The HFE p.H63D (p.His63Asp) Polymorphism Is a Modifier of ALS Outcome in Italian and French Patients with SOD1 Mutations
Abstract
:1. Introduction
2. Materials and Methods
2.1. Patients
2.2. Genotyping
2.3. Statistical Analysis
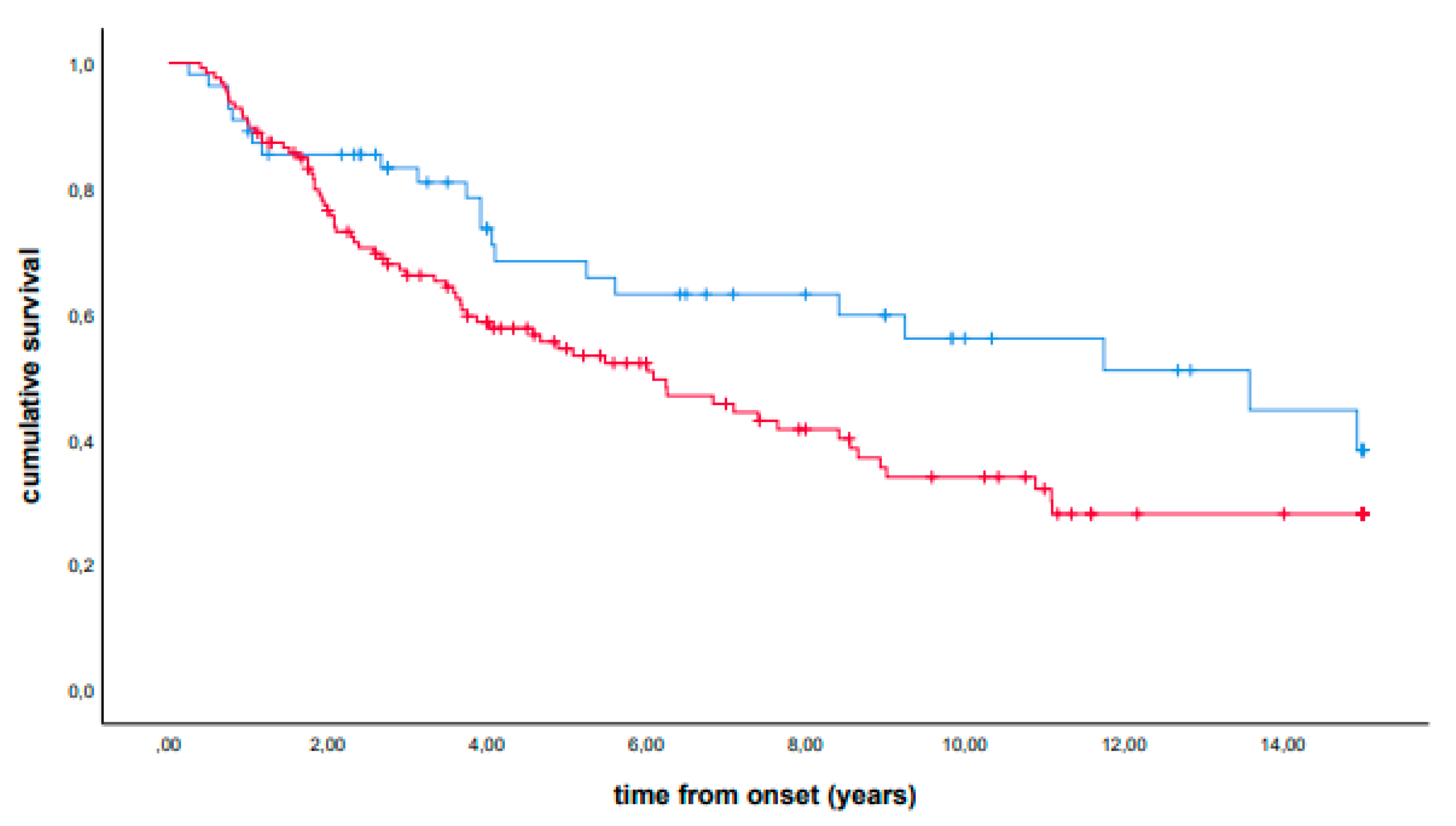
3. Results
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Renton, A.E.; Chiò, A.; Traynor, B.J. State of play in amyotrophic lateral sclerosis genetics. Nat. Neurosci. 2014, 17, 17–23. [Google Scholar] [CrossRef]
- Diekstra, F.P.; van Vught, P.W.J.; van Rheenen, W.; Koppers, M.; Pasterkamp, R.J.; van Es, M.A.; Schelhaas, H.J.; de Visser, M.; Robberecht, W.; Van Damme, P.; et al. UNC13A is a modifier of survival in amyotrophic lateral sclerosis. Neurobiol. Aging 2012, 33, 630.e3–630.e8. [Google Scholar] [CrossRef]
- Chiò, A.; Mora, G.; Restagno, G.; Brunetti, M.; Ossola, I.; Barberis, M.; Ferrucci, L.; Canosa, A.; Manera, U.; Moglia, C.; et al. UNC13A influences survival in Italian amyotrophic lateral sclerosis patients: A population-based study. Neurobiol. Aging 2013, 34, 357.e1–357.e5. [Google Scholar] [CrossRef] [Green Version]
- Blauw, H.M.; van Rheenen, W.; Koppers, M.; Van Damme, P.; Waibel, S.; Lemmens, R.; van Vught, P.W.J.; Meyer, T.; Schulte, C.; Gasser, T.; et al. NIPA1 polyalanine repeat expansions are associated with amyotrophic lateral sclerosis. Hum. Mol. Genet. 2012, 21, 2497–2502. [Google Scholar] [CrossRef] [Green Version]
- Chiò, A.; Calvo, A.; Moglia, C.; Canosa, A.; Brunetti, M.; Barberis, M.; Restagno, G.; Conte, A.; Bisogni, G.; Marangi, G.; et al. ATXN2 polyQ intermediate repeats are a modifier of ALS survival. Neurology 2015, 84, 251–258. [Google Scholar] [CrossRef] [PubMed]
- Nandar, W.; Connor, J.R. HFE gene variants affect iron in the brain. J. Nutr. 2011, 141, 729S–739S. [Google Scholar] [CrossRef] [Green Version]
- van Rheenen, W.; Diekstra, F.P.; van Doormaal, P.T.C.; Seelen, M.; Kenna, K.; McLaughlin, R.; Shatunov, A.; Czell, D.; van Es, M.A.; van Vught, P.W.J.; et al. H63D polymorphism in HFE is not associated with amyotrophic lateral sclerosis. Neurobiol. Aging 2013, 34, 1517.e5–1517.e7. [Google Scholar] [CrossRef] [PubMed]
- Li, M.; Wang, L.; Wang, W.; Qi, X.L.; Tang, Z.Y.; Li, M.; Wang, L.; Wang, W.; Qi, X.L.; Tang, Z.Y. Mutations in the HFE gene and sporadic amyotrophic lateral sclerosis risk: A meta-analysis of observational studies. Braz. J. Med. Biol. Res. 2014, 47, 215–222. [Google Scholar] [CrossRef] [Green Version]
- Chiò, A.; Mora, G.; Sabatelli, M.; Caponnetto, C.; Lunetta, C.; Traynor, B.J.; Johnson, J.O.; Nalls, M.A.; Calvo, A.; Moglia, C.; et al. HFE p.H63D polymorphism does not influence ALS phenotype and survival. Neurobiol. Aging 2015, 36, 2906.e7–2906.e11. [Google Scholar] [CrossRef] [Green Version]
- Chiò, A.; Calvo, A.; Mazzini, L.; Cantello, R.; Mora, G.; Moglia, C.; Corrado, L.; D’Alfonso, S.; Majounie, E.; Renton, A.; et al. Extensive genetics of ALS: A population-based study in Italy. Neurology 2012, 79, 1983–1989. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wang, X.-S.; Lee, S.; Simmons, Z.; Boyer, P.; Scott, K.; Liu, W.; Connor, J. Increased incidence of the Hfe mutation in amyotrophic lateral sclerosis and related cellular consequences. J. Neurol. Sci. 2004, 227, 27–33. [Google Scholar] [CrossRef]
- Goodall, E.F.; Greenway, M.J.; van Marion, I.; Carroll, C.B.; Hardiman, O.; Morrison, K.E. Association of the H63D polymorphism in the hemochromatosis gene with sporadic ALS. Neurology 2005, 65, 934–937. [Google Scholar] [CrossRef] [PubMed]
- Restagno, G.; Lombardo, F.; Ghiglione, P.; Calvo, A.; Cocco, E.; Sbaiz, L.; Mutani, R.; Chiò, A. HFE H63D polymorphism is increased in patients with amyotrophic lateral sclerosis of Italian origin. J. Neurol. Neurosurg. Psychiatry 2007, 78, 327. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- He, X.; Lu, X.; Hu, J.; Xi, J.; Zhou, D.; Shang, H.; Liu, L.; Zhou, H.; Yan, B.; Yu, L.; et al. H63D polymorphism in the hemochromatosis gene is associated with sporadic amyotrophic lateral sclerosis in China. Eur. J. Neurol. 2011, 18, 359–361. [Google Scholar] [CrossRef] [PubMed]
- Sutedja, N.A.; Sinke, R.J.; Van Vught, P.W.J.; Van der Linden, M.W.; Wokke, J.H.J.; Van Duijn, C.M.; Njajou, O.T.; Van der Schouw, Y.T.; Veldink, J.H.; Van den Berg, L.H. The association between H63D mutations in HFE and amyotrophic lateral sclerosis in a Dutch population. Arch. Neurol. 2007, 64, 63–67. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yen, A.A.; Simpson, E.P.; Henkel, J.S.; Beers, D.R.; Appel, S.H. HFE mutations are not strongly associated with sporadic ALS. Neurology 2004, 62, 1611–1612. [Google Scholar] [CrossRef]
- Praline, J.; Blasco, H.; Vourc’h, P.; Rat, V.; Gendrot, C.; Camu, W.; Andres, C.R. French ALS Study Group Study of the HFE gene common polymorphisms in French patients with sporadic amyotrophic lateral sclerosis. J. Neurol. Sci. 2012, 317, 58–61. [Google Scholar] [CrossRef] [PubMed]
- Su, X.W.; Lee, S.Y.; Mitchell, R.M.; Stephens, H.E.; Simmons, Z.; Connor, J.R. H63D HFE polymorphisms are associated with increased disease duration and decreased muscle superoxide dismutase-1 expression in amyotrophic lateral sclerosis patients. Muscle Nerve 2013, 48, 242–246. [Google Scholar] [CrossRef]
- Nandar, W.; Neely, E.B.; Simmons, Z.; Connor, J.R. H63D HFE genotype accelerates disease progression in animal models of amyotrophic lateral sclerosis. Biochim. Biophys. Acta 2014, 1842, 2413–2426. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hollerer, I.; Bachmann, A.; Muckenthaler, M.U. Pathophysiological consequences and benefits of HFE mutations: 20 years of research. Haematologica 2017, 102, 809–817. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Naugler, C. Hemochromatosis: A Neolithic adaptation to cereal grain diets. Med. Hypotheses 2008, 70, 691–692. [Google Scholar] [CrossRef] [PubMed]
| Mutation | Number of Carriers | Percentage | Mutation | Number of Carriers | Percentage |
|---|---|---|---|---|---|
| p.Gly94Asp | 25 | 13.7 | p.Ile150Thr | 5 | 2.7 |
| p.Asp91Ala (heterozygous) | 14 | 7.7 | p.Gly94Cys | 4 | 2.2 |
| p.Leu145Phe | 14 | 7.7 | p.Ser135Asn † | 4 | 2.2 |
| p.Leu85Phe † | 11 | 6.0 | p.Thr138Ala | 3 | 1.6 |
| p.Gly42Ser † | 10 | 5.5 | p.Gly11Arg | 2 | 1.1 |
| p.Ala5Val † | 9 | 4.9 | p.Ala96Gly | 2 | 1.1 |
| p.Asn66Ser | 7 | 3.8 | p.Asp110Tyr | 2 | 1.1 |
| p.Asp12Tyr | 6 | 3.3 | p.Ile114Thr | 2 | 1.1 |
| p.Glu134del | 6 | 3.3 | p.Asp125Gly | 2 | 1.1 |
| p.Asn20Ser | 5 | 2.7 | p.Gly148Asp | 2 | 1.1 |
| p.Gly73Ser † | 5 | 2.7 | p.Gly148Ser | 2 | 1.1 |
| p.Asp91Ala (homozygous) | 5 | 2.7 | Others ‡ | 36 | 19.7 |
| Whole Series | Mild Mutation Carriers | Severe Mutation Carriers | p | |
|---|---|---|---|---|
| Sex Female (%) Male (%) | 98 (53.6%) 85 (46.4%) | 75 (56.8%) 57 (43.2%) | 23 (45.1%) 28 (54.9%) | p = 0.15 |
| Familial Status FALS (%) SALS (%) | 111 (60.7%) 72 (39.3%) | 74 (56.1%) 58 (43.9%) | 37 (72.5%) 14 (27.5%) | p = 0.04 |
| Age at onset, mean [SD] | 53.7 [12.1] | 53.6 [12.1] | 53.8 [12.2] | p = 0.93 |
| Site of onset Bulbar (%) Spinal (%) | 10 (5.5%) 173 (94.5%) | 8 (6.1%) 124 (93.9%) | 2 (3.9%) 49 (96.1%) | p = 0.57 |
| Survival (years), median [IQR] | 7.41 [2.58-NA] | 9.02 [4.06-NA] | 1.99 [1–4.03] | p < 0.001 |
| Total | 183 | 132 | 51 |
| HFE Genotypes | Observed | Expected |
|---|---|---|
| CC | 127 | 127.1 |
| CG | 51 | 50.8 |
| GG | 5 | 5.1 |
| Minor allele frequency | 0.17 | |
| χ2 test p-value | 0.96 with 1 degree of freedom |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Canosa, A.; Calvo, A.; Mora, G.; Moglia, C.; Brunetti, M.; Barberis, M.; Borghero, G.; Caponnetto, C.; Trojsi, F.; Spataro, R.; et al. The HFE p.H63D (p.His63Asp) Polymorphism Is a Modifier of ALS Outcome in Italian and French Patients with SOD1 Mutations. Biomedicines 2023, 11, 704. https://doi.org/10.3390/biomedicines11030704
Canosa A, Calvo A, Mora G, Moglia C, Brunetti M, Barberis M, Borghero G, Caponnetto C, Trojsi F, Spataro R, et al. The HFE p.H63D (p.His63Asp) Polymorphism Is a Modifier of ALS Outcome in Italian and French Patients with SOD1 Mutations. Biomedicines. 2023; 11(3):704. https://doi.org/10.3390/biomedicines11030704
Chicago/Turabian StyleCanosa, Antonio, Andrea Calvo, Gabriele Mora, Cristina Moglia, Maura Brunetti, Marco Barberis, Giuseppe Borghero, Claudia Caponnetto, Francesca Trojsi, Rossella Spataro, and et al. 2023. "The HFE p.H63D (p.His63Asp) Polymorphism Is a Modifier of ALS Outcome in Italian and French Patients with SOD1 Mutations" Biomedicines 11, no. 3: 704. https://doi.org/10.3390/biomedicines11030704








