Pilot Study: Immune Checkpoints Polymorphisms in Greek Primary Breast Cancer Patients
Abstract
:1. Introduction
2. Materials and Methods
2.1. Human Breast Cancer Samples
2.2. DNA Extraction from Fresh-Frozen Samples
2.3. SNP Selection
2.4. Immunofluorescence
2.5. Data Analysis
3. Results
3.1. Clinicopathological and Demographic Parameters of Breast Cancer Patients
3.2. Association of PD-1, PD-L1, and PD-L2 Polymorphisms with Breast Cancer
3.3. Association of PD-1 PD-L1 and PD-L2 Polymorphisms with Cancer Subtypes
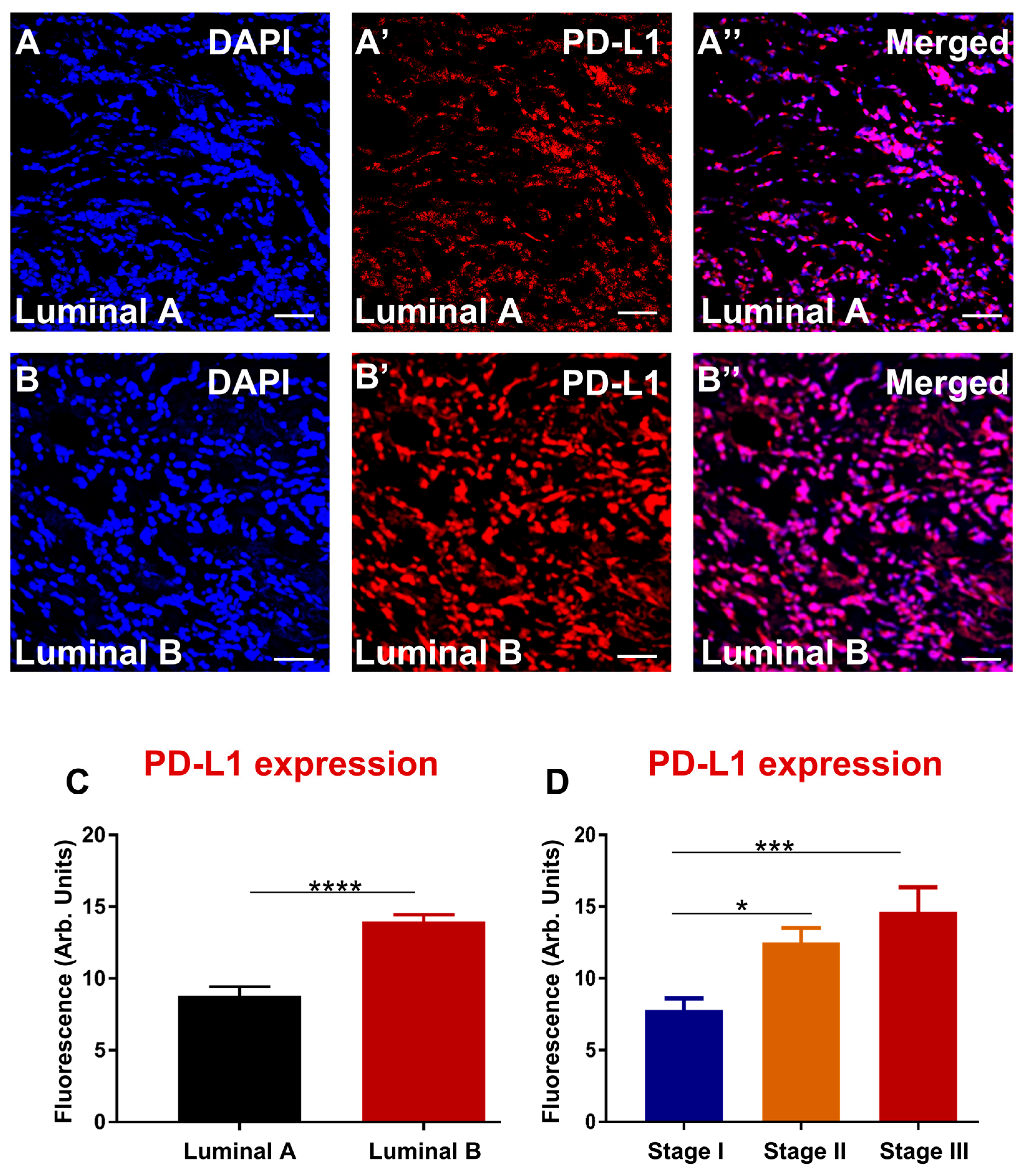
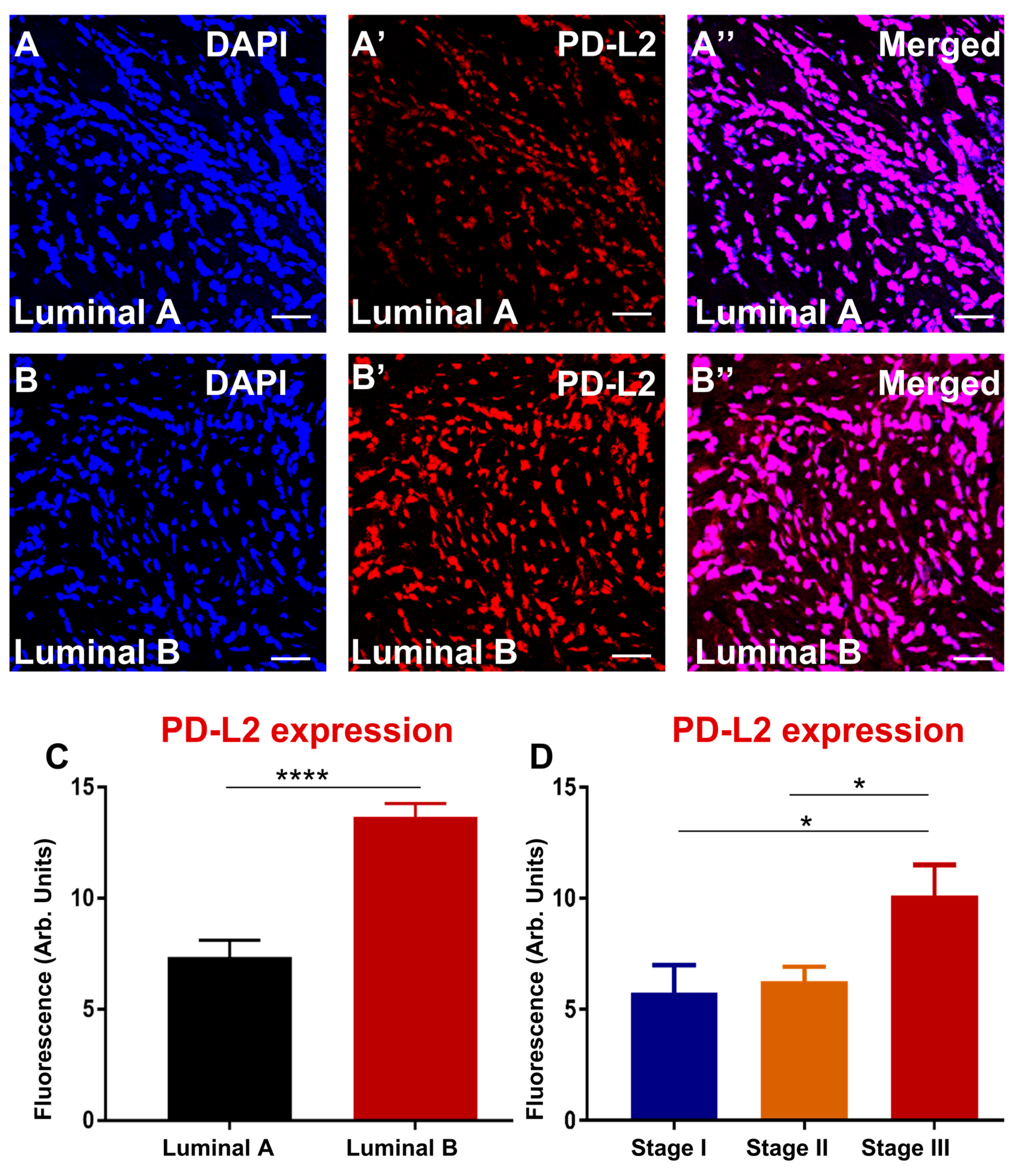
3.4. Enhanced Expression of PD-L1 and PD-L2 Associates with Luminal B and Advanced Stage in Greek Primary Breast Cancer Cohort
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Acknowledgments
Conflicts of Interest
References
- DeSantis, C.E.; Bray, F.; Ferlay, J.; Lortet-Tieulent, J.; Anderson, B.O.; Jemal, A. International variation in female breast cancer incidence and mortality rates. Cancer Epidemiol. Biomark. Prev. 2015, 24, 1495–1506. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ferlay, J.; Soerjomataram, I.; Ervik, M.; Dikshit, R.; Eser, S.; Mathers, C.; Rebelo, M.; Parkin, D.; Forman, D.; Bray, F. GLOBOCAN 2012 v1.0, Cancer Incidence and Mortality Worldwide: IARC CancerBase No. 11. International Agency for Research on Cancer, Lyon. 2013. Available online: https://gco.iarc.fr/ (accessed on 10 October 2014).
- Gonzalez-Angulo, A.M.; Morales-Vasquez, F.; Hortobagyi, G.N. Overview of resistance to systemic therapy in patients with breast cancer. In Breast Cancer Chemosensitivity. Advances in Experimental Medicine and Biology; Springer: New York, NY, USA, 2007; Volume 608, pp. 1–22. [Google Scholar]
- Barriga, V.; Kuol, N.; Nurgali, K.; Apostolopoulos, V. The Complex Interaction between the Tumor Micro-Environment and Immune Checkpoints in Breast Cancer. Cancers 2019, 11, 1205. [Google Scholar] [CrossRef] [Green Version]
- Sorlie, T.; Tibshirani, R.; Parker, J.; Hastie, T.; Marron, J.S.; Nobel, A.; Deng, S.; Johnsen, H.; Pesich, R.; Geisler, S.; et al. Repeated observation of breast tumor subtypes in independent gene expression data sets. Proc. Natl. Acad. Sci. USA 2003, 100, 8418–8423. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Voduc, K.D.; Cheang, M.C.U.; Tyldesley, S.; Gelmon, K.; Nielsen, T.O.; Kennecke, H. Breast Cancer Subtypes and the Risk of Local and Regional Relapse. J. Clin. Oncol. 2010, 28, 1684–1691. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Harbeck, N.; Thomssen, C.; Gnant, M. St. Gallen 2013: Brief preliminary summary of the consensus discussion. Breast Care 2013, 8, 102–109. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sørlie, T.; Perou, C.M.; Tibshirani, R.; Aas, T.; Geisler, S.; Johnsen, H.; Hastie, T.; Eisen, M.B.; van de Rijn, M.; Jeffrey, S.S.; et al. Gene expression patterns of breast carcinomas distinguish tumor subclasses with clinical implications. Proc. Natl. Acad. Sci. USA 2001, 98, 10869–10874. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tran, B.; Bedard, P.L. Luminal-B breast cancer and novel therapeutic targets. Breast Cancer Res. 2011, 13, 221. [Google Scholar] [CrossRef] [Green Version]
- Metzger-Filho, O.; Sun, Z.; Viale, G.; Price, K.N.; Crivellari, D.; Snyder, R.D.; Gelber, R.D.; Castiglione-Gertsch, M.; Coates, A.S.; Goldhirsch, A.; et al. Patterns of Recurrence and outcome according to breast cancer subtypes in lymph node-negative disease: Results from international breast cancer study group trials VIII and IX. J. Clin. Oncol. 2013, 31, 3083–3090. [Google Scholar] [CrossRef] [Green Version]
- Han, Y.; Liu, D.; Li, L. PD-1/PD-L1 pathway: Current research in cancer. Am. J. Cancer Res. 2020, 10, 727–742. [Google Scholar]
- Kuol, N.; Stojanovska, L.; Nurgali, K.; Apostolopoulos, V. PD-1/PD-L1 in disease. Immunotherapy 2018, 10, 149–160. [Google Scholar] [CrossRef]
- Liu, C.; Jiang, J.; Gao, L.; Hu, X.; Wang, F.; Shen, Y.; Yu, G.; Zhao, Z.; Zhang, X. A Promoter Region Polymorphism in PDCD-1 Gene Is Associated with Risk of Rheumatoid Arthritis in the Han Chinese Population of Southeastern China. Int. J. Genom. 2014, 2014, 247637. [Google Scholar]
- Zhang, W.; Song, Y.; Zhang, X. Relationship of Programmed Death-1 (PD-1) and Programmed Death Ligand-1 (PD-L1) Polymorphisms with Overall Cancer Susceptibility: An Updated Meta-Analysis of 28 Studies with 60 612 Subjects. Med. Sci. Monit. 2021, 27, e932146. [Google Scholar] [CrossRef]
- Haghshenas, M.R.; Naeimi, S.; Talei, A.; Ghaderi, A.; Erfani, N. Program death 1 (PD1) haplotyping in patients with breast carcinoma. Mol. Biol. Rep. 2011, 38, 4205–4210. [Google Scholar] [CrossRef]
- Hashemi, M.; Karami, S.; Sarabandi, S.; Moazeni-Roodi, A.; Małecki, A.; Ghavami, S.; Wiechec, E. Association between PD-1 and PD-L1 Polymorphisms and the Risk of Cancer: A Meta-Analysis of Case-Control Studies. Cancers 2019, 11, 1150. [Google Scholar] [CrossRef] [Green Version]
- Zhang, J.; Zhao, T.; Xu, C.; Huang, J.; Yu, H. The association between polymorphisms in the PDCD1 gene and the risk of cancer: A PRISMA-compliant meta-analysis. Medicine 2016, 95, e4423. [Google Scholar] [CrossRef]
- Dong, W.; Gong, M.; Shi, Z.; Xiao, J.; Zhang, J.; Peng, J. Programmed Cell Death-1 Polymorphisms Decrease the Cancer Risk: A Meta-Analysis Involving Twelve Case-Control Studies. PLoS ONE 2016, 11, e0152448. [Google Scholar] [CrossRef] [Green Version]
- Karami, S.; Sattarifard, H.; Kiumarsi, M.; Sarabandi, S.; Taheri, M.; Hashemi, M.; Bahari, G.; Ghavami, S. Evaluating the Possible Association between PD-1 (Rs11568821, Rs2227981, Rs2227982) and PD-L1 (Rs4143815, Rs2890658) Polymorphisms and Susceptibility to Breast Cancer in a Sample of Southeast Iranian Women. Asian Pac. J. Cancer Prev. 2020, 21, 3115–3123. [Google Scholar] [CrossRef]
- Mamat, U.; Arkinjan, M. Association of programmed death-1 gene polymorphism rs2227981 with tumor: Evidence from a meta analysis. Int. J. Clin. Exp. Med. 2015, 8, 13282–13288. [Google Scholar]
- Kuol, N.; Stojanovska, L.; Nurgali, K.; Apostolopoulos, V. The mechanisms tumor cells utilize to evade the host’s immune system. Maturitas 2017, 105, 8–15. [Google Scholar] [CrossRef]
- Qin, W.; Hu, L.; Zhang, X.; Jiang, S.; Li, J.; Zhang, Z.; Wang, X. The Diverse Function of PD-1/PD-L Pathway Beyond Cancer. Front. Immunol. 2019, 10, 2298. [Google Scholar] [CrossRef]
- Gao, L.; Guo, Q.; Li, X.; Yang, X.; Ni, H.; Wang, T.; Zhao, Q.; Liu, H.; Xing, Y.; Xi, T.; et al. MiR-873/PD-L1 axis regulates the stemness of breast cancer cells. EBioMedicine 2019, 41, 395–407. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ashizawa, M.; Okayama, H.; Ishigame, T.; Thar Min, A.K.; Saito, K.; Ujiie, D.; Murakami, Y.; Kikuchi, T.; Nakayama, Y.; Noda, M.; et al. miRNA-148a-3p Regulates Immunosuppression in DNA Mismatch Repair–Deficient Colorectal Cancer by Targeting PD-L1. Mol. Cancer Res. 2019, 17, 1403–1413. [Google Scholar] [CrossRef] [Green Version]
- Kataoka, K.; Shiraishi, Y.; Takeda, Y.; Sakata, S.; Matsumoto, M.; Nagano, S.; Maeda, T.; Nagata, Y.; Kitanaka, A.; Mizuno, S.; et al. Aberrant PD-L1 expression through 3′-UTR disruption in multiple cancers. Nature 2016, 534, 402–406. [Google Scholar] [CrossRef] [PubMed]
- Mezzadra, R.; Sun, C.; Jae, L.T.; Gomez-Eerland, R.; de Vries, E.; Wu, W.; Logtenberg, M.E.W.; Slagter, M.; Rozeman, E.A.; Hofland, I.; et al. Identification of CMTM6 and CMTM4 as PD-L1 protein regulators. Nature 2017, 549, 106–110. [Google Scholar] [CrossRef]
- Zhao, X.; Peng, Y.; Li, X.; Zhang, S.; Peng, Y.; Shan, J.; Li, M.; Dai, N.; Feng, Y.; Xu, C.; et al. The association of PD-L1 gene polymorphisms with non-small-cell lung cancer susceptibility and clinical outcomes in a Chinese population. Int. J. Clin. Exp. Pathol. 2020, 13, 2130–2136. [Google Scholar] [PubMed]
- Yang, H.; Zhou, X.; Sun, L.; Mao, Y. Correlation Between PD-L2 Expression and Clinical Outcome in Solid Cancer Patients: A Meta-Analysis. Front. Oncol. 2019, 9, 47. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wu, Y.; Zhao, T.; Jia, Z.; Cao, D.; Cao, X.; Pan, Y.; Zhao, D.; Zhang, B.; Jiang, J. Polymorphism of the programmed death-ligand 1 gene is associated with its protein expression and prognosis in gastric cancer. J. Gastroenterol. Hepatol. 2019, 34, 1201–1207. [Google Scholar] [CrossRef] [PubMed]
- Mikkelsen, K.; Prakash, M.D.; Kuol, N.; Nurgali, K.; Stojanovska, L.; Apostolopoulos, V. Anti-Tumor Effects of Vitamin B2, B6 and B9 in Promonocytic Lymphoma Cells. Int. J. Mol. Sci. 2019, 20, 3763. [Google Scholar] [CrossRef] [Green Version]
- He, H.X.; Gao, Y.; Fu, J.C.; Zhou, Q.H.; Wang, X.X.; Bai, B.; Li, P.F.; Huang, C.; Rong, Q.X.; Ping, L.Q.; et al. VISTA and PD-L1 synergistically predict poor prognosis in patients with extranodal natural killer/T-cell lymphoma. Oncoimmunology 2021, 10, 1907059. [Google Scholar] [CrossRef]
- Pasternack, H.; Kuempers, C.; Deng, M.; Watermann, I.; Olchers, T.; Kuehnel, M.; Jonigk, D.; Kugler, C.; Stellmacher, F.; Goldmann, T.; et al. Identification of molecular signatures associated with early relapse after complete resection of lung adenocarcinomas. Sci. Rep. 2021, 11, 9532. [Google Scholar] [CrossRef]
- Guo, P.D.; Sun, Z.W.; Lai, H.J.; Yang, J.; Wu, P.P.; Guo, Y.D.; Sun, J. Clinicopathological analysis of PD-L2 expression in colorectal cancer. Onco. Targets Ther. 2018, 11, 7635–7642. [Google Scholar] [CrossRef] [Green Version]
- Stavely, R.; Robinson, A.M.; Miller, S.; Boyd, R.; Sakkal, S.; Nurgali, K. Human adult stem cells derived from adipose tissue and bone marrow attenuate enteric neuropathy in the guinea-pig model of acute colitis. Stem Cell Res. Ther. 2015, 6, 1–21. [Google Scholar] [CrossRef] [Green Version]
- Brion, C.; Lutz, S.M.; Albert, F.W. Simultaneous quantification of mRNA and protein in single cells reveals post-transcriptional effects of genetic variation. eLife 2020, 9, e60645. [Google Scholar] [CrossRef]
- Baghban, R.; Roshangar, L.; Jahanban-Esfahlan, R.; Seidi, K.; Ebrahimi-Kalan, A.; Jaymand, M.; Kolahian, S.; Javaheri, T.; Zare, P. Tumor microenvironment complexity and therapeutic implications at a glance. Cell Commun. Signal. 2020, 18, 59. [Google Scholar] [CrossRef] [Green Version]
- Whiteside, T.L. The tumor microenvironment and its role in promoting tumor growth. Oncogene 2008, 27, 5904–5912. [Google Scholar] [CrossRef] [Green Version]
- Topalian, S.L.; Hodi, F.S.; Brahmer, J.R.; Gettinger, S.N.; Smith, D.C.; McDermott, D.F.; Powderly, J.D.; Carvajal, R.D.; Sosman, J.A.; Atkins, M.B.; et al. Safety, activity, and immune correlates of anti-PD-1 antibody in cancer. N. Engl. J. Med. 2012, 366, 2443–2454. [Google Scholar] [CrossRef]
- Pardoll, D.M. The blockade of immune checkpoints in cancer immunotherapy. Nat. Rev. Cancer 2012, 12, 252–264. [Google Scholar] [CrossRef] [Green Version]
- Bates, J.P.; Derakhshandeh, R.; Jones, L.; Webb, T.J. Mechanisms of immune evasion in breast cancer. BMC Cancer 2018, 18, 556. [Google Scholar] [CrossRef] [Green Version]
- Yang, Q.; Liu, Y.; Liu, D.; Zhang, Y.; Mu, K. Association of polymorphisms in the programmed cell death 1 (PD-1) and PD-1 ligand genes with ankylosing spondylitis in a Chinese population. Clin. Exp. Rheumatol. 2011, 29, 13–18. [Google Scholar]
- Nomizo, T.; Ozasa, H.; Tsuji, T.; Funazo, T.; Yasuda, Y.; Yoshida, H.; Yagi, Y.; Sakamori, Y.; Nagai, H.; Hirai, T.; et al. Clinical Impact of Single Nucleotide Polymorphism in PD-L1 on Response to Nivolumab for Advanced Non-Small-Cell Lung Cancer Patients. Sci. Rep. 2017, 7, 45124. [Google Scholar] [CrossRef]
- Ge, J.; Zhu, L.; Zhou, J.; Li, G.; Li, Y.; Li, S.; Wu, Z.; Rong, J.; Yuan, H.; Liu, Y.; et al. Association between co-inhibitory molecule gene tagging single nucleotide polymorphisms and the risk of colorectal cancer in Chinese. J. Cancer Res. Clin. Oncol. 2015, 141, 1533–1544. [Google Scholar] [CrossRef] [PubMed]
- Hua, Z.; Li, D.; Xiang, G.; Xu, F.; Jie, G.; Fu, Z.; Jie, Z.; Da, P.; Li, D. PD-1 polymorphisms are associated with sporadic breast cancer in Chinese Han population of Northeast China. Breast Cancer Res. Treat. 2011, 129, 195–201. [Google Scholar] [CrossRef] [PubMed]
- Ren, H.T.; Li, Y.M.; Wang, X.J.; Kang, H.F.; Jin, T.B.; Ma, X.B.; Liu, X.H.; Wang, M.; Liu, K.; Xu, P.; et al. PD-1 rs2227982 Polymorphism Is Associated With the Decreased Risk of Breast Cancer in Northwest Chinese Women: A Hospital-Based Observational Study. Medicine 2016, 95, e3760. [Google Scholar] [CrossRef] [PubMed]
- Dong, H.; Strome, S.E.; Salomao, D.R.; Tamura, H.; Hirano, F.; Flies, D.B.; Roche, P.C.; Lu, J.; Zhu, G.; Tamada, K.; et al. Tumor-associated B7-H1 promotes T-cell apoptosis: A potential mechanism of immune evasion. Nat. Med. 2002, 8, 793–800. [Google Scholar] [CrossRef]
- Kim, H.R.; Ha, S.-J.; Hong, M.H.; Heo, S.J.; Koh, Y.W.; Choi, E.C.; Kim, E.K.; Pyo, K.H.; Jung, I.; Seo, D.; et al. PD-L1 expression on immune cells, but not on tumor cells, is a favorable prognostic factor for head and neck cancer patients. Sci. Rep. 2016, 6, 36956. [Google Scholar] [CrossRef] [Green Version]
- Wolchok, J.D.; Kluger, H.; Callahan, M.K.; Postow, M.A.; Rizvi, N.A.; Lesokhin, A.M.; Segal, N.H.; Ariyan, C.E.; Gordon, R.A.; Reed, K.; et al. Nivolumab plus ipilimumab in advanced melanoma. N. Engl. J. Med. 2013, 369, 122–133. [Google Scholar] [CrossRef] [Green Version]
- Reck, M.; Rodriguez-Abreu, D.; Robinson, A.G.; Hui, R.; Csoszi, T.; Fulop, A.; Gottfried, M.; Peled, N.; Tafreshi, A.; Cuffe, S.; et al. Pembrolizumab versus Chemotherapy for PD-L1-Positive Non-Small-Cell Lung Cancer. N. Engl. J. Med. 2016, 375, 1823–1833. [Google Scholar] [CrossRef] [Green Version]
- Antonia, S.J.; Villegas, A.; Daniel, D.; Vicente, D.; Murakami, S.; Hui, R.; Yokoi, T.; Chiappori, A.; Lee, K.H.; de Wit, M.; et al. Durvalumab after Chemoradiotherapy in Stage III Non-Small-Cell Lung Cancer. N. Engl. J. Med. 2017, 377, 1919–1929. [Google Scholar] [CrossRef] [Green Version]
- Wimberly, H.; Brown, J.R.; Schalper, K.; Haack, H.; Silver, M.R.; Nixon, C.; Bossuyt, V.; Pusztai, L.; Lannin, D.R.; Rimm, D.L. PD-L1 Expression Correlates with Tumor-Infiltrating Lymphocytes and Response to Neoadjuvant Chemotherapy in Breast Cancer. Cancer Immunol. Res. 2015, 3, 326. [Google Scholar] [CrossRef] [Green Version]
- Ghebeh, H.; Barhoush, E.; Tulbah, A.; Elkum, N.; Al-Tweigeri, T.; Dermime, S. FOXP3+ Tregs and B7-H1+/PD-1+ T lymphocytes co-infiltrate the tumor tissues of high-risk breast cancer patients: Implication for immunotherapy. BMC Cancer 2008, 8, 57. [Google Scholar] [CrossRef] [Green Version]
- Li, F.; Ren, Y.; Wang, Z. Programmed death 1 Ligand 1 expression in breast cancer and its association with patients’ clinical parameters. J. Cancer Res. Ther. 2018, 14, 150–154. [Google Scholar]
- Jung, H.I.; Jeong, D.; Ji, S.; Ahn, T.S.; Bae, S.H.; Chin, S.; Chung, J.C.; Kim, H.C.; Lee, M.S.; Baek, M.J. Overexpression of PD-L1 and PD-L2 Is Associated with Poor Prognosis in Patients with Hepatocellular Carcinoma. Cancer Res. Treat. 2017, 49, 246–254. [Google Scholar] [CrossRef] [Green Version]
- Wang, H.; Yao, H.; Li, C.; Liang, L.; Zhang, Y.; Shi, H.; Zhou, C.; Chen, Y.; Fang, J.Y.; Xu, J. PD-L2 expression in colorectal cancer: Independent prognostic effect and targetability by deglycosylation. Oncoimmunology 2017, 6, e1327494. [Google Scholar] [CrossRef] [Green Version]
- Shin, S.J.; Jeon, Y.K.; Kim, P.J.; Cho, Y.M.; Koh, J.; Chung, D.H.; Go, H. Clinicopathologic Analysis of PD-L1 and PD-L2 Expression in Renal Cell Carcinoma: Association with Oncogenic Proteins Status. Ann. Surg. Oncol. 2016, 23, 694–702. [Google Scholar] [CrossRef]
| Parameters | No. of Cases | Percentage (%) |
|---|---|---|
| Total | 123 | 100 |
| Tumour size | ||
| <2 | 74 | 60.2 |
| >2 | 49 | 39.8 |
| Age | ||
| <65 | 75 | 61 |
| >65 | 48 | 39 |
| Stage | ||
| 0 | 3 | 2.4 |
| I | 59 | 48 |
| II | 33 | 26.8 |
| III | 19 | 15.4 |
| Missing | 9 | 7.3 |
| Subtype | ||
| Luminal A | 81 | 65.9 |
| Luminal B | 42 | 34.1 |
| rs2227981 G>A | rs7421861 A>G | rs11568821 C>T | |
|---|---|---|---|
| Chromosome | 2 | 2 | 2 |
| Position | 241851121 | 241853198 | 241851760 |
| Region | 2KB Upstream | Intron | Intron |
| MAF in Europeans | 0.4013 | 0.3517 | 0.1086 |
| p-Value for HWE | 0.20 | 0.88 | 0.52 |
| APR | 92.75% | 96.38% | 98.70% |
| rs10815225 G>C | rs2282055 T>G | rs1009759 A>G | rs6476985 C>T | |
|---|---|---|---|---|
| Chromosome | 9 | 9 | 9 | 9 |
| Position | 5450497 | 5455732 | 5556786 | 5517559 |
| Region | 2KB Upstream | Intron | Intron | Intron |
| MAF in Europeans | 0.1347 | 0.2605 | 0.3101 | 0.1126 |
| p-Value for HWE | 0.07 | 0.82 | 0.99 | 0.86 |
| APR | 98.55% | 98.55% | 94.38% | 98.70% |
| Genes | Alleles | Breast Cancer | European Population | OR Caner to Population (95% CI) | p | ||
|---|---|---|---|---|---|---|---|
| n | % | n | % | ||||
| PD-1 | rs2227981 G>A | ||||||
| G | 142 | 55.47% | 5600 | 59.87% | 1.20 (0.93–1.54) | 0.175 | |
| A | 114 | 44.53% | 3754 | 40.13% | |||
| rs7421861 A>G | |||||||
| A | 184 | 69.17% | 61,022 | 64.83% | 1.22 (0.94–1.58) | 0.157 | |
| G | 82 | 30.83% | 33,102 | 35.17 | |||
| rs11568821 C>T | |||||||
| C | 247 | 90.15% | 10,322 | 89.14% | 1.12 (0.75–1.66) | 0.694 | |
| T | 27 | 9.85% | 1258 | 10.86% | |||
| PD-L1 | rs10815225 G>C | ||||||
| G | 230 | 84.56% | 1793 | 86.53% | 1.17 (0.82–1.67) | 0.398 | |
| C | 42 | 15.44% | 279 | 13.47% | |||
| rs2282055 T>G | |||||||
| T | 204 | 75.00% | 28,314 | 73.95% | 0.95 (0.72–1.25) | 0.729 | |
| G | 68 | 25.00% | 9972 | 26.05% | |||
| PD-L2 | rs1009759 A>G | ||||||
| A | 165 | 62.98% | 50,016 | 68.99% | 0.77 (0.60–0.98) | 0.038 * | |
| G | 97 | 37.02% | 22,478 | 31.01% | |||
| rs6476985 C>T | |||||||
| C | 239 | 87.23% | 34,559 | 88.74% | 0.87 (0.61–1.25) | 0.442 | |
| T | 35 | 12.77% | 4383 | 11.26% | |||
| Genes | Alleles | Luminal B | Luminal A | OR Luminal B to A (95% CI) | p | ||
|---|---|---|---|---|---|---|---|
| n | % | n | % | ||||
| PD-1 | rs2227981 G>A | ||||||
| G | 48 | 63.16% | 76 | 50.67% | 1.67 (0.95–2.93) | 0.090 | |
| A | 28 | 36.84% | 74 | 49.33% | |||
| rs7421861 A>G | |||||||
| A | 51 | 63.75% | 114 | 72.15% | 0.68 (0. 9–1.20) | 0.234 | |
| G | 29 | 36.25% | 44 | 27.85% | |||
| rs11568821 C>T | |||||||
| C | 71 | 86.59% | 149 | 91.98% | 0.56 (0.24–1.26) | 0.254 | |
| T | 11 | 13.41% | 13 | 8.02% | |||
| PD-L1 | rs10815225 G>C | ||||||
| G | 70 | 87.50% | 135 | 83.33% | 1.40 (0.64–2.94) | 0.452 | |
| C | 10 | 12.50% | 25 | 16.67% | |||
| rs2282055 T>G | |||||||
| T | 66 | 82.50% | 111 | 68.52% | 2.17 (1.10–4.19) | 0.022 * | |
| G | 14 | 17.50% | 51 | 31.48% | |||
| PD-L2 | rs1009759 A>G | ||||||
| A | 54 | 67.50% | 92 | 59.74% | 1.40 (0.79–2.49) | 0.259 | |
| G | 26 | 32.50% | 62 | 40.26% | |||
| rs6476985 C>T | |||||||
| C | 74 | 90.24% | 140 | 86.42% | 1.45 (0.62–3.24) | 0.536 | |
| T | 8 | 9.76% | 22 | 13.58% | |||
| Genes | Genotype | Luminal B | Luminal A | p | ||
|---|---|---|---|---|---|---|
| n | % | n | % | |||
| PD-1 | rs2227981 G>A | |||||
| GG | 17 | 44.74% | 21 | 28.00% | 0.197 | |
| GA | 14 | 36.84% | 34 | 45.33% | ||
| AA | 7 | 18.42% | 20 | 26.67% | ||
| rs7421861 A>G | ||||||
| AA | 18 | 45.00% | 39 | 49.37% | 0.084 | |
| AG | 15 | 37.50% | 36 | 45.57% | ||
| GG | 7 | 17.50% | 4 | 5.06% | ||
| rs11568821 C>T | ||||||
| CC | 32 | 78.05% | 68 | 83.95% | NV | |
| CT | 7 | 17.07% | 13 | 16.05% | ||
| TT | 2 | 4.88% | 0 | 0 | ||
| PD-L1 | rs10815225 G>C | |||||
| GG | 31 | 77.50% | 58 | 71.60% | NV | |
| GC | 8 | 20.00% | 19 | 23.46% | ||
| CC | 1 | 2.50% | 4 | 4.94% | ||
| rs2282055 T>G | ||||||
| TT | 26 | 65.00% | 38 | 46.91% | 0.049 * | |
| TG | 14 | 35.00% | 35 | 43.21% | ||
| GG | 0 | 0 | 8 | 9.88% | ||
| PD-L2 | rs1009759 A>G | |||||
| AA | 19 | 47.50% | 28 | 36.37% | 0.494 | |
| AG | 16 | 40.00% | 36 | 46.75% | ||
| GG | 5 | 12.50% | 13 | 16.88% | ||
| rs6476985 C>T | ||||||
| CC | 33 | 80.49% | 61 | 75.31% | 0.549 | |
| CT | 8 | 19.51% | 18 | 22.22% | ||
| TT | 0 | 0 | 2 | 2.47% | ||
| Genes | Genotype | Luminal B | Luminal A | OR Luminal B to A (95% CI) | p | ||
|---|---|---|---|---|---|---|---|
| n | n | ||||||
| PD-1 | rs2227981 G>A | ||||||
| GG vs. AA | 17 | 7 | 21 | 20 | 2.31 (0.76–6.28) | 0.192 | |
| GG+GA vs. AA | 31 | 7 | 55 | 20 | 1.61 (0.60–4.07) | 0.362 | |
| GG vs. GA+AA | 17 | 21 | 21 | 54 | 2.08 (0.95–4.56) | 0.093 | |
| rs7421861 A>G | |||||||
| AA vs. GG | 18 | 7 | 39 | 4 | 0.26 (0.08–1.04) | 0.084 | |
| AA+AG vs. GG | 33 | 7 | 75 | 4 | 0.25 (0.08–0.89) | 0.042 * | |
| AA vs. AG +GG | 18 | 22 | 39 | 40 | 0.84 (0.38–1.79) | 0.700 | |
| rs11568821 C>T | |||||||
| CC vs. TT | 32 | 2 | 68 | 0 | 0 | 0.109 | |
| CC+CT vs. TT | 39 | 2 | 81 | 0 | 0 | 0.111 | |
| CC vs. CT+TT | 32 | 9 | 68 | 13 | 0.68 (0.27–1.66) | 0.460 | |
| PD-L1 | rs10815225 G>C | ||||||
| GG vs. CC | 31 | 1 | 58 | 4 | 2.14 (0.33–26.91) | 0.658 | |
| GG+GC vs. CC | 39 | 1 | 77 | 4 | 2.03 (0.32–25.36) | >0.999 | |
| GG vs. GC+CC | 31 | 9 | 58 | 23 | 1.37 (0.58–3.20) | 0.521 | |
| rs2282055 T>G | |||||||
| TT vs. GG | 26 | 0 | 38 | 8 | Infinity | 0.044 * | |
| TT+TG vs. GG | 40 | 0 | 73 | 8 | Infinity | 0.051 | |
| TT vs. TG+GG | 26 | 14 | 38 | 43 | 2.10 (0.98–4.53) | 0.081 | |
| PD-L2 | rs1009759 A>G | ||||||
| AA vs. GG | 19 | 5 | 28 | 13 | 1.76 (0.55–5.17) | 0.401 | |
| AA+AG vs. GG | 35 | 5 | 64 | 13 | 1.42 (0.46–3.84) | 0.600 | |
| AA vs. AG+GG | 19 | 21 | 28 | 49 | 1.58 (0.72–3.50) | 0.320 | |
| rs6476985 C>T | |||||||
| CC vs. TT | 33 | 0 | 61 | 2 | Infinity | 0.544 | |
| CC+CT vs. TT | 41 | 0 | 79 | 2 | Infinity | 0.550 | |
| CC vs. CT+TT | 33 | 8 | 61 | 20 | 1.35 (0.54–3.35) | 0.650 | |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kuol, N.; Yan, X.; Barriga, V.; Karakkat, J.; Vassilaros, S.; Fyssas, I.; Tsimpanis, A.; Fraser, S.; Nurgali, K.; Apostolopoulos, V. Pilot Study: Immune Checkpoints Polymorphisms in Greek Primary Breast Cancer Patients. Biomedicines 2022, 10, 1827. https://doi.org/10.3390/biomedicines10081827
Kuol N, Yan X, Barriga V, Karakkat J, Vassilaros S, Fyssas I, Tsimpanis A, Fraser S, Nurgali K, Apostolopoulos V. Pilot Study: Immune Checkpoints Polymorphisms in Greek Primary Breast Cancer Patients. Biomedicines. 2022; 10(8):1827. https://doi.org/10.3390/biomedicines10081827
Chicago/Turabian StyleKuol, Nyanbol, Xu Yan, Vanessa Barriga, Jimsheena Karakkat, Stamatis Vassilaros, Ioannis Fyssas, Anastasios Tsimpanis, Sarah Fraser, Kulmira Nurgali, and Vasso Apostolopoulos. 2022. "Pilot Study: Immune Checkpoints Polymorphisms in Greek Primary Breast Cancer Patients" Biomedicines 10, no. 8: 1827. https://doi.org/10.3390/biomedicines10081827
APA StyleKuol, N., Yan, X., Barriga, V., Karakkat, J., Vassilaros, S., Fyssas, I., Tsimpanis, A., Fraser, S., Nurgali, K., & Apostolopoulos, V. (2022). Pilot Study: Immune Checkpoints Polymorphisms in Greek Primary Breast Cancer Patients. Biomedicines, 10(8), 1827. https://doi.org/10.3390/biomedicines10081827









