A Photo-Controllable DNAzyme-Based Nanosensor for miRNA Imaging in Living Cells
Abstract
1. Introduction
2. Materials and Methods
2.1. Materials
2.2. Polyacrylamide Gel Electrophoresis
2.3. Preparation of hMNs and UTD@hMNs
2.4. miR-21 Detection In Vitro
2.5. Cell Incubation
2.6. MTT Assay
2.7. Stability of UTD@hMNs
2.8. CLSM Imaging
3. Results and Discussion
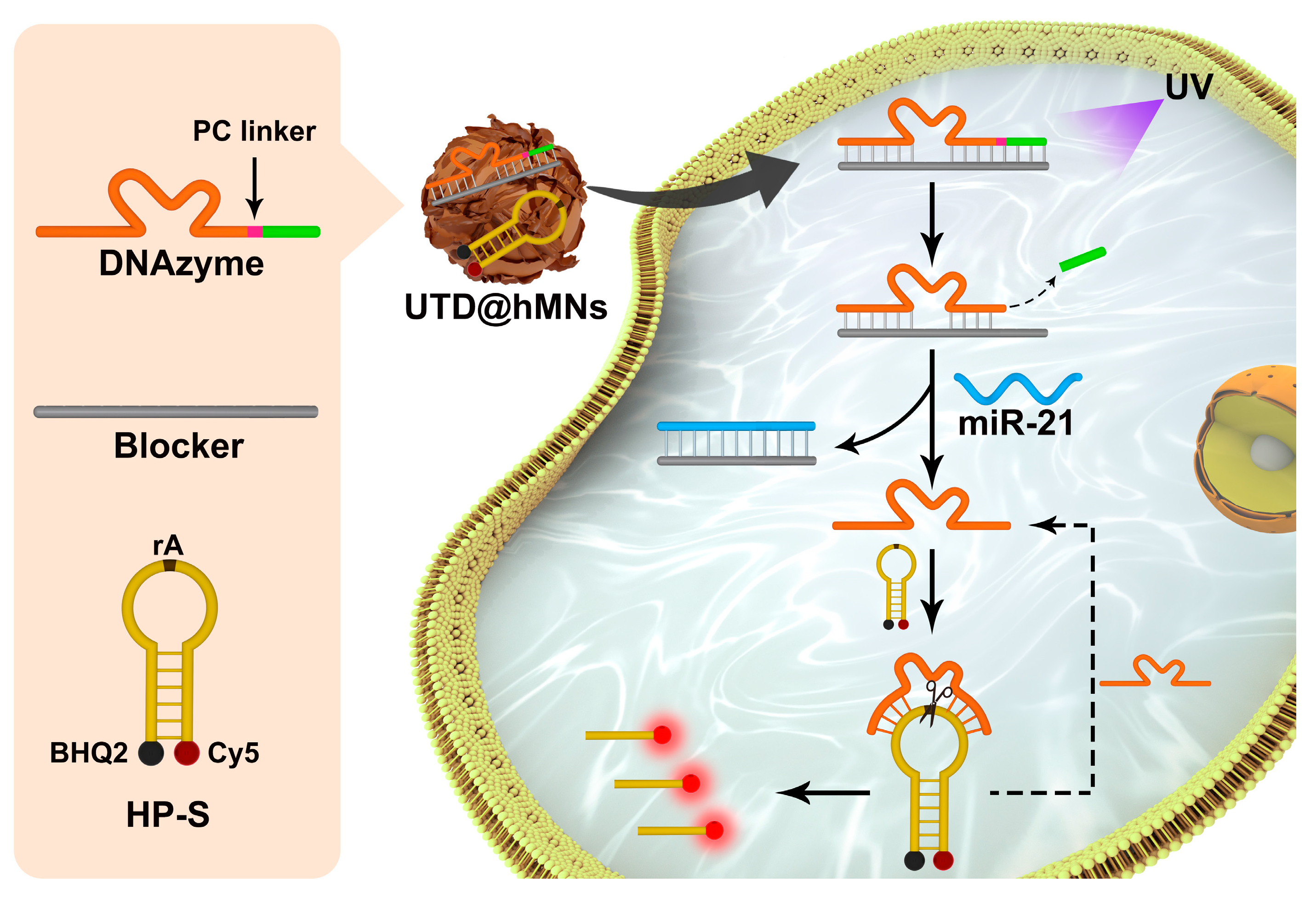
3.1. Principle of the UTD Nanodevices
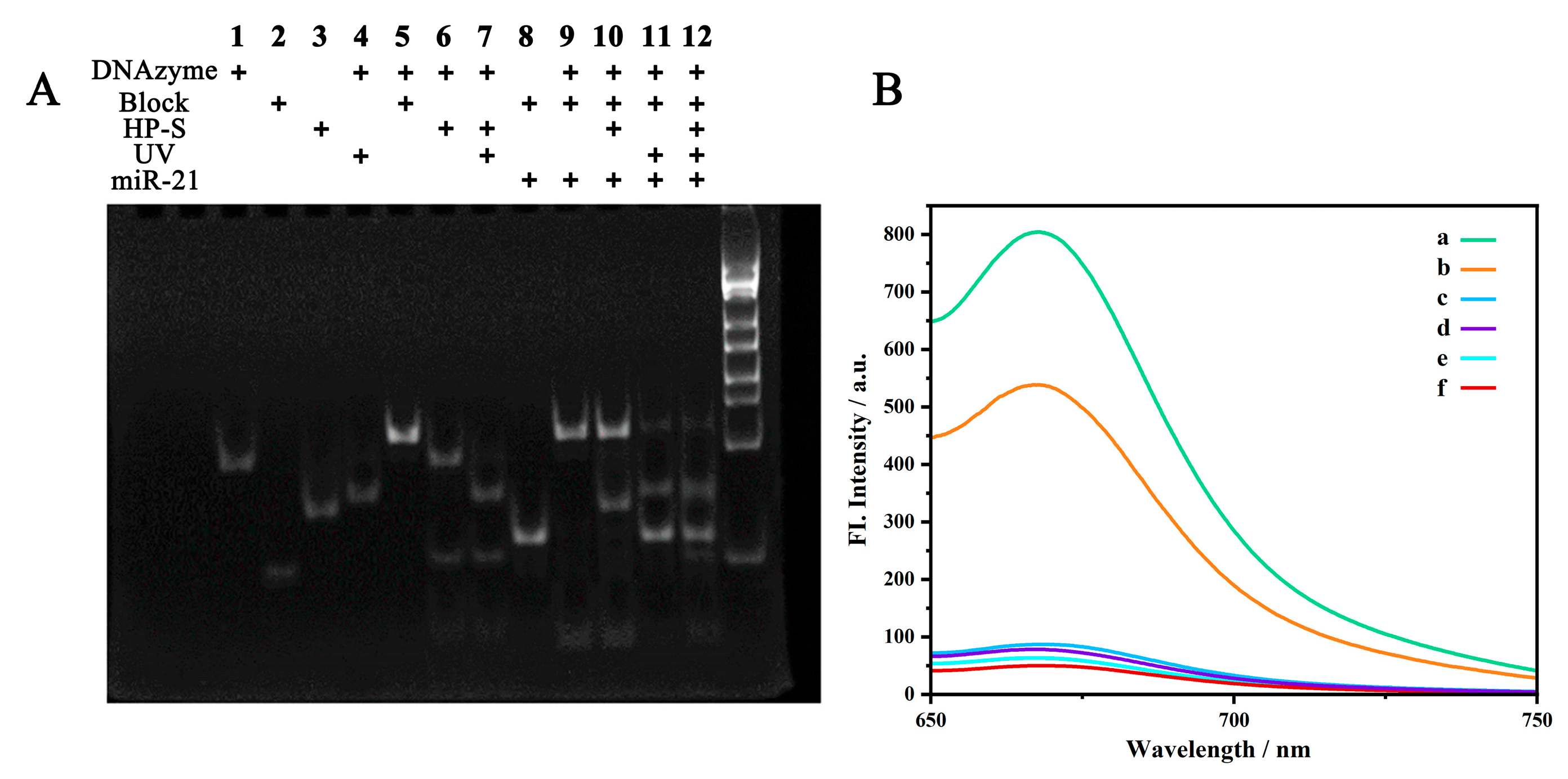
3.2. Feasibility of the UTD Nanosystems
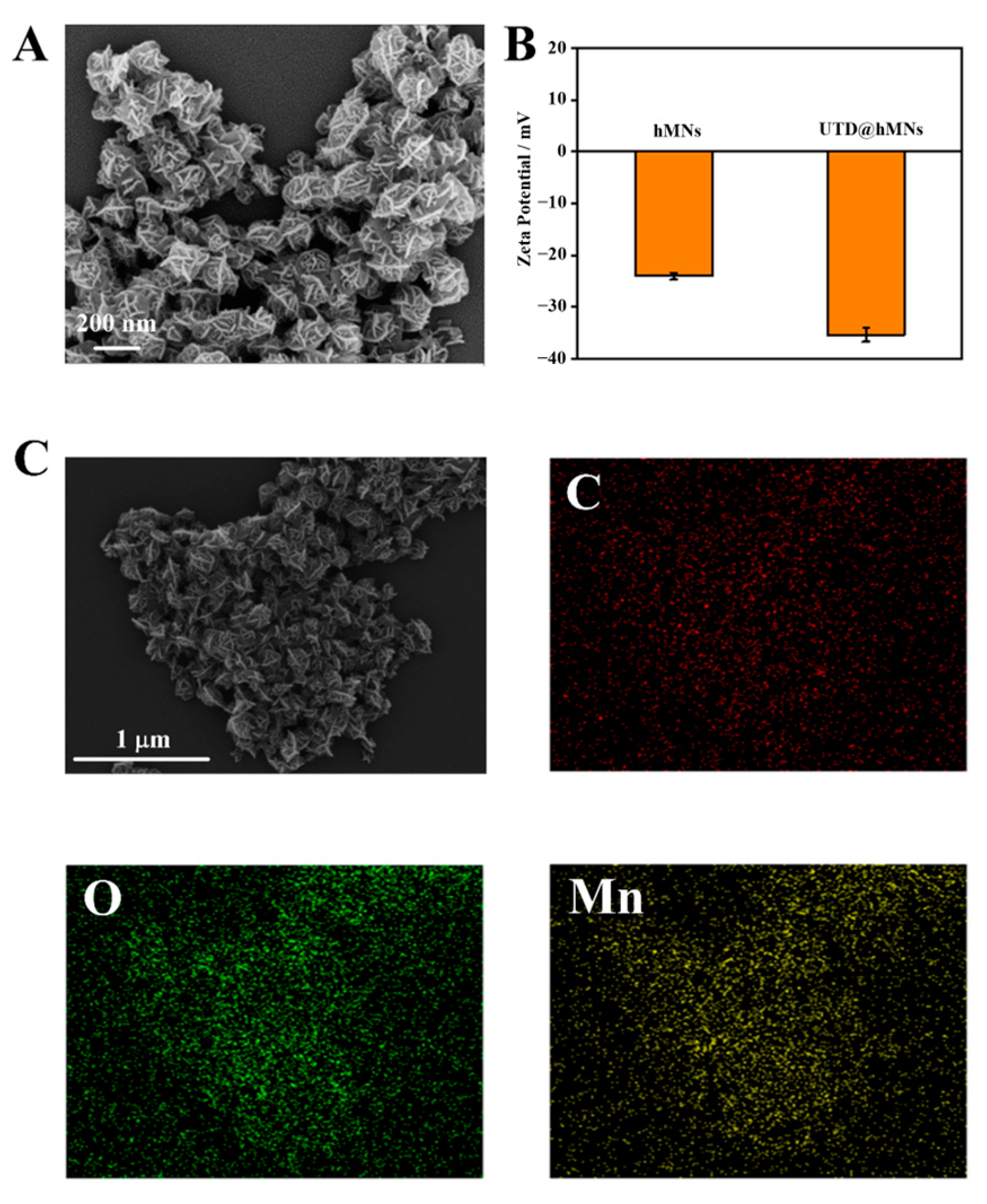
3.3. Characterization of hMNs and UTD@hMNs
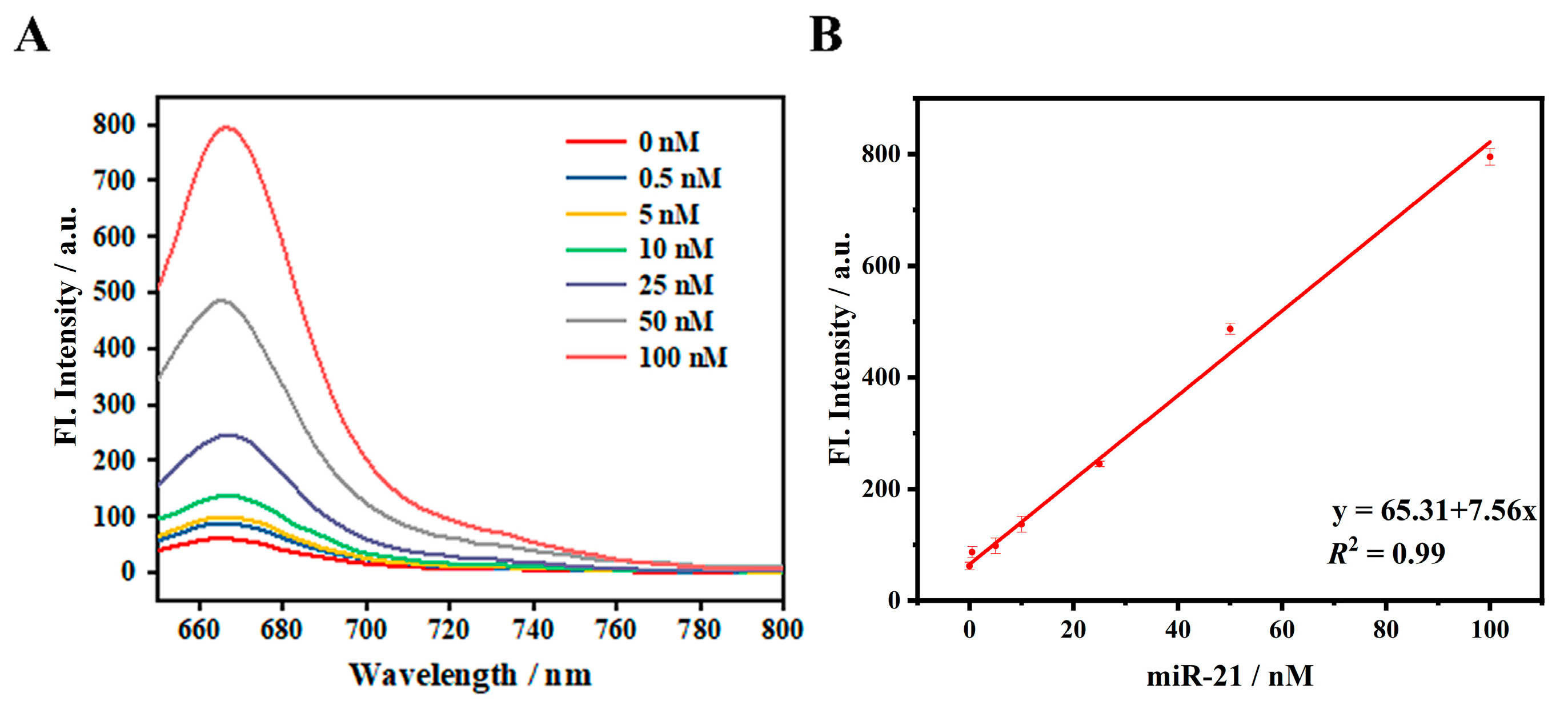
3.4. miRNA Detection In Vitro
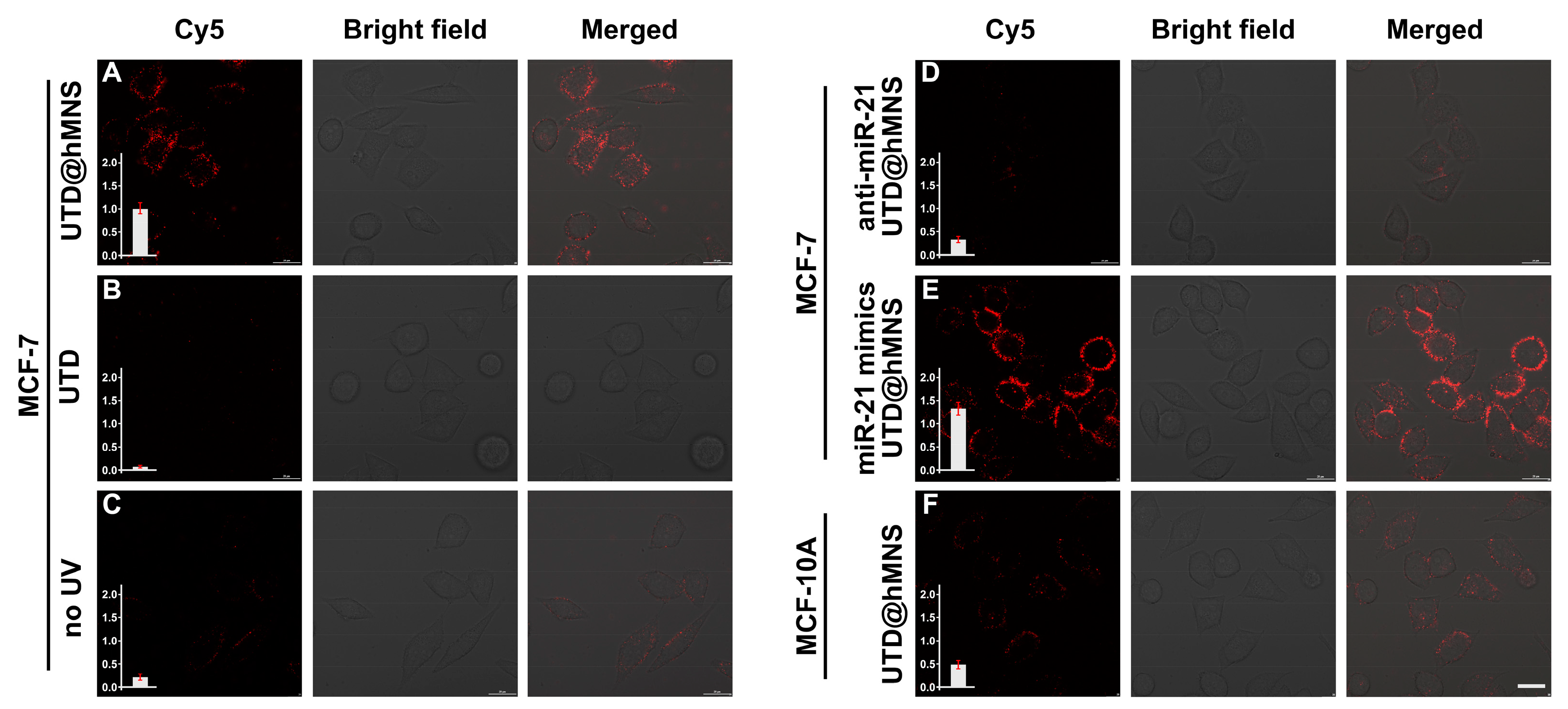
3.5. miRNA Imaging In Vivo
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Abbreviations
| UTD | UV-triggered DNAzyme |
| hMNs | honeycomb MnO2 nanoflowers |
| CLSM | confocal laser scanning microscopy |
References
- You, X.; He, R.; Liu, S.-Y.; Dai, Z. Advances in fluorescence imaging of RNAs in living cells based on functional nucleic acid probes. Biomed. Anal. 2024, 1, 1–27. [Google Scholar]
- Armand-Labit, V.; Pradines, A. Circulating cell-free microRNAs as clinical cancer biomarkers. Biomol. Concepts 2017, 8, 61–81. [Google Scholar]
- Rupaimoole, R.; Calin, G.; Lopez-Berestein, G.; Sood, A. miRNA Deregulation in Cancer Cells and the Tumor Microenvironment. Cancer Discov. 2016, 6, 235–246. [Google Scholar]
- Hu, Z.; Zhang, J.; Li, Y.; Song, S.; Feng, C.; Wang, J.; Zhang, F.; Wang, J.; Liu, X. G-quadruplex-deficient precursor hairpin probes for ultra-low background dual-mode detection of miRNAs. Talanta 2023, 253, 123954. [Google Scholar]
- Yin, M.; Jiao, J.; Lu, L.; Hu, B.; Xue, L.; Dai, F.; Wang, X.; Wang, Z.; Wang, T.; Chen, Q. A simultaneous strategy with multiple-signal amplification and self-calibration for ultrasensitive assay of miRNA-21 based on 3D MNPs-IL-rGO-AuNPs. Biosens. Bioelectron. 2024, 249, 116009. [Google Scholar] [CrossRef]
- Siddika, T.; Heinemann, I.U. Bringing MicroRNAs to Light: Methods for MicroRNA Quantification and Visualization in Live Cells. Front. Bioeng. Biotechnol. 2021, 8, 619583. [Google Scholar]
- Zhou, M.; Teng, X.; Li, Y.; Deng, R.; Li, J. Cascade Transcription Amplification of RNA Aptamer for Ultrasensitive MicroRNA Detection. Anal. Chem. 2019, 91, 5295–5302. [Google Scholar]
- Zhou, Q.; Peng, J.; Liu, S.; Huang, T.; Luo, J.; Bai, L.; Xie, B.; Chen, J.; Li, C.; Chen, J. A target-triggered mimic nucleic acid enzyme enables versatile and sequence-independent biosensing. Biomed. Anal. 2024, 1, 28–35. [Google Scholar]
- Nie, K.; Jiang, Y.; Wang, N.; Wang, Y.; Li, D.; Zhan, L.; Huang, C.; Li, C. Programmable, Universal DNAzyme Amplifier Supporting Pancreatic Cancer-Related miRNAs Detection. Chemosensors 2022, 10, 276. [Google Scholar] [CrossRef]
- Wang, M.; Liu, Z.; Liu, C.; He, W.; Qin, D.; You, M. DNAzyme-based ultrasensitive immunoassay: Recent advances and emerging trends. Biosens. Bioelectron. 2024, 251, 116122. [Google Scholar]
- Shi, C.; Yang, D.; Ma, X.; Pan, L.; Shao, Y.; Arya, G.; Ke, Y.; Zhang, C.; Wang, F.; Zuo, X.; et al. A Programmable DNAzyme for the Sensitive Detection of Nucleic Acids. Angew. Chem. Int. Ed. 2024, 63, e202320179. [Google Scholar]
- Zhou, W.; Liang, W.; Li, D.; Yuan, R.; Xiang, Y. Dual-color encoded DNAzyme nanostructures for multiplexed detection of intracellular metal ions in living cells. Biosens. Bioelectron. 2016, 85, 573–579. [Google Scholar] [PubMed]
- Zhou, Y.; Ma, W.; Sun, R.; Liu, B.; Zhang, X.; Yang, H. Upconverting nanoparticles based nanodevice for DNAzymes amplified miRNAs detection and artificially controlled chemo-gene therapy. Biosens. Bioelectron. 2022, 214, 114549. [Google Scholar]
- Chen, L.; Liu, Y.; Guo, W.; Liu, Z. Light responsive nucleic acid for biomedical application. Exploration 2022, 2, 20210099. [Google Scholar]
- Huang, S.; Zhu, J.-J. Linkage Pathways of DNA–Nanoparticle Conjugates and Biological Applications. Chemosensors 2023, 11, 444. [Google Scholar]
- Zhou, H.; Jiang, Y.; Zhao, W.; Zhang, S. Light-Activated Nanodevice for On-Demand Imaging of miRNA in Living Cells via Logic Assembly. ACS Appl. Mater. Interfaces 2022, 14, 13070–13078. [Google Scholar]
- Qiu, L.; Wu, C.; You, M.; Han, D.; Chen, T.; Zhu, G.; Jiang, J.; Yu, R.; Tan, W. A Targeted, Self-Delivered, and Photocontrolled Molecular Beacon for mRNA Detection in Living Cells. J. Am. Chem. Soc. 2013, 135, 12952–12955. [Google Scholar]
- Zhao, J.; Gao, J.; Xue, W.; Di, Z.; Xing, H.; Lu, Y.; Li, L. Upconversion Luminescence-Activated DNA Nanodevice for ATP Sensing in Living Cells. J. Am. Chem. Soc. 2018, 140, 578–581. [Google Scholar]
- Wang, W.; Satyavolu, N.S.R.; Wu, Z.; Zhang, J.-R.; Zhu, J.-J.; Lu, Y. Near-Infrared Photothermally Activated DNAzyme-Gold Nanoshells for Imaging Metal Ions in Living Cells. Angew. Chem. Int. Ed. 2017, 56, 6798–6802. [Google Scholar]
- Stafford, M.; Willoughby, C.; Walsh, C.; McKenna, D. Prognostic value of miR-21 for prostate cancer: A systematic review and meta-analysis. Biosci. Rep. 2022, 42, SR20211972. [Google Scholar]
- Hu, M.; Qiu, Z.; Bi, Z.; Tian, T.; Jiang, Y.; Zhou, X. Photocontrolled crRNA activation enables robust CRISPR-Cas12a diagnostics. Proc. Natl. Acad. Sci. USA 2022, 119, e2202034119. [Google Scholar] [PubMed]
- Zhao, J.; Di, Z.; Li, L. Spatiotemporally Selective Molecular Imaging via Upconversion Luminescence-Controlled, DNA-Based Biosensor Technology. Angew. Chem. (Int. Ed. Engl.) 2022, 61, e202204277. [Google Scholar] [PubMed]
- Chen, H.; He, J.; Zhang, C.; He, H. Self-Assembly of Novel Mesoporous Manganese Oxide Nanostructures and Their Application in Oxidative Decomposition of Formaldehyde. J. Phys. Chem. C 2007, 111, 18033–18038. [Google Scholar]
- Xue, H.; Yu, M.; He, K.; Liu, Y.; Cao, Y.; Shui, Y.; Li, J.; Farooq, M.; Wang, L. A novel colorimetric and fluorometric probe for biothiols based on MnO2 NFs-Rhodamine B system. Anal. Chim. Acta 2020, 1127, 39–48. [Google Scholar]
- Wei, J.; Wang, H.; Wu, Q.; Gong, X.; Ma, K.; Liu, X.; Wang, F. A Smart, Autocatalytic, DNAzyme Biocircuit for In Vivo, Amplified, MicroRNA Imaging. Angew. Chem. Int. Ed. 2020, 59, 5965–5971. [Google Scholar] [CrossRef]
- Fan, Z.; Yan, J.; Wei, T.; Zhi, L.; Ning, G.; Li, T.; Wei, F. Asymmetric Supercapacitors Based on Graphene/MnO2 and Activated Carbon Nanofiber Electrodes with High Power and Energy Density. Adv. Funct. Mater. 2011, 21, 2366–2375. [Google Scholar]
- Rama, A.R.; Lara, P.; Mesas, C.; Quiñonero, F.; Vélez, C.; Melguizo, C.; Prados, J. Circular Sponge against miR-21 Enhances the Antitumor Activity of Doxorubicin against Breast Cancer Cells. Int. J. Mol. Sci. 2022, 23, 14803. [Google Scholar] [CrossRef]
| Strand | Sequence (5′-3′) |
|---|---|
| DNAzyme (D) | CTTATCAGAC/iPCLink/TGATGTTGATCTTCTCTTCTCCGAGCCGGTCGAAATAGTAGCTTA |
| blocker (B) | AAGAGAAGATCAACATCAGTCTGATAAGCTA |
| HP-S | BHQ2-ACGTAAAGCAAAGAAGCTACTAT/rA/GGAAGAGAAGACTAGCTTACCATGCTTTACGT-Cy5 |
| Cy5-HP-S | ACGTAAAGCAAAGAAGCTACTAT/rA/GGAAGAGAAGACTAGCTTACCATGCTTTACGT-Cy5 |
| mismatch-1 (mis-1) | TAACTTATCAGACTGATGTTGA |
| mismatch-2 (mis-2) | TAGCTTATCAGACTGATAATGA |
| mismatch-3 (mis-3) | TAGCTTATCAGACTGATAAAGA |
| miR-205 | TCCTTCATTCCACCGGAGTCTG |
| miR-155 | TTAATGCTAATCGTGATAGGGG |
| anti-miR-21 | TCAACATCAGTCTGATAAGCTA |
| miR-21 | UAGCUUAUCAGACUGAUGUUGA |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhang, Y.; Zhang, Y.; Ouyang, R.; Dai, Z.; Liu, S.-Y. A Photo-Controllable DNAzyme-Based Nanosensor for miRNA Imaging in Living Cells. Chemosensors 2025, 13, 123. https://doi.org/10.3390/chemosensors13040123
Zhang Y, Zhang Y, Ouyang R, Dai Z, Liu S-Y. A Photo-Controllable DNAzyme-Based Nanosensor for miRNA Imaging in Living Cells. Chemosensors. 2025; 13(4):123. https://doi.org/10.3390/chemosensors13040123
Chicago/Turabian StyleZhang, Yanfei, Yiling Zhang, Runqi Ouyang, Zong Dai, and Si-Yang Liu. 2025. "A Photo-Controllable DNAzyme-Based Nanosensor for miRNA Imaging in Living Cells" Chemosensors 13, no. 4: 123. https://doi.org/10.3390/chemosensors13040123
APA StyleZhang, Y., Zhang, Y., Ouyang, R., Dai, Z., & Liu, S.-Y. (2025). A Photo-Controllable DNAzyme-Based Nanosensor for miRNA Imaging in Living Cells. Chemosensors, 13(4), 123. https://doi.org/10.3390/chemosensors13040123







