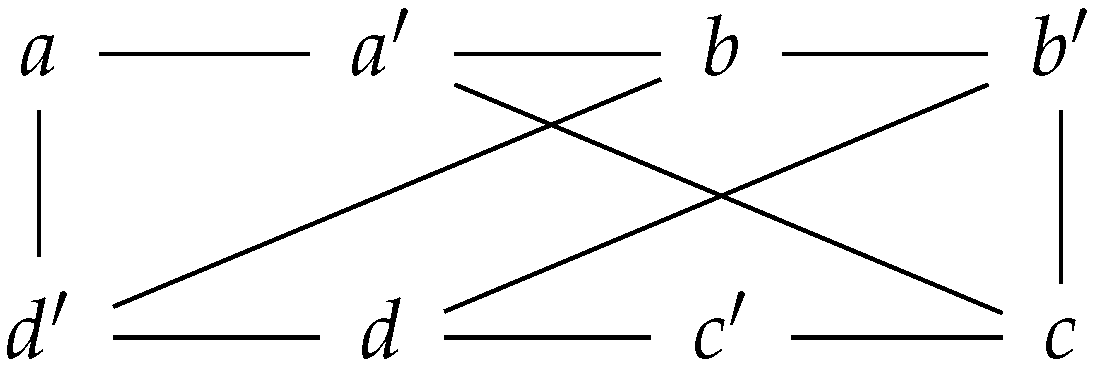Abstract
We use Category Theory to construct a ‘bridge’ relating directed graphs with undirected graphs, such that the notion of direction is preserved. Specifically, we provide an isomorphism between the category of simple directed graphs and a category we call ‘prime graphs category’; this has as objects labeled undirected bipartite graphs (which we call prime graphs), and as morphisms undirected graph morphisms that preserve the labeling (which we call prime graph morphisms). This theoretical bridge allows us to extend undirected graph techniques to directed graphs by converting the directed graphs into prime graphs. To give a proof of concept, we show that our construction preserves topological features when applied to the problems of network alignment and spectral graph clustering.
Keywords:
undirected graphs; directed graphs; spectral clustering; network alignment; category theory MSC:
00A69; 05C50; 18A99; 68R10
1. Introduction
Networks naturally arise in many real-world situations. Examples vary from macro-structures, such as social networks [1] and economic trade networks [2], to microscopic structures involving protein–protein interactions [3], transcriptional regulation networks [4], gene regulatory networks [5], and both biological and artificial neural networks [6,7]. While real-world networks are often modeled as directed graphs, their computational analysis is not only challenging but far more cumbersome and restricted. Thus, it would be helpful to analyze and solve certain classes of problems for directed graphs from an undirected graph framework. Here, we tackle this problem by using notions and principles of Category Theory (CT) within a graph context. Put simply, CT studies abstract structures and their relations. These structures, or categories, are composed of a collection of things we called “objects”, and a collection of relations between two objects that we call “morphisms”. Originally, CT developed within pure mathematics; much more recently, it started to be used broadly across the natural sciences and engineering, including applications in machine learning and artificial neural networks [8], biological networks [9,10], and social networks [11].
The way we bridge a directed graph framework with an undirected one is by first considering a category of undirected graphs that encode the notion of direction. This category, which we call the prime graphs category, has as objects undirected graphs equipped by a ‘prime labeling’, and as morphisms undirected graphs morphisms that preserve the prime labeling. Indeed, it is the prime labeling that provides a notion of direction over the structure of the undirected graph, allowing us to define a unique direction when transforming to a directed graph, and vice versa. With this in mind, we construct a bijective functor that relates the category of simple directed graphs with the category of prime graphs, such that the notion of direction is preserved. It is worth mentioning that one can always relate a directed graph to an undirected graph in a trivial way: by simply considering the underlying structure and ‘forgetting’ the direction. This correspondence gives rise to a ’forgetful’ functor which is not invertible. In [12], Miller provides one of the first nontrivial transformations between simple directed graphs and undirected graphs by the construction of “gadgets”. These gadgets are used to encode the notion of direction, just as our prime label does. However, each gadget adds seven nodes to the corresponding undirected graphs, while in our framework, we are just adding a prime label node. Additionally, we are the first to address the problem of converting directed graph morphisms into undirected graphs, while preserving the notion of direction. This latter aspect is crucial for both of our applications; in network alignment, the labeling and its preservation through prime graph morphisms ensures the mapping between appropriate nodes through the node similarity metric, while in spectral graph clustering, the labeling plays a role in determining edge weights.
Network alignment is a technique that allows us to compare two networks. This is performed by “putting one on top of the other” in such a way that the structure—or topology—between the networks being compared is preserved as much as possible, and the similarity between the networks is quantified. To date, several network alignment tools exist for undirected networks (see [13,14,15,16]), but to the best of our knowledge, none exist for directed graphs. Our framework, hence, proposes to perform network alignment on directed graphs via their corresponding prime graphs (which are undirected). Within this line of applications, we show the efficacy of our approach empirically by using synthetically generated pairs of networks whose pairwise similarity is known and controlled by the graph generator’s pairwise correlation coefficient. Our results in Section 3.1.1 show that there is a strong statistical correspondence between the generated networks and their resulting pairwise network similarity scores.
Spectral clustering is a widely used and robust technique that considers the spectrum—or eigenvalues and eigenvectors—of the graph Laplacian matrix to partition the nodes of a graph into clusters. More precisely, one can cluster the nodes of a graph by sorting the components of the eigenvector corresponding to the second-smallest eigenvalue of the characteristic polynomial of the graph Laplacian matrix. Initially, spectral clustering was developed for undirected graphs using their adjacency and Laplacian matrices [17]. Later, the technique was extended to directed graphs [18], where a transition probability matrix—or random walk—is used to overcome the asymmetry found in their adjacency and Laplacian matrices. There are heuristic techniques that circumvent the latter construction by making the adjacency matrix of a directed graph symmetric; therefore, they can define a symmetric graph Laplacian matrix [19], which they then use to find the cuts of the directed graph. However, while they are able to show empirically that the resulting cuts are the same, they do not prove equivalence, as we do. Precisely, we prove that our framework preserves minimum cuts, and consequently, it preserves clusters in directed graphs and their respective prime graph counterparts.
The paper is organized as follows. Section 2 gives preliminary notions about directed and undirected graphs from a category theory perspective. We describe the category of prime graphs and prime graph morphisms. We also present the concepts and theoretical results used in the application of our framework. In Section 3, we discuss the applications of network alignment and spectral clustering for directed graphs via functoriality. Finally, Section 4 gives some concluding arguments.
2. Methods
Mathematically, a graph is a structure that consist of a set of vertices V (nodes) and a set , which we call edges (connections). Within this context, we say that a graph is undirected if its set of edges are connections without directionality. A graph is directed if all its edges are connections with a direction. Now, given an undirected graph, we represent an edge between the nodes u and v by the unordered pair , equivalently . Instead, for a directed graph, an edge from node u to node v is represented by the ordered pair ; conceptually, we can think of this edge as an arrow with initial node u and terminal node v. Also, we say that the nodes u and v are neighbors if there is at least one edge connecting nodes u and v. Throughout this work, we will consider simple directed graphs, this is, directed graphs with no multiple edges and no self loops.
Intuitively, a graph morphism is a function between the vertex sets that preserves the structure or topology of the graphs; this is, it preserves the edges under transformation:
Definition 1.
Let and be two undirected graphs. An undirected graph morphism is a function that maps adjacent vertices in into adjacent vertices in . Algebraically, this means that for any pair of vertices with , we have .
For two directed graphs and , we say that a function is a directed graph morphism if f maps initial nodes into initial nodes, and terminal nodes into terminal nodes. One can verify that the composition of two graph morphism always yields another graph morphism, in both undirected and directed cases. Further, the composition is an associative operation that has a neutral element, called the identity morphism (which coincides with the identity map). With this in mind, one can show that the collection of all undirected graphs and all undirected graph morphisms () forms a category. Similarly, the collection of all simple directed graphs and all directed graph morphisms () forms a category. As we see next, the category is isomorphic to a subcategory of ; this subcategory, in fact, is the category of prime graphs () that we define.
Before describing the isomorphism between and , we will detail the category of prime graphs and its connection to simple directed graphs. Conceptually, a prime graph is an undirected graph which admits a ’prime labeling’ on its set of nodes (Figure 1). By allowing a “prime labeling”, we mean that there exists a labeling function on the vertex set with the following two properties; any prime labeled node has only non-prime labeled nodes as neighbors (and vice versa), and any prime labeled node is always adjacent to its non-prime labeled counterpart node.
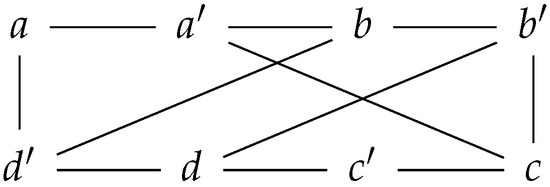
Figure 1.
A prime graph with eight nodes; four nodes have a prime labeling, while the other four nodes have a non-prime labeling.
Definition 2.
Let be an undirected graph with vertices, and let . We say that admits a prime labeling if there exists a bijective function such that for , one has the following three cases:
- (i)
- If and , then for each neighbor u of v, for some . We visualize this as
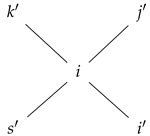
- (ii)
- If and , then for each neighbor u of v, for some . We visualize this as
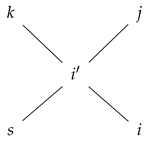
- (iii)
- For each , if and , then there exists a neighbor of v such that and .
For illustrative purposes, consider the following graphs:


The graph on the left is not a prime graph, as the prime labeled nodes are connected to each other. For the same reasons, the graph on the right is not a prime graph; however, this last graph becomes a prime graph if we endow it with the following prime labeling:
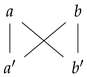
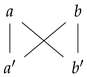
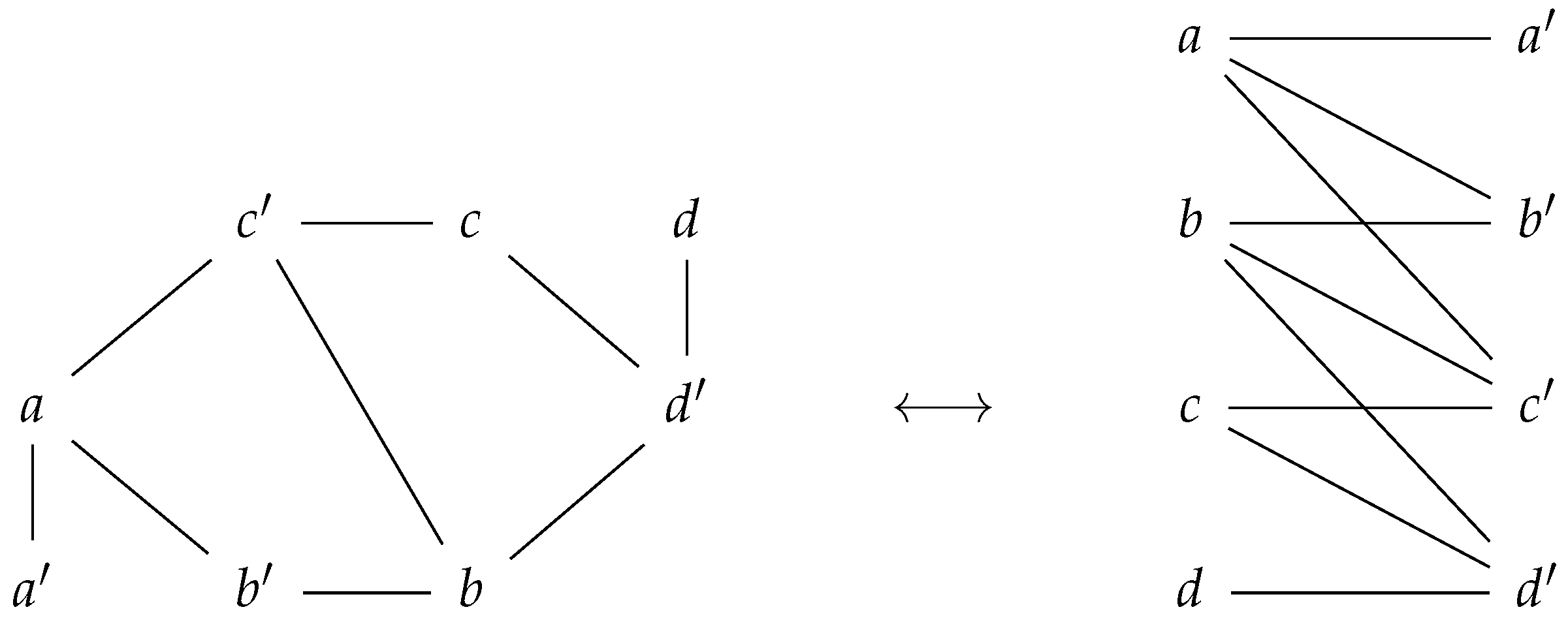
We also observe that one can visualize a prime graph as an undirected bipartite graph. This is a consequence of the definition of a prime labeling, as prime labeled node only have non-prime labeled nodes as neighbors, and vice versa. Please see in Figure 2.
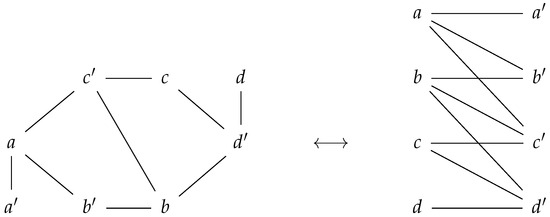
Figure 2.
A prime graph corresponds to a bipartite graph. The set of nodes of a prime graph can be written as the disjoint union of prime labeled nodes and the non-prime labeled nodes.
Now, due to condition , we can naturally induce a directed graph from a prime graph. In this case, the direction of an arrow will be given by the ’prime’ label vertex. For instance, if we consider the linear undirected graph G
 we can give the prime labeling
we can give the prime labeling
 which induces the directed graph
which induces the directed graph
 having a directed edge from an initial vertex to a terminal vertex .
having a directed edge from an initial vertex to a terminal vertex .



In terms of morphisms, we can think of a prime graph morphism as an undirected graph morphism that preserves the prime and non-prime labelings. Formally:
Definition 3.
Let and be two prime graphs, with labeling functions and , respectively. A prime graph morphism is an undirected graph morphism that satisfies the following conditions:
- (i)
- (Non-prime label preservation) If with for some , then is such thatfor .
- (ii)
- (Prime label preservation) If with for some , then is such thatfor .
- (iii)
- If are adjacent vertices with and , then one always has that
The above can be rephrased by saying that a prime graph morphism is an undirected graph morphism compatible with the non-prime and prime labelings. From an algebraic perspective, this compatibility condition means that, for each prime graph morphism , there exists a function making
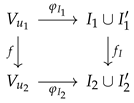 a commutative diagram; that is, .
a commutative diagram; that is, .
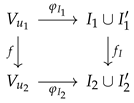
Furthermore, the composition of two prime graph morphisms results in a prime graph morphism itself. This follows from the fact that the composition of undirected graph morphisms is a closed operation, and also from the fact that the composition of prime graph morphisms preserves the prime and non-prime labeling conditions. Expressed in diagrams, this latter aspect is equivalent to saying that the commutativity of the large diagram is a consequence of the commutativity of the smaller diagrams:
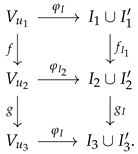
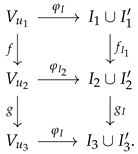
Additionally, as the composition operation of undirected graph morphisms is an associative operation, it follows that the composition of prime graph morphisms is also an associative operation. Further, for any prime graph , its identity prime graph morphism coincides with the identity map defined on the undirected graph . Considering the above, one has that the collection of prime graphs, along with the collection of prime graph morphisms, form a category.
Theorem 1.
The collection of all prime graphs, and all prime graph morphisms, defines a category denoted by PGraph.
It is worth noting that not every undirected graph morphism induces a prime graph morphism. To see this, let us consider the following prime graphs:
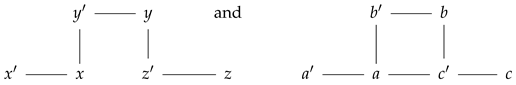
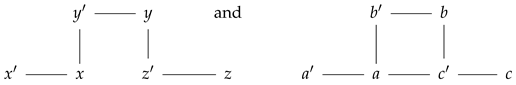
Then, the function with correspondence rule , , , , , and is an undirected graph morphism which is not a prime graph morphism. In fact, this illustrates that the category is not a full subcategory of UndGraph. Consequently, both the graph isomorphism problem and the subgraph isomorphism problem for prime graphs differentiates that for undirected graphs, which are known to be open and NP-complete, respectively.
2.1. and Are Isomorphic Categories
The first part of this subsection describes the functors and . The second part of this subsection shows that and are inverse of each other.
2.1.1. The Functors and
The following two propositions describe the assignment on objects and morphisms of functor :
Proposition 1.
Let be a directed graph in . Then, induces a prime graph in .
Proof of Proposition 1.
Let be a finite directed graph in DGraph. Without loss of generality, let us suppose that is represented by the set . Thus, by denoting for each , the prime graph has vertex set
Now, to define the edge set , we will consider the following two cases:
- (i)
- for each , the tuple defines an edge in ;
- (ii)
- for in , we have that defines an edge in if, and only if, there exists a directed edge .
In other words,
We claim that admits an I-labeling. Let and be the non-prime and prime sets, respectively. We define the labeling function as follows. For each ,
Clearly, is a bijective function. Moreover, based on how is defined, condition of Definition 2 is satisfied. Thus, it suffices to show that equipped by satisfies conditions and . Now, if is such that , then for . Hence, as the incident edges to vertex have the form —that is, when —or , it follows that the neighbors of also have the form , with , or . In any case, one has that for any possible neighbor of v. Likewise, if for some , then . Again, based on how the undirected graph is defined, one has that the incident edges to vertex in are either of the form (namely, when ) or . Thus, the neighbors of can be either (for some ) or , which in turn implies that for any possible neighbor of v. Therefore, defines a prime labeling function on . □
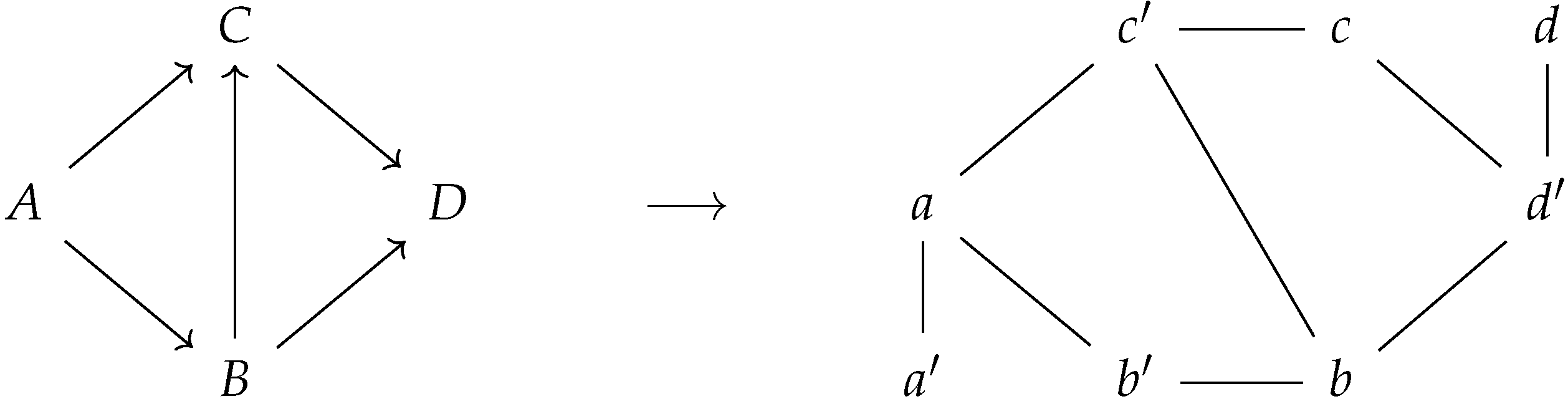
Figure 3 displays the correspondence between a directed graph and its corresponding prime graph . Notice that the prime labeled nodes encode for the notion of incoming edges of the corresponding directed graph.
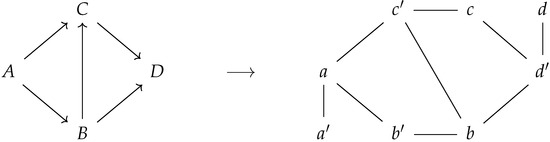
Figure 3.
A directed graph and its corresponding prime graph .
Proposition 2.
Let and be two directed graphs inDGraph, and let be a directed graph morphism. Then, f induces a prime graph morphism between the prime graphs and .
Proof of Proposition 2.
Let , and let us take , a directed graph morphism. Without loss of generality, assume that the vertex sets are and , respectively. Then, by Proposition 1, the vertex set of the corresponding prime graphs and are given by
and
respectively. With this in mind, we define by
where is the prime labeled vertex in such that . We claim that is a prime graph morphism. Indeed, for any edge , with , it corresponds to a directed edge in . Thus, as f preserves adjacencies—in the directed case—we have that , which in turn implies that . Considering this, one has that
Moreover, for edges of the form , one obtains
Thus, preserves adjacencies. Now, based on how is defined, it is clear that is compatible with the labeling, as it preserves the prime and non-prime labelings: . Therefore, is a prime graph morphism. □
To better visualize Proposition 2, let us consider a directed graph morphism f that maps a directed edge into a directed edge :
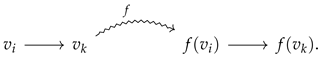

Then, in the prime graph context, one has a prime graph morphism mapping
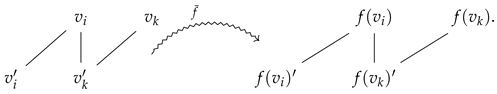
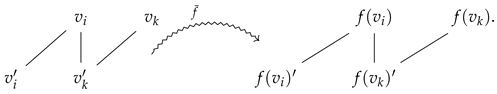
Remark 1.
If is a directed graph, then its corresponding prime graph satisfies that
This follows from the fact that an edge in has either the form —corresponding to a directed edge in —or the form .
Considering the results from above, we have the following:
Theorem 2.
The map that assigns to each object the object and to each morphism the morphism defines a functor from DGraph to PGraph.
Proof Theorem 2.
respectively.
Hence, and have the same correspondence rule, and hence, . Consequently,
that is, preserves compositions of morphisms. □
Observe that the object and morphism assignments of functor are exactly Proposition 1 and Proposition 2, respectively. Thus, it suffices to show that preserves identity morphisms and composition of morphisms.
The fact that preserves identity morphisms follows from Proposition 2 as coincides with the identity map . To see that preserves compositions, we will show that, given the directed graph morphisms and , one has that . Following the notation so far, we will denote the vertex set of the directed graph by , the vertex set of the directed graph by , and the vertex set of the directed graph by . Then, by Proposition 10, the vertex sets of their corresponding prime graphs are given by
Now, on the one hand, the image under functor of the directed graph morphism defined by the composition is the prime graph morphism , whose correspondence rule is given by
Here, denotes the prime labeled vertex on such that .
On the other hand, the image of f under is the prime graph morphism given by
while the image of g under is the prime graph morphism given by
Note that, as prime graph morphisms preserve the prime vertex and non-prime vertex labelings, one then has that and . This way, when considering the composite morphism , one has that, for each ,
whereas for each ,
Remark 2.
The functor preserves topological features. For instance, connectivity is preserved under . Also, if is a complete directed graph, then its corresponding prime graph is a complete bipartite graph.
On the other end, the next two propositions describe the assignment on objects and morphisms of functor :
Proposition 3.
Let be a prime graph. Then, induces a simple directed graph .
Proof of Proposition 3.
□
Let be a prime graph, and let be its labeling function, with . If the vertex set of is given by
then, we define the vertex set of the directed graph , induced by , as
In order to define the edge set , we must consider all incident edges to the set of prime label vertices in . For instance, if is adjacent to in (with ), then we will obtain a directed edge in from vertex towards vertex . In case is the only adjacent vertex to in , then the corresponding vertex in will not have incoming directed edges. In other words, the set is given by
Proposition 4.
Let and let be a prime graph morphism. Then, g induces a directed graph morphism between their corresponding directed graphs and .
Proof of Proposition 4.
showing that is a directed graph morphism. □
Let , and let be a prime graph morphism. Without loss of generality, let us assume that and . Then, by Proposition 3, the vertex sets and are given by and , respectively. With this in mind, we define the map as follows. For each , we set . We claim that preserves adjacencies. Indeed, given , we have that . Thus, as f is a prime graph morphism, we obtain that . Moreover, since f is compatible with the prime and non-prime labelings, it follows that , from which we have that . Therefore,
Considering the above, we have the following:
Theorem 3.
The map that assigns to each object the object , and to each morphism the morphism is a functor.
Proof of Theorem 3.
By Propositions 3 and 4, it suffices to show that the map preserves identity morphisms and composition of morphisms.
The fact that preserves identity morphisms follows from Proposition 4, as coincides with the identity map . To see that preserves compositions, we will prove that, given the prime graph morphisms and , one has that . Without loss of generality, let us suppose that
Following along with the notation, we will denote the vertex sets of the directed graphs and by , and , respectively. Now, on the one hand, the image under of the prime graph morphism is the directed graph morphism , whose correspondence rule is given by
On the other hand, the directed graph morphism is defined by , for all . Likewise, the directed graph morphism is defined by , for all . Therefore, the composition is a directed graph morphism such that, for each vertex ,
The above shows that and have the same correspondence rule. Since both functions have the same domain and codomain, we obtain that . Therefore,
that is, preserves compositions of morphisms. □
2.1.2. The Functors and Are Isomorphisms
Recall that an isomorphism between two categories and is a functor that is a bijection on both objects and morphisms. In other words, a functor is an isomorphism if, and only if, there exists a functor for which the compositions and are the identity functors and , respectively. In this case, we say that the categories and are isomorphic.
Proposition 5.
Let be a directed graph inDGraph. Then
Proof of Proposition 5.
and the edge set
If we now consider the directed graph induced by , that is, , then, the vertex set is given by and its edge by
In this way, as if, and only if, , it follows that if, and only if, . Moreover, since and have the same vertex set , we can conclude that . Therefore,
□
Let . Without loss of generality, we can assume that . Then, the prime graph has the vertex set
Proposition 6.
Let , and let be a directed graph morphism. Then,
Proof of Proposition 6.
Let and be directed graphs, and let be a directed graph morphism. Without loss of generality, assume that . Then, by Theorem 2, we obtain the prime graphs and , along with the prime graph morphism defined by
Here, . Now, as
for , it suffices to show that the functions and f have the same correspondence rule, since both have the same domain and codomain sets. By considering , we see that , for each vertex . Therefore, , and thus,
□
Corollary 1.
The functor and the functor satisfy
On the other end, we have the following results:
Proposition 7.
Let be a prime graph. Then
Proof of Proposition 7.
Now, if we denote by the prime graph induced by , that is, , then, , and
Notice that if, and only if, , which holds if, and only if, . Since both graphs and have as vertex set the set , we conclude that . Therefore,
□
Let . Without loss of generality, assume that the vertex set is given by the set . Then is the directed graph with vertex set , and edge set
Proposition 8.
Let be a prime graph morphism. Then
Proof of Proposition 8.
for , it suffices to show that and f have the same correspondence rule, as these have the same domain and codomain sets. By considering that is such that
we see that and f have the same correspondence rule. Thus, , which in turn implies that
□
Let be a prime graph morphism, and let us suppose that . By Theorem 3, we have the directed graphs and , along with the directed graph morphism defined by , for each . Now, as
Corollary 2.
The functor and the functor satisfy
Therefore, by Corollaries 1 and 2, we have the following:
Theorem 4.
The categories DGraph and PGraph are isomorphic.
We highlight that the property of functoriality on graph morphisms is what allowed us to extend network alignment techniques for undirected graphs to directed graphs. As previously mentioned, an alignment of two networks consists of a mapping between the nodes of the compared networks. In doing this, one aims to preserve as much of the structure (or topology) between the considered networks as possible. Thus, when transforming two directed graphs into their corresponding prime graphs, the way we relate these prime graphs is by defining a prime graph morphism. This prime graph morphism preserves the topology and the labeling conditions, which ultimately gives the notion of direction. Consequently, by functoriality, these prime graph morphisms correspond to the directed graph morphisms used when aligning the compared directed graphs.
2.2. Prime Transformation Algorithm
This section outlines an algorithm (Algorithm 1) to convert a directed graph to a prime graph, reflecting definitions and constructions found in the previous section. Then, we discuss the time and space complexity for creating and storing a prime graph from a directed graph. Note that the algorithm does not consider edge weights because weighting schemes can vary based on application.
The input to Algorithm 1 is a directed graph, and the output is its corresponding prime graph. The algorithm operates on each edge of the directed graph. As such, the algorithm’s time complexity is , where e is the number of edges and n is the number of nodes in the directed graph. While additional space is required to store the prime graph, the space necessary scales according to . The extra space needed to store a prime graph results from the additional nodes and edges it contains relative to its directed graph counterpart. For prime graphs, the number of nodes is always double that of directed graphs because each node in the directed graph spawns an additional prime node. Additionally, an edge is created between each pair of prime and non-prime nodes; therefore, additional n edges are required.
| Algorithm 1 Prime Transformation Algorithm | |
| Input DGraph | |
| Output PGraph | |
| |
| ▹ Initialize edge list |
| ▹ Initialize node list |
| ▹ Initialize an empty prime graph |
| |
| ▹ store the label of node n |
| ▹ create a label for the n’s prime node |
| ▹ add prime node |
| ▹ add non-prime node |
| ▹ add an edge between prime and non-prime node pairs |
| |
| ▹ store edge head label |
| ▹ store edge tail label |
| ▹ store the tail node’s prime label |
| ▹ add edge to PGraph using labels |
| ▹ add edge to PGraph using labels |
| |
3. Numerical Results
This section contains the two applications of the isomorphisms between the category of simple directed graphs DGraph and the category of prime graphs PGraph. To perform the computations, we relied on two things: the explicit description of the objects and morphisms of the category PGraph, and the explicit definition of functors and .
3.1. Network Alignment
Network alignment is a technique that consists of mapping the nodes of the compared networks in such a way that the structure, or topology, is preserved as much as possible. While mapping the nodes, the alignment looks to maximize node similarity and preserve the edge topology. Network alignment algorithms use an objective function—which is defined over candidate alignments—to find the best possible alignment between network pairs. This objective function represents a score, usually based on node and edge similarity. By considering this score, the aligner follows a “search algorithm" to find the fittest network map that maximizes the objective function. For a deeper understanding of network alignment and its applications, we recommend the reader see [13,14,15].
3.1.1. Synthetic Network Model
To demonstrate the efficacy of our correspondence between directed graphs and prime graphs, we generated a synthetic dataset that allowed us to precisely control the ground truth network similarity by generating correlated graph pairs. In this way, by using a network aligner, we calculated the network similarity scores between pairs of correlated prime graphs that were generated from pairs of correlated directed graphs (where correlation values for each pair were known). Our results indicate that the alignment scores between pairs of prime graphs are closely related to the correlation coefficients used to generate the corresponding directed graphs.
To generate the synthetic dataset, we used a stochastic block model (SBM) network topology [20] with the following parameters and values. The directed network consisted of 150 total nodes, organized into 3 blocks with 50 nodes per block, as well as 10 to 20 percent connection density inter-block and 1 percent intra-block connection density. For each discrete value of graph pair correlation coefficients , we generated 50 pairs of directed graphs. It is the correlation coefficient that determines the similarity between graphs. This way, we first created a directed graph from an edge probability matrix, which is defined by the previously mentioned inter- and intra-block connection density, and then, we use the coefficient to adjust the probability matrix for a second directed graph in the pair. Observe that the closer is to 1, the more similar the probability matrices will be, which means that it is more likely that the two generated graphs will be similar. Our results iterate through discrete values of in the range of at increments.
In summary, after generating all the directed graph pairs, we converted them to prime graphs, ran the network alignment algorithm for each prime graph pair, and plotted the similarity score as a function of using the highest similarity score out of the 5 alignment attempts per pair.
3.1.2. Numerical Result
We used both edge and node similarity scores to optimize network alignments. The optimizer uses a cost function that consists of an equally weighted linear combination of edge and node similarity scores. We used EC, ICS, and S3 edge similarity scores along with a graphlet orbit degree-based node similarity score (see the references [13,14,15,21,22] for details on each score’s definition). Finally, we used SANA [13], an off-the-shelf network aligner, and the aforementioned similarity scores for alignment optimization.
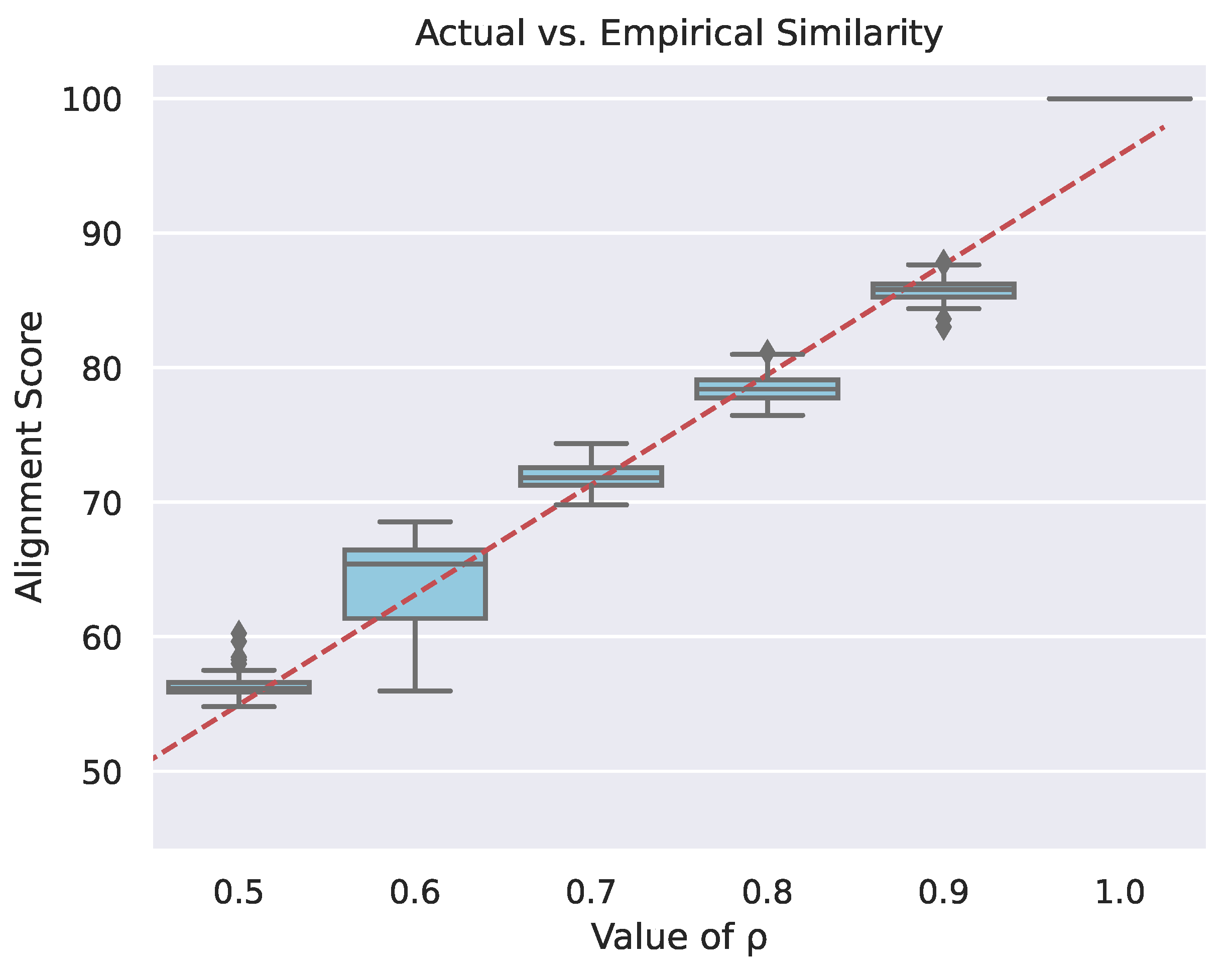
Figure 4 is a box and whisker plot (see [23]), which contains empirical alignment scores for varying values of the graph correlation coefficient. When the graph pairs are perfectly correlated (), that is, they are exact copies, but the mapping between nodes is unknown, the alignment tool finds the perfect mapping between the graphs. As the values decrease, the alignment scores decrease. The variance in the alignment scores primarily results from the network alignment algorithm’s random initialization, stochastic optimization routines, finite number of optimization steps, as well as because the optimization function is not guaranteed to be convex. A secondary source of alignment score variability is from the stochastic graph pair generation process (see Section 3.1.1). Variation in alignment scores results in an [24] value of with a corresponding p-value of . These results indicate that calibrating for the slope yields a usable network alignment metric.
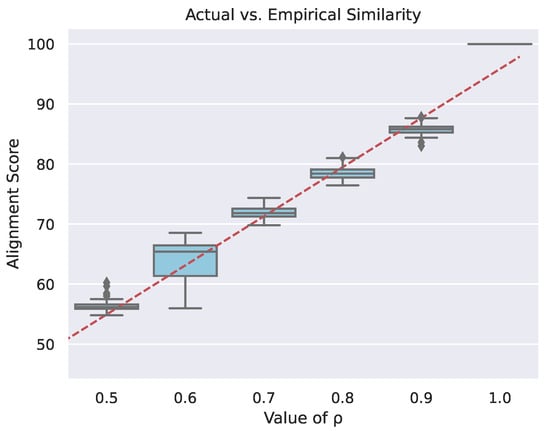
Figure 4.
Box and whisker plot of network alignment scores of prime graphs as a result of varying the underlying similarity between directed graph pairs. Each column in the plot corresponds to a specific value of , which is the correlation coefficient between a generated pair of graphs. The red line is the linear regression of all of the boxes together with respect to the values. It has an value of with a corresponding p-value of .
3.2. Minimum Cuts and Spectral Clustering in Prime Graphs
We start by showing the connection between cuts in a directed graph and cuts in its corresponding prime graph. Afterwards, we see that the functorial correspondence preserves clusters. This will allow us to use spectral clustering and show that a weighted prime graph maintains the same minimum cuts as its directed graph counterpart.
Recall that a cut in an undirected graph is a partition of the vertex set into a subset S and its complement . Likewise, one defines a cut in a directed graph as a partition of the vertex set . Now, for each weighted directed graph with weight values given by , there is a weighted prime graph whose weight values are given as follows. For each edge in of the form , we set the weight value equal to . For the remaining edges in of the form , we give a weight value satisfying
With this in mind, we have a correspondence between cuts in and cuts in with lesser volume. Recall that, given a weighted undirected graph , we define the volume of a cut (where ) as
Notice that considers those edges whose endpoints belong to different components C and . Furthermore, as the graph is undirected, it follows that . Considering this, for the weighted prime graph , with weight values described as above, we have the following:
Proposition 9.
Any cut in the weighted prime graph can be reduced to a cut of the form with lower volume. Here, S and are subsets that partition the non-prime nodes of , and and denote their prime node counterparts, which partition the set of prime nodes of .
Proof of Proposition 9.
By continuing with this process, we will obtain a cut of the form , where S and are subsets of that partition the non-prime vertices, whereas and are their corresponding prime nodes which partition the prime vertices of . Note that, by construction, this last cut has lower volume than the initial cut . □
Suppose that is a cut of the prime graph such that and . This implies that the weight of the edge adds to the volume of the cut. Now, if we consider the cut defined by the sets and , the weight of the edge will no longer add to the volume of the cut; instead, we will be adding those weights corresponding to the edges that are adjacent to , which turns out to be at most
The above proposition shows that the cuts with less volume in the weighted prime graph are those with form . These cuts, in turn, are clearly in a one-to-one correspondence between cuts of the form in the directed graph . We now show that the volume of a directed graph is preserved when transformed to its corresponding weighted prime graph .
Definition 4.
for every vertex .
Let be a directed graph. A circulation on is a function that assigns each directed edge to a non-negative value such that
Intuitively, a circulation is a flow in the directed graph that is conserved at each vertex; that is, the flow into each vertex equals the flow out of each vertex. We define the volume crossing the cut in as
We now prove that the volume crossing a cut in a directed graph is the same as the volume of the cut in its corresponding weighted prime graph . To that end, we will assume that the weights of the directed graph are given by a circulation , and the weights of its corresponding prime graph are given as where previously stated.
Proposition 10.
Let be a weighted directed graph with weights given by a circulation F, and let be its corresponding weighted prime graph. Then, for any cut of , the corresponding cut in the prime graph satisfies
Proof of Proposition 10.
Let be a weighted directed graph whose weights are given by the circulation . In this case, . Let us denote by its corresponding weighted prime graph, and by its weight values. Now, for the sake of clarity, we will denote by the cut in the weighted directed graph , and denote by the corresponding cut in the weighted prime graph . Then, by definition, we have that
Notice that, by the way prime graphs are constructed, there are no edges between nodes in S and nodes in . The same can be said between nodes in and nodes in . Hence,
□
Consider now any vertex w in lying in a cluster. Then, in its corresponding prime graph , the nodes and belong to the same cluster. This latter aspect follows because we have assigned a high weight value to the edge between and in . Now, from the local topology around the vertex w in , and the local topology around the vertices and in :
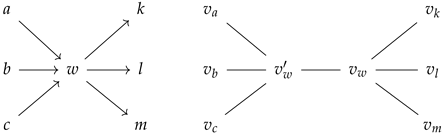 we can see that and , where and denote the indegree and outdegree of vertex , respectively. Further, observe that for any vertex in within the same cluster as w, we obtain that its corresponding prime and non-prime nodes belong to the same cluster as (and ), as the functor preserves connectivity. Therefore, by functoriality, each cluster in a directed graph induces a cluster in its corresponding prime graph with twice the number of nodes, namely the prime and non-prime vertices associated to the vertices of the cluster in the directed graph.
we can see that and , where and denote the indegree and outdegree of vertex , respectively. Further, observe that for any vertex in within the same cluster as w, we obtain that its corresponding prime and non-prime nodes belong to the same cluster as (and ), as the functor preserves connectivity. Therefore, by functoriality, each cluster in a directed graph induces a cluster in its corresponding prime graph with twice the number of nodes, namely the prime and non-prime vertices associated to the vertices of the cluster in the directed graph.

Proposition 11.
If C is a cluster in the directed graph , then its corresponding cluster in has twice the number of nodes as C.
Numerical Example: Spectrum of Graph Laplacian
For the technical application, we considered a method involving the graph Laplacian for optimal clustering. For further references, we recommend the reader to see [17,18].
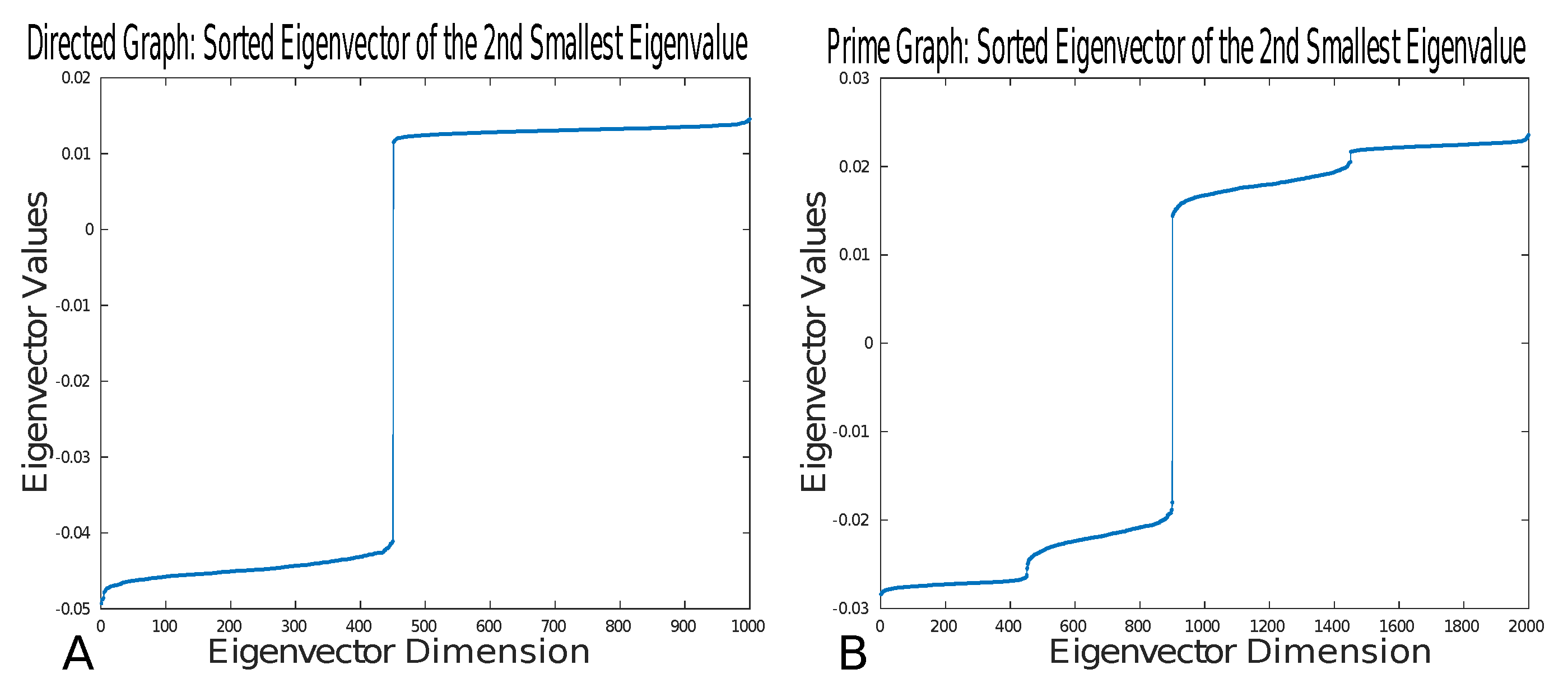
To illustrate that prime graphs preserve the cuts of a simple directed graph, we numerically compute minimum cuts of directed graphs and prime graphs using the graph Laplacian. The directed graph consists of 1000 nodes split into two clusters. The first cluster consists of 450 nodes, and the second cluster consists of 550 nodes. The edge connectivity parameters of the directed graph are as follows. In the first cluster, the inter-cluster connectivity was , that is, there was a chance of connection between nodes; in the second cluster, the inter-cluster connectivity was ; finally, the intra-cluster connectivity was . Upon constructing the directed graph, we generated an associated prime graph. In Figure 5, we show the values of the sorted eigenvector associated with the second-smallest eigenvalue. This numerical example thus shows that the minimum cut is preserved between directed graphs and prime graphs.
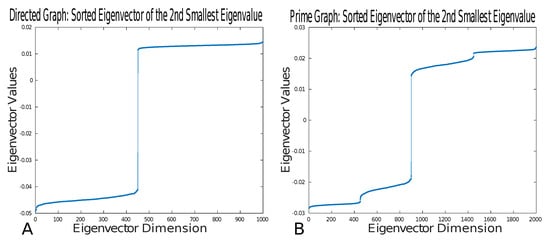
Figure 5.
(A) Plot of the sorted values in the eigenvector associated with the second-smallest eigenvalue for the directed graph. (B) Plot of the sorted values in the eigenvector associated with the second-smallest eigenvalue for the prime graph derived from the directed graph generated in (A).
4. Discussion
This work shows a novel construction that reinforces the power of CT as a tool to formalize structures and their relations. In this case, we use CT to bridge a directed graph framework to an undirected graph framework, so that not only directionality is preserved but also several topological features. This bridge enables the use of undirected graph techniques to obtain information from systems that are represented as directed graphs. Both the computational and space complexity of the transformations are , where N is the number of nodes in the network. As an empirical demonstration, we provide a new option to perform network alignment for directed graphs. This is relevant since network alignment tools do not exist for a directed graph setting. Furthermore, our transformation does preserve network similarity between directed graphs and their prime graph counterparts; we attained an value of (with a corresponding p-value of ) between the network aligner results, i.e., the similarity metric, a known graph generation correlation coefficient. Because we proved that our construction leads to an invertible transformation, there is only one prime graph that describes a simple directed graph and vice versa; as such, and in that sense, our transformation is error-free. Be that as it may, our transformation does not mitigate errors inherent in postprocessing the resultant graphs, for example, not achieving an value of 1 in the network alignment task.
Although the process of making an adjacency matrix of a directed graph symmetric is not new [19], nor is transforming a directed graph into an undirected graph [12], our framework is an advance. We proved that the minimum cuts are preserved when going from a directed graph framework to a prime graph framework and vice versa. These results, in turn, imply that clusters are preserved when moving from one setting to the other. As a proof of concept, we proved cluster preservation by generating a directed SBM network with known intra-block and inter-block connectivity.
While this work is a step towards a new application of existing network alignment tools, there is much left in this area to be explored in future work. Adoption of this technique may be limited by and rely upon showing additional mathematical proofs for commonly used techniques on graphs, for example, answering how the existing undirected node and edge similarity metrics might be skewed by the prime graph transformation. Another avenue for the application of prime graphs is to take advantage of their bipartite nature in problems such as the graph isomorphism problems for directed graphs. It is worthwhile studying the complexity of checking for equivalence between arbitrarily labeled DGraphs and PGraphs. Lastly, a categorical bridge, now between a multidirected graphs setting to a prime graph setting, might unlock new ways to study high complex data.
yes
Author Contributions
Conceptualization, S.P.-G. and J.R.; methodology, S.P.-G. and V.K.G.; software, V.K.G. and V.M.; validation, S.P.-G., V.K.G., V.M., J.R. and G.A.S.; formal analysis, S.P.-G.; investigation, S.P.-G. and V.K.G.; resources, G.A.S.; data curation, V.K.G. and V.M.; writing—original draft preparation, S.P.-G.; writing—review and editing, S.P.-G. and G.A.S.; visualization, S.P.-G.; supervision, G.A.S.; project administration, G.A.S.; funding acquisition, G.A.S. All authors have read and agreed to the published version of the manuscript.
Funding
This research received unrestricted funds to the Center for Engineered Natural Intelligence at the University of California San Diego.
Data Availability Statement
All data and figures of this paper were generated synthetically. The code used in this work can be found in the link https://github.com/vgeorgeucsd/prime_graphs, accessed on 8 April 2024.
Conflicts of Interest
The authors declare no conflicts of interest.
References
- Scott, J. Social network analysis. Sociology 1988, 22, 109–127. [Google Scholar] [CrossRef]
- Dimitrova, T.; Petrovski, K.; Kocarev, L. Graphlets in Multiplex Networks. Sci. Rep. 2020, 10, 1928. [Google Scholar] [CrossRef] [PubMed]
- Pržulj, N.; Wigle, D.A.; Jurisica, I. Functional topology in a network of protein interactions. Bioinformatics 2004, 20, 340–348. [Google Scholar] [CrossRef] [PubMed]
- Shen-Orr, S.S.; Milo, R.; Mangan, S.; Alon, U. Network motifs in the transcriptional regulation network of Escherichia coli. Nat. Genet. 2022, 31, 64–68. [Google Scholar] [CrossRef] [PubMed]
- Chu, B.K.; Tse, M.J.; Sato, R.R.; Read, E.L. Markov State Models of gene regulatory networks. Syst. Biol. 2017, 11, 1–17. [Google Scholar] [CrossRef] [PubMed]
- Buibas, M.; Silva, G.A. A framework for simulating and estimating the state and functional topology of complex dynamic geometric networks. Neural Comput. 2011, 23, 183–214. [Google Scholar] [CrossRef] [PubMed]
- Silva, G.A. The effect of signaling latencies and refractory node states on the dynamics of networks. Neural Comput. 2019, 31, 2492–2522. [Google Scholar] [CrossRef] [PubMed]
- Fong, B.; Spivak, D.; Tuyéras, R. Backprop as functor: A compositional perspective on supervised learning. In Proceedings of the 2019 34th Annual ACM/IEEE Symposium on Logic in Computer Science (LICS), Vancouver, BC, Canada, 24–27 June 2019; pp. 1–13. [Google Scholar]
- Haruna, T. Theory of interface: Category theory, directed networks and evolution of biological networks. Biosystems 2013, 114, 125–148. [Google Scholar] [CrossRef]
- Northoff, G.; Tsuchiya, N.; Saigo, H. Mathematics and the Brain: A Category Theoretical Approach to Go Beyond the Neural Correlates of Consciousness. Entropy 2019, 21, 1234. [Google Scholar] [CrossRef]
- Otter, N.; Porter, M.A. A unified framework for equivalences in social networks. arXiv 2020, arXiv:2006.10733. [Google Scholar]
- Miller, G.L. Graph isomorphism, general remarks. J. Comput. Syst. Sci. 1979, 18, 128–142. [Google Scholar] [CrossRef][Green Version]
- Mamano, N.; Hayes, W.B. SANA: Simulated annealing far outperforms many other search algorithms for biological network alignment. Bioinformatics 2017, 33, 2156–2164. [Google Scholar] [CrossRef]
- Vijayan, V.; Saraph, V.; Milenković, T. MAGNA++: Maximizing accuracy in global network alignment via both node and edge conservation. Bioinformatics 2015, 31, 2409–2411. [Google Scholar] [CrossRef] [PubMed]
- Sun, Y.; Crawford, J.; Tang, J.; Milenković, T. Simultaneous optimization of both node and edge conservation in network alignment via WAVE. In Proceedings of the International Workshop on Algorithms in Bioinformatics, Atlanta, GA, USA, 10–12 September 2015; pp. 16–39. [Google Scholar]
- Trung, H.T.; Toan, N.T.; Van Vinh, T.; Dat, H.T.; Thang, D.C.; Hung, N.Q.V.; Sattar, A. A comparative study on network alignment techniques. Expert Syst. Appl. 2020, 140, 112883. [Google Scholar] [CrossRef]
- Chung, F.R.; Graham, F.C. Spectral grAph Theory; Number 92; American Mathematical Society: Providence, RI, USA, 1997. [Google Scholar]
- Chung, F. Laplacians and the Cheeger inequality for directed graphs. Ann. Comb. 2005, 9, 1–19. [Google Scholar] [CrossRef]
- Satuluri, V.; Parthasarathy, S. Symmetrizations for clustering directed graphs. In Proceedings of the 14th International Conference on Extending Database Technology, Uppsala, Sweden, 21–24 March 2011; pp. 343–354. [Google Scholar]
- Chung, J.; Pedigo, B.D.; Bridgeford, E.W.; Varjavand, B.K.; Helm, H.S.; Vogelstein, J.T. GraSPy: Graph Statistics in Python. J. Mach. Learn. Res. 2019, 20, 1–7. [Google Scholar]
- Hayes, W.B. An introductory guide to aligning networks using sana, the simulated annealing network aligner. In Protein-Protein Interaction Networks; Springer: Berlin/Heidelberg, Germany, 2020; pp. 263–284. [Google Scholar]
- Milenković, T.; Pržulj, N. Uncovering biological network function via graphlet degree signatures. Cancer Inform. 2008, 6, CIN–S680. [Google Scholar] [CrossRef]
- Frigge, M.; Hoaglin, D.C.; Iglewicz, B. Some implementations of the boxplot. Am. Stat. 1989, 43, 50–54. [Google Scholar] [CrossRef]
- Wright, S. Correlation and causation. J. Agric. Res. 1921, 20, 557–585. [Google Scholar]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).