287 | 0.198 | 0.658 | 0.021 | ||
| IFN.gamma | No covariates | 898 | 0.199 | 0.220 | 0.906 | 0.365 | 0.511 | 0.001 |
| Age | 0.189 | 0.224 | 0.844 | 0.399 | 0.520 | 0.001 | ||
| Age + Age2 | 0.183 | 0.223 | 0.821 | 0.412 | 0.581 | 0.003 | ||
| Age + Age2 + log(BMI) | 0.178 | 0.230 | 0.774 | 0.439 | 0.645 | 0.019 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Cherny, S.S.; Livshits, G.; Williams, F.M.K. A Genetic and Environmental Analysis of Inflammatory Factors in Chronic Widespread Pain Using the TwinsUK Cohort. Biomolecules 2025, 15, 155. https://doi.org/10.3390/biom15020155
Cherny SS, Livshits G, Williams FMK. A Genetic and Environmental Analysis of Inflammatory Factors in Chronic Widespread Pain Using the TwinsUK Cohort. Biomolecules. 2025; 15(2):155. https://doi.org/10.3390/biom15020155
Chicago/Turabian StyleCherny, Stacey S., Gregory Livshits, and Frances M. K. Williams. 2025. "A Genetic and Environmental Analysis of Inflammatory Factors in Chronic Widespread Pain Using the TwinsUK Cohort" Biomolecules 15, no. 2: 155. https://doi.org/10.3390/biom15020155
APA StyleCherny, S. S., Livshits, G., & Williams, F. M. K. (2025). A Genetic and Environmental Analysis of Inflammatory Factors in Chronic Widespread Pain Using the TwinsUK Cohort. Biomolecules, 15(2), 155. https://doi.org/10.3390/biom15020155








