Molecular Identification of Parasitic Protozoa Sarcocystis in Water Samples
Abstract
:Simple Summary
Abstract
1. Introduction
2. Materials and Methods
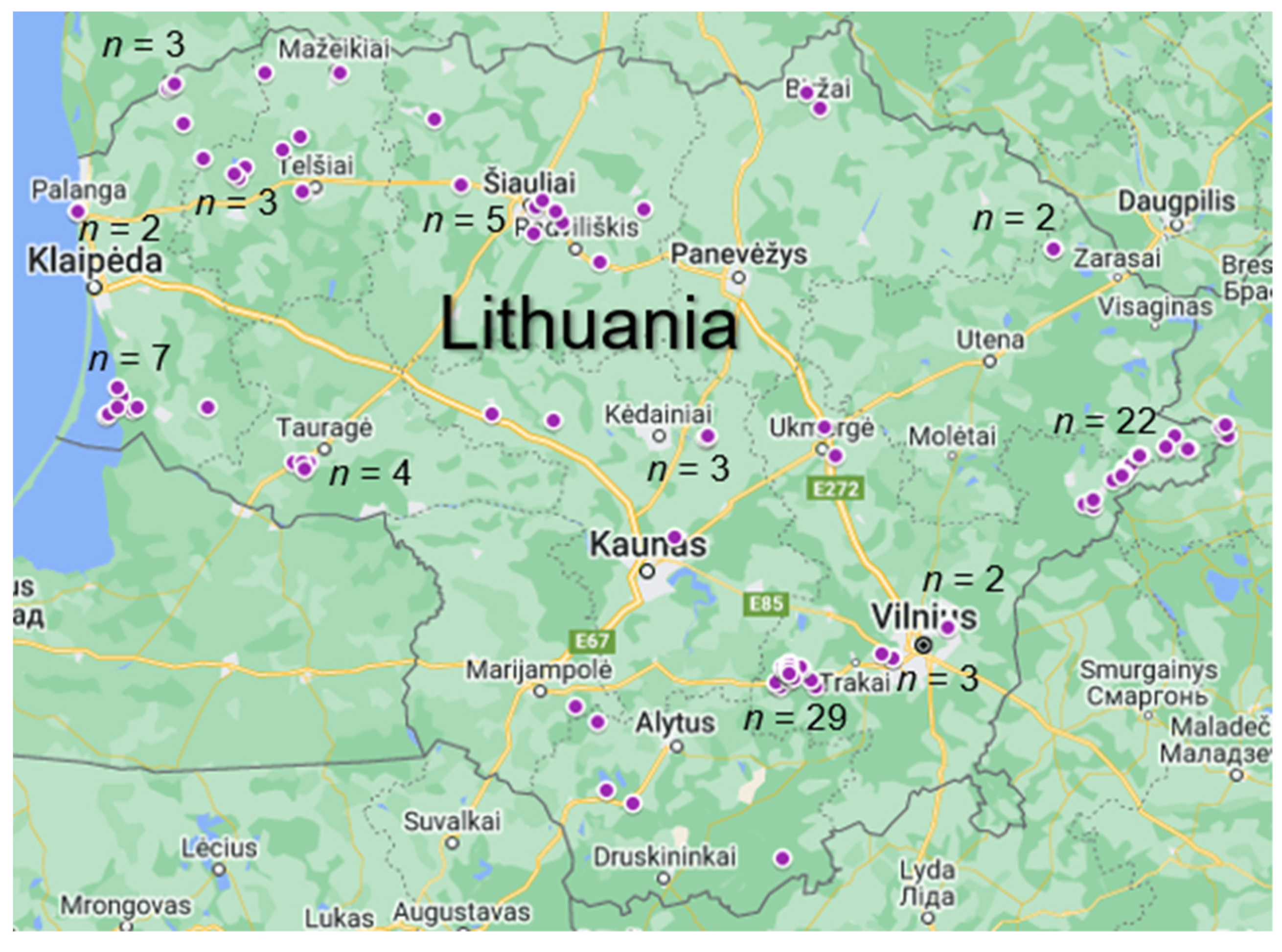
2.1. Sample Collection
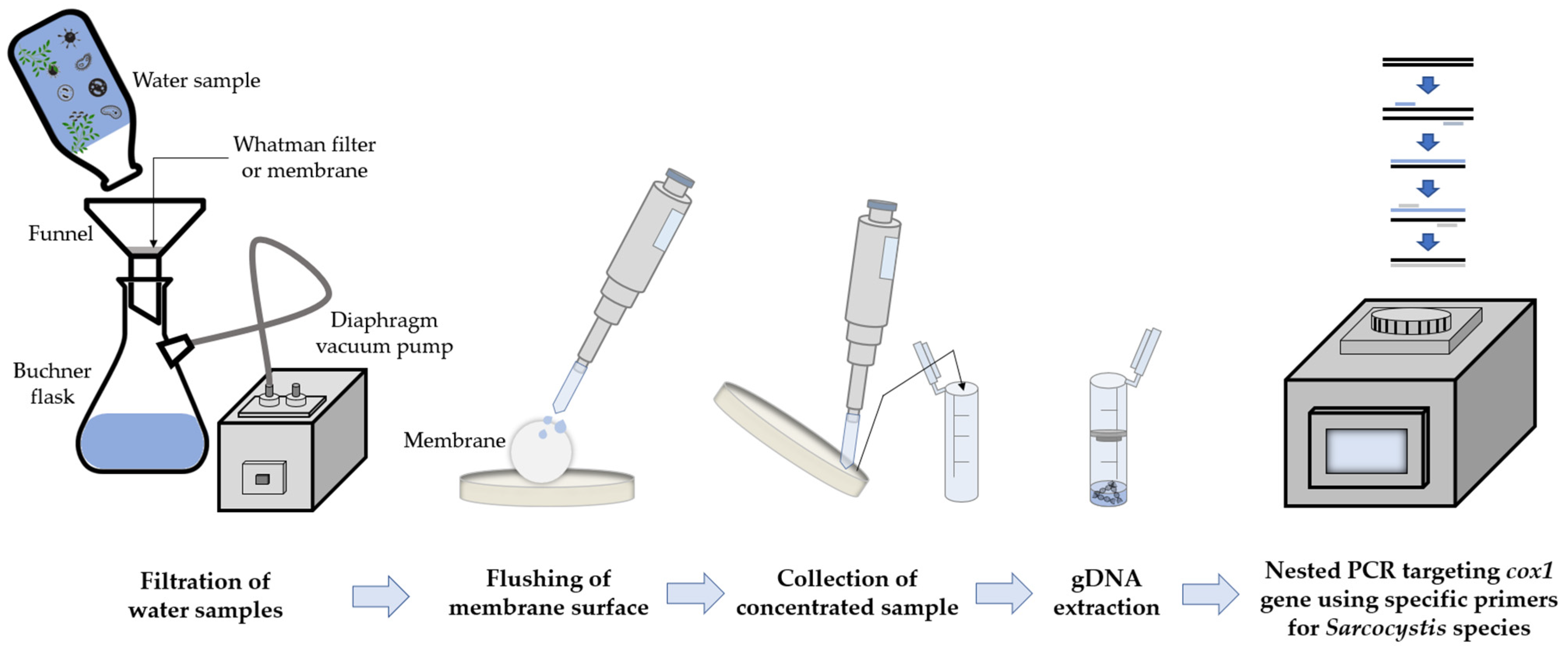
2.2. Isolation of Sporocysts and Genomic DNA Extraction
2.3. Identification of Sarcocystis Species Using PCR Procedures
2.4. Statistical Analysis
3. Results
3.1. Selection of PCR Conditions
3.2. Identification of Sarcocystis spp. from in Water Bodies
3.3. Summary of Molecular-Based Sarcocystis Identification from Water Samples
3.4. Sarcocystis spp. Occurrence Rates in the Analyzed Water Samples
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Dubey, J.P.; Calero-Bernal, R.; Rosenthal, B.M.; Speer, C.A.; Fayer, R. Sarcocystosis of Animals and Humans, 2nd ed.; CRC Press: Boca Raton, FL, USA, 2016. [Google Scholar]
- Florin-Christensen, M.; Schnittger, L. Sarcocystis. In Parasitic Protozoa of Farm Animals and Pets; Florin-Christensen, M., Schnittger, L., Eds.; Springer Science and Business Media LLC: Berlin, Germany, 2018; pp. 103–124. [Google Scholar]
- Fayer, R. Sarcocystis spp. in Human Infections. Clin. Microbiol. Rev. 2004, 17, 894–902. [Google Scholar]
- Fayer, R.; Esposito, D.H.; Dubey, J.P. Human infections with Sarcocystis species. Clin. Microbiol. Rev. 2015, 28, 295–311. [Google Scholar] [CrossRef] [Green Version]
- Basso, W.; Alvarez Rojas, C.A.; Buob, D.; Ruetten, M.; Deplazes, P. Sarcocystis infection in red deer (Cervus elaphus) with eosinophilic myositis/fasciitis in Switzerland and involvement of red foxes (Vulpes vulpes) and hunting dogs in the transmission. Int. J. Parasitol. Parasites Wildl. 2020, 13, 130–141. [Google Scholar] [CrossRef]
- Juozaitytė-Ngugu, E.; Švažas, S.; Šneideris, D.; Rudaitytė-Lukošienė, E.; Butkauskas, D.; Prakas, P. The role of birds of the family corvidae in transmitting Sarcocystis protozoan parasites. Animals 2021, 11, 3258. [Google Scholar] [CrossRef]
- Shahari, S.; Tengku-Idris, T.I.N.; Fong, M.Y.; Lau, Y.L. Molecular evidence of Sarcocystis nesbitti in water samples of Tioman Island, Malaysia. Parasites Vectors 2016, 9, 598. [Google Scholar] [CrossRef] [Green Version]
- Lee, F.C.H. Finding Sarcocystis spp. on the Tioman island: 28S rRNA gene next-generation sequencing reveals nine new Sarcocystis species. J. Water Health 2019, 17, 416–427. [Google Scholar] [CrossRef] [Green Version]
- Verma, S.K.; Lindsay, D.S.; Grigg, M.E.; Dubey, J.P. Isolation, culture and cryopreservation of Sarcocystis species. Curr. Protoc. Microbiol. 2017, 45, 11–127. [Google Scholar] [CrossRef] [Green Version]
- Ma, C.L.; Ye, Y.L.; Wen, T.; Huang, Z.M.; Pan, J.; Hu, J.J.; Tao, J.P.; Song, J.L. Prevalence and morphological and molecular characteristics of Sarcocystis bertrami in horses in China. Parasite 2020, 27, 1. [Google Scholar] [CrossRef]
- Prakas, P.; Strazdaitė-Žielienė, Ž.; Januškevičius, V.; Chiesa, F.; Baranauskaitė, A.; Rudaitytė-Lukošienė, E.; Servienė, E.; Petkevičius, S.; Butkauskas, D. Molecular identification of four Sarcocystis species in cattle from Lithuania, including S. hominis, and development of a rapid molecular detection method. Parasites Vectors 2020, 13, 610. [Google Scholar] [CrossRef]
- Cheun, H.I.; Kim, C.H.; Cho, S.H.; Ma, D.W.; Goo, B.L.; Na, M.S.; Youn, S.K.; Lee, W.J. T The first outbreak of Giardiasis with drinking water in Korea. Osong Public Health Res. Perspect. 2013, 4, 89–92. [Google Scholar] [CrossRef] [Green Version]
- Hassan, E.M.; Örmeci, B.; DeRosa, M.C.; Dixon, B.R.; Sattar, S.A.; Iqbal, A. A review of Cryptosporidium spp. and their detection in water. Water Sci. Technol. 2021, 83, 1–25. [Google Scholar] [CrossRef]
- Januskevicius, V.; Januskeviciene, G.; Prakas, P.; Butkauskas, D.; Petkevicius, S. Prevalence and intensity of Sarcocystis spp. infection in animals slaughtered for food in Lithuania. Vet. Med. Czech 2019, 64, 149–157. [Google Scholar] [CrossRef]
- Gjerde, B. Phylogenetic relationships among Sarcocystis species in cervids, cattle and sheep inferred from the mitochondrial cytochrome c oxidase subunit I gene. Int. J. Parasitol. 2013, 43, 579–591. [Google Scholar] [CrossRef]
- Gjerde, B. Sarcocystis species in red deer revisited: With a re-description of two known species as Sarcocystis elongata n. sp. and Sarcocystis truncata n. sp. based on mitochondrial cox1 sequences. Parasitology 2014, 141, 441–452. [Google Scholar] [CrossRef]
- Prakas, P.; Rudaitytė, E.; Butkauskas, D.; Kutkienė, L. Sarcocystis entzerothi n. sp. from the European roe deer (Capreolus capreolus). Parasitol. Res. 2017, 116, 271–279. [Google Scholar] [CrossRef]
- Prakas, P.; Rehbein, S.; Rudaitytė-Lukošienė, E.; Butkauskas, D. Molecular identification of Sarcocystis species in diaphragm muscle tissue of European mouflon (Ovis Gmelini Musimon) from Austria. Parasitol. Res. 2021, 120, 2695–2702. [Google Scholar] [CrossRef]
- Prakas, P.; Balčiauskas, L.; Juozaitytė-Ngugu, E.; Butkauskas, D. The role of mustelids in the transmission of Sarcocystis spp. using cattle as intermediate hosts. Animals 2021, 11, 822. [Google Scholar] [CrossRef]
- Untergasser, A.; Nijveen, H.; Rao, X.; Bisseling, T.; Geurts, R.; Leunissen, J.A.M. Primer3Plus, an enhanced web interface to Primer3. Nucleic Acids Res. 2007, 35, W71–W74. [Google Scholar] [CrossRef] [Green Version]
- Rózsa, L.; Reiczigel, J.; Majoros, G. Quantifying parasites in samples of hosts. J. Parasitol. 2000, 86, 228–232. [Google Scholar] [CrossRef]
- Reiczigel, J. Confidence intervals for the binomial parameter: Some new considerations. Stat. Med. 2003, 22, 611–621. [Google Scholar] [CrossRef]
- Omarova, A.; Tussupova, K.; Berndtsson, R.; Kalishev, M.; Sharapatova, K. Protozoan parasites in drinking water: A system approach for improved water, sanitation and hygiene in developing countries. Int. J. Environ. Res. Public Health 2018, 15, 495. [Google Scholar] [CrossRef] [Green Version]
- Moreno, Y.; Moreno-Mesonero, L.; Amorós, I.; Pérez, R.; Morillo, J.A.; Alonso, J.L. Multiple identification of most important waterborne protozoa in surface water used for irrigation purposes by 18S rRNA amplicon-based metagenomics. Int. J. Hyg. Environ. Health 2018, 221, 102–111. [Google Scholar] [CrossRef]
- Prystajecky, N.; Huck, P.M.; Schreier, H.; Isaac-Renton, J.L. Assessment of Giardia and Cryptosporidium spp. as a microbial source tracking tool for surface water: Application in a mixed-use watershed. Appl. Environ. Microbiol. 2014, 80, 2328–2336. [Google Scholar] [CrossRef] [Green Version]
- Schets, F.M.; Engels, G.B.; During, M.; De Roda Husman, A.M. Detection of infectious Cryptosporidium oocysts by cell culture immunofluorescence assay: Applicability to environmental samples. Appl. Environ. Microbiol. 2005, 71, 6793–6798. [Google Scholar] [CrossRef] [Green Version]
- Doležel, D.; Koudela, B.; Jirků, M.; Hypša, V.; Oborník, M.; Votýpka, J.; Modrý, D.; Šlapeta, J.R.; Lukeš, J. Phylogenetic analysis of Sarcoystis spp. of mammals and reptiles supports the coevolution of Sarcocystis spp. with their final hosts. Int. J. Parasitol. 1999, 29, 795–798. [Google Scholar] [CrossRef]
- Lee, F.C.H.; Muthu, V. From 18S to 28S rRNA gene: An improved targeted Sarcocystidae PCR amplification, species identification with long DNA sequences. Am. J. Trop. Med. Hyg. 2021, 104, 1388–1393. [Google Scholar] [CrossRef]
- Gjerde, B. Molecular characterisation of Sarcocystis bovifelis, Sarcocystis bovini n. sp., Sarcocystis hirsuta and Sarcocystis cruzi from cattle (Bos taurus) and Sarcocystis sinensis from water buffaloes (Bubalus bubalis). Parasitol. Res. 2016, 115, 1473–1492. [Google Scholar] [CrossRef]
- Gjerde, B.; de la Fuente, C.; Alunda, J.M.; Luzón, M. Molecular characterisation of five Sarcocystis species in domestic sheep (Ovis aries) from Spain. Parasitol. Res. 2020, 119, 2145–2150. [Google Scholar] [CrossRef]
- Prakas, P.; Butkauskas, D.; Rudaitytė, E.; Kutkienė, L.; Sruoga, A.; Pūraitė, I. Morphological and molecular characterization of Sarcocystis taeniata and Sarcocystis pilosa n. sp. from the sika deer (Cervus nippon) in Lithuania. Parasitol. Res. 2016, 115, 3021–3032. [Google Scholar] [CrossRef]
- Huang, Z.; Ye, Y.; Zhang, H.; Deng, S.; Tao, J.; Hu, J.; Yang, Y. Morphological and molecular characterizations of Sarcocystis miescheriana and Sarcocystis suihominis in domestic pigs (Sus scrofa) in China. Parasitol. Res. 2019, 118, 3491–3496. [Google Scholar] [CrossRef]
- Metwally, D.M.; Al-Damigh, M.A.; Al-Turaiki, I.M.; El-Khadragy, M.F. Molecular characterization of Sarcocystis species isolated from sheep and goats in Riyadh, Saudi Arabia. Animals 2019, 9, 256. [Google Scholar] [CrossRef] [Green Version]
- El-Morsey, A.; Abdo, W.; Sultan, K.; Elhawary, N.M.; AbouZaid, A.A. Ultrastructural and molecular identification of the sarcocysts of Sarcocystis tenella and Sarcocystis arieticanis infecting domestic sheep (Ovis aries) from Egypt. Acta Parasitol. 2019, 64, 501–513. [Google Scholar] [CrossRef]
- Odening, K.; Wesemeier, H.H.; Walter, G.; Bockhardt, I. The wisent (Bison Bonasus, Bovidae) as an intermediate host of three Sarcocystis species (Apicomplexa: Sarcocystidae) of cattle. Folia Parasitol. 1994, 41, 115–121. [Google Scholar]
- Cabaj, W.; Grzelak, S.; Moskwa, B.; Bień-Kalinowska, J. Sarcocystis cruzi infection in free-living European bison (Bison bonasus bonasus L.) from the Białowieża forest, Poland—A molecular analysis based on the cox1 gene. Int. J. Parasitol. Parasites Wildl. 2021, 16, 59–63. [Google Scholar] [CrossRef]
- Balčiauskas, L.; Trakimas, G.; Juškaitis, R.; Ulevičius, A.; Balčiauskienė, L. Atlas of Lithuanian Mammals, Amphibians and Reptiles, 2nd ed.; Akstis: Vilnius, Lithuania, 1999. [Google Scholar]

| Species | Primers | Ta, °C | Extension Time, s | Product Size (bp) | Reference | ||
|---|---|---|---|---|---|---|---|
| Name | Orientation | Sequence (5′–3′) | |||||
| Primers for Direct and Nested PCR | |||||||
| Sarcocystis spp. | SF1 | Forward | ATGGCGTACAACAATCATAAAGAA | [15] | |||
| SR8D | Reverse | CATTGCCCATDACTACGCC | 55 | 70 | 1072 | [15] | |
| SR9 | Reverse | ATATCCATACCRCCATTGCCCAT | 60 | 70 | 1085 | [16] | |
| SR12H | Reverse | AAATACCTTGGTGCCCGTAG | 56 | 60 | 952 | [17] | |
| S. bovifelis | GaBfEF | Forward | ATCAACTTCCTAGGTACAGCGGTATT | 56 | 45 | 523 | [11] |
| GaBfER | Reverse | CCACATCATTGGTGCTTAGTCTAGTA | [11] | ||||
| S. cruzi | GaCrEF | Forward | GCTATGTATCTACTTACGGCAGGTATC | 56 | 45 | 608 | [11] |
| GaCrER | Reverse | GAATATAATGGCCCAGGTAAATAATG | [11] | ||||
| S. hirsuta | GaHiEF | Forward | GTTGTGCGGTATGAATTATCAACCT | 56 | 45 | 513 | [11] |
| GaHiER | Reverse | GGTAAGAACTGGAATGGTTAATATCAG | [11] | ||||
| S. arieticanis | GsSariF | Forward | TTCTTGGTATGGCTATTCTTGGACT | 65 | 45 | 586 | [18] |
| GsSariR | Reverse | GATATGTCAATCCAGAGATCGGTAG | [18] | ||||
| S. tenella | GsStenF | Forward | TACTCGGAGCGGTGAACTTCTTA | 63 | 35 | 451 | [18] |
| GsStenR | Reverse | ATAGTCACGGCAGAGAAGTAGGAC | [18] | ||||
| S. capracanis | GsScapF | Forward | AGCGGTAAACTTCCTGGGTACT | 63 | 35 | 467 | [18] |
| GsScapR | Reverse | GCCTATCCAGTTGAATATCTTGGT | [18] | ||||
| Primers for multiplex-nested PCR | |||||||
| 1st round | |||||||
| Sarcocystis spp. | SF1 | Forward | ATGGCGTACAACAATCATAAAGAA | [15] | |||
| S. arieticanis, S. cruzi | SsunR2 | Reverse | GTGCCTCCCAGGCTGAAYAG | 56 | 70 | 1055 | [19] |
| S. bertrami, S. tenella, S. capracanis | SsunR1 | Reverse | GTACCGCCCAGGCTGAAYAG | 56 | 70 | 1055 | [18] |
| S. bovifelis, S. hirsuta | SkatR | Reverse | CAGGCTGAACAGHABTACGA | 56 | 70 | 1042 | [19] |
| S. miescheriana | SmieF | Forward | ACGCTGTATGCACCACTGAG | 56 | 45 | 658 | Present study |
| SmieR | Reverse | CTGAACAGCGCTACAAATGC | |||||
| 2nd round | |||||||
| S. bertrami | GsSberF | Forward | GTATGAACTGTCAACGGATGGAGTA | 65 | 35 | 482 | Present study |
| GsSberR | Reverse | TCAACATTAGCGAGGTAAATACTATC | |||||
| S. miescheriana | GsSmieF | Forward | GTTCCTCGGTATTAGCAGCGTACT | 65 | 35 | 509 | Present study |
| GsSmieR | Reverse | AGTTAAATATTTTAGTGCCCGTTGGA | |||||
| Primers for semi-nested PCR | |||||||
| S. bovifelis | VoboF | Forward | GATCGGTATTACTGTTGCACTCATT | 58 | 45 | 701 | Present study |
| VoboR | Reverse | AGGCCACATCATTGGTGCTTA | |||||
| GaBfEF | Forward | ATCAACTTCCTAGGTACAGCGGTATT | 57 | 35 | 521 | [11] | |
| S. tenella | VoteF1 | Forward | AGCGGTGAACTTCTTAGGAACC | 59 | 35 | 526 | Present study |
| VoteR | Reverse | AATAATCCGCTGTTAACGTATGC | Present study | ||||
| VoteF2 | Forward | CATTGTAATGCTCCTCGACGATA | 59 | 30 | 401 | Present study | |
| S. capracanis | VocaF | Forward | GTAAACTTCCTGGGTACTGTGCTGT | 60 | 35 | 526 | Present study |
| VocaR1 | Reverse | CCAGTAATCCGCTGTCAAGATAC | Present study | ||||
| VocaR2 | Reverse | AGTACCCATCACGGTGCCTATC | 63 | 35 | 500 | Present study | |
| Species-specific primers for nested PCR | |||||||
| S. bovifelis | V2bo1 | Forward | AACTTCCTAGGTACAGCGGTATTCG | 60 | 40 | 556 | Present study |
| V2bo2 | Reverse | TGAACAGCAGTACGAAGGCAAC | Present study | ||||
| V2bo3 | Forward | ATATTTACCGGTGCCGTACTTATGTT | 60 | 30 | 410 | Present study | |
| V2bo4 | Reverse | GCCACATCATTGGTGCTTAGTCT | Present study | ||||
| S. cruzi | V2cr1 | Forward | TACAATGTGCTGTTTACGCTCCA | 57 | 50 | 776 | Present study |
| V2cr2 | Reverse | GCAATCATGATAGTTACGGCAGA | Present study | ||||
| V2cr3 | Forward | ACCATCCTGTTCTGTGGTGCTATG | 65 | 30 | 298 | Present study | |
| V2cr4 | Reverse | AAACTACTTTACTGCCTACGGTACTC | Present study | ||||
| S. hirsuta | V2hi5 | Forward | TATGTTGGTTCTGCCGAAGTCAT | 60 | 45 | 686 | Present study |
| V2hi6 | Reverse | GGTATGGCAATCATTATGGTTACAG | Present study | ||||
| V2hi7 | Forward | GCACCGTAATATTTCAGGGATGT | 60 | 30 | 299 | Present study | |
| V2hi8 | Reverse | AACCTGCTTGCCGGAGTAAGTA | Present study | ||||
| S. arieticanis | V2arie1 | Forward | CTCTTTGCCGTAGATTCGCTAGTTA | 63 | 55 | 884 | Present study |
| V2arie2 | Reverse | CAAAGATCGGTAGATATCCAATGC | Present study | ||||
| V2arie3 | Forward | TAGTTCTTGGCCTGGCTATTCTT | 59 | 25 | 371 | Present study | |
| V2arie4 | Reverse | CTGACCTCCAAAAACTGGCTTAC | Present study | ||||
| S. tenella | V2te1 | Forward | GAGCGGTGAACTTCTTAGGAACC | 60 | 40 | 537 | Present study |
| V2te2 | Reverse | CCCAATAATCCGCTGTTAACGTA | Present study | ||||
| V2te3b | Forward | ATTGTAATGCTCCTCGACGATATG | 57 | 30 | 314 | Present study | |
| V2te4 | Reverse | ATAGTCACGGCAGAGAAGTAGGAC | Present study | ||||
| S. capracanis | VocaF | Forward | GTAAACTTCCTGGGTACTGTGCTGT | 60 | 40 | 531 | Present study |
| VocaR1 | Reverse | CCAGTAATCCGCTGTCAAGATAC | Present study | ||||
| V2cap3 | Forward | ATACCGATCTTTACGGGAGCAGTA | 63 | 30 | 330 | Present study | |
| V2cap4 | Reverse | GGTCACCGCAGAGAAGTACGAT | Present study | ||||
| S. bertrami | V2ber1 | Forward | GTATGAACTGTCAACGGATGGAGTA | 58 | 60 | 883 | Present study |
| V2ber2 | Reverse | AGAAGCCATGTTCGTGACTACC | Present study | ||||
| V2ber3 | Forward | GTACTACCTCCTTCCAGTCGGTTC | 57 | 40 | 600 | Present study | |
| V2ber4 | Reverse | CGGGTATCCACTTCAAGTCCAG | Present study | ||||
| S. miescheriana | V2mie1 | Forward | TGCTGCGGTATGAACTATCTACCT | 61 | 60 | 922 | Present study |
| V2mie2 | Reverse | GCCCAGAGATCCAAATCCAG | Present study | ||||
| V2mie3 | Forward | CTTGGTTCAACGTTACTCCTCCA | 61 | 30 | 474 | Present study | |
| V2mie4 | Reverse | CTTCGATCCAGCTGAACTAAAGC | Present study | ||||
| Method | Sporocysts Isolation | gDNA Extraction | Number of Water Samples (n) | Positive Samples * | Cases | Species Detected **** | |
|---|---|---|---|---|---|---|---|
| Positive ** | False Positive *** | ||||||
| Direct PCR (species-specific primers) | Sucrose gradient | “GeneJET Genomic DNA Purification Kit” | 20 | 0 (0.0%) | 0 | 0 | 0 |
| Nested PCR (species-specific primers only in 2nd round of PCR) | Sucrose gradient | “GeneJET Genomic DNA Purification Kit” | 35 | 8 (22.9%) | 12 | 2 | 2 |
| Sedimentation | “Genomic DNA Purification Kit” | 35 | 3 (8.6%) | 4 | 1 | 2 | |
| Filtration | “GeneJET Genomic DNA Purification Kit” | 35 | 14 (40.0%) | 15 | 1 | 3 | |
| Multiplex-nested PCR (species-specific primers only in 2nd round of PCR) | Filtration | “GeneJET Genomic DNA Purification Kit” | 35 | 7 (20.0%) | 10 | 3 | 2 |
| Semi-nested PCR (species-specific primers) | Filtration | “GeneJET Genomic DNA Purification Kit” | 35 | 0 (0.0%) | 0 | 0 | 0 |
| Nested PCR (species-specific primers in both rounds) | Filtration | “GeneJET Genomic DNA Purification Kit” | 35 | 34 (97.1%) | 84 | 0 | 8 |
| Species | GenBank Acc. No. (Length, bp) | Sequence Similarity, % | ||
|---|---|---|---|---|
| Comparing Obtained Sequences with Each Other | Comparing Obtained Sequences and Those of the Same Species Available in GenBank | Comparing Obtained Sequences with Those of Most Closely Related Species | ||
| S. bovifelis | ON211315-18 (361) | 99.7–100 | 99.5–100 | S. bovini 93.1–94.5 |
| S. cruzi | ON211319-22 (248) | 98.8–100 | 98.0–99.6 | S. alceslatrans 85.1–86.8 |
| S. hirsuta | ON211323-26 (254) | 99.6–100 | 98.4–100 | S. buffalonis 92.8–93.2 |
| S. arieticanis | ON211327-30 (325) | 99.7–100 | 92.3–99.4 | S. hircicanis 86.2–87.4 |
| S. tenella | ON211331-34 (263) | 99.2–100 | 96.2–100 | S. capracanis 91.3–93.2 |
| S. capracanis | ON211335-38 (284) | 99.7–100 | 96.8–99.7 | S. tenella 90.4–92.9 |
| S. bertrami | ON211339-42 (554) | 99.1–99.8 | 97.5–99.8 | S. bovifelis 77.0–78.4 |
| S. miescheriana | ON211343-46 (428) | 99.5–100 | 93.9–99.5 | S. rangiferi 79.6–80.2 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Strazdaitė-Žielienė, Ž.; Baranauskaitė, A.; Butkauskas, D.; Servienė, E.; Prakas, P. Molecular Identification of Parasitic Protozoa Sarcocystis in Water Samples. Vet. Sci. 2022, 9, 412. https://doi.org/10.3390/vetsci9080412
Strazdaitė-Žielienė Ž, Baranauskaitė A, Butkauskas D, Servienė E, Prakas P. Molecular Identification of Parasitic Protozoa Sarcocystis in Water Samples. Veterinary Sciences. 2022; 9(8):412. https://doi.org/10.3390/vetsci9080412
Chicago/Turabian StyleStrazdaitė-Žielienė, Živilė, Agnė Baranauskaitė, Dalius Butkauskas, Elena Servienė, and Petras Prakas. 2022. "Molecular Identification of Parasitic Protozoa Sarcocystis in Water Samples" Veterinary Sciences 9, no. 8: 412. https://doi.org/10.3390/vetsci9080412








