Prediction Models for Intrauterine Growth Restriction Using Artificial Intelligence and Machine Learning: A Systematic Review and Meta-Analysis
Abstract
1. Introduction
- describe the most widely used AI/ML algorithms applied in IUGR diagnosis
- evaluate the performance of AI/ML models in terms of accuracy
- determine whether there is a combination of these methods that has been shown to provide better accuracy in early diagnosis.
2. Materials and Methods
2.1. Search Strategy
- MEDLINE (PubMed): “Fetal Growth Retardation [Mesh]) Machine Learning [Mesh]”, “Fetal growth restriction machine learning”, “Machine learning IUGR”, “(Artificial intelligence) AND (IUGR)”, “(Artificial Intelligence) AND (Fetal growth restriction)”, “(Artificial Intelligence) AND (Fetal growth retardation)”
- EMBASE: “‘artificial intelligence’/exp AND ‘intrauterine growth retardation’/exp EMTREE”, “‘machine learning’/exp AND ‘fetal growth retardation’/exp” AND “‘machine learning’/exp AND ‘fetal growth restriction’/exp”
- Scopus: “Fetal Growth Retardation Machine Learning”, “Fetal growth restriction machine learning”, “Machine learning IUGR”, “(Artificial intelligence) AND (IUGR)”, “(Artificial Intelligence) AND (Fetal growth restriction)” and “(Artificial Intelligence) AND (Fetal growth retardation)”
- CINAHL: “Fetal Growth Retardation Machine Learning”, “Fetal growth restriction Machine Learning”, “Machine learning IUGR”, “(Artificial intelligence) AND (IUGR)”, “(Artificial Intelligence) AND (Fetal growth restriction)”, “(Artificial Intelligence) AND (Fetal growth retardation)”
- Web of Science: “Machine learning IUGR”, “Fetal growth retardation machine learning”, “Fetal growth retardation machine learning”, “(artificial intelligence) and IUGR”, “(artificial intelligence) and (fetal growth restriction)”, “(artificial intelligence) and (fetal growth retardation)”
- Cochrane Database of Systematic Reviews: “Fetal Growth Retardation/restriction/IUGR AND Machine Learning”, “Fetal Growth Retardation/restriction/ IUGR AND Artificial intelligence”
2.2. Inclusion and Exclusion Criteria
2.3. Selection Criteria
2.4. Quality Assessments of Studies
2.5. Data Extraction
2.6. Statistical Analysis
3. Results
3.1. Study Selection
3.2. Study Characteristics
| Author | Study Type | Setting | Sample (n) | Preg Time (weeks) | Methods | AI/ML Model | Outcomes | Measures (Accuracy) |
|---|---|---|---|---|---|---|---|---|
| Guo, Z. [50] | Obs. retr. case-control | China | 2199 | 12–28 | DNA profiling | LR | Using ML to predict FGR and BW | 79% |
| Dahdoud, S. [58] | Obs. retr. case-control | USA | 80 | 18–39 | MRI | RUSBoost | Using ML to predict FGR and BW | 86% |
| Lunghi, F. [45] | Obs. retr. case-control | Italy | 909 | 30–35 | FHR by CTG | SVM | Realizing an automatic system for identified FGR | 84% |
| Magenes, G. [48] | Obs. retr. case-control | Italy | 100 | 30–35 | FHR by CTG | SVM | Realizing an automatic system for identified FGR | 78% |
| Signorini, M. [60] | Obs. retr. case-control | Italy | 120 | 30–35 | FHR by CTG | RF (best) | Find the best classification ML model for identifying IUGR | 91% |
| Crockart, I.C. [27] | Obs. prosp. case-control | USA and S. Africa | 6004 | 20–29 | FHR by CTG | Stochastic Gradient Descent, LR & RF | Using ML to predict FGR and BW | 91% |
| Bahado–Singh, R. [46] | Obs. retr. case-control | USA | 80 | Delivery | Biochemical | SVM | Find the best classification ML model for identifying IUGR | 80% |
| Pini, N. [47] | Obs. retr. case-control | Italy | 262 | 36–37 | FHR by CTG | RBF-SVM | Build a ML screener for late IUGR | 93% |
| Magenes, G. [51] | Obs. retr. case-control | Italy | 122 | 30–35 | FHR by CTG | RF & LR | Find the best classification ML model for identifying IUGR | RF = 85%; LR = 83% |
| Xu, C. [52] | Obs. retr. nested case-control | China | 810 | 12–27 | DNA profiling | SVM & LR | Find the best classification ML model for identifying IUGR | 83% |
| Buscema, M. [54] | Obs. retr. case-control | Italy | 46 | Delivery | Biochemical | ACM & ACS | Find the best classification ML model for identifying IUGR | 87% |
| Foltran, F. [55] | Obs. prosp. case-control | Italy | 46 | 20–32 | Biochemical | BN | Realizing an automatic system for identified FGR | 90% |
| Street, M.E. [56] | Obs. retr. case-control | Italy | 48 | 20–32 | Biochemical | ANNS | Find the best classification ML model for identifying IUGR | 89% |
| Ferrario, M. [57] | Obs. retr. case-control | Italy | 59 | 27–34 | FHR by CTG | LZ complexity | Realizing an automatic system for identified FGR | 91% |
| Deval, R. [49] | Obs. retr. case-control | India | 214 | - | Biochemical | SVM, MLP | Using ML models to predict IUGR | SMO = 95.5%; MLP = 8.5% |
| Gómez–Jemes, L. [53] | Obs. retr. case-control | Slovenia | 95 | 24–38 | Doppler indices: UA, sFIt-1, and PIGF values | Multi-models (extra-trees, RF) | Using ML models to predict pre-Eclampsia, IUGR | Extra trees = 78%, RF = 73% |
| Sufriyana, H. [59] | Obs. prosp. case-control | Slovenia | 95 | 24–37 | Doppler indices: UA, sFIt–1, and PIGF values | CVR | Using ML models to predict pre-Eclampsia, IUGR | CVR = 90.6% |
| Aslam, N. [61] | Obs. retr. case-control | Italy | 382 | 30–37 | FHR by CTG | SVM & RF | Using ML models to predict IUGR | RF = 97% |
| Gürgen, F. [62] | Obs. retr. case-control | Turkey | 44 | <38 | Doppler indices: PI & RI of UA, MCA, DV, and AFI | SVM | Using ML models to predict IUGR | SVM = 81% |
| Van, S.N. [25] | Obs. prosp. case-control | China | 75 | - | Physiological, clinical, and socioeconomic | Seven ML algorithms | Identify the latent risk clinical attributes using the ML algorithms. | 94.73% |
3.3. Performance of ML Models for IUGR Prediction: Qualitative Synthesis
3.3.1. Prediction of IUGR from Biochemical and Clinical Parameters
3.3.2. Prediction of IUGR from DNA Profiling
3.3.3. Prediction of IUGR from MRI Data
3.3.4. Prediction of IUGR from FHR Parameters by CTG
3.3.5. Prediction of IUGR from Doppler Indices
3.3.6. Detection of Latent Clinical Attributes for Children Born under IUGR Condition
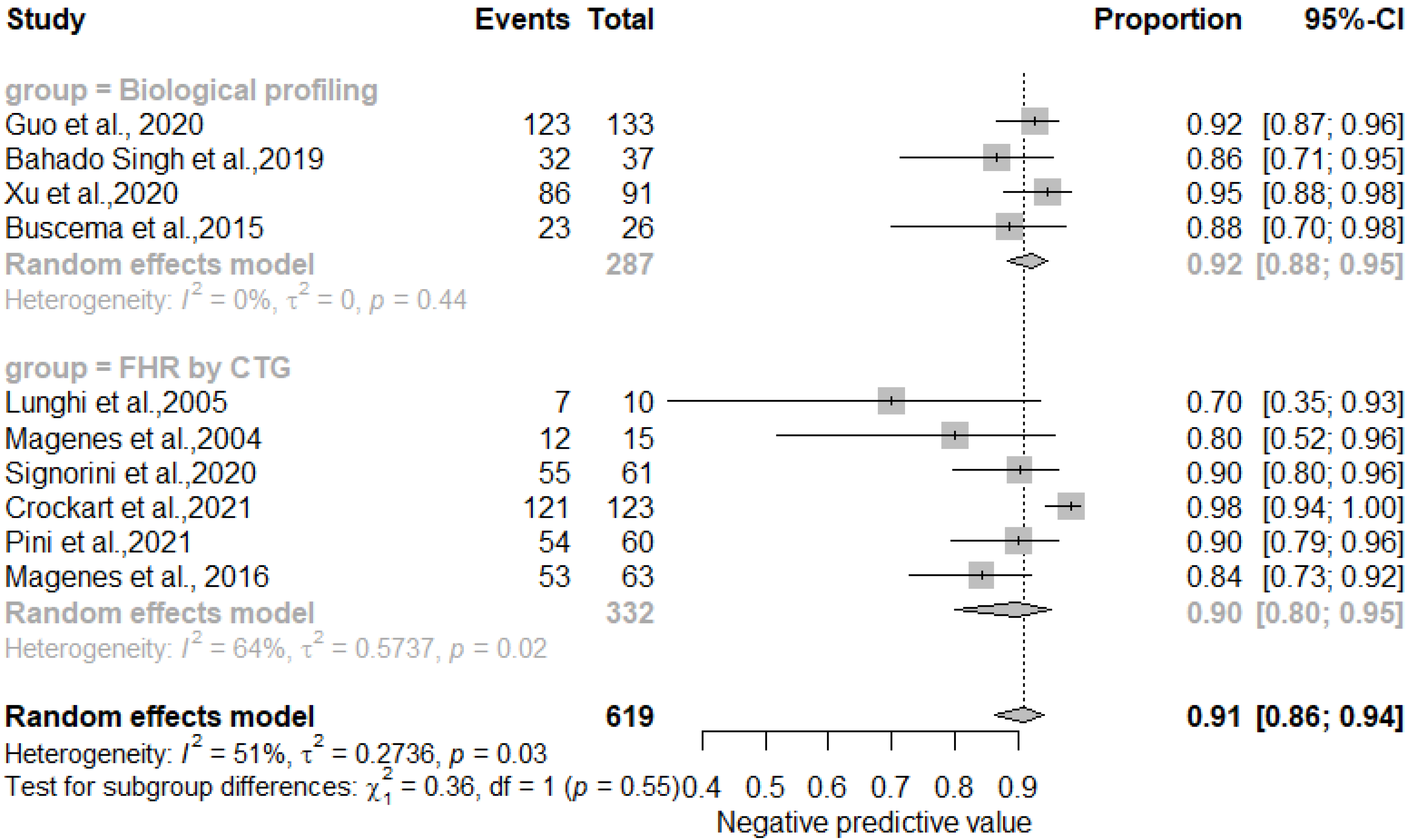
3.4. Meta-Analysis
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Abbreviations
| SR | Systematic Review |
| MA | Meta Analysis |
| AI | Artificial Intelligence |
| ML | Machine Learning |
| IUGR | Intra Uterine Growth Restriction |
| SGA | Small for Gestational Age |
| CTG | Cardiotocography |
| FHR | Fetal Heart Rate |
| RUSBoost | Random under sampling boosting |
| ACM ACS | Auto Contractive Map and Activation and Competition System |
| TNF-alfa | Tumor necrosis factor- Alfa |
| IGF | Insulin-like growth factor |
| UtA | Uterine artery |
| sFIt-1 | soluble fms-like tyrosine kinase receptor-1 |
| PIGF | Placental growth factor |
| PI /RI | Pulsitility Index/Resistance Index |
| UA | umbelical artery |
| MCA/DV | middle celebral artery/ductus venosus |
| AFI | amniotic fluid index |
| MLP | multilayer perceptron |
| CVR | classification via regression |
References
- Gordijn, S.J.; Beune, I.M.; Thilaganathan, B.; Papageorghiou, A.; Baschat, A.A.; Baker, P.N.; Silver, R.M.; Wynia, K.; Ganzevoort, W. Consensus definition of fetal growth restriction: A Delphi procedure. Ultrasound Obstet. Gynecol. Off. J. Int. Soc. Ultrasound Obstet. Gynecol. 2016, 48, 333–339. [Google Scholar] [CrossRef] [PubMed]
- Sharma, D.; Farahbakhsh, N.; Shastri, S.; Sharma, P. Intrauterine growth restriction—Part 2. J. Matern.-Fetal Neonatal Med. Off. J. Eur. Assoc. Perinat. Med. Fed. Asia Ocean. Perinat. Soc. Int. Soc. Perinat. Obstet. 2016, 29, 4037–4048. [Google Scholar] [CrossRef]
- Sharma, D.; Shastri, S.; Farahbakhsh, N.; Sharma, P. Intrauterine growth restriction—Part 1. J. Matern.-Fetal Neonatal Med. Off. J. Eur. Assoc. Perinat. Med. Fed. Asia Ocean. Perinat. Soc. Int. Soc. Perinat. Obstet. 2016, 29, 3977–3987. [Google Scholar] [CrossRef]
- Mandruzzato, G.; Antsaklis, A.; Botet, F.; Chervenak, F.A.; Figueras, F.; Grunebaum, A.; Puerto, B.; Skupski, D.; Stanojevic, M.; WAPM. Intrauterine restriction (IUGR). J. Perinat. Med. 2008, 36, 277–281. [Google Scholar] [CrossRef]
- Sovio, U.; White, I.R.; Dacey, A.; Pasupathy, D.; Smith, G.C. Screening for fetal growth restriction with universal third trimester ultrasonography in nulliparous women in the Pregnancy Outcome Prediction (POP) study: A prospective cohort study. Lancet 2015, 386, 2089–2097. [Google Scholar] [CrossRef] [PubMed]
- Vesterinen, H.M.; Morello-Frosch, R.; Sen, S.; Zeise, L.; Woodruff, T.J. Cumulative effects of prenatal-exposure to exogenous chemicals and psychosocial stress on fetal growth: Systematic-review of the human and animal evidence. PLoS ONE 2017, 12, e0176331. [Google Scholar] [CrossRef] [PubMed]
- Abraham, M.; Alramadhan, S.; Iniguez, C.; Duijts, L.; Jaddoe, V.W.; Dekker, H.T.; Crozier, S.; Godfrey, K.M.; Hindmarsh, P.; Vik, T.; et al. A systematic review of maternal smoking during pregnancy and fetal measurements with meta-analysis. PLoS ONE 2017, 12, e0170946. [Google Scholar] [CrossRef]
- Salomon, L.; Alfirevic, Z.; Da Silva Costa, F.; Deter, R.; Figueras, F.; Ghi, T.; Glanc, P.; Khalil, A.; Lee, W.; Napolitano, R.; et al. ISUOG Practice Guidelines: Ultrasound assessment of fetal biometry and growth. Ultrasound Obstet. Gynecol. 2019, 53, 715–723. [Google Scholar] [CrossRef]
- Lees, C.C.; Stampalija, T.; Baschat, A.A.; Silva Costa, F.; Ferrazzi, E.; Figueras, F.; Hecher, K.; Kingdom, J.; Poon, L.C.; Salomon, L.J.; et al. ISUOG Practice Guidelines: Diagnosis and management of small-for-gestational-age fetus and fetal growth restriction. Ultrasound Obstet. Gynecol. 2020, 56, 298–312. [Google Scholar] [CrossRef]
- McCowan, L.M.; Thompson, J.M.; Taylor, R.S.; North, R.A.; Poston, L.; Baker, P.N.; Myers, J.; Roberts, C.T.; Dekker, G.A.; Simpson, N.A.; et al. Clinical Prediction in Early Pregnancy of Infants Small for Gestational Age by Customised Birthweight Centiles: Findings from a Healthy Nulliparous Cohort. PLoS ONE 2013, 8, e0070917. [Google Scholar] [CrossRef]
- Velauthar, L.; Plana, M.N.; Kalidindi, M.; Zamora, J.; Thilaganathan, B.; Illanes, S.E.; Khan, K.S.; Aquilina, J.; Thangaratinam, S. First-trimester uterine artery Doppler and adverse pregnancy outcome: A meta-analysis involving 55,974 women. Ultrasound Obstet. Gynecol. 2014, 43, 500–507. [Google Scholar] [CrossRef]
- Parry, S.; Sciscione, A.; Haas, D.M.; Grobman, W.A.; Iams, J.D.; Mercer, B.M.; Silver, R.M.; Simhan, H.N.; Wapner, R.J.; Wing, D.A.; et al. Role of early second-trimester uterine artery Doppler screening to predict small-for-gestational-age babies in nulliparous women. Am. J. Obstet. Gynecol. 2017, 217, 594.e1–594.e10. [Google Scholar] [CrossRef] [PubMed]
- Tarca, A.L.; Hernandez-Andrade, E.; Ahn, H.; Garcia, M.; Xu, Z.; Korzeniewski, S.J.; Saker, H.; Chaiworapongsa, T.; Hassan, S.S.; Yeo, L.; et al. Single and serial fetal biometry to detect preterm and term small- and large-for-gestational-age neonates: A longitudinal cohort study. PLoS ONE 2016, 11, e0164161. [Google Scholar] [CrossRef] [PubMed]
- Mendez-Figueroa, H.; Truong, V.T.T.; Pedroza, C.; Khan, A.M.; Chauhan, S.P. Small-for-gestational-age infants among uncomplicated pregnancies at term: A secondary analysis of 9 Maternal-Fetal Medicine Units Network studies. Am. J. Obstet. Gynecol. 2016, 215, 628.e1–628.e7. [Google Scholar] [CrossRef] [PubMed]
- Poon, L.C.; Lesmes, C.; Gallo, D.M.; Akolekar, R.; Nicolaides, K.H. Prediction of small-for-gestational-age neonates: Screening by biophysical and biochemical markers at 19–24 weeks. Ultrasound Obstet. Gynecol. 2015, 46, 437–445. [Google Scholar] [CrossRef]
- Conde-Agudelo, A.; Papageorghiou, A.T.; Kennedy, S.H.; Villar, J. Novel biomarkers for predicting intrauterine growth restriction: A systematic review and meta-analysis. BJOG Int. J. Obstet. Gynaecol. 2013, 120, 681–694. [Google Scholar] [CrossRef]
- Zhong, Y.; Zhu, F.; Ding, Y. Serum screening in first trimester to predict pre-eclampsia, small for gestational age and preterm delivery: Systematic review and meta-analysis. BMC Pregnancy Childbirth 2015, 15, 191. [Google Scholar] [CrossRef]
- Crovetto, F.; Crispi, F.; Scazzocchio, E.; Mercade, I.; Meler, E.; Figueras, F.; Gratacos, E. First-trimester screening for early and late small-for-gestational-age neonates using maternal serum biochemistry, blood pressure and uterine artery Doppler. Ultrasound Obstet. Gynecol. 2014, 43, 34–40. [Google Scholar] [CrossRef]
- Papastefanou, I.; Nowacka, U.; Syngelaki, A.; Mansukhani, T.; Karamanis, G.; Wright, D.; Nicolaides, K.H. Competing risks model for prediction of small-for-gestational-age neonates from biophysical markers at 19 to 24 weeks’ gestation. Am. J. Obstet. Gynecol. 2021, 225, 530.e1–530.e19. [Google Scholar] [CrossRef]
- Crovetto, F.; Triunfo, S.; Crispi, F.; Rodriguez-Sureda, V.; Dominguez, C.; Figueras, F.; Gratacos, E. Differential performance of first-trimester screening in predicting small-for-gestational-age neonate or fetal growth restriction. Ultrasound Obstet. Gynecol. 2017, 49, 349–356. [Google Scholar] [CrossRef]
- Zhang, S.; Regnault, T.R.; Barker, P.L.; Botting, K.J.; McMillen, I.C.; McMillan, C.M.; Roberts, C.T.; Morrison, J.L. Placental adaptations in growth restriction. Nutrients 2015, 7, 360–389. [Google Scholar] [CrossRef] [PubMed]
- Hurtado, S.; García-Nieto, J.; Popov, A.; Navas-Delgado, I. Human Activity Recognition From Sensorised Patient’s Data in Healthcare: A Streaming Deep Learning-Based Approach. Int. J. Interact. Multimed. Artif. Intell. 2022, 8, 23–37. [Google Scholar] [CrossRef]
- Gerlache, H.A.M.; Ger, P.M.; Valentín, L.d.l.F. Towards the Grade’s Prediction. A Study of Different Machine Learning Approaches to Predict Grades from Student Interaction Data. Int. J. Interact. Multimed. Artif. Intell. 2022, 7, 196–204. [Google Scholar] [CrossRef]
- Ejiyi, C.J.; Qin, Z.; Salako, A.A.; Happy, M.N.; Nneji, G.U.; Ukwuoma, C.C.; Chikwendu, I.A.; Gen, J. Comparative Analysis of Building Insurance Prediction Using Some Machine Learning Algorithms. Int. J. Interact. Multimed. Artif. Intell. 2022, 7, 75–85. [Google Scholar] [CrossRef]
- Nguyen Van, S.; Lobo Marques, J.; Biala, T.; Li, Y. Identification of Latent Risk Clinical Attributes for Children Born Under IUGR Condition Using Machine Learning Techniques. Comput. Methods Programs Biomed. 2021, 200, 105842. [Google Scholar] [CrossRef]
- Maity, N.G.; Das, S. Machine learning for improved diagnosis and prognosis in healthcare. In Proceedings of the 2017 IEEE Aerospace Conference, Big Sky, MT, USA, 4–11 March 2017; pp. 1–9. [Google Scholar] [CrossRef]
- Crockart, I.C.; Brink, L.T.; du Plessis, C.; Odendaal, H.J. Classification of intrauterine growth restriction at 34–38 weeks gestation with machine learning models. Informatics Med. Unlocked 2021, 23, 100533. [Google Scholar] [CrossRef]
- Alzubaidi, M.; Agus, M.; Alyafei, K.; Althelaya, K.A.; Shah, U.; Abd-Alrazaq, A.; Anbar, M.; Makhlouf, M.; Househ, M. Toward deep observation: A systematic survey on artificial intelligence techniques to monitor fetus via ultrasound images. iScience 2022, 25, 104713. [Google Scholar] [CrossRef]
- Choi, R.Y.; Coyner, A.S.; Kalpathy-Cramer, J.; Chiang, M.F.; Campbell, J.P. Introduction to machine learning, neural networks, and deep learning. Transl. Vis. Sci. Technol. 2020, 9, 14. [Google Scholar]
- Wosiak, A.; Zamecznik, A.; Niewiadomska-Jarosik, K. Supervised and Unsupervised Machine Learning for Improved Identification of Intrauterine Growth Restriction Types. In Proceedings of the 2016 Federated Conference on Computer Science and Information Systems (FEDCSIS), Gdansk, Poland, 11–14 September 2016; Ganzha, M., Maciaszek, L., Paprzycki, M., Eds.; Volume 8, pp. 323–329. [Google Scholar] [CrossRef]
- Ganaie, M.; Hu, M.; Malik, A.; Tanveer, M.; Suganthan, P. Ensemble deep learning: A review. Eng. Appl. Artif. Intell. 2022, 115, 105151. [Google Scholar] [CrossRef]
- Macrohon, J.J.E.; Villavicencio, C.N.; Inbaraj, X.A.; Jeng, J.H. A Semi-Supervised Machine Learning Approach in Predicting High-Risk Pregnancies in the Philippines. Diagnostics 2022, 12, 2782. [Google Scholar] [CrossRef]
- Tonkovic, P.; Kalajdziski, S.; Zdravevski, E.; Lameski, P.; Corizzo, R.; Pires, I.M.; Garcia, N.M.; Loncar-Turukalo, T.; Trajkovik, V. Literature on applied machine learning in metagenomic classification: A scoping review. Biology 2020, 9, 453. [Google Scholar] [CrossRef]
- Bote-Curiel, L.; Muñoz-Romero, S.; Gerrero-Curieses, A.; Rojo-álvarez, J.L. Deep learning and big data in healthcare: A double review for critical beginners. Appl. Sci. 2019, 9, 2331. [Google Scholar] [CrossRef]
- Ngiam, K.Y.; Khor, I.W. Big data and machine learning algorithms for health-care delivery. Lancet Oncol. 2019, 20, e262–e273. [Google Scholar] [CrossRef] [PubMed]
- Oprescu, A.M.; Miró-Amarante, G.; García-Díaz, L.; Beltrán, L.M.; Rey, V.E.; Romero-Ternero, M. Artificial intelligence in pregnancy: A scoping review. IEEE Access 2020, 8, 181450–181484. [Google Scholar] [CrossRef]
- Page, M.J.; McKenzie, J.E.; Bossuyt, P.M.; Boutron, I.; Hoffmann, T.C.; Mulrow, C.D.; Shamseer, L.; Tetzlaff, J.M.; Akl, E.A.; Brennan, S.E.; et al. The PRISMA 2020 statement: An updated guideline for reporting systematic reviews. BMJ 2021, 372, 71. [Google Scholar] [CrossRef]
- Covidence Inc., 446 Collins St, Melbourne VIC 3000, Australia. Covidence Tool. 2022. Available online: https://www.covidence.org/ (accessed on 15 May 2022).
- Joanna Briggs Institute. Joanna Briggs Institute Checklists. Available online: https://jbi.global/critical-appraisal-tools (accessed on 8 August 2022).
- CASP. Critical Appraisals Skills Programme (CASP). Available online: https://casp-uk.net/casp-tools-checklists/ (accessed on 9 August 2022).
- R Core Team. R: A Language and Environment for Statistical Computing. 2022. Available online: https://www.R-project.org/ (accessed on 9 April 2022).
- RStudio Team. RStudio: Integrated Development Environment for R. 2020. Available online: https://www.rstudio.com/ (accessed on 9 April 2022).
- Shim, S.R.; Kim, S.J.; Lee, J. Diagnostic test accuracy: Application and practice using R software. Epidemiol. Health 2019, 41, e2019007. [Google Scholar] [CrossRef]
- Higgins, J.P.T.; Thompson, S.G.; Deeks, J.J.; Altman, D.G. Measuring inconsistency in meta-analyses. BMJ Clinical Res. Ed. 2003, 327, 557–560. [Google Scholar] [CrossRef]
- Lunghi, F.; Magenes, G.; Pedrinazzi, L.; Signorini, M. Detection of fetal distress though a support vector machine based on fetal heart rate parameters. In Proceedings of the Computers in Cardiology, Lyon, France, 25–28 September 2005; pp. 247–250. [Google Scholar] [CrossRef]
- Bahado-Singh, R.O.; Yilmaz, A.; Bisgin, H.; Turkoglu, O.; Kumar, P.; Sherman, E.; Mrazik, A.; Odibo, A.; Graham, S.F. Artificial intelligence and the analysis of multi-platform metabolomics data for the detection of intrauterine growth restriction. PLoS ONE 2019, 14, e0214121. [Google Scholar] [CrossRef]
- Pini, N.; Lucchini, M.; Esposito, G.; Tagliaferri, S.; Campanile, M.; Magenes, G.; Signorini, M.G. A Machine Learning Approach to Monitor the Emergence of Late Intrauterine Growth Restriction. Front. Artif. Intell. 2021, 4, 622616. [Google Scholar] [CrossRef]
- Magenes, G.; Pedrinazzi, L.; Signorini, M.G. Identification of fetal sufferance antepartum through a multiparametric analysis and a support vector machine. In Proceedings of the Annual International Conference of the IEEE Engineering in Medicine and Biology Society, New York, NY, USA, 30 August–3 September 2006; pp. 462–465. [Google Scholar] [CrossRef]
- Deval, R.; Saxena, P.; Pradhan, D.; Mishra, A.K.; Jain, A.K. A Machine Learning–Based Intrauterine Growth Restriction (IUGR) Prediction Model for Newborns. Indian J. Pediatr. 2022, 89, 1140–1143. [Google Scholar] [CrossRef]
- Guo, Z.; Yang, F.; Zhang, J.; Zhang, Z.; Li, K.; Tian, Q.; Hou, H.; Xu, C.; Lu, Q.; Ren, Z.; et al. Whole-Genome Promoter Profiling of Plasma DNA Exhibits Diagnostic Value for Placenta-Origin Pregnancy Complications. Adv. Sci. 2020, 7, 1901819. [Google Scholar] [CrossRef] [PubMed]
- Magenes, G.; Bellazzi, R.; Malovini, A.; Signorini, M.G. Comparison of data mining techniques applied to fetal heart rate parameters for the early identification of IUGR fetuses. In Proceedings of the 2016 38th Annual International Conference of the IEEE Engineering in Medicine and Biology Society (EMBC), Orlando, FL, USA, 16–20 August 2016; pp. 916–919. [Google Scholar] [CrossRef]
- Xu, C.; Guo, Z.; Zhang, J.; Lu, Q.; Tian, Q.; Liu, S.; Li, K.; Wang, K.; Tao, Z.; Li, C.; et al. Non-invasive prediction of fetal growth restriction by whole-genome promoter profiling of maternal plasma DNA: A nested case-control study. BJOG Int. J. Obstet. Gynaecol. 2021, 128, 458–466. [Google Scholar] [CrossRef] [PubMed]
- Gómez-Jemes, L.; Oprescu, A.M.; Chimenea-Toscano, Á.; García-Díaz, L.; Romero-Ternero, M.d.C. Machine Learning to Predict Pre-Eclampsia and Intrauterine Growth Restriction in Pregnant Women. Electronics 2022, 11, 3240. [Google Scholar] [CrossRef]
- Buscema, M.; Grossi, E.; Montanini, L.; Street, M.E. Data Mining of Determinants of Intrauterine Growth Retardation Revisited Using Novel Algorithms Generating Semantic Maps and Prototypical Discriminating Variable Profiles. PLoS ONE 2015, 10, e0126020. [Google Scholar] [CrossRef]
- Foltran, F.; Berchialla, P.; Bernasconi, S.; Grossi, E.; Gregori, D.; Street, M.E. A systems biology approach: New insights into fetal growth restriction using Bayesian Networks. J. Biol. Regul. Homeost. Agents 2011, 25, 269–277. [Google Scholar] [PubMed]
- Street, M.E.; Grossi, E.; Volta, C.; Faleschini, E.; Bernasconi, S. Placental determinants of fetal growth: Identification of key factors in the insulin-like growth factor and cytokine systems using artificial neural networks. BMC Pediatr. 2008, 8, 24. [Google Scholar] [CrossRef]
- Ferrario, M.; Signorini, M.G.; Magenes, G. Comparison between fetal heart rate standard parameters and complexity indexes for the identification of severe intrauterine growth restriction. Methods Inf. Med. 2007, 46, 186–190. [Google Scholar]
- Dahdouh, S.; Andescavage, N.; Yewale, S.; Yarish, A.; Lanham, D.; Bulas, D.; du Plessis, A.J.; Limperopoulos, C. In vivo placental MRI shape and textural features predict fetal growth restriction and postnatal outcome. J. Magn. Reson. Imaging JMRI 2018, 47, 449–458. [Google Scholar] [CrossRef] [PubMed]
- Sufriyana, H.; Wu, Y.W.; Su, E.C.Y. Prediction of Preeclampsia and Intrauterine Growth Restriction: Development of Machine Learning Models on a Prospective Cohort. JMIR Med. Inform. 2020, 8, e15411. [Google Scholar] [CrossRef]
- Signorini, M.G.; Pini, N.; Malovini, A.; Bellazzi, R.; Magenes, G. Integrating machine learning techniques and physiology based heart rate features for antepartum fetal monitoring. Comput. Methods Programs Biomed. 2020, 185, 105015. [Google Scholar] [CrossRef]
- Aslam, N.; Khan, I.U.; Aljishi, R.F.; Alnamer, Z.M.; Alzawad, Z.M.; Almomen, F.A.; Alramadan, F.A. Explainable Computational Intelligence Model for Antepartum Fetal Monitoring to Predict the Risk of IUGR. Electronics 2022, 11, 593. [Google Scholar] [CrossRef]
- Gürgen, F.; Zengin, Z.; Varol, F. Intrauterine growth restriction (IUGR) risk decision based on support vector machines. Expert Syst. Appl. 2012, 39, 2872–2876. [Google Scholar] [CrossRef]
- Caly, H.; Rabiei, H.; Coste-Mazeau, P.; Hantz, S.; Alain, S.; Eyraud, J.L.; Chianea, T.; Caly, C.; Makowski, D.; Hadjikhani, N.; et al. Machine learning analysis of pregnancy data enables early identification of a subpopulation of newborns with ASD. Sci. Rep. 2021, 11, 1–14. [Google Scholar] [CrossRef] [PubMed]
- Javaid, M.; Haleem, A.; Pratap Singh, R.; Suman, R.; Rab, S. Significance of machine learning in healthcare: Features, pillars and applications. Int. J. Intell. Netw. 2022, 3, 58–73. [Google Scholar] [CrossRef]
- Alkhaleefah, M.; Wu, C.C. A Hybrid CNN and RBF-Based SVM Approach for Breast Cancer Classification in Mammograms. In Proceedings of the 2018 IEEE International Conference on Systems, Man, and Cybernetics, SMC 2018, Miyazaki, Japan, 7–10 October 2018; pp. 894–899. [Google Scholar] [CrossRef]
- Liu, K.; Fu, Q.; Liu, Y.; Wang, C. An integrative bioinformatics analysis of microarray data for identifying hub genes as diagnostic biomarkers of preeclampsia. Biosci. Rep. 2019, 39, 1–10. [Google Scholar] [CrossRef]
- Malacova, E.; Tippaya, S.; Bailey, H.D.; Chai, K.; Farrant, B.M.; Gebremedhin, A.T.; Leonard, H.; Marinovich, M.L.; Nassar, N.; Phatak, A.; et al. Stillbirth risk prediction using machine learning for a large cohort of births from Western Australia, 1980–2015. Sci. Rep. 2020, 10, 5354. [Google Scholar] [CrossRef]
- Bertini, A.; Salas, R.; Chabert, S.; Sobrevia, L.; Pardo, F. Using Machine Learning to Predict Complications in Pregnancy: A Systematic Review. Front. Bioeng. Biotechnol. 2022, 9, 780389. [Google Scholar] [CrossRef]
- Holzinger, A.; Langs, G.; Denk, H.; Zatloukal, K.; Müller, H. Causability and explainability of artificial intelligence in medicine. Wiley Interdiscip. Rev. Data Min. Knowl. Discov. 2019, 9, e1312. [Google Scholar] [CrossRef]
- Molnar, P. Technology on the margins: AI and global migration management from a human rights perspective. Camb. Int. Law J. 2019, 8, 305–330. [Google Scholar] [CrossRef]




Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Rescinito, R.; Ratti, M.; Payedimarri, A.B.; Panella, M. Prediction Models for Intrauterine Growth Restriction Using Artificial Intelligence and Machine Learning: A Systematic Review and Meta-Analysis. Healthcare 2023, 11, 1617. https://doi.org/10.3390/healthcare11111617
Rescinito R, Ratti M, Payedimarri AB, Panella M. Prediction Models for Intrauterine Growth Restriction Using Artificial Intelligence and Machine Learning: A Systematic Review and Meta-Analysis. Healthcare. 2023; 11(11):1617. https://doi.org/10.3390/healthcare11111617
Chicago/Turabian StyleRescinito, Riccardo, Matteo Ratti, Anil Babu Payedimarri, and Massimiliano Panella. 2023. "Prediction Models for Intrauterine Growth Restriction Using Artificial Intelligence and Machine Learning: A Systematic Review and Meta-Analysis" Healthcare 11, no. 11: 1617. https://doi.org/10.3390/healthcare11111617
APA StyleRescinito, R., Ratti, M., Payedimarri, A. B., & Panella, M. (2023). Prediction Models for Intrauterine Growth Restriction Using Artificial Intelligence and Machine Learning: A Systematic Review and Meta-Analysis. Healthcare, 11(11), 1617. https://doi.org/10.3390/healthcare11111617








