Leptospira interrogans Serogroup Pomona in a Dairy Cattle Farm in a Multi-Host Zootechnical System
Abstract
:1. Introduction
2. Materials and Methods
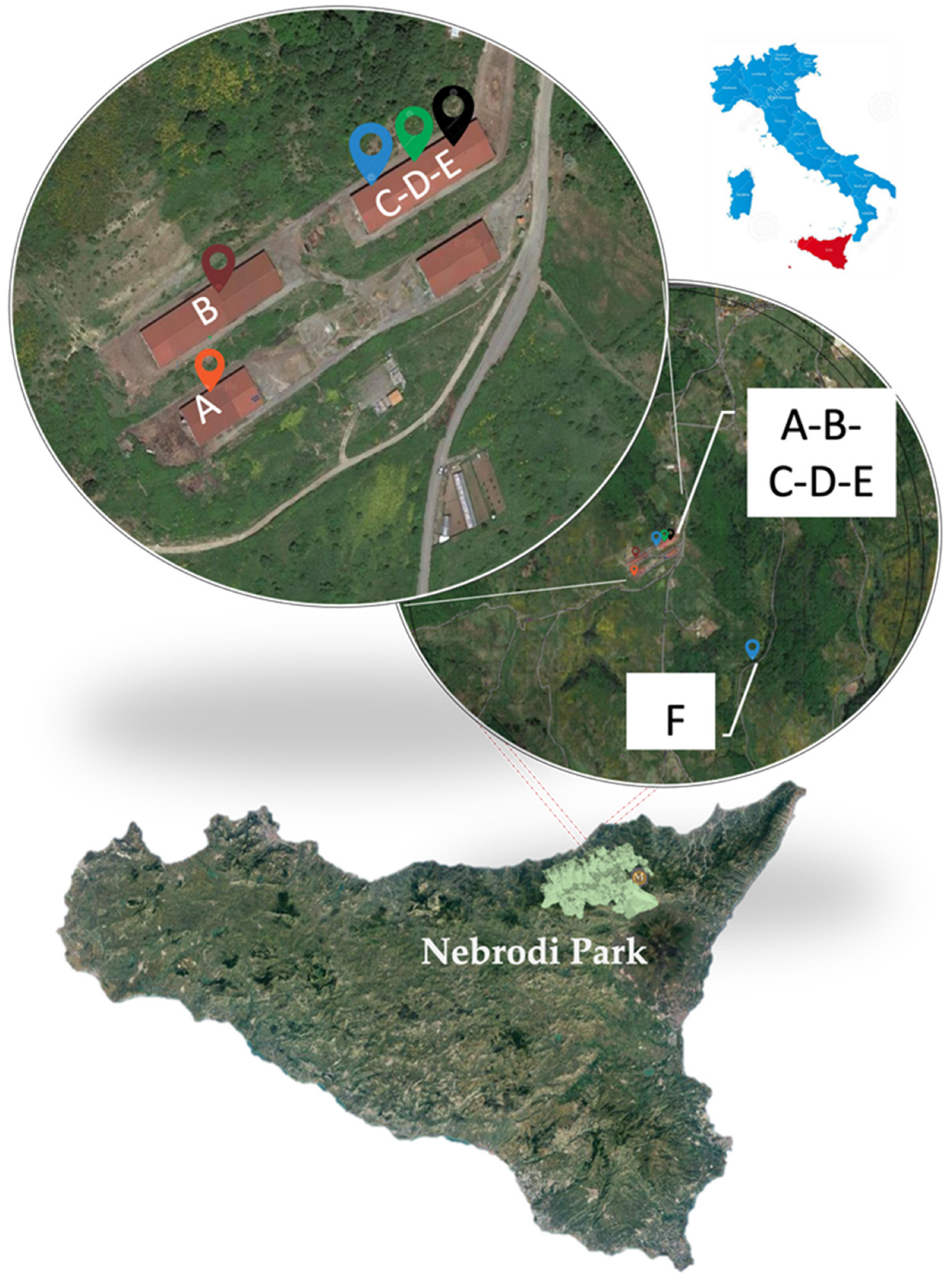
2.1. Farms
2.2. Sampling
2.3. Differential Diagnosis
2.4. Serological Test for Leptospirosis
2.5. Culture Examination/Isolation of Leptospira spp.
2.6. Molecular Tests for Leptospirosis
2.7. MLST and Phylogenetic Analyses
3. Results
3.1. Differential Diagnosis
3.2. Serological Results for Leptospirosis
3.3. Leptospiral Molecular Detection and Isolation
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Pappas, G.; Papadimitriou, P.; Siozopoulou, V.; Christou, L.; Akritidis, N. The globalization of leptospirosis: Worldwide incidence trends. Int. J. Infect. Dis. 2008, 12, 351–357. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Grippi, F.; Giudice, E.; Pietro, S.D.; Sciacca, C.; Santangelo, F.; Galluzzo, P.; Barreca, S.; Guercio, A. Leptospira Interrogans Serogroup Sejroe Serovar Hardjo in Aborting Cows: Two Herd Cases in Sicily (Italy). J. Vet. Res. 2020, 64, 73–78. [Google Scholar] [CrossRef] [PubMed]
- Bryson, D.G.; Ellis, W.A. Leptospirosis in a British domestic cat. J. Small Anim. Pract. 1976, 17, 459–465. [Google Scholar] [CrossRef] [PubMed]
- Ellis, W.A.; O’Brien, J.J.; Cassells, J.A.; Montgomery, J. Leptospiral infection in horses in Northen Ireland: Serological and microbiological findings. Equine Vet. J. 1983, 15, 317–320. [Google Scholar] [CrossRef]
- Ellis, W.A. Bovine leptospirosis in the tropics: Prevalence, pathogenesis and control. Prev. Vet. Med. 1984, 2, 411–421. [Google Scholar] [CrossRef]
- Gravekamp, C.; Korver, H.; Montgomery, J.; Everdard, C.O.R.; Carrington, D.; Ellis, W.A.; Terpstra, W.J. Leptospires isolated from Toads and Frogs on the Island of Barbados. Zent. Bakteriol. 1991, 275, 403–411. [Google Scholar] [CrossRef]
- White, F.H.; Sulzer, K.R.; Engel, R.W. Isolations of Leptospira Interrogans serovars hardjo, balcanica, and pomona from cattle at slaughter. Am. J. Vet. Res. 1982, 43, 1172–1173. [Google Scholar]
- Miotto, B.A.; Guilloux, A.; Tozzi, B.F.; Moreno, L.Z.; da Hora, A.S.; Dias, R.A.; Heinemann, M.B.; Moreno, A.M.; Filho, A.; Lilenbaum, W.; et al. Prospective study of canine leptospirosis in shelter and stray dog populations: Identification of chronic carriers and different Leptospira species infecting dogs. PLoS ONE 2018, 13, e0200384. [Google Scholar] [CrossRef] [Green Version]
- Mohammed, H.; Nozha, C.; Hakim, K.; Abdelaziz, F. Leptospira: Morphology, classification and pathogenesis. J. Bacteriol Parasitol. 2011, 2, 6. [Google Scholar] [CrossRef]
- Schuller, S.; Francey, T.; Hartman, K.; Hugonnard, M.; Kohn, B.; Nally, J.E.; Sykes, J. European consensus statement on leptospirosis in dogs and cats. J. Small Anim. Pract. 2015, 56, 159–179. [Google Scholar] [CrossRef]
- Cilia, G.; Bertelloni, F.; Mignone, W.; Spina, S.; Berio, E.; Razzuoli, E.; Vencia, W.; Franco, V.; Cecchi, F.; Bogi, S.; et al. Molecular detection of Leptospira spp. in wild boar (Sus scrofa) hunted in Liguria region (Italy). Comp. Immunol. Microbiol. Infect. Dis. 2020, 68, 101410. [Google Scholar] [CrossRef]
- Cerri, D.; Ebani, V.V.; Fratini, F.; Pinzauti, P.; Andreani, E. Epidemiology of leptospirosis: Observations on serological data obtained by a “diagnostic laboratory for leptospirosis” from 1995 to 2001. New Microbiol. 2003, 26, 383–389. [Google Scholar] [PubMed]
- Cilia, G.; Bertelloni, F.; Coppola, F.; Turchi, B.; Biliotti, C.; Poli, A.; Parisi, F.; Felicioli, A.; Cerri, D.; Fratini, F. Isolation of Leptospira serovar Pomona from a crested porcupine (Hystrix cristata L., 1758). Vet. Med. Sci. 2020, 6, 985–991. [Google Scholar] [CrossRef] [PubMed]
- Blasdell, K.R.; Morand, S.; Perera, D.; Firth, C. Association of rodent-borne Leptospira spp. with urban environments in Malaysian Borneo. PLoS Negl. Trop. Dis. 2019, 13, e0007141. [Google Scholar] [CrossRef] [PubMed]
- Mori, M.; Bourhy, P.; Le Guyader, M.; Van Esbroeck, M.; Djelouadji, Z.; Septfons, A.; Kodjo, A.; Picardeau, M. Pet rodents as possible risk for leptospirosis, Belgium and France, 2009 to 2016. Euro Surveill. 2017, 22, 16-00792. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Coppola, F.; Cilia, G.; Bertelloni, F.; Casini, L.; D’Addio, E.; Fratini, F.; Cerri, D.; Felicioli, A. Crested Porcupine (Hystrix cristata L.): A New Potential Host for Pathogenic Leptospira among Semi-Fossorial Mammals. Comp. Immunol. Microbiol. Infect. Dis. 2020, 70, 101472. [Google Scholar] [CrossRef]
- Bertelloni, F.; Mazzei, M.; Cilia, G.; Forzan, M.; Felicioli, A.; Sagona, S.; Bandecchi, P.; Turchi, B.; Cerri, D.; Fratini, F. Serological Survey on Bacterial and Viral Pathogens in Wild Boars Hunted in Tuscany. Ecohealth 2020, 17, 85–93. [Google Scholar] [CrossRef] [PubMed]
- Cilia, G.; Bertelloni, F.; Piredda, I.; Ponti, M.N.; Turchi, B.; Cantile, C.; Parisi, F.; Pinzauti, P.; Armani, A.; Palmas, B.; et al. Presence of pathogenic leptospira spp. In the reproductive system and fetuses of wild boars (Sus scrofa) in Italy. PLoS Negl. Trop. Dis 2020, 14, e0008982. [Google Scholar] [CrossRef]
- Cilia, G.; Bertelloni, F.; Angelini, M.; Cerri, D.; Fratini, F. Leptospira Survey in Wild Boar (Sus scrofa) Hunted in Tuscany, Central Italy. Pathogens 2020, 9, 377. [Google Scholar] [CrossRef]
- Vale-Gonçalves, H.M.; Cabral, J.A.; Faria, M.C.; Nunes-Pereira, M.; Faria, A.S.; Veloso, O.; Vieira, M.L.; Paiva-Cardoso, M.N. Prevalence of Leptospira antibodies in wild boars (Sus scrofa) from Northern Portugal: Risk factor analysis. Epidemiol. Infect. 2015, 143, 2126–2130. [Google Scholar] [CrossRef] [Green Version]
- Arent, Z.; Frizzell, C.; Gilmore, C.; Allen, A.; Ellis, W.A. Leptospira interrogans serovars Bratislava and Muenchen animal infections: Implications for epidemiology and control. Vet. Microbiol. 2016, 190, 19–26. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rocha, T.; Ellis, W.A.; Montgomery, J.; Gilmore, C.; Regalla, J.; Brem, S. Microbiological and serological study of leptospirosis in horses at slaughter: First isolations. Res. Vet. Sci. 2004, 76, 199–202. [Google Scholar] [CrossRef] [PubMed]
- Farina, R.; Cerri, D.; Renzoni, G.; Andreani, E.; Mani, P.; Ebani, V.; Pedrini, A.; Nuvoloni, R. Leptospira interrogans in the genital tract of sheep. Research on ewes and rams experimentally infected with serovar hardjo (hardjobovis). New Microbiol. 1996, 19, 235–242. [Google Scholar]
- Tagliabue, S.; Figarolli, B.M.; D’Incau, M.; Foschi, G.; Gennero, M.S.; Giordani, R.; Giordani, R.; Natale, A.; Papa, P.; Ponti, N.; et al. Serological surveillance of Leptospirosis in Italy: Two-year national data (2010–2011). Vet. Ital. 2016, 52, 129–138. [Google Scholar] [CrossRef]
- Heymann, D. Control of Communicable Diseases Manual, 19th ed.; American Public Health Association: Washington, DC, USA, 2008. [Google Scholar]
- Klarenbeek, A.; Schuffner, W.A.P. Appearance in Holland of Leptospira differing from Weil Strain. Ned. Tijdschr. Voor Geneeskd. 1933, 77, 4271–4276. [Google Scholar]
- Ellis, W.A. Control of canine leptospirosis in Europe: Time for a change? Vet. Rec. 2010, 167, 602–605. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Guglielmini, J.; Bourhy, P.; Schiettekatte, O.; Zinini, F.; Brisse, S.; Picardeau, M. Correction: Genus-wide Leptospira core genome multilocus sequence typing for strain taxonomy and global surveillance. PLoS Negl. Trop. Dis. 2020, 14, e0008673, Erratum in PLoS Negl. Trop. Dis. 2019, 13, e0007374. [Google Scholar] [CrossRef]
- Ismail, Z.B.; Abutarbush, S.M.; Al-Majali, A.M.; Gharaibeh, M.H.; Al-Khateeb, B. Seroprevalence and risk factors of Leptospira serovar Pomona and Leptospira serovar Hardjo infection in dairy cows in Jordan. J. Infect. Dev. Ctries. 2019, 30, 473–479. [Google Scholar] [CrossRef]
- Ellis, W.A. Animal leptospirosis. Curr Top Microbiol. Immunol. 2015, 387, 99–137. [Google Scholar] [CrossRef]
- Nally, J.E.; Hornsby, R.L.; Alt, D.P.; Bayles, D.; Wilson-Welder, J.H.; Palmquist, D.E.; Bauer, N.E. Isolation and characterization of pathogenic leptospires associated with cattle. Vet. Microbiol. 2018, 218, 25–30. [Google Scholar] [CrossRef]
- Leonard, F.; Quinn, P.J.; Ellis, W.A. Possible effect of pH on the survival of leptospires in cattle urine. Vet. Rec. 1992, 131, 53–54. [Google Scholar] [CrossRef] [PubMed]
- Delooz, L.; Czaplicki, G.; Gregoire, F.; Dal Pozzo, F.; Pez, F.; Kodjo, A.; Saegerman, C. Serogroups and genotypes of Leptospira spp. strains from bovine aborted foetuses. Transbound Emerg. Dis. 2018, 65, 158–165. [Google Scholar] [CrossRef] [PubMed]
- Bertelloni, F.; Cilia, G.; Turchi, B.; Pinzauti, P.; Cerri, D.; Fratini, F. Epidemiology of leptospirosis in North-Central Italy: Fifteen years of serological data (2002–2016). Comp. Immunol. Microbiol. Infect. Dis. 2019, 65, 14–22. [Google Scholar] [CrossRef] [PubMed]
- World Organization for Animal Health. Brucellosis (Brucella abortus, B. melitensis, B. suis) (infection with B. abortus, B. melitensis and B. suis). Manual of Diagnostic Tests and Vaccines for Terrestrial Animals; Chapter 2.1.4; Office International des Epizooties: Paris, France, 2017; Available online: http://www.oie.int/en/international-standard-setting/terrestrial-manual/access-online/ (accessed on 10 May 2020).
- World Organization for Animal Health. Leptospirosis. Manual of Diagnostic Tests and Vaccines for Terrestrial Animals; Chapter 3.1.12; Office International des Epizooties: Paris, France, 2018; pp. 503–516. Available online: https://www.oie.int/fileadmin/Home/eng/Health_standards/tahm/3.01.12_LEPTO.pdf (accessed on 10 May 2020).
- Stoddard, R.A.; Gee, J.E.; Wilkins, P.P.; McCaustland, K.; Hoffmaster, A.R. Detection of pathogenic Leptospira spp. through TaqMan polymerase chain reaction targeting the LipL32 gene. Diagn. Microbiol. Infect. Dis. 2009, 64, 247–255. [Google Scholar] [CrossRef] [PubMed]
- Bertasio, C.; Papetti, A.; Scaltriti, E.; Tagliabue, S.; D’Incau, M.; Boniotti, M.B. Serological Survey and Molecular Typing Reveal New Leptospira Serogroup Pomona Strains among Pigs of Northern Italy. Pathogens 2020, 9, 332. [Google Scholar] [CrossRef]
- Boonsilp, S.; Thaipadungpanit, J.; Amornchai, P.; Wuthiekanun, V.; Bailey, M.S.; Holden, M.T.; Zhang, C.; Jiang, X.; Koizumi, N.; Taylor, K.; et al. A single multilocus sequence typing (MLST) scheme for seven pathogenic Leptospira species. PLoS Negl. Trop. Dis. 2013, 7, e1954. [Google Scholar] [CrossRef] [Green Version]
- Salaün, L.; Mérien, F.; Gurianova, S.; Baranton, G.; Picardeau, M. Application of multilocus variable-number tandem-repeat analysis for molecular typing of the agent of leptospirosis. J. Clin. Microbiol. 2006, 44, 3954–3962. [Google Scholar] [CrossRef] [Green Version]
- Weiss, S.; Menezes, A.; Woods, K.; Chanthongthip, A.; Dittrich, S.; Opoku-Boateng, A.; Kimuli, M.; Chalker, V. An Extended Multilocus Sequence Typing (MLST) Scheme for Rapid Direct Typing of Leptospira from Clinical Samples. PLoS Negl. Trop. Dis. 2016, 10, e0004996. [Google Scholar] [CrossRef]
- Saitou, N.; Nei, M. The neighbor-joining method: A new method for reconstructing phylogenetic trees. Mol. Biol. Evol. 1987, 4, 406–425. [Google Scholar]
- Felsenstein, J. Confidence limits on phylogenies: An approach using the bootstrap. Evolution 1985, 39, 783–791. [Google Scholar] [CrossRef]
- Tamura, K.; Nei, M.; Kumar, S. Prospects for inferring very large phylogenies by using the neighbor-joining method. Proc. Natl. Acad. Sci. USA 2004, 101, 11030–11035. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kumar, S.; Stecher, G.; Li, M.; Knyaz, C.; Tamura, K. MEGA X: Molecular Evolutionary Genetics Analysis across computing platforms. Mol. Biol. Evol. 2018, 35, 1547–1549. [Google Scholar] [CrossRef] [PubMed]
- Hovingh, E. Abortions in Dairy Cattle-I Common Causes of Abortions. Va. Coop. Ext. 2009, 404-288. Available online: https://vtechworks.lib.vt.edu/bitstream/handle/10919/48400/404-288_pdf.pdf?sequence=1&isAllowed=y (accessed on 10 May 2020).
- Garcia-Ispierto, I.; Tutusaus, J.; López-Gatius, F. Does Coxiella burnetii affect reproduction in cattle? A clinical update. Reprod. Domest. Anim. 2014, 49, 529–535. [Google Scholar] [CrossRef]
- Angelakis, E.; Raoult, D. Q fever. Vet. Microbiol. 2010, 140, 297–309. [Google Scholar] [CrossRef] [Green Version]
- Levett, P.N. Leptospirosis. Clin. Microbiol Rev. 2001, 14, 296–326. [Google Scholar] [CrossRef] [Green Version]
- Faine, S.; Adler, B.; Bolin, C.; Perolat, P. Leptospira and Leptospirosis, 2nd ed.; MediSci: Melbourne, Australia, 1999. [Google Scholar]
- Guedes, I.B.; de Souza, G.O.; de Paula Castro, J.F.; Cavalini, M.B.; de Souza Filho, A.F.; Maia, A.L.P.; Dos Reis, E.A.; Cortez, A.; Heinemann, M.B. Leptospira interrogans serogroup Pomona strains isolated from river buffaloes. Trop. Anim. Health Prod. 2021, 53, 194. [Google Scholar] [CrossRef]
- Mackintosh, C.G.; Blackmore, D.K.; Marshal, R.B. Isolation of Leptospira interrogans serovars Tarassovi and Pomona from dogs. N. Z. Vet. J. 1980, 28, 100. [Google Scholar] [CrossRef]
- Zarantonelli, L.; Suanes, A.; Meny, P.; Buroni, F.; Nieves, C.; Salaberry, X.; Briano, C.; Ashfield, N.; Da Silva Silveira, C.; Dutra, F.; et al. Isolation of pathogenic Leptospira strains from naturally infected cattle in Uruguay reveals high serovar diversity, and uncovers a relevant risk for human leptospirosis. PLoS Negl. Trop. Dis. 2018, 12, e0006694. [Google Scholar] [CrossRef] [Green Version]
- Hamond, C.; Silveira, C.S.; Buroni, F.; Suanes, A.; Nieves, C.; Salaberry, X.; Aráoz, V.; Costa, R.A.; Rivero, R.; Giannitti, F.; et al. Leptospira interrogans serogroup Pomona serovar Kennewicki infection in two sheep flocks with acute leptospirosis in Uruguay. Transbound. Emerg. Diseases. 2019, 66, 1186–1194. [Google Scholar] [CrossRef]
- Rocha, B.R.; Martins, G.; Lilenbaum, W. An historical view of the experimental leptospiral infection in ruminants. Comp. Immunol. Microbiol. Infect. Dis. 2020, 73, 101532. [Google Scholar] [CrossRef] [PubMed]
- Vitale, M.; Vitale, F.; Di Marco, V.; Curró, V.; Vesco, G.; Caracappa, S. Polymerase chain reaction method for leptospirosis, analysis on samples from an autochthon swine population in Sicily, Italy. Rev. Cubana Med. Trop. 2005, 57, 25–27. [Google Scholar] [PubMed]
- Cilia, G.; Bertelloni, F.; Albini, S.; Fratini, F. Insight into the Epidemiology of Leptospirosis: A Review of Leptospira Isolations from “Unconventional” Hosts. Animals 2021, 11, 191. [Google Scholar] [CrossRef]
- Żmudzki, J.; Arent, Z.; Jabłoński, A.; Nowak, A.; Zębek, S.; Stolarek, A.; Bocian, Ł.; Brzana, A.; Pejsak, Z. Seroprevalence of 12 serovars of pathogenic Leptospira in red foxes (Vulpes vulpes) in Poland. Acta Vet. Scand. 2018, 60, 34. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Slavica, A.; Cvetnić, Ž.; Milas, Z.; Janicki, Z.; Turk, N.; Konjević, D.; Severin, K.; Tončić, J.; Lipej, Z. Incidence of leptospiral antibodies in different game species over a 10-year period (1996–2005) in Croatia. Eur. J. Wildl. Res. 2008, 54, 305–311. [Google Scholar] [CrossRef]

| Infectious Agent | ELISA Test Kit | Manufacturer | Antigens | Se * | Sp * |
|---|---|---|---|---|---|
| Neospora caninum | ID Screen® Neospora caninum Indirect Multi-species | ID.vet Innovative Diagnostics, Grabels, France | Purified extract of Neospora caninum | 100% (CI95%: 98.8–100%) | 100% (CI95%: 99.41–100%) |
| Coxiella burnetii | ID Screen® Q fever indirect Multi-species | ID.vet Innovative Diagnostics, Grabels, France | phase I and phase II antigens Coxiella burnetii | 100% (CI95%: 89.28–100%) | 100% (CI95%: 97.75–100%) |
| Infectious Bovine Rhinotracheitis Virus | Infectious Bovine Rhinotracheitis Virus (BHV1) gB Antibody Test Kit | IDEXX Laboratories, Inc. | Bovine herpesvirus-1 specific Glycoprotein-B (gB) | 99.6% (CI95% 98.0–99.9%) | 100.0% (CI95% 99.5–100.0%) |
| Infectious Bovine Rhinotracheitis Virus (BHV-1) gE Antibody Test Kit | Bovine herpesvirus-1 specific Glycoprotein-E (gE) | ||||
| Bovine Viral Diarrhoea | Bovine Viral Diarrhoea Virus (BVDV) Antigen Test Kit/Serum Plus | IDEXX Laboratories, Inc. | Bovine Viral Diarrhoea Virus E-antigen | 95.9% (CI95% 92.3–97.9%) | 100% (CI95% 97.7–100%) |
| Oligonucleotide | Target | Sequence 5′–3′ | Refererence |
|---|---|---|---|
| LipL32-45F | lipL32 | 5′-AAGCATTACCGCTTGTGGTG-3′ | [37] |
| LipL32-286R | lipL32 | 5′-GAACTCCCATTTCAGCGATT-3′ | |
| LipL32-189P | lipL32 | FAM-5′-AAAGCCAGGACAAGCGCCG-3′-BHQ1 |
| ID | T0 | ANTIB | T1 | T2 | |||||
|---|---|---|---|---|---|---|---|---|---|
| Serogroup | Titer | Serogroup | Titer | ↑↓ | Serogroup | Titer | ↑↓ | ||
| 1A | Australis Icterohem. Pomona | 1:400 1:400 1:200 | Yes | Pomona | 1:6400 | ↑ | Pomona | 1:1600 | ↓ |
| 2A | Icterohem. Pomona | 1:100 1:100 | Yes | Pomona | 1:3200 | ↑ | Pomona | 1:1600 | ↓ |
| 3A | Pomona | 1:100 | Yes | Pomona | 1:800 | ↑ | Pomona | 1:400 | ↓ |
| 4A | Pomona | 1:100 | Yes | Pomona | 1:1600 | ↑ | Pomona | 1:1600 | = |
| 5A | Australis Icterohem. Pomona | 1:100 1:400 1:400 | Yes | Pomona | 1:800 | ↑ | Pomona | 1:3200 | ↑ |
| 6A | Icterohem. Pomona | 1:100 1:200 | Yes | Pomona | 1:400 | ↑ | Neg | ↓ | |
| 7A | Icterohem. | 1:200 | Yes | Pomona | 1:100 | I | Pomona | 1:6400 | ↑ |
| 8A | Tarassovi | 1:200 | Yes | Pomona | 1:200 | I | Pomona | 1:200 | = |
| 9A | Australis Icterohem. Pomona | 1:200 1:200 1:400 | Yes | Pomona | 1:6400 | ↑ | Pomona | 1:3200 | ↓ |
| 10A | Pomona | 1:200 | Yes | Pomona | 1:100 | ↓ | Neg | ↓ | |
| 11A | Icterohem. | 1:400 | Yes | Neg | Pomona | 1:800 | ↑ | ||
| 12A | Neg | Pomona | 1:6400 | I | Pomona | 1:400 | ↓ | ||
| 13A | Neg | Pomona | 1:1600 | I | Pomona | 1:800 | ↓ | ||
| 14A | Neg | Pomona | 1:200 | I | Pomona | 1:800 | ↑ | ||
| 15A | Neg | Pomona | 1:100 | I | Pomona | 1:1600 | ↑ | ||
| 16A | Neg | Pomona | 1:6400 | I | Pomona | 1:3200 | ↓ | ||
| 17A | Neg | Pomona | 1:800 | I | Pomona | 1:200 | ↓ | ||
| 18A | Neg | Pomona | 1:200 | I | Pomona | 1:400 | ↑ | ||
| 19A | Neg | Neg | = | Neg | = | ||||
| 20A | Neg | Neg | = | Neg | = | ||||
| 21A | Neg | Dead | |||||||
| 22A | Neg | Pomona | 1:200 | I | Neg | ↓ | |||
| 23A | Neg | Pomona | 1:100 | I | Pomona | 1:100 | = | ||
| 24 M | Neg. | Neg. | = | Neg | = | ||||
| 25A | Neg | Pomona | 1:200 | I | Neg | ↓ | |||
| 26A | Neg | Pomona | 1:800 | I | Pomona | 1:200 | ↓ | ||
| 27A | Neg | Pomona | 1:800 | I | Pomona | 1:800 | = | ||
| 28A | Neg | Pomona | 1:100 | I | Neg | ↓ | |||
| 29A | Neg | Pomona | 1:800 | I | Pomona | 1:800 | = | ||
| 30A | Neg | Pomona | 1:6400 | I | Pomona | 1:800 | ↓ | ||
| 31A | Neg | Neg | = | Neg | = | ||||
| 32A | Neg | Neg | = | Pomona | 1:200 | ↑ | |||
| 33A | Neg | Neg | = | Neg | = | ||||
| ID | T0 | T1 | ↑↓ | ||
|---|---|---|---|---|---|
| Serogroup | Titer | Serogroup | Titer | ↑ | |
| 1B | Australis Icterohaem Sejroe Pomona | 1:400 1:800 1:100 1:3200 | Pomona | 1:6400 | |
| 2B | Pomona | 1:1600 | Pomona | 1:800 | ↓ |
| 3B | Pomona Icterohaem Sejroe | 1:400 1:400 1:100 | Pomona | 1:800 | ↑ |
| 4B | Pomona | 1:800 | Pomona | 1:800 | ↓ |
| 5B | Sejroe | 1:100 | Neg | ↓ | |
| 6B | Neg | Pomona | 1:200 | I | |
| 7B | Neg | Pomona | 1:200 | I | |
| Farm | ID | Serogroup | Titer |
|---|---|---|---|
| C | 1C | Ballum | 1:100 |
| C | 2C | Pomona | 1:200 |
| C | 3C | Pomona | 1:800 |
| D | 1D | Ballum | 1:100 |
| E | 1E | Sejroe | 1:200 |
| F | 1F | Grippotyp/Sejroe/Tarassovi | 1:400/1:200/1:100 |
| F | 2F | Grippotyp/Sejroe/Tarassovi | 1:100/1:200/1:400 |
| F | 3F | Grippotyp/Sejroe/Tarassovi/Pomona | 1:400/1:100/1:100/1:400 |
| F | 4F | Grippotyp/Sejroe | 1:400/1:800 |
| F | 5F | Grippotyp/Tarassovi | 1:200/1:400 |
| F | 6F | Grippotyp/Sejroe | 1:400/1:400 |
| F | 7F | Grippotyp/Sejroe/Tarassovi | 1:100/1:200/1:100 |
| F | 8F | Grippotyp/Sejroe/Tarassovi | 1:1600/1:1600/1:800 |
| F | 9F | Grippotyp/Sejroe | 1:400/1:400 |
| F | 10F | Grippotyp | 1:800 |
| F | 11F | Grippotyp/Tarassovi/Pomona | 1:100/1:200/1:800 |
| F | 12F | Grippotyp/Tarassovi | 1:800/1:100 |
| F | 13F | Pomona/Sejroe/Tarassovi | 1:400/1:200/1:400 |
| F | 14F | Pomona/Sejroe | 1:100/1:400 |
| F | 15F, 23F | Pomona | 1:400 |
| F | 16F | Tarassovi/Pomona | 1:400/1:200 |
| F | 17F, 22F | Pomona | 1:800 |
| F | 18F | Pomona/Sejroe | 1:400/1:200 |
| F | 19F | Pomona | 1:1600 |
| F | 20F | Pomona/Sejroe | 1:200/1:100 |
| F | 21F | Tarassovi/Pomona | 1:200/1:3200 |
| F | 24F, 26F | Sejroe | 1:100 |
| F | 25F, 30F, 32F, 33F | Sejroe | 1:200 |
| F | 27F | Tarassovi/Sejroe | 1:400/1:400 |
| F | 28F, 31F | Sejroe | 1:400 |
| F | 29F | Tarassovi/Sejroe | 1:200/1:400 |
| F | 34F, 37F, 40F | Tarassovi | 1:400 |
| F | 35F, 38F, 41F, 42F, 43F, 44F | Tarassovi | 1:200 |
| F | 36F, 39F | Tarassovi | 1:100 |
| Farm | ID | Real Time-PCR | MLST | Isolation |
|---|---|---|---|---|
| A | 1A | Pos | Pos | Pos |
| A | 2A | Pos | / | Neg |
| A | 3A | Pos | Pos | Pos |
| A | 4A | Pos | / | Neg |
| A | 5A | Pos | Pos | Neg |
| A | 6A | Pos | Pos | Neg |
| A | 7A | Pos | / | Neg |
| A | 8A | Pos | Pos | Neg |
| A | 9A | Pos | Pos | Neg |
| A | 10A | Pos | Pos | Neg |
| A | 11A | Pos | / | Neg |
| B | 1B | Neg | / | Pos |
| B | 2B | Neg | / | Neg |
| B | 3B | Neg | / | Neg |
| B | 4B | Neg | / | Neg |
| B | 5B | Neg | / | Neg |
| C | 1C | Neg | / | Pos |
| C | 2C | / | / | Neg |
| C | 3C | Neg | / | Neg |
| D | 1D | Neg | / | Cs |
| E | 1E | Neg | / | Cs |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Aliberti, A.; Blanda, V.; Di Marco Lo Presti, V.; Macaluso, G.; Galluzzo, P.; Bertasio, C.; Sciacca, C.; Arcuri, F.; D’Agostino, R.; Ippolito, D.; et al. Leptospira interrogans Serogroup Pomona in a Dairy Cattle Farm in a Multi-Host Zootechnical System. Vet. Sci. 2022, 9, 83. https://doi.org/10.3390/vetsci9020083
Aliberti A, Blanda V, Di Marco Lo Presti V, Macaluso G, Galluzzo P, Bertasio C, Sciacca C, Arcuri F, D’Agostino R, Ippolito D, et al. Leptospira interrogans Serogroup Pomona in a Dairy Cattle Farm in a Multi-Host Zootechnical System. Veterinary Sciences. 2022; 9(2):83. https://doi.org/10.3390/vetsci9020083
Chicago/Turabian StyleAliberti, Antonino, Valeria Blanda, Vincenzo Di Marco Lo Presti, Giusi Macaluso, Paola Galluzzo, Cristina Bertasio, Carmela Sciacca, Francesca Arcuri, Rosalia D’Agostino, Dorotea Ippolito, and et al. 2022. "Leptospira interrogans Serogroup Pomona in a Dairy Cattle Farm in a Multi-Host Zootechnical System" Veterinary Sciences 9, no. 2: 83. https://doi.org/10.3390/vetsci9020083







