Potential Prognostic Biomarkers of OSBPL Family Genes in Patients with Pancreatic Ductal Adenocarcinoma
Abstract
:1. Introduction
2. Materials and Methods
2.1. Oncomine Gene Analysis of OSBPL Family Members in PDAC
2.2. The Gene Expression Profiling Interactive Analysis 2 (GEPIA 2) Database Analysis of Clinicopathological Factors of the OSBPL Gene Family
2.3. Cancer Cell Line Encyclopedia (CCLE) Analysis
2.4. Kaplan–Meier (KM) Plotter Evaluates the Influence of Expressions of OSBPL Family Transcription Levels in PDAC
2.5. Analysis of Genetic Alterations by cBioPortal
2.6. GeneMANIA Analysis for Functions and Interactions of OSBPL Gene Family Members
2.7. STRING Analysis for OSBPL Family Members and Other Related Proteins
2.8. Co-Expression Materials Obtained from DAVID through cBioPortal
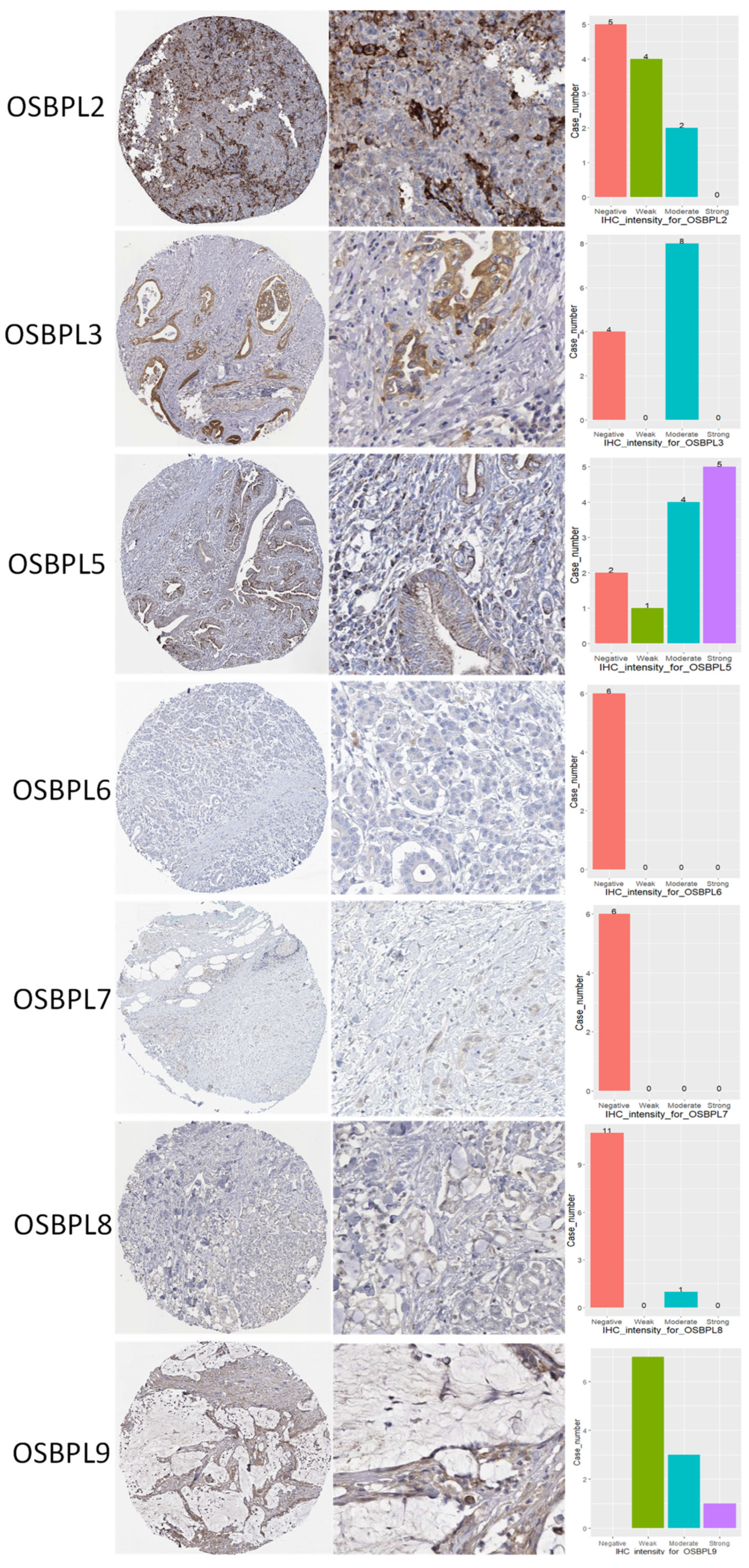
2.9. Analysis of Protein Expressions in Clinical Human Specimens
2.10. Tumor Immune Estimation Resource (TIMER) 2.0 contains Materials of Immune-Infiltration of OSBPL Gene Family Members
2.11. Statistical Analysis
3. Results
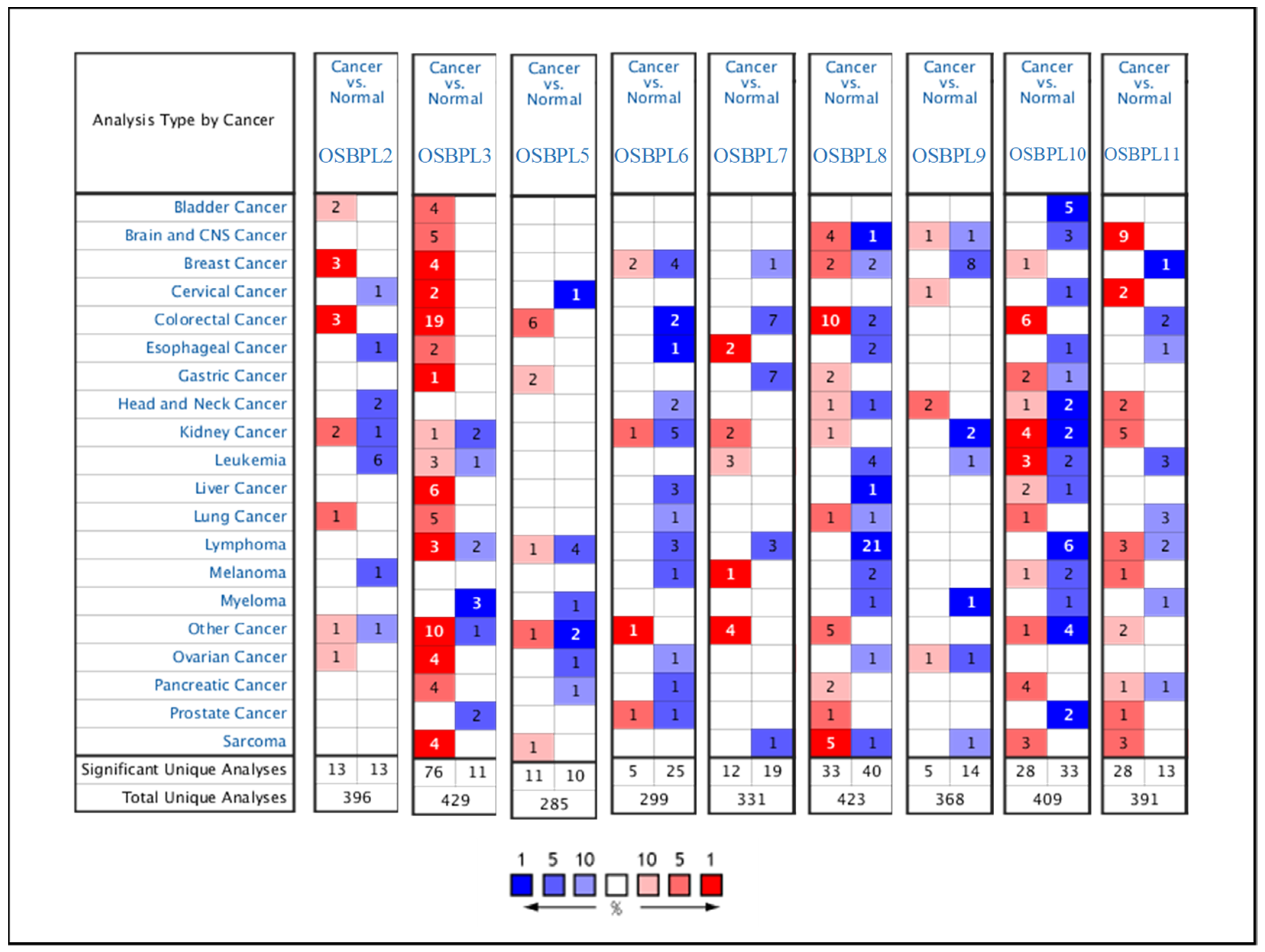
3.1. OSBPL Gene Expressions in PDAC
3.2. Associations of Transcription Levels with Clinicopathological Factors of OSBPL Family Members in Patients with PDAC
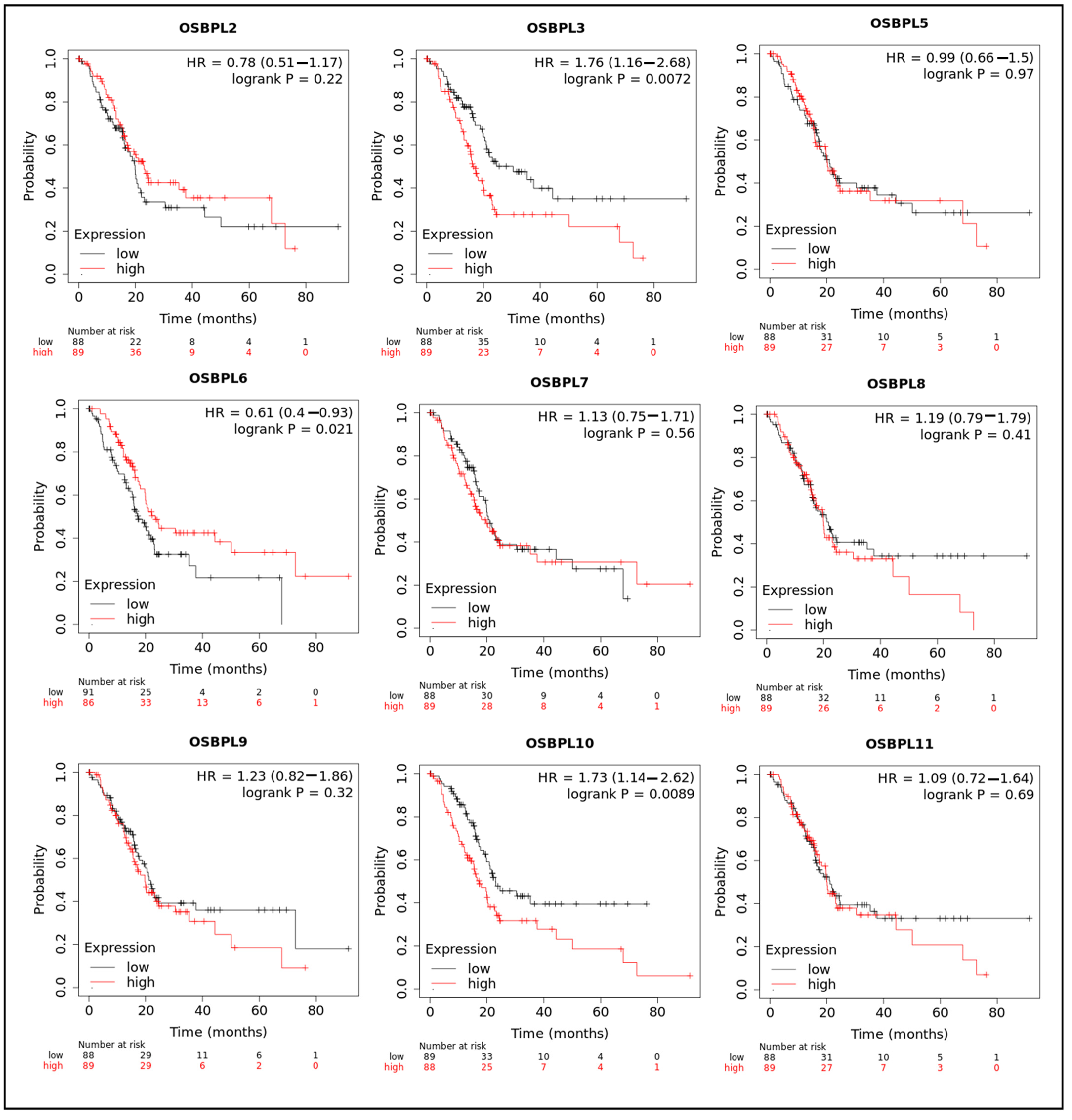
3.3. Prognostic Values of OSBPL Family Members in PDAC
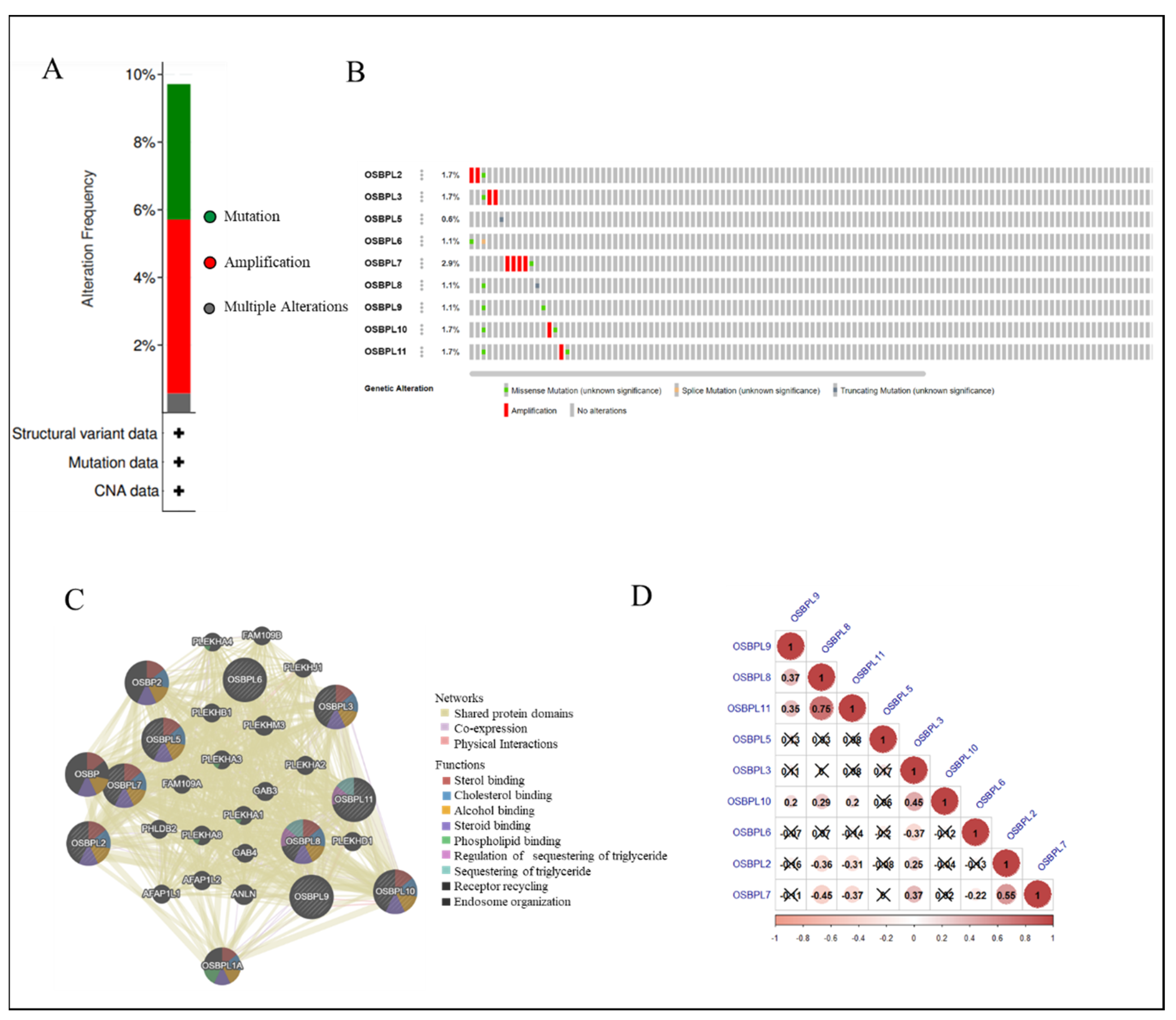
3.4. Analyses of Genetic Alterations, Co-Expressions, and Interactions of OSBPL Family Members in PDAC
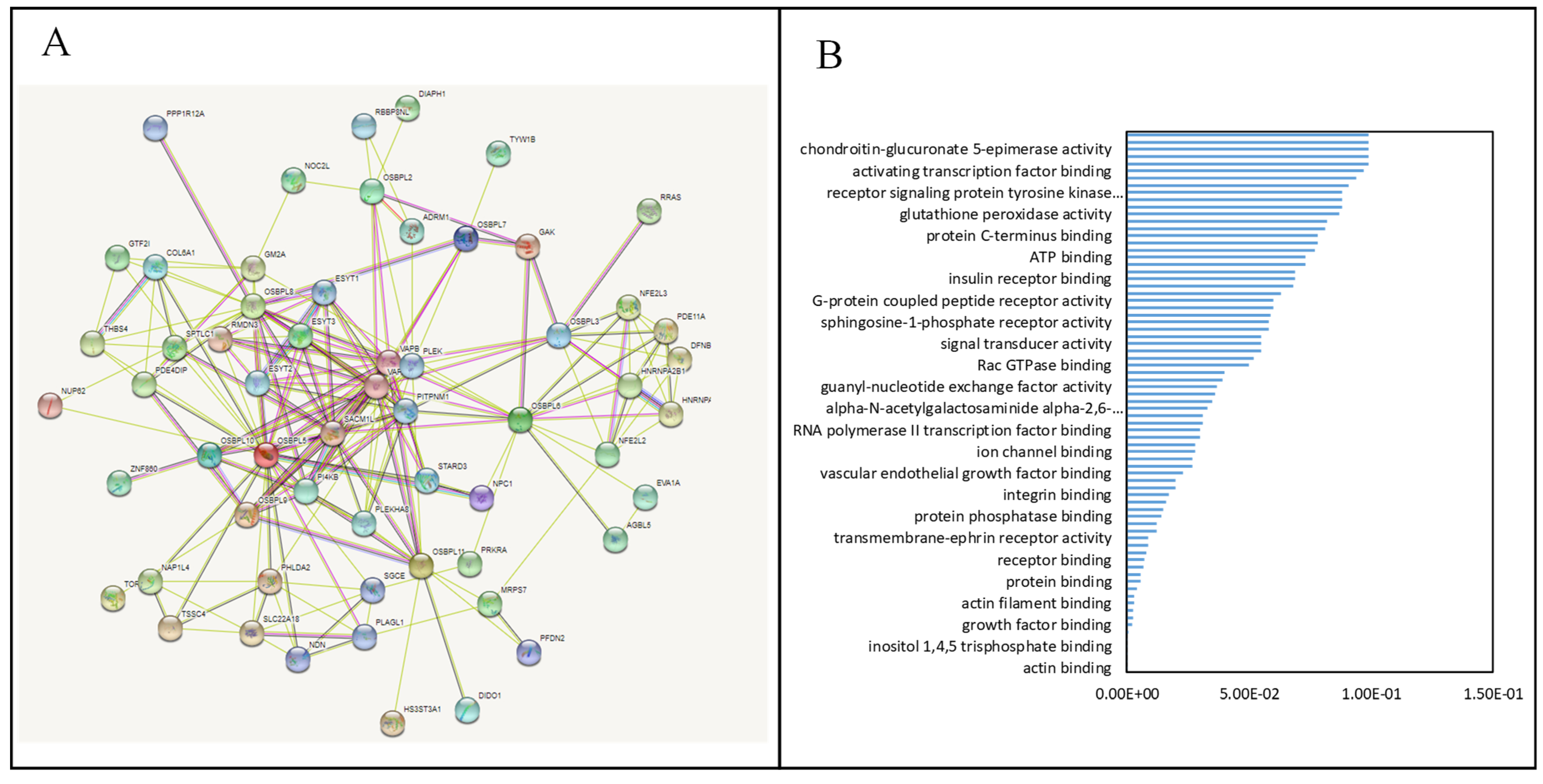
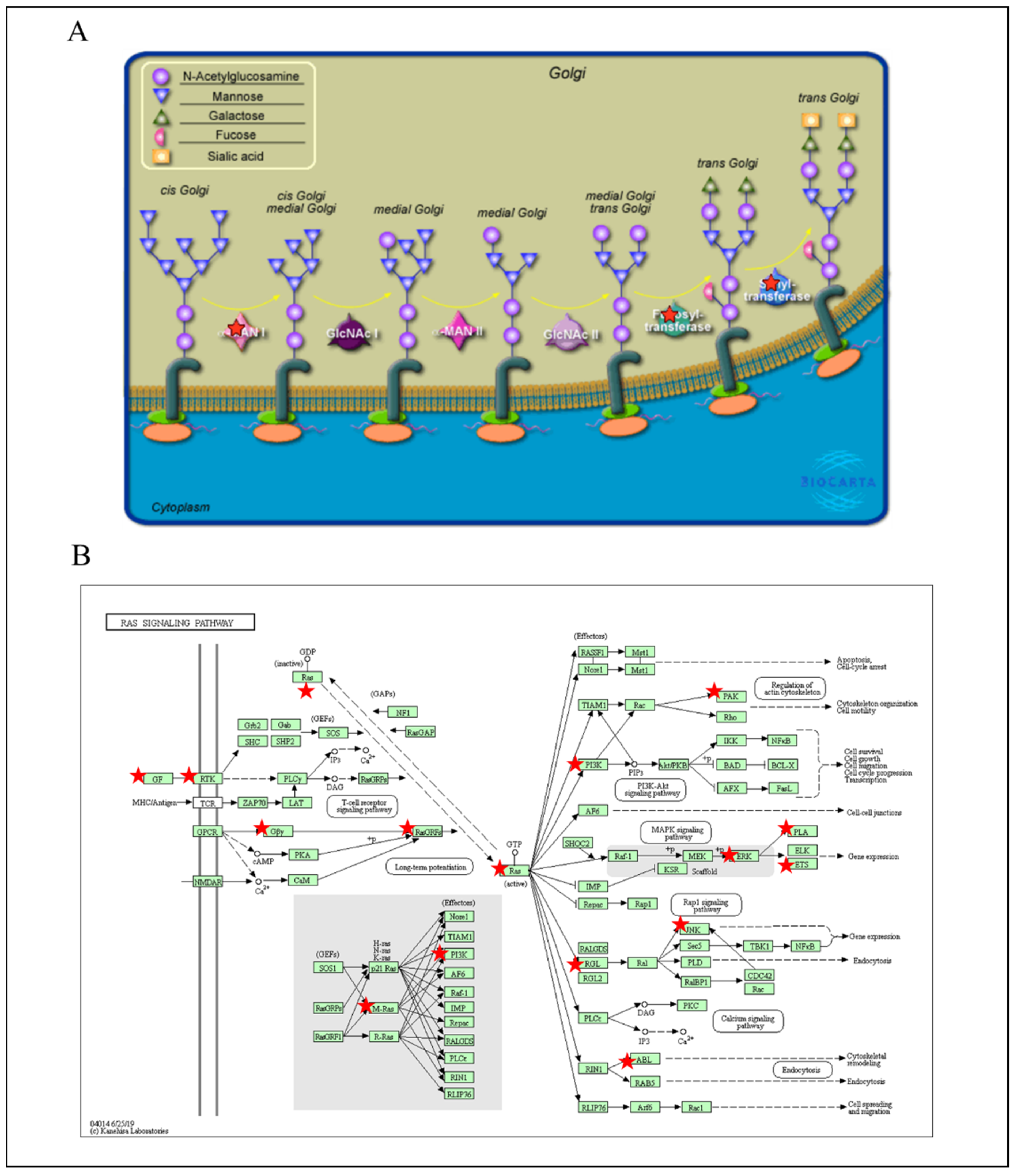
3.5. PPIs and Co-Expression for Pathway Enrichment Analysis of OSBP Family Members in Patients with PDAC
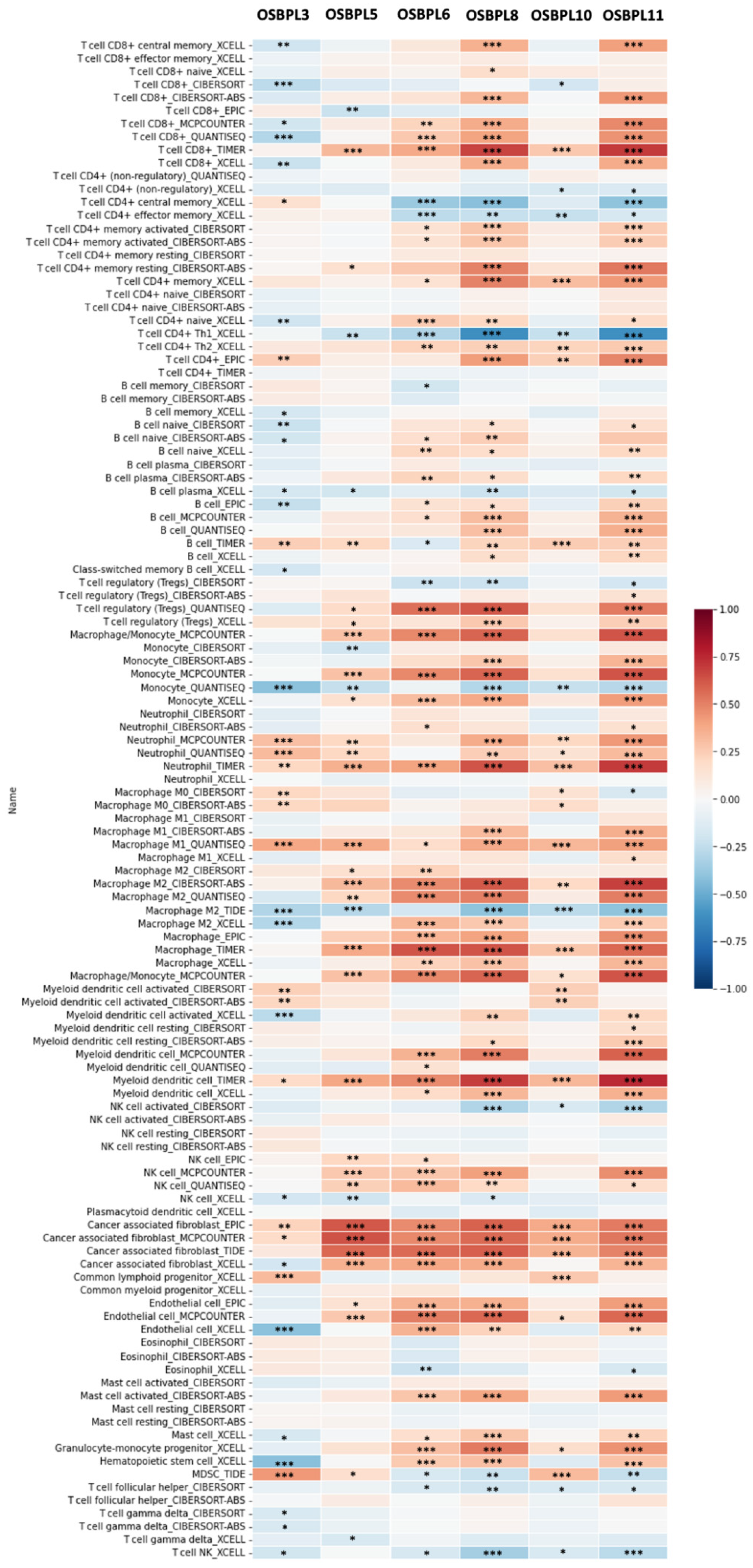
3.6. Associations of OSBPL Family Gene Transcriptional Levels and Immune-Infiltration in PDAC
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Olkkonen, V.M. OSBP-Related Protein Family in Lipid Transport Over Membrane Contact Sites. Lipid Insights 2015, 8, 1–9. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Jiao, H.L.; Weng, B.S.; Yan, S.S.; Lin, Z.M.; Wang, S.Y.; Chen, X.P.; Liang, G.H.; Li, X.Q.; Zhao, W.Y.; Huang, J.Y.; et al. Upregulation of OSBPL3 by HIF1A promotes colorectal cancer progression through activation of RAS signaling pathway. Cell Death Dis. 2020, 11, 571. [Google Scholar] [CrossRef] [PubMed]
- Olkkonen, V.M.; Koponen, A.; Arora, A. OSBP-related protein 2 (ORP2): Unraveling its functions in cellular lipid/carbohydrate metabolism, signaling and F-actin regulation. J. Steroid Biochem. Mol. Biol. 2019, 192, 105298. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Roberts, B.L.; Severance, Z.C.; Bensen, R.C.; Le-McClain, A.T.; Malinky, C.A.; Mettenbrink, E.M.; Nunez, J.I.; Reddig, W.J.; Blewett, E.L.; Burgett, A.W.G. Differing activities of oxysterol-binding protein (OSBP) targeting anti-viral compounds. Antivir. Res. 2019, 170, 104548. [Google Scholar] [CrossRef] [PubMed]
- Santos, N.C.; Girik, V.; Nunes-Hasler, P. ORP5 and ORP8: Sterol Sensors and Phospholipid Transfer Proteins at Membrane Contact Sites? Biomolecules 2020, 10, 928. [Google Scholar] [CrossRef] [PubMed]
- Jang, J.; To, C.; De Clercq, D.J.H.; Park, E.; Ponthier, C.M.; Shin, B.H.; Mushajiang, M.; Nowak, R.P.; Fischer, E.S.; Eck, M.J.; et al. Mutant-Selective Allosteric EGFR Degraders are Effective Against a Broad Range of Drug-Resistant Mutations. Angew. Chem. Int. Ed. Engl. 2020, 59, 14481–14489. [Google Scholar] [CrossRef] [PubMed]
- Vozianov, S.O.; Kashuba, V.I.; Grygorenko, V.M.; Gordiyuk, V.V.; Danylets, R.O.; Bondarenko, Y.M.; Vikarchuk, M.V. Identification of a New Diagnostic Markers of Prostatic Cancer, Using Noti-Microchips. Klin Khir 2016, 54–57. [Google Scholar]
- Loilome, W.; Wechagama, P.; Namwat, N.; Jusakul, A.; Sripa, B.; Miwa, M.; Kuver, R.; Yongvanit, P. Expression of oxysterol binding protein isoforms in opisthorchiasis-associated cholangiocarcinoma: A potential molecular marker for tumor metastasis. Parasitol. Int. 2012, 61, 136–139. [Google Scholar] [CrossRef] [PubMed]
- Wang, S.; Zhang, X.; Li, Z.; Wang, W.; Li, B.; Huang, X.; Sun, G.; Xu, J.; Li, Q.; Xu, Z.; et al. Circular RNA profile identifies circOSBPL10 as an oncogenic factor and prognostic marker in gastric cancer. Oncogene 2019, 38, 6985–7001. [Google Scholar] [CrossRef] [PubMed]
- Dobashi, A.; Togashi, Y.; Tanaka, N.; Yokoyama, M.; Tsuyama, N.; Baba, S.; Mori, S.; Hatake, K.; Yamaguchi, T.; Noda, T.; et al. TP53 and OSBPL10 alterations in diffuse large B-cell lymphoma: Prognostic markers identified via exome analysis of cases with extreme prognosis. Oncotarget 2018, 9, 19555–19568. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Peng, H.; Yan, Z.; Zeng, X.; Zhang, S.; Jiang, H.; Huang, H.; Zhuo, H. Serum and tissue proteomic signatures of patients with hepatocellular carcinoma using 2D gel electrophoresis. Mol. Med. Rep. 2019, 20, 1025–1038. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hou, G.X.; Liu, P.; Yang, J.; Wen, S. Mining expression and prognosis of topoisomerase isoforms in non-small-cell lung cancer by using Oncomine and Kaplan-Meier plotter. PLoS ONE 2017, 12, e0174515. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Thorat, M.A.; Balasubramanian, R. Breast cancer prevention in high-risk women. Best Pract. Res. Clin. Obstet. Gynaecol. 2020, 65, 18–31. [Google Scholar] [CrossRef] [PubMed]
- Lin, C.Y.; Lee, C.H.; Chuang, Y.H.; Lee, J.Y.; Chiu, Y.Y.; Wu Lee, Y.H.; Jong, Y.J.; Hwang, J.K.; Huang, S.H.; Chen, L.C.; et al. Membrane protein-regulated networks across human cancers. Nat. Commun. 2019, 10, 3131. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tsai, H.T.; Huang, C.S.; Tu, C.C.; Liu, C.Y.; Huang, C.J.; Ho, Y.S.; Tu, S.H.; Tseng, L.M.; Huang, C.C. Multi-gene signature of microcalcification and risk prediction among Taiwanese breast cancer. Sci. Rep. 2020, 10, 18276. [Google Scholar] [CrossRef]
- Nguyen, H.D.; Liao, Y.C.; Ho, Y.S.; Chen, L.C.; Chang, H.W.; Cheng, T.C.; Liu, D.; Lee, W.R.; Shen, S.C.; Wu, C.H.; et al. The α9 Nicotinic Acetylcholine Receptor Mediates Nicotine-Induced PD-L1 Expression and Regulates Melanoma Cell Proliferation and Migration. Cancers 2019, 11, 1991. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lee, K.L.; Kuo, Y.C.; Ho, Y.S.; Huang, Y.H. Triple-Negative Breast Cancer: Current Understanding and Future Therapeutic Breakthrough Targeting Cancer Stemness. Cancers 2019, 11, 1334. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tang, Z.; Kang, B.; Li, C.; Chen, T.; Zhang, Z. GEPIA2: An enhanced web server for large-scale expression profiling and interactive analysis. Nucleic Acids Res. 2019, 47, W556–W560. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sun, C.C.; Li, S.J.; Hu, W.; Zhang, J.; Zhou, Q.; Liu, C.; Li, L.L.; Songyang, Y.Y.; Zhang, F.; Chen, Z.L.; et al. Comprehensive Analysis of the Expression and Prognosis for E2Fs in Human Breast Cancer. Mol. Ther. 2019, 27, 1153–1165. [Google Scholar] [CrossRef] [PubMed]
- Wang, D.; Hu, B.; Hu, C.; Zhu, F.; Liu, X.; Zhang, J.; Wang, B.; Xiang, H.; Cheng, Z.; Xiong, Y.; et al. Clinical Characteristics of 138 Hospitalized Patients With 2019 Novel Coronavirus-Infected Pneumonia in Wuhan, China. JAMA 2020, 323, 1061–1069. [Google Scholar] [CrossRef] [PubMed]
- Barretina, J.; Caponigro, G.; Stransky, N.; Venkatesan, K.; Margolin, A.A.; Kim, S.; Wilson, C.J.; Lehar, J.; Kryukov, G.V.; Sonkin, D.; et al. The Cancer Cell Line Encyclopedia enables predictive modelling of anticancer drug sensitivity. Nature 2012, 483, 603–607. [Google Scholar] [CrossRef] [PubMed]
- Ghandi, M.; Huang, F.W.; Jané-Valbuena, J.; Kryukov, G.V.; Lo, C.C.; McDonald, E.R.; Barretina, J.; Gelfand, E.T.; Bielski, C.M.; Li, H. Next-generation characterization of the cancer cell line encyclopedia. Nature 2019, 569, 503–508. [Google Scholar] [CrossRef]
- Wang, C.Y.; Chiao, C.C.; Phan, N.N.; Li, C.Y.; Sun, Z.D.; Jiang, J.Z.; Hung, J.H.; Chen, Y.L.; Yen, M.C.; Weng, T.Y.; et al. Gene signatures and potential therapeutic targets of amino acid metabolism in estrogen receptor-positive breast cancer. Am. J. Cancer Res. 2020, 10, 95–113. [Google Scholar]
- Anuraga, G.; Tang, W.C.; Phan, N.N.; Ta, H.D.K.; Liu, Y.H.; Wu, Y.F.; Lee, K.H.; Wang, C.Y. Comprehensive Analysis of Prognostic and Genetic Signatures for General Transcription Factor III (GTF3) in Clinical Colorectal Cancer Patients Using Bioinformatics Approaches. Curr. Issues Mol. Biol. 2021, 43, 2. [Google Scholar] [CrossRef] [PubMed]
- Khoa Ta, H.D.; Tang, W.C.; Phan, N.N.; Anuraga, G.; Hou, S.Y.; Chiao, C.C.; Liu, Y.H.; Wu, Y.F.; Lee, K.H.; Wang, C.Y. Analysis of LAGEs Family Gene Signature and Prognostic Relevance in Breast Cancer. Diagnostics 2021, 11, 726. [Google Scholar] [CrossRef] [PubMed]
- Ning, G.; Huang, Y.L.; Zhen, L.M.; Xu, W.X.; Jiao, Q.; Yang, F.J.; Wu, L.N.; Zheng, Y.Y.; Song, J.; Wang, Y.S.; et al. Transcriptional expressions of Chromobox 1/2/3/6/8 as independent indicators for survivals in hepatocellular carcinoma patients. Aging 2018, 10, 3450–3473. [Google Scholar] [CrossRef]
- Wang, X.; Liu, R.; Qu, X.; Yu, H.; Chu, H.; Zhang, Y.; Zhu, W.; Wu, X.; Gao, H.; Tao, B.; et al. α-Ketoglutarate-Activated NF-κB Signaling Promotes Compensatory Glucose Uptake and Brain Tumor Development. Mol. Cell 2019, 76, 148–162.e7. [Google Scholar] [CrossRef]
- Chen, Y.L.; Wang, C.Y.; Fang, J.H.; Hsu, H.P. Serine/threonine-protein kinase 24 is an inhibitor of gastric cancer metastasis through suppressing CDH1 gene and enhancing stemness. Am. J. Cancer Res. 2021, 11, 4277–4293. [Google Scholar]
- Ta, H.D.K.; Wang, W.J.; Phan, N.N.; An Ton, N.T.; Anuraga, G.; Ku, S.C.; Wu, Y.F.; Wang, C.Y.; Lee, K.H. Potential Therapeutic and Prognostic Values of LSM Family Genes in Breast Cancer. Cancers 2021, 13, 4902. [Google Scholar] [CrossRef]
- Kao, T.J.; Wu, C.C.; Phan, N.N.; Liu, Y.H.; Ta, H.D.K.; Anuraga, G.; Wu, Y.F.; Lee, K.H.; Chuang, J.Y.; Wang, C.Y. Prognoses and genomic analyses of proteasome 26S subunit, ATPase (PSMC) family genes in clinical breast cancer. Aging 2021, 13, 17970. [Google Scholar] [CrossRef]
- Choy, T.K.; Wang, C.Y.; Phan, N.N.; Khoa Ta, H.D.; Anuraga, G.; Liu, Y.H.; Wu, Y.F.; Lee, K.H.; Chuang, J.Y.; Kao, T.J. Identification of Dipeptidyl Peptidase (DPP) Family Genes in Clinical Breast Cancer Patients via an Integrated Bioinformatics Approach. Diagnostics 2021, 11, 1204. [Google Scholar] [CrossRef]
- Chen, P.S.; Hsu, H.P.; Phan, N.N.; Yen, M.C.; Chen, F.W.; Liu, Y.W.; Lin, F.P.; Feng, S.Y.; Cheng, T.L.; Yeh, P.H.; et al. CCDC167 as a potential therapeutic target and regulator of cell cycle-related networks in breast cancer. Aging 2021, 13, 4157–4181. [Google Scholar] [CrossRef]
- Ramezani, M.; Baharzadeh, F.; Almasi, A.; Sadeghi, M. A Systematic Review and Meta-Analysis: Evaluation of the beta-Human Papillomavirus in Immunosuppressed Individuals with Cutaneous Squamous Cell Carcinoma. Biomedicine 2020, 10, 1–10. [Google Scholar] [CrossRef]
- Jiao, X.D.; He, X.; Qin, B.D.; Liu, K.; Wu, Y.; Liu, J.; Hou, T.; Zang, Y.S. The prognostic value of tumor mutation burden in EGFR-mutant advanced lung adenocarcinoma, an analysis based on cBioPortal data base. J. Thorac. Dis. 2019, 11, 4507–4515. [Google Scholar] [CrossRef]
- Cerami, E.; Gao, J.; Dogrusoz, U.; Gross, B.E.; Sumer, S.O.; Aksoy, B.A.; Jacobsen, A.; Byrne, C.J.; Heuer, M.L.; Larsson, E.; et al. The cBio cancer genomics portal: An open platform for exploring multidimensional cancer genomics data. Cancer Discov. 2012, 2, 401–404. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Liu, F.; Wu, H. CC Chemokine Receptors in Lung Adenocarcinoma: The Inflammation-Related Prognostic Biomarkers and Immunotherapeutic Targets. J. Inflamm. Res. 2021, 14, 267–285. [Google Scholar] [CrossRef]
- Szklarczyk, D.; Morris, J.H.; Cook, H.; Kuhn, M.; Wyder, S.; Simonovic, M.; Santos, A.; Doncheva, N.T.; Roth, A.; Bork, P.; et al. The STRING database in 2017: Quality-controlled protein-protein association networks, made broadly accessible. Nucleic Acids Res. 2017, 45, D362–D368. [Google Scholar] [CrossRef]
- Wang, C.Y.; Chao, Y.J.; Chen, Y.L.; Wang, T.W.; Phan, N.N.; Hsu, H.P.; Shan, Y.S.; Lai, M.D. Upregulation of peroxisome proliferator-activated receptor-α and the lipid metabolism pathway promotes carcinogenesis of ampullary cancer. Int. J. Med. Sci. 2021, 18, 256–269. [Google Scholar] [CrossRef] [PubMed]
- Wu, P.S.; Yen, J.H.; Wang, C.Y.; Chen, P.Y.; Hung, J.H.; Wu, M.J. 8-Hydroxydaidzein, an Isoflavone from Fermented Soybean, Induces Autophagy, Apoptosis, Differentiation, and Degradation of Oncoprotein BCR-ABL in K562 Cells. Biomedicines 2020, 8, 506. [Google Scholar] [CrossRef]
- Wu, C.C.; Ekanem, T.I.; Phan, N.N.; Loan, D.T.T.; Hou, S.Y.; Lee, K.H.; Wang, C.Y. Gene signatures and prognostic analyses of the Tob/BTG pituitary tumor-transforming gene (PTTG) family in clinical breast cancer patients. Int. J. Med. Sci. 2020, 17, 3112–3124. [Google Scholar] [CrossRef] [PubMed]
- Uhlén, M.; Fagerberg, L.; Hallström, B.M.; Lindskog, C.; Oksvold, P.; Mardinoglu, A.; Sivertsson, Å.; Kampf, C.; Sjöstedt, E.; Asplund, A. Tissue-based map of the human proteome. Science 2015, 347, 1260419. [Google Scholar] [CrossRef] [PubMed]
- Phan, N.N.; Liu, S.; Wang, C.Y.; Hsu, H.P.; Lai, M.D.; Li, C.Y.; Chen, C.F.; Chiao, C.C.; Yen, M.C.; Sun, Z.; et al. Overexpressed gene signature of EPH receptor A/B family in cancer patients-comprehensive analyses from the public high-throughput database. Int. J. Clin. Exp. Pathol. 2020, 13, 1220–1242. [Google Scholar] [PubMed]
- Hagerling, C.; Owyong, M.; Sitarama, V.; Wang, C.Y.; Lin, C.; van den Bijgaart, R.J.E.; Koopman, C.D.; Brenot, A.; Nanjaraj, A.; Wärnberg, F.; et al. LGR5 in breast cancer and ductal carcinoma in situ: A diagnostic and prognostic biomarker and a therapeutic target. BMC Cancer 2020, 20, 542. [Google Scholar] [CrossRef] [PubMed]
- Li, T.; Fan, J.; Wang, B.; Traugh, N.; Chen, Q.; Liu, J.S.; Li, B.; Liu, X.S. TIMER: A Web Server for Comprehensive Analysis of Tumor-Infiltrating Immune Cells. Cancer Res. 2017, 77, e108–e110. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Galon, J.; Bruni, D. Approaches to treat immune hot, altered and cold tumours with combination immunotherapies. Nat. Rev. Drug Discov. 2019, 18, 197–218. [Google Scholar] [CrossRef] [PubMed]
- Segara, D.; Biankin, A.V.; Kench, J.G.; Langusch, C.C.; Dawson, A.C.; Skalicky, D.A.; Gotley, D.C.; Coleman, M.J.; Sutherland, R.L.; Henshall, S.M. Expression of HOXB2, a retinoic acid signaling target in pancreatic cancer and pancreatic intraepithelial neoplasia. Clin. Cancer Res. 2005, 11, 3587–3596. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Pei, H.; Li, L.; Fridley, B.L.; Jenkins, G.D.; Kalari, K.R.; Lingle, W.; Petersen, G.; Lou, Z.; Wang, L. FKBP51 affects cancer cell response to chemotherapy by negatively regulating Akt. Cancer Cell 2009, 16, 259–266. [Google Scholar] [CrossRef] [Green Version]
- Badea, L.; Herlea, V.; Dima, S.O.; Dumitrascu, T.; Popescu, I. Combined gene expression analysis of whole-tissue and microdissected pancreatic ductal adenocarcinoma identifies genes specifically overexpressed in tumor epithelia. Hepatogastroenterology 2008, 55, 2016–2027. [Google Scholar] [PubMed]
- Iacobuzio-Donahue, C.A.; Maitra, A.; Olsen, M.; Lowe, A.W.; van Heek, N.T.; Rosty, C.; Walter, K.; Sato, N.; Parker, A.; Ashfaq, R.; et al. Exploration of global gene expression patterns in pancreatic adenocarcinoma using cDNA microarrays. Am. J. Pathol. 2003, 162, 1151–1162. [Google Scholar] [CrossRef]
- Buchholz, M.; Braun, M.; Heidenblut, A.; Kestler, H.A.; Kloppel, G.; Schmiegel, W.; Hahn, S.A.; Luttges, J.; Gress, T.M. Transcriptome analysis of microdissected pancreatic intraepithelial neoplastic lesions. Oncogene 2005, 24, 6626–6636. [Google Scholar] [CrossRef] [Green Version]
- Grutzmann, R.; Pilarsky, C.; Ammerpohl, O.; Luttges, J.; Bohme, A.; Sipos, B.; Foerder, M.; Alldinger, I.; Jahnke, B.; Schackert, H.K.; et al. Gene expression profiling of microdissected pancreatic ductal carcinomas using high-density DNA microarrays. Neoplasia 2004, 6, 611–622. [Google Scholar] [CrossRef] [Green Version]
- Karamitopoulou, E. The Tumor Microenvironment of Pancreatic Cancer. Cancers 2020, 12, 3076. [Google Scholar] [CrossRef] [PubMed]
- Berkel, C.; Cacan, E. GAB2 and GAB3 are expressed in a tumor stage-, grade- and histotype-dependent manner and are associated with shorter progression-free survival in ovarian cancer. J. Cell Commun. Signal. 2021, 15, 57–70. [Google Scholar] [CrossRef]
- Olkkonen, V.M. The emerging roles of OSBP-related proteins in cancer: Impacts through phosphoinositide metabolism and protein-protein interactions. Biochem. Pharmacol. 2021, 114455. [Google Scholar] [CrossRef] [PubMed]
- Pan, S.; Brentnall, T.A.; Chen, R. Glycoproteins and glycoproteomics in pancreatic cancer. World J. Gastroenterol. 2016, 22, 9288–9299. [Google Scholar] [CrossRef] [PubMed]
- Ishikawa, S.; Nagai, Y.; Masuda, T.; Koga, Y.; Nakamura, T.; Imamura, Y.; Takamori, H.; Hirota, M.; Funakosi, A.; Fukushima, M.; et al. The role of oxysterol binding protein-related protein 5 in pancreatic cancer. Cancer Sci. 2010, 101, 898–905. [Google Scholar] [CrossRef]
- Khoshghamat, N.; Jafari, N.; Toloue-Pouya, V.; Azami, S.; Mirnourbakhsh, S.H.; Khazaei, M.; Ferns, G.A.; Rajabian, M.; Avan, A. The therapeutic potential of renin-angiotensin system inhibitors in the treatment of pancreatic cancer. Life Sci. 2021, 270, 119118. [Google Scholar] [CrossRef] [PubMed]
- Yarchoan, M.; Hopkins, A.; Jaffee, E.M. Tumor Mutational Burden and Response Rate to PD-1 Inhibition. N. Engl. J. Med. 2017, 377, 2500–2501. [Google Scholar] [CrossRef] [PubMed]
- Ino, Y.; Yamazaki-Itoh, R.; Shimada, K.; Iwasaki, M.; Kosuge, T.; Kanai, Y.; Hiraoka, N. Immune cell infiltration as an indicator of the immune microenvironment of pancreatic cancer. Br. J. Cancer 2013, 108, 914–923. [Google Scholar] [CrossRef] [PubMed]
- Leinwand, J.; Miller, G. Regulation and modulation of antitumor immunity in pancreatic cancer. Nat. Immunol. 2020, 21, 1152–1159. [Google Scholar] [CrossRef]
- Zhou, Q.; Tao, X.; Xia, S.; Guo, F.; Pan, C.; Xiang, H.; Shang, D. T Lymphocytes: A Promising Immunotherapeutic Target for Pancreatitis and Pancreatic Cancer? Front. Oncol. 2020, 10, 382. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Jin, L.; Kim, H.S.; Shi, J. Neutrophil in the Pancreatic Tumor Microenvironment. Biomolecules 2021, 11, 1170. [Google Scholar] [CrossRef] [PubMed]




Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Chou, C.-W.; Hsieh, Y.-H.; Ku, S.-C.; Shen, W.-J.; Anuraga, G.; Khoa Ta, H.D.; Lee, K.-H.; Lee, Y.-C.; Lin, C.-H.; Wang, C.-Y.; et al. Potential Prognostic Biomarkers of OSBPL Family Genes in Patients with Pancreatic Ductal Adenocarcinoma. Biomedicines 2021, 9, 1601. https://doi.org/10.3390/biomedicines9111601
Chou C-W, Hsieh Y-H, Ku S-C, Shen W-J, Anuraga G, Khoa Ta HD, Lee K-H, Lee Y-C, Lin C-H, Wang C-Y, et al. Potential Prognostic Biomarkers of OSBPL Family Genes in Patients with Pancreatic Ductal Adenocarcinoma. Biomedicines. 2021; 9(11):1601. https://doi.org/10.3390/biomedicines9111601
Chicago/Turabian StyleChou, Cheng-Wei, Yu-Hsiu Hsieh, Su-Chi Ku, Wan-Jou Shen, Gangga Anuraga, Hoang Dang Khoa Ta, Kuen-Haur Lee, Yu-Cheng Lee, Cheng-Hsien Lin, Chih-Yang Wang, and et al. 2021. "Potential Prognostic Biomarkers of OSBPL Family Genes in Patients with Pancreatic Ductal Adenocarcinoma" Biomedicines 9, no. 11: 1601. https://doi.org/10.3390/biomedicines9111601
APA StyleChou, C.-W., Hsieh, Y.-H., Ku, S.-C., Shen, W.-J., Anuraga, G., Khoa Ta, H. D., Lee, K.-H., Lee, Y.-C., Lin, C.-H., Wang, C.-Y., & Wang, W.-J. (2021). Potential Prognostic Biomarkers of OSBPL Family Genes in Patients with Pancreatic Ductal Adenocarcinoma. Biomedicines, 9(11), 1601. https://doi.org/10.3390/biomedicines9111601









