MicroRNA-138-5p Targets Pro-Apoptotic Factors and Favors Neural Cell Survival: Analysis in the Injured Spinal Cord
Abstract
:1. Introduction
2. Materials and Methods
2.1. Surgical Procedures
2.2. Histology, Fluorescent in Situ Hybridization (FISH), Immunofluorescence, and Image Analysis
2.3. Cell Culture
2.4. Computational Prediction of miR-138-5p Targets
2.5. Microarray Data
2.6. RT-qPCR Analysis
2.7. Cell Death Assays
2.7.1. Calcein/Propidium Iodide Assay
2.7.2. Terminal Deoxynucleotidyl Transferase-Mediated Deoxyuridine Triphosphate (dUTP) Nick End-Labeling (TUNEL) Assay
2.7.3. MTT Assay
2.7.4. Quantification of Cell Death Using Flow Cytometry
2.8. Dual Luciferase Reporter Gene Construction and 3′UTR Luciferase Reporter Assays
2.9. Immunoblot Assay
2.10. Measurement of Caspase-3/7 Activity
2.11. Data Analysis
3. Results
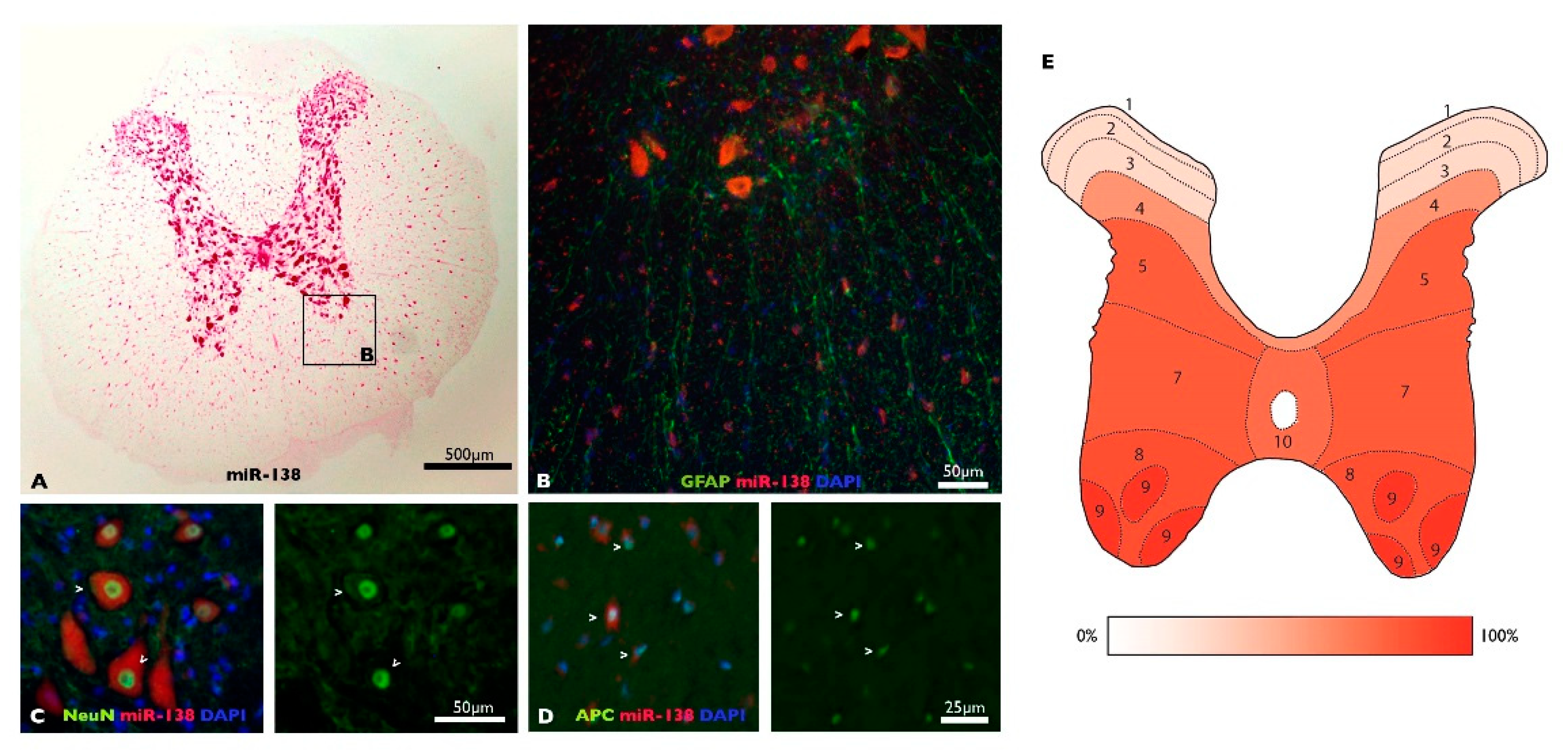
3.1. MiR-138-5p Becomes Downregulated among Spinal Neurons during Subacute SCI
3.2. MiR-138-5p Protects Neurons In Vitro
3.3. Predicting Novel Targets of miR-138-5p among the Genes Dysregulated after SCI
3.4. Reporter Assays Validate miR-138-5p Targeting on Casp3, Casp7, and Bak1 3′UTRs
3.5. MiR-138-5p Reduces the Gene and Protein Expression of Caspase 3, Caspase 7, and BAK
3.6. MiR-138-5p Attenuates Caspase-Dependent Apoptosis
3.7. Expression of Caspases 3 and 7 Increases after Subacute SCI and Negatively Correlates with the Downregulation of miR-18-5p
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Ahuja, C.S.; Wilson, J.R.; Nori, S.; Kotter, M.R.N.; Druschel, C.; Curt, A.; Fehlings, M.G. Traumatic spinal cord injury. Nat. Rev. Dis. Primers. 2017, 3, 17018. [Google Scholar] [CrossRef] [PubMed]
- Crowe, M.J.; Bresnahan, J.C.; Shuman, S.L.; Masters, J.N.; Beattie, M.S. Apoptosis and delayed degeneration after spinal cord injury in rats and monkeys. Nat. Med. 1997, 3, 73–76. [Google Scholar] [CrossRef] [PubMed]
- Grossman, S.D.; Rosenberg, L.J.; Wrathall, J.R. Temporal–Spatial Pattern of Acute Neuronal and Glial Loss after Spinal Cord Contusion. Exp. Neurol. 2001, 168, 273–282. [Google Scholar] [CrossRef] [PubMed]
- Liu, X.Z.; Xu, X.M.; Hu, R.; Du, C.; Zhang, S.X.; McDonald, J.W.; Dong, H.X.; Wu, Y.J.; Fan, G.S.; Jacquin, M.F.; et al. Neuronal and glial apoptosis after traumatic spinal cord injury. J. Neurosci. 1997, 17, 5395–5406. [Google Scholar] [CrossRef]
- Lou, J.; Lenke, L.G.; Ludwig, F.J.; O’Brien, M.F. Apoptosis as a mechanism of neuronal cell death following acute experimental spinal cord injury. Spinal Cord. 1998, 36, 683–690. [Google Scholar] [CrossRef]
- Galluzzi, L.; Vitale, I.; Aaronson, S.A.; Abrams, J.M.; Adam, D.; Agostinis, P.; Alnemri, E.S.; Altucci, L.; Amelio, I.; Andrews, D.W.; et al. Molecular mechanisms of cell death: Recommendations of the Nomenclature Committee on Cell Death 2018. Cell Death Differ. 2018, 25, 486–541. [Google Scholar] [CrossRef]
- Aimone, J.; Leasure, J.; Perreau, V.; Thallmair, M. The Christopher Reeve Paralysis Foundation. Spatial and temporal gene expression profiling of the contused rat spinal cord. Exp. Neurol. 2004, 189, 204–221. [Google Scholar] [CrossRef]
- Ahn, Y.H.; Bae Yeon, Y.; Lee, G.; Kang Mee, K.; Kang, S.K. Molecular insights of the injured lesions of rat spinal cords: Inflammation, apoptosis, and cell survival. Biochem. Biophys. Res. Commu. 2006, 348, 560–570. [Google Scholar] [CrossRef]
- Di Giovanni, S.; De Biase, A.; Yakovlev, A.; Finn, T.; Beers, J.; Hoffman, E.P.; Faden, A.I. In Vivo and in Vitro Characterization of el Neuronal Plasticity Factors Identified following Spinal Cord Injury. J. Biol. Chem. 2005, 21, 2084–2091. [Google Scholar] [CrossRef]
- Barbon, A.; Fumagalli, F.; Caracciolo, L.; Madaschi, L.; Lesma, E.; Mora, C.; Carelli, S.; Slotkin, T.A.; Racagni, G.; Di Giulio, A.M.; et al. Acute spinal cord injury persistently reduces R/G RNA editing of AMPA receptors: AMPA R/G editing is reduced by SCI. J. Neurochem. 2010, 114, 397–407. [Google Scholar] [CrossRef]
- Nakae, A.; Nakai, K.; Tanaka, T.; Hosokawa, K.; Mashimo, T. Serotonin 2C receptor alternative splicing in a spinal cord injury model. Neurosci. Lett. 2013, 532, 49–54. [Google Scholar] [CrossRef] [PubMed]
- Liu, N.K.; Wang, X.F.; Lu, Q.B.; Xu, X.M. Altered microRNA expression following traumatic spinal cord injury. Exp. Neurol. 2009, 219, 424–429. [Google Scholar] [CrossRef] [PubMed]
- Strickland, E.R.; Hook, M.A.; Balaraman, S.; Huie, J.R.; Grau, J.W. Miranda RC. MicroRNA dysregulation following spinal cord contusion: Implications for neural plasticity and repair. Neuroscience 2011, 186, 146–160. [Google Scholar] [CrossRef] [PubMed]
- Yunta, M.; Nieto-Diaz, M.; Esteban, F.J.; Caballero-Lopez, M.; Navarro-Ruiz, R.; Reigada, D.; Pita-Thomas, D.W.; Águila, Á.D.; Munoz-Galdeano, T.; Maza, R.M. MicroRNA Dysregulation in the Spinal Cord following Traumatic Injury. Di Giovanni S, editor. PLoS ONE 2012, 7, e34534. [Google Scholar]
- Kosik, K.S. MicroRNAs and Cellular Phenotypy. Cell 2010, 143, 21–26. [Google Scholar] [CrossRef]
- Bhalala, O.G.; Srikanth, M.; Kessler, J.A. The emerging roles of microRNAs in CNS injuries. Nat. Rev. Neurol. 2013, 9, 328–339. [Google Scholar] [CrossRef]
- Meza-Sosa, K.F.; Valle-García, D.; Pedraza-Alva, G.; Pérez-Tínez, L. Role of microRNAs in central nervous system development and pathology. J. Neurosci. Res. 2012, 90, 1–12. [Google Scholar] [CrossRef]
- Saugstad, J.A. MicroRNAs as Effectors of Brain Function with Roles in Ischemia and Injury, Neuroprotection, and Neurodegeneration. J. Cereb. Blood Flow Metabol. 2010, 30, 1564–1576. [Google Scholar] [CrossRef]
- Liu, X.J.; Zheng, X.P.; Zhang, R.; Guo, Y.L.; Wang, J.H. Combinatorial effects of miR-20a and miR-29b on neuronal apoptosis induced by spinal cord injury. Int. J. Clin. Exp. Pathol. 2015, 8, 3811–3818. [Google Scholar]
- Yi, X.; Jiang, D. miR-124 mediated neuron apoptosis in spinal cord injury by targeting the 3’ UTR end of calpain. Int. J. Clin. Exp. Pathol. 2016, 15, 5989–5997. [Google Scholar]
- Tang, Y.; Fu, R.; Ling, Z.M.; Liu, L.L.; Yu, G.Y.; Li, W.; Fang, X.Y.; Zhu, Z.; Wu, W.T.; Zhou, L.H. MiR-137-3p rescue motoneuron death by targeting calpain-2. Nitric Oxide 2018, 74, 74–85. [Google Scholar] [CrossRef] [PubMed]
- Siegel, G.; Obernosterer, G.; Fiore, R.; Oehmen, M.; Bicker, S.; Christensen, M.; Khudayberdiev, S.; Leuschner, P.F.; Busch, C.J.; Kane, C.; et al. A functional screen implicates microRNA-138-dependent regulation of the depalmitoylation enzyme APT1 in dendritic spine morphogenesis. Nat. Cell Biol. 2009, 11, 705–716. [Google Scholar] [CrossRef] [PubMed]
- Liu, C.M.; Wang, R.Y.; Jiao, Z.X.; Zhang, B.Y.; Zhou, F.Q. MicroRNA-138 and SIRT1 form a mutual negative feedback loop to regulate mammalian axon regeneration. Genes Dev. 2013, 7, 27. [Google Scholar] [CrossRef]
- Qian, B.J.; You, L.; Shang, F.F.; Liu, J.; Dai, P.; Lin, N.; He, M.; Liu, R.; Zhang, Y.; Xu, Y.; et al. Vimentin Regulates Neuroplasticity in Transected Spinal Cord Rats Associated with miRNA138. Mol. Neurobiol. 2015, 51, 437–447. [Google Scholar] [CrossRef] [PubMed]
- Chan, X.H.D.; Nama, S.; Gopal, F.; Rizk, P.; Ramasamy, S.; Meenakshisundaram, G.; Ow, G.S.; Vladimirovna, I.A.; Tanavde, V.; Haybaeck, J.; et al. Targeting Glioma Stem Cells by Functional Inhibition of a Prosurvival OncomiR-138 in Malignant Gliomas. Cell Rep. 2012, 2, 591–602. [Google Scholar] [CrossRef]
- Basso, D.M.; Beattie, M.S.; Bresnahan, J.C. A sensitive and reliable locomotor rating scale for open field testing in rats. J. Neurotrauma 1995, 12, 1–21. [Google Scholar] [CrossRef] [PubMed]
- Søe, M.J.; Møller, T.; Dufva, M.; Holmstrøm, K. A sensitive alternative for microRNA in situ hybridizations using probes of 2′-O-methyl RNA + LNA. J. Histochem. Cytochem. 2011, 59, 661–672. [Google Scholar] [CrossRef]
- Schneider, C.A.; Rasband, W.S.; Eliceiri, K.W. NIH Image to ImageJ: 25 years of image analysis. Nat. Methods 2012, 9, 671–675. [Google Scholar] [CrossRef]
- Agarwal, V.; Bell, G.W.; Nam, J.W.; Bartel, D.P. Predicting effective microRNA target sites in mammalian mRNAs. Elife 2015, 4, 101. [Google Scholar] [CrossRef]
- Vejnar, C.E.; Blum, M.; Zdob, E.M. miRmap web: Comprehensive microRNA target prediction online. Nucleic Acids Res. 2013, 1, 41. [Google Scholar] [CrossRef]
- Betel, D.; Wilson, M.; Gabow, A.; Marks, D.S.; Sander, C. The microRNA.org resource: Targets and expression. Nucleic Acids Res. 2008, 1, 36. [Google Scholar] [CrossRef] [PubMed]
- Dweep, H.; Gretz, N. miRWalk2.0: A comprehensive atlas of microRNA-target interactions. Nat. Publ. Group 2015, 12, 697. [Google Scholar] [CrossRef]
- Barrett, T.; Wilhite, S.E.; Ledoux, P.; Evangelista, C.; Kim, I.F.; Tomashevsky, M.; Marshall, K.A.; Phillippy, K.H.; Sherman, P.M.; Holko, M.; et al. NCBI GEO: Archive for functional genomics data sets—update. Nucleic Acids Res. 2012, 26, 41. [Google Scholar] [CrossRef] [PubMed]
- Di Giovanni, S.; Knoblach, S.M.; Brandoli, C.; Aden, S.A.; Hoffman, E.P.; Faden, A.I. Gene profiling in spinal cord injury shows role of cell cycle in neuronal death. Ann. Neurol. 2003, 53, 454–468. [Google Scholar] [CrossRef] [PubMed]
- Duan, H.; Ge, W.; Zhang, A.; Xi, Y.; Chen, Z.; Luo, D.; Cheng, Y.; Fan, K.S.; Horvath, S.; Sofroniew, M.V.; et al. Transcriptome analyses reveal molecular mechanisms underlying functional recovery after spinal cord injury. Proc. Natl. Acad. Sci. USA 2015, 112, 13360–13365. [Google Scholar] [CrossRef]
- Huang, D.W.; Sherman, B.T.; Lempicki, R.A. Systematic and integrative analysis of large gene lists using DAVID Bioinformatics Resources. Nat. Protoc. 2009, 4, 44–57. [Google Scholar] [CrossRef]
- Bioinformatics: Volume I: Data, Sequence Analysis, and Evolution. In Methods in Molecular Biology, 2nd ed.; Keith, J.M. (Ed.) Humana Press: Berlin/Heidelberg, Germany, 2017; p. 1525. [Google Scholar]
- Kertesz, M.; Iovino, N.; Unnerstall, U.; Gaul, U.; Segal, E. The role of site accessibility in microRNA target recognition. Nat. Genet. 2007, 39, 1278–1284. [Google Scholar] [CrossRef]
- Lorenz, R.; Bernhart, S.H.; Höner zu Siederdissen, C.; Tafer, H.; Flamm, C.; Stadler, P.F.; Hofacker, I.L. ViennaRNA Package 2.0. Algorithms Mol. Biol. 2011, 6, 26. [Google Scholar] [CrossRef]
- Livak, K.J.; Schmittgen, T.D. Analysis of Relative Gene Expression Data Using Real-Time Quantitative PCR and the 2−ΔΔCT Method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef]
- Headrick, T.C. Statistical Simulation: Power Method Polynomials and Other Transformations; Chapman & Hall/CRC: Boca Raton, FL, USA, 2010; 166p. [Google Scholar]
- Hwang, D.W.; Son, S.; Jang, J.; Youn, H.; Lee, S.; Lee, D.; Lee, Y.S.; Jeong, J.M.; Kim, W.J.; Lee, D.S. A brain-targeted rabies virus glycoprotein-disulfide linked PEI nanocarrier for delivery of neurogenic microRNA. Biomaterials 2011, 32, 4968–4975. [Google Scholar] [CrossRef] [PubMed]
- Maza, R.M.; Reinecke, H.; Muñoz-Galdeano, T.; Reigada, D.; Elvira, C.; Tínez-Campos, E.; Gallardo, A.; Bosch, P.; Rodriguez-Hernandez, J.; Nieto-Diaz, M. Evaluation of RVG29-Functionalized polyethylenimines for neural cell transfection. In Proceedings of the Poster Communication to the 10th Cajal Conference, Cuenca, Spain; Organized by SENC. 2018. Available online: https://osf.io/yqn58/ (accessed on 24 May 2022).
- R Core Team. R: A language and environment for statistical computing. In Proceedings of the R Foundation for Statistical Computing, Vienna, Austria; 2017. Available online: https://wwwR-project.org/ (accessed on 10 February 2015).
- Landgraf, P.; Rusu, M.; Sheridan, R.; Sewer, A.; Iovino, N.; Aravin, A.; Pfeffer, S.; Rice, A.; Kamphorst, A.O.; Landthaler, M.; et al. A mammalian microRNA expression atlas based on small RNA library sequencing. Cell 2007, 129, 1401–1414. [Google Scholar] [CrossRef] [PubMed]
- Obernosterer, G.; Leuschner, P.J.; Alenius, M.; Martinez, J. Post-transcriptional regulation of microRNA expression. RNA 2006, 12, 1161–1167. [Google Scholar] [CrossRef] [PubMed]
- Pan, D.; Flores, O.; Umbach, J.L.; Pesola, J.M.; Bentley, P.; Rosato, P.C.; Leib, D.A.; Cullen, B.R.; Coen, D.M. A neuron-specific host microRNA targets herpes simplex virus-1 ICP0 expression and promotes latency. Cell Host Microbe 2014, 15, 446–456. [Google Scholar] [CrossRef] [PubMed]
- Westphal, D.; Dewson, G.; Czabotar, P.E.; Kluck, R.M. Molecular biology of Bax and Bak activation and action. Biochim. Biophys. Acta (BBA) Mol. Cell Res. 2011, 1813, 521–531. [Google Scholar] [CrossRef] [PubMed]
- Juźwik, C.A.; S. Drake, S.; Zhang, Y.; Paradis-Isler, N.; Sylvester, A.; Zifkin, A.A.; Douglas, C.; Morquette, B.; Moore, C.S.; Fournier, A.E. microRNA dysregulation in neurodegenerative diseases: A systematic review. Prog. Neurobiol. 2019, 182, 101664. [Google Scholar] [CrossRef] [PubMed]
- Nieto-Diaz, M.; Esteban, F.J.; Reigada, D.; Muñoz-Galdeano, T.; Yunta, M.; Caballero-López, M.; Navarro-Ruiz, R.; Del Águila, A.; Maza, R.M. MicroRNA dysregulation in spinal cord injury: Causes, consequences and therapeutics. Front. Cell. Neurosci. 2014, 8, 53. [Google Scholar] [CrossRef]
- Ludwig, N.; Leidinger, P.; Becker, K.; Backes, C.; Fehlmann, T.; Pallasch, C.; Rheinheimer, S.; Meder, B.; Stähler, C.; Meese, E.; et al. Distribution of miRNA expression across human tissues. Nucleic Acids Res. 2016, 44, 3865–3877. [Google Scholar] [CrossRef]
- Bicker, S.; Lackinger, M.; Weiß, K.; Schratt, G. MicroRNA-132, -134, and -138: A microRNA troika rules in neuronal dendrites. Cell. Mol. Life Sci. 2014, 71, 3987–4005. [Google Scholar] [CrossRef]
- Carmel, J.B.; Galante, A.; Soteropoulos, P.; Tolias, P.; Recce, M.; Young, W.; Hart, R. Gene expression profiling of acute spinal cord injury reveals spreading inflammatory signals and neuron loss. Physiol. Genomics. 2001, 7, 201–213. [Google Scholar] [CrossRef]
- Citron, B.A.; Arnold, P.M.; Sebastian, C.; Qin, F.; Malladi, S.; Ameenuddin, S.; Landis, M.E.; Festoff, B.W. Rapid Upregulation of Caspase-3 in Rat Spinal Cord after Injury: mRNA, Protein, and Cellular Localization Correlates with Apoptotic Cell Death. Exp. Neurol. 2000, 166, 213–226. [Google Scholar] [CrossRef]
- Tatro, E.T.; Risbrough, V.; Soontornniyomkij, B.; Young, J.; Shumaker-Armstrong, S.; Jeste, D.V.; Achim, C.L. Short-term recognition memory correlates with regional CNS expression of microRNA-138 in mice. Am. J. Geriatr. Psychiatry 2013, 15, 461–473. [Google Scholar] [CrossRef] [PubMed]
- Uceta-García, M. Therapeutic Potential of miR-135a-5p and miR-138-5p for the Treatment of Spinal Cord Injury: Analsysis of Expression Changes. Master’s Thesis, UCM-HNP. 2020. Available online: https://osf.io/yqn58/ (accessed on 4 December 2020).
- Daswani, R.; Gilardi, C.; Soutschek, M.; Nanda, P.; Weiss, K.; Bicker, S.; Fiore, R.; Dieterich, C.; Germain, P.L.; Winterer, J.; et al. MicroRNA-138 controls hippocampal interneuron function and short-term memory in mice. Elife 2022, 11, e74056. [Google Scholar] [CrossRef] [PubMed]
- Chang, S.H.; Su, Y.C.; Chang, M.; Chen, J.A. MicroRNAs mediate precise control of spinal interneuron populations to exert delicate sensory-to-motor outputs. Elife 2021, 10, e63768. [Google Scholar] [CrossRef]
- Dugas, J.C.; Cuellar, T.L.; Scholze, A.; Ason, B.; Ibrahim, A.; Emery, B.; Zamanian, J.L.; Foo, L.C.; McManus, M.T.; Barres, B.A. Dicer1 and miR-219 Are required for normal oligodendrocyte differentiation and myelination. Neuron 2010, 65, 597–611. [Google Scholar] [CrossRef] [PubMed]
- Wang, W.; Su, Y.; Tang, S.; Li, H.; Xie, W.; Chen, J.; Shen, L.; Pan, X.; Ning, B. Identification of noncoding RNA expression profiles and regulatory interaction networks following traumatic spinal cord injury by sequence analysis. Aging 2019, 11, 2352–2368. [Google Scholar] [CrossRef]
- Tian, F.; Yuan, C.; Yue, H. MiR-138/SIRT1 axis is implicated in impaired learning and memory abilities of cerebral ischemia/reperfusion injured rats. Exp. Cell Res. 2018, 367, 232–240. [Google Scholar] [CrossRef] [PubMed]
- Schober, K.; Ondruschka, B.; Dreßler, J.; Abend, M. Detection of hypoxia, kers in the cerebellum after a traumatic frontal cortex injury: A human postmortem gene expression analysis. Int. J. Leg. Med. 2015, 129, 701–707. [Google Scholar] [CrossRef]
- Lei, P.; Li, Y.; Chen, X.; Yang, S.; Zhang, J. Microarray based analysis of microRNA expression in rat cerebral cortex after traumatic brain injury. Brain Res. 2009, 1284, 191–201. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Tang, C.; Wang, D. LncRNA H19 promotes inflammatory response induced by cerebral ischemia-reperfusion injury through regulating the miR-138-5p-p65 axis. Biochem. Cell Biol. 2020, 98, 525–536. [Google Scholar] [CrossRef]
- Chen, J.; Qin, R. MicroRNA-138-5p regulates the development of spinal cord injury by targeting SIRT1. Mol. Med. Rep. 2020, 22, 328–336. [Google Scholar] [CrossRef]
- Schröder, J.; Ansaloni, S.; Schilling, M.; Liu, T.; Radke, J.; Jaedicke, M.; Schjeide, B.M.; Mashychev, A.; Tegeler, C.; Radbruch, H.; et al. MicroRNA-138 is a potential regulator of memory performance in humans. Front. Hum. Neurosci. 2014, 8, 501. [Google Scholar] [CrossRef] [PubMed]
- Feng, X.; Zhan, F.; Luo, D.; Hu, J.; Wei, G.; Hua, F.; Xu, G. LncRNA 4344 promotes NLRP3-related neuroinflammation and cognitive impairment by targeting miR-138-5p. Brain Behav. Immun. 2021, 98, 283–298. [Google Scholar] [CrossRef] [PubMed]
- Kan, A.A.; van Erp, S.; Derijck, A.A.; de Wit, M.; Hessel, E.V.; O’Duibhir, E.; de Jager, W.; Van Rijen, P.C.; Gosselaar, P.H.; de Graan, P.N.; et al. Genome-wide microRNA profiling of human temporal lobe epilepsy identifies modulators of the immune response. Cell. Mol. Life Sci. 2012, 69, 3127–3145. [Google Scholar] [CrossRef] [PubMed]
- Weiss, K.; Treiber, T.; Meister, G.; Schratt, G. The nuclear matrix protein Matr3 regulates processing of the synaptic microRNA-138-5p. Neurobiol. Learn. Mem. 2019, 159, 36–45. [Google Scholar] [CrossRef] [PubMed]
- Hu, J.Z.; Huang, J.H.; Zeng, L.; Wang, G.; Cao, M.; Lu, H.B. Anti-Apoptotic Effect of MicroRNA-21 after Contusion Spinal Cord Injury in Rats. J. Neurotrauma 2013, 30, 1349–1360. [Google Scholar] [CrossRef]
- Yu, D.S.; Lv, G.; Mei, X.F.; Cao, Y.; Wang, Y.F.; Bi, Y.L. MiR-200c regulates ROS-induced apoptosis in murine BV-2 cells by targeting FAP-1. Spinal Cord. 2015, 53, 182–189. [Google Scholar] [CrossRef]
- McEwen, M.L.; Springer, J.E. A Mapping Study of Caspase-3 Activation Following Acute Spinal Cord Contusion in Rats. J. Histochem. Cytochem. 2005, 53, 809–819. [Google Scholar] [CrossRef]
- Yu, W.R.; Fehlings, M.G. Fas/FasL-mediated apoptosis and inflammation are key features of acute human spinal cord injury: Implications for translational, clinical application. Acta Neuropathol. 2011, 122, 747–761. [Google Scholar] [CrossRef]
- Springer, J.E.; Azbill, R.D.; Knapp, P.E. Activation of the caspase-3 apoptotic cascade in traumatic spinal cord injury. Nat. Med. 1999, 5, 943–946. [Google Scholar] [CrossRef]
- Hudson, R.S.; Yi, M.; Esposito, D.; A Glynn, S.; Starks, A.M.; Yang, Y.; Schetter, A.J.; Watkins, S.K.; A Hurwitz, A.; Dorsey, T.H.; et al. MicroRNA-106b-25 cluster expression is associated with early disease recurrence and targets caspase-7 and focal adhesion in human prostate cancer. Oncogene 2013, 32, 4139–4147. [Google Scholar] [CrossRef]
- Park, J.; Doseff, A.; Schmittgen, T. MicroRNAs Targeting Caspase-3 and -7 in PANC-1 Cells. IJMS 2018, 19, 1206. [Google Scholar] [CrossRef] [PubMed]
- Shi, X.-B.; Xue, L.; Yang, J.; Ma, A.-H.; Zhao, J.; Xu, M.; Tepper, C.G.; Evans, C.P.; Kung, H.-J.; de Vere White, R.W. An androgen-regulated miRNA suppresses Bak1 expression and induces androgen-independent growth of prostate cancer cells. Proc. Natl. Acad. Sci. USA 2007, 104, 19983–19988. [Google Scholar] [CrossRef] [PubMed]
- Deng, Y.; Chen, D.; Gao, F.; Lv, H.; Zhang, G.; Sun, X.; Liu, L.; Mo, D.; Ma, N.; Song, L.; et al. Exosomes derived from microRNA-138-5p-overexpressing bone, row-derived mesenchymal stem cells confer neuroprotection to astrocytes following ischemic stroke via inhibition of LCN2. J. Biol. Eng. 2019, 13, 71. [Google Scholar] [CrossRef] [PubMed]
- Tang, X.J.; Yang, M.H.; Cao, G.; Lu, J.T.; Luo, J.; Dai, L.J.; Huang, K.M.; Zhang, L.I. Protective effect of microRNA-138 against cerebral ischemia/reperfusion injury in rats. Exp. Ther. Med. 2016, 11, 1045–1050. [Google Scholar] [CrossRef]
- Stojcheva, N.; Schechtmann, G.; Sass, S.; Roth, P.; Florea, A.-M.; Stefanski, A.; Stühler, K.; Wolter, M.; Müller, N.; Theis, F.; et al. MicroRNA-138 promotes acquired alkylator resistance in glioblastoma by targeting the Bcl-2-interacting mediator BIM. Oncotarget 2016, 7, 12937–12950. [Google Scholar] [CrossRef]
- Li, S.; Ran, Y.; Zhang, D.; Chen, J.; Li, S.; Zhu, D. microRNA-138 plays a role in hypoxic pulmonary vascular remodelling by targeting Mst1. Biochem. J. 2013, 452, 281–291. [Google Scholar] [CrossRef]
- He, S.; Liu, P.; Jian, Z.; Li, J.; Zhu, Y.; Feng, Z.; Xiao, Y. miR-138 protects cardiomyocytes from hypoxia-induced apoptosis via MLK3/JNK/c-, pathway. Biochem. Biophys. Res. Commun. 2013, 441, 763–769. [Google Scholar] [CrossRef]
- Ren, R.; Chen, S.D.; Fan, J.; Zhang, G.; Li, J.B. miRNA-138 regulates MLK3/JNK/MAPK pathway to protect BV-2 cells from H2O2-induced apoptosis. BLL 2018, 119, 284–288. [Google Scholar] [CrossRef]
- Zheng, Z.; Zhao, B. Astragalus polysaccharide protects hypoxia-induced injury by up-regulation of miR-138 in rat neural stem cells. Biomed. Pharmacoth. 2018, 102, 295–301. [Google Scholar] [CrossRef]
- Zhang, M.; Tao, W.; Yuan, Z.; Liu, Y. Mst-1 deficiency promotes post-traumatic spinal motor neuron survival via enhancement of autophagy flux. J. Neurochem. 2017, 143, 244–256. [Google Scholar] [CrossRef]
- Kole, A.J.; Annis, R.P.; Deshmukh, M. Mature neurons: Equipped for survival. Cell Death Dis. 2013, 4, e689. [Google Scholar] [CrossRef] [PubMed]
- Xu, C.; Fu, H.; Gao, L.; Wang, L.; Wang, W.; Li, J.; Li, Y.; Dou, L.; Gao, X.; Luo, X.; et al. BCR-ABL/GATA1/miR-138 mini circuitry contributes to the leukemogenesis of chronic myeloid leukemia. Oncogene 2014, 33, 44–54. [Google Scholar] [CrossRef] [PubMed]
- Wang, W.; Zhao, L.J.; Tan, Y.X.; Ren, H.; Qi, Z.T. MiR-138 induces cell cycle arrest by targeting cyclin D3 in hepatocellular carcinoma. Carcinogenesis 2012, 33, 1113–1120. [Google Scholar] [CrossRef] [PubMed]
- Qiu, S.; Huang, D.; Yin, D.; Li, F.; Li, X.; Kung, H.F.; Peng, Y. Suppression of tumorigenicity by MicroRNA-138 through inhibition of EZH2-CDK4/6-pRb-E2F1 signal loop in glioblastoma multiforme. BBA Mol. Basis Dis. 2013, 1832, 1697–1707. [Google Scholar] [CrossRef]
- Wu, J.; Sabirzhanov, B.; A Stoica, B.; Lipinski, M.M.; Zhao, Z.; Zhao, S.; Ward, N.; Yang, D.; I Faden, A. Ablation of the transcription factors E2F1-2 limits neuroinflammation and associated neurological deficits after contusive spinal cord injury. Cell Cycle 2015, 14, 3698–3712. [Google Scholar] [CrossRef]
- Yadav, R.; Weng, H.R. EZH2 regulates spinal neuroinflammation in rats with neuropathic pain. Neuroscience 2017, 349, 106–117. [Google Scholar] [CrossRef]
- Ji, Y.; Xiao, F.; Sun, L.; Qin, J.; Shi, S.; Yang, J.; Liu, Y.; Zhou, D.; Zhao, J.; Shen, A. Increased expression of CDK11p58 and cyclin D3 following spinal cord injury in rats. Mol. Cell. Biochem. 2008, 309, 49–60. [Google Scholar] [CrossRef]
- Byrnes, K.R.; Stoica, B.A.; Fricke, S.; Di Giovanni, S.; Faden, A.I. Cell cycle activation contributes to post-mitotic cell death and secondary damage after spinal cord injury. Brain 2007, 130, 2977–2992. [Google Scholar] [CrossRef]
- Tian, S.; Guo, X.; Yu, C.; Sun, C.; Jiang, J. miR-138-5p suppresses autophagy in pancreatic cancer by targeting SIRT1. Oncotarget 2017, 14, 8. [Google Scholar] [CrossRef]
- Muñoz-Galdeano, T.; Reigada, D.; Del Aguila, A.; Velez, I.; Caballero-López, M.J.; Maza, R.M.; Nieto-Díaz, M. Cell Specific Changes of Autophagy in a Mouse Model of Contusive Spinal Cord Injury. Front. Cell. Neurosci. 2018, 12, 164. [Google Scholar] [CrossRef]
- Zhao, X.; Yang, L.; Hu, J.; Ruan, J. miR-138 might reverse multidrug resistance of leukemia cells. Leuk. Res. 2010, 34, 1078–1082. [Google Scholar] [CrossRef] [PubMed]
- Dulin, J.N.; Moore, M.L.; Grill, R.J. The Dual Cyclooxygenase/5-Lipoxygenase Inhibitor Licofelone Attenuates P-Glycoprotein-Mediated Drug Resistance in the Injured Spinal Cord. J. Neurotrauma 2013, 30, 211–226. [Google Scholar] [CrossRef] [PubMed]







Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Maza, R.M.; Barreda-Manso, M.A.; Reigada, D.; Silván, Á.; Muñoz-Galdeano, T.; Soto, A.; del Águila, Á.; Nieto-Díaz, M. MicroRNA-138-5p Targets Pro-Apoptotic Factors and Favors Neural Cell Survival: Analysis in the Injured Spinal Cord. Biomedicines 2022, 10, 1559. https://doi.org/10.3390/biomedicines10071559
Maza RM, Barreda-Manso MA, Reigada D, Silván Á, Muñoz-Galdeano T, Soto A, del Águila Á, Nieto-Díaz M. MicroRNA-138-5p Targets Pro-Apoptotic Factors and Favors Neural Cell Survival: Analysis in the Injured Spinal Cord. Biomedicines. 2022; 10(7):1559. https://doi.org/10.3390/biomedicines10071559
Chicago/Turabian StyleMaza, Rodrigo M., María Asunción Barreda-Manso, David Reigada, Ágata Silván, Teresa Muñoz-Galdeano, Altea Soto, Ángela del Águila, and Manuel Nieto-Díaz. 2022. "MicroRNA-138-5p Targets Pro-Apoptotic Factors and Favors Neural Cell Survival: Analysis in the Injured Spinal Cord" Biomedicines 10, no. 7: 1559. https://doi.org/10.3390/biomedicines10071559





