Mating-Type Genes Play an Important Role in Fruiting Body Development in Morchella sextelata
Abstract
:1. Introduction
2. Materials and Methods
2.1. Strains and Culture Conditions
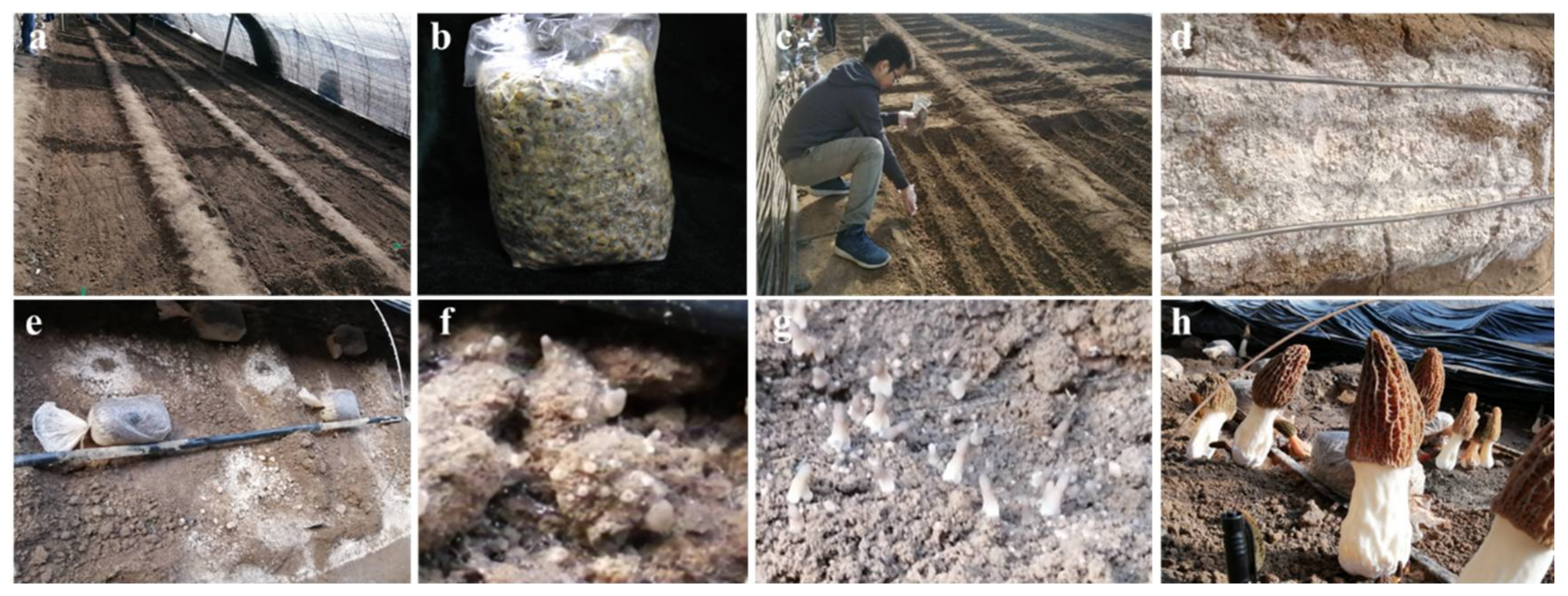
2.2. Fruiting Body Cultivation and Sample Collection
2.3. Morphological Observation and Measurement
2.4. Construction of RNAi Plasmids for mat1-2-1
2.5. Agrobacterium-Mediated Transformation of M. sextelata
2.6. DNA Extraction and PCR
2.7. RNA Extraction and Reverse-Transcription (RT)-PCR
3. Results
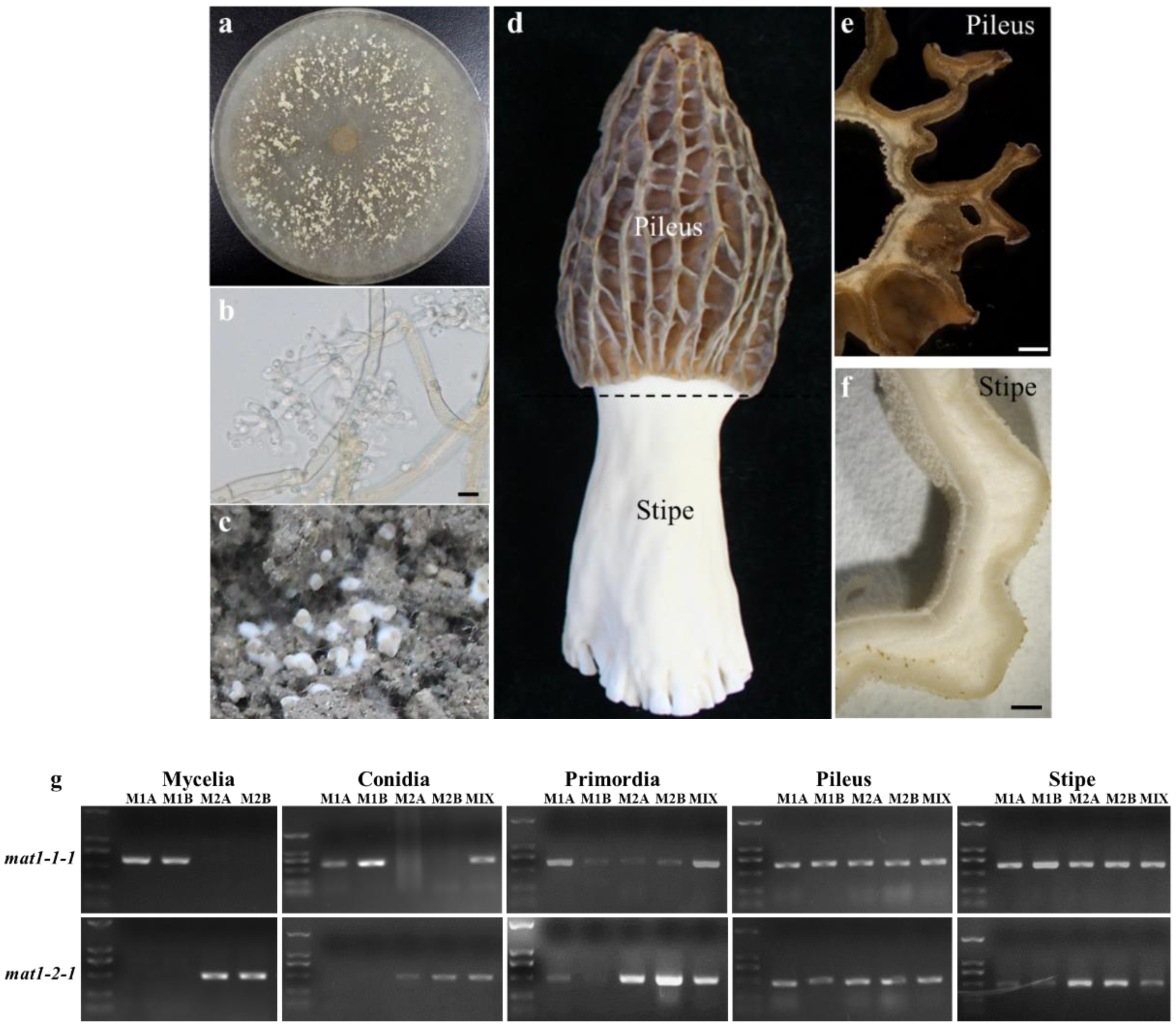
3.1. Mating Type Detection at the Different Stages
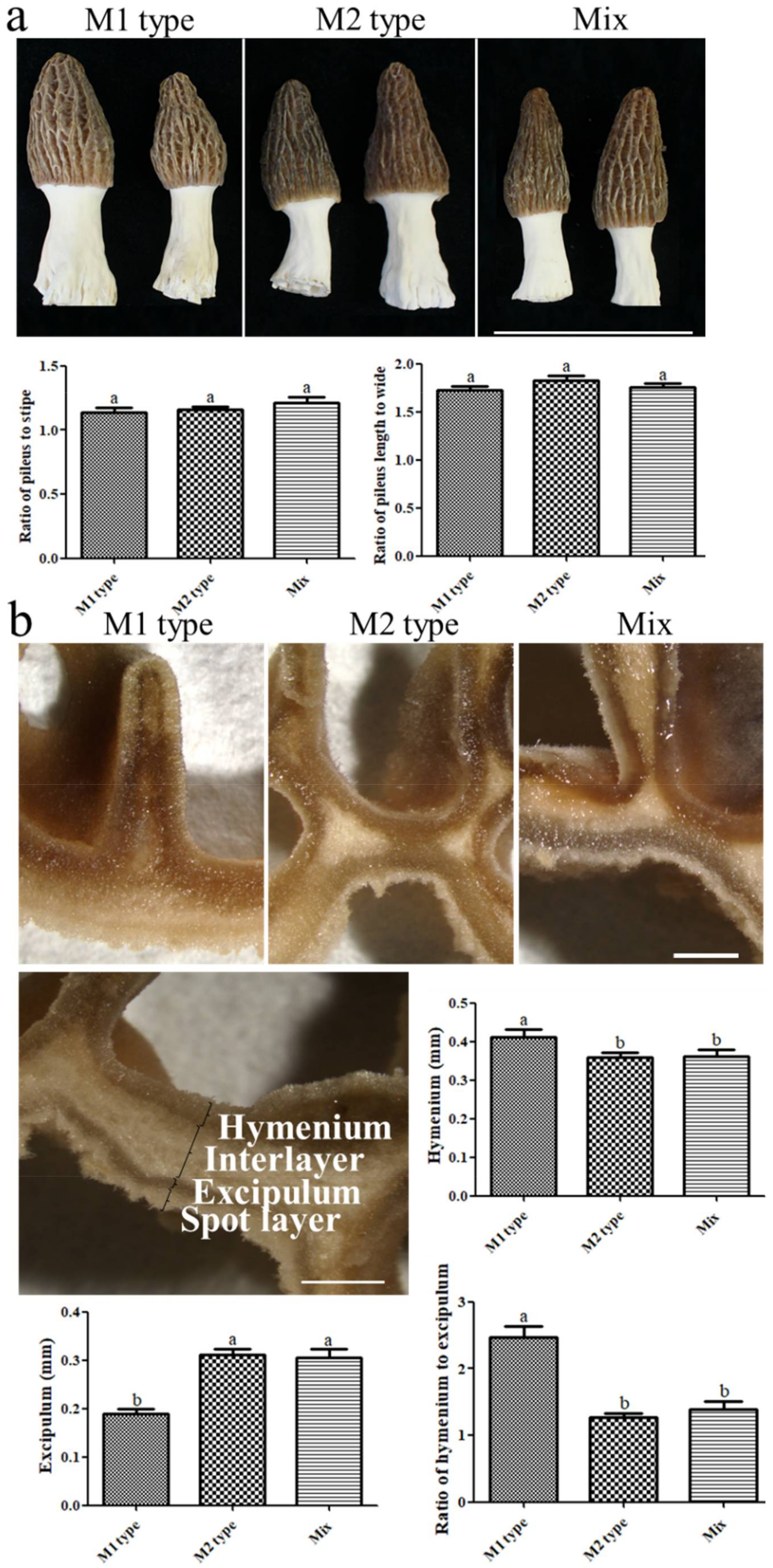
3.2. Different Mating Types Generate Divergent Physiological Structures of Fruiting Bodies
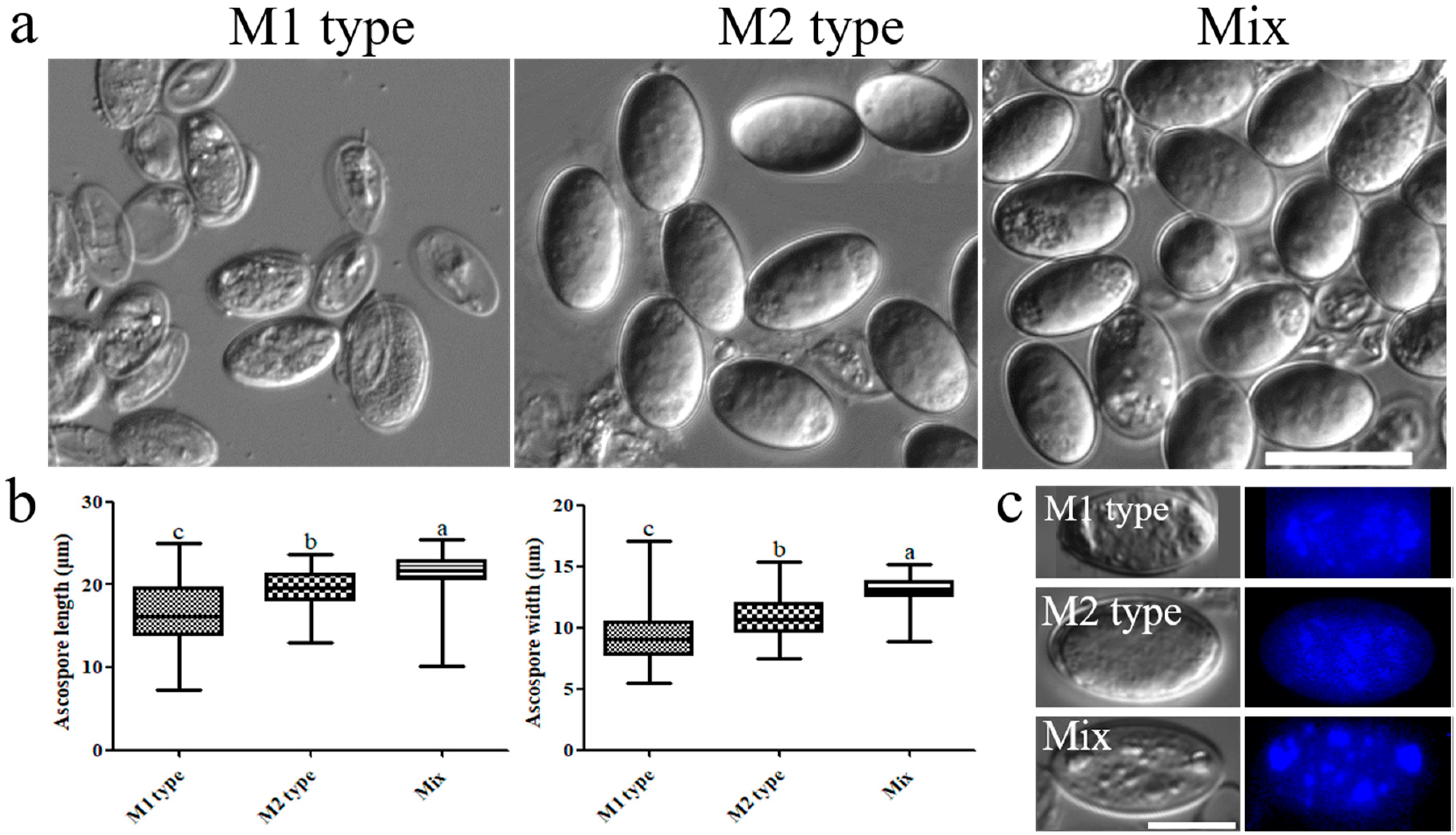
3.3. Mating Type Influences Ascus and Ascospore Development
3.4. Transcription Levels of Mating-Type Genes and Ascus Development-Related Genes in Pileus
3.5. mat1-2-1 Regulated Ascus Development-Related Genes at the Mycelial Stage
4. Discussion
4.1. Is It Mating-Type Switching?
4.2. Mating-Type Genes Regulate Fruiting Body Development
4.3. A Conservative Mechanism of Controlling Target Gene Expression by Mating-Type Genes
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Wu, F.; Zhou, L.W.; Yang, Z.L.; Bau, T.; Li, T.H.; Dai, Y.C. Resource diversity of Chinese macrofungi: Edible, medicinal and poisonous species. Fungal Divers. 2019, 98, 1–76. [Google Scholar] [CrossRef]
- Kirk, P.M.; Cannon, P.F.; Minter, D.W.; Stalpers, J.A. Ainsworth and Bisby’s Dictionary of Fungi, 10th ed.; CAB International: Wallingford, UK, 2008; p. 438. [Google Scholar] [CrossRef]
- Liu, Q.Z.; Ma, H.S.; Zhang, Y.; Dong, C.H. Artificial cultivation of true morels: Current state, issues and perspectives. Crit. Rev. Biotechnol. 2018, 38, 259–271. [Google Scholar] [CrossRef] [PubMed]
- Metzenberg, R.L.; Glass, N.L. Mating type and mating strategies in Neurospora. BioEssays 1990, 12, 53–59. [Google Scholar] [CrossRef] [PubMed]
- Ni, M.; Feretzaki, M.; Sun, S.; Wang, X.Y.; Heitman, J. Sex in Fungi. Annu. Rev. Genet. 2011, 45, 405–430. [Google Scholar] [CrossRef]
- Liu, K.H.; Shen, W.C. Mating differentiation in Cryptococcus neoformans is negatively regulated by the Crk1 protein kinase. Fungal Genet. Biol. 2011, 48, 225–240. [Google Scholar] [CrossRef]
- Wang, Z.; Kin, K.; Lopez-Giraldez, F.; Johannesson, H.; Townsend, J.P. Sex-specific gene expression during asexual development of Neurospora crassa. Fungal Genet. Biol. 2012, 49, 533–543. [Google Scholar] [CrossRef] [Green Version]
- Wang, Q.; Wang, S.; Xiong, C.L.; James, T.Y.; Zhang, X.G. Mating-type genes of the anamorphic fungus Ulocladium botrytis affect both asexual sporulation and sexual reproduction. Sci. Rep. 2017, 7, 7932. [Google Scholar] [CrossRef] [Green Version]
- Rodenburg, S.Y.A.; Terhem, R.B.; Veloso, J.; Stassen, J.H.M.; van Kan, J.A.L. Functional analysis of mating type genes and transcriptome analysis during fruiting body development of Botrytis cinerea. mBio 2018, 9, e01939-17. [Google Scholar] [CrossRef] [Green Version]
- Doughan, B.; Rollins, J.A. Characterization of mat gene functions in the life cycle of sclerotinia sclerotiorum reveals a lineage-specific mat gene functioning in apothecium morphogenesis. Fungal Biol. 2016, 120, 1105–1117. [Google Scholar] [CrossRef]
- Liu, W.; Chen, L.F.; Cai, Y.L.; Zhang, Q.; Bian, Y.B. Opposite polarity monospore genome de novo sequencing and comparative analysis reveal the possible heterothallic life cycle of Morchella importuna. Int. J. Mol. Sci. 2018, 19, 2525. [Google Scholar] [CrossRef]
- Chai, H.M.; Chen, W.M.; Zhang, X.L.; Su, K.M.; Zhao, Y.C. Structural variation and phylogenetic analysis of the mating-type locus in the genus Morchella. Mycologia 2019, 111, 551–562. [Google Scholar] [CrossRef] [PubMed]
- Du, X.H.; Yang, Z.L. Mating systems in true morels (Morchella). Microbiol. Mol. Biol. Rev. 2021, 85, e00220-20. [Google Scholar] [CrossRef] [PubMed]
- Du, X.H.; Zhao, Q.; Xia, E.H.; Gao, L.Z.; Richard, F.; Yang, Z.L. Mixed-reproductive strategies, competitive mating-type distribution and life cycle of fourteen black morel species. Sci. Rep. 2017, 7, 1493. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tan, H.; Liu, T.H.; Yu, Y.; Tang, J.; Jiang, L.; Martin, F.M.; Peng, W.H. Morel production related to soil microbial diversity and evenness. Microbiol. Spectrum. 2021, 9, e00229-21. [Google Scholar] [CrossRef]
- Longley, R.; Benucci, G.M.N.; Mills, G.; Bonito, G. Fungal and bacterial community dynamics in substrates during the cultivation of morels (Morchella rufobrunnea) indoors. FEMS Microbiol. Lett. 2019, 366, fnz215. [Google Scholar] [CrossRef]
- Tan, H.; Kohler, A.; Miao, R.Y.; Liu, T.H.; Zhang, Q.; Zhang, B.; Jiang, L.; Wang, Y.; Xie, L.Y.; Tang, J.; et al. Multi-omic analyses of exogenous nutrient bag decomposition by the black morel Morchella importuna reveal sustained carbon acquisition and transferring. Environ. Microbiol. 2019, 21, 3909–3926. [Google Scholar] [CrossRef] [Green Version]
- Du, X.H.; Zhao, Q.; Yang, Z.L.; Hansen, K.; Taskin, H.; Buyukalaca, S.; Dewsbury, D.; Moncalvo, J.M.; Douhan, G.W.; Robert, V.A.R.G.; et al. How well do ITS rDNA sequences differentiate species of true morels (Morchella)? Mycologia 2012, 104, 1351–1368. [Google Scholar] [CrossRef] [Green Version]
- Lv, S.Y.; Chen, X.; Mou, C.Y.; Dai, S.H.; Bian, Y.B. Agrobacterium-mediated transformation of the ascomycete mushroom Morchella importuna using polyubiquitin and glyceraldehyde-3-phosphate dehydrogenase promoter-based binary vectors. World J. Microbiol. Biotechnol. 2018, 34, 148. [Google Scholar] [CrossRef]
- Du, X.H.; Wu, D.; Kang, H.; Wang, H.; Xu, N.; Li, T.; Chen, K. Heterothallism and potential hybridization events inferred for twenty-two yellow morel species. IMA Fungus 2020, 11, 4. [Google Scholar] [CrossRef] [Green Version]
- Cao, F.; Cheng, J.T.; Chen, X.A.; Li, Y.Q.; Mao, X.M. Development of an efficient genetic system in a gene cluster-rich endophytic fungus Calcarisporium arbuscula NRRL 3705. J. Microbiol. Methods 2018, 151, 1–6. [Google Scholar] [CrossRef]
- Doyle, J.J.; Doyle, J.L. A rapid DNA isolation procedure for small quantities of fresh leaf tissue. Phytochem. Bull. 1987, 19, 11–15. [Google Scholar] [CrossRef]
- Liu, Q.Z.; He, G.Q.; Wei, J.K.; Dong, C.H. Comparative transcriptome analysis of cells from different areas reveals ROS responsive mechanism at sclerotial initiation stage in Morchella importuna. Sci. Rep. 2021, 11, 9418. [Google Scholar] [CrossRef] [PubMed]
- Kim, H.K.; Jo, S.M.; Kim, G.Y.; Kim, D.W.; Kim, Y.K.; Yun, S.H. A large-scale functional analysis of putative target genes of mating-type loci provides insight into the regulation of sexual development of the cereal pathogen Fusarium graminearum. PLoS Genet. 2015, 11, e1005486. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Murat, C.; Payen, T.; Noel, B.; Kuo, A.; Morin, E.; Chen, J.; Kohler, A.; Krizsán, K.; Balestrini, R.; Da Silva, C.; et al. Pezizomycetes genomes reveal the molecular basis of ectomycorrhizal truffle lifestyle. Nat. Ecol. Evol. 2018, 2, 1956–1965. [Google Scholar] [CrossRef] [Green Version]
- Liu, W.; Cai, Y.L.; He, P.X.; Bian, Y.B. Cultivation of monosporic and hybrid populations and polarity analysis of Morchella importuna. J. Fungal Res. 2019, 17, 43–49. [Google Scholar] [CrossRef]
- Hanson, S.J.; Wolfe, K.H. An evolutionary perspective on yeast mating-type switching. Genetics 2017, 206, 9–32. [Google Scholar] [CrossRef] [Green Version]
- Zhang, J.J.; Liu, M.Q.; Yang, T.; Gan, X.; Liu, W.Y.; Dong, C.H. Mating type of wild and cultivated Cordyceps militaris. Acta Edulis Fungi 2021, 28, 38–46. [Google Scholar] [CrossRef]
- Tan, H.; Yu, Y.; Tang, J.; Liu, T.H.; Miao, R.Y.; Huang, Z.Q.; Martin, F.M.; Peng, W.H. Build your own mushroom soil: Microbiota succession and nutritional accumulation in semi-synthetic substratum drive the fructification of a soil-saprotrophic morel. Front. Microbiol. 2021, 12, 656656. [Google Scholar] [CrossRef]
- Chai, H.M.; Ma, Y.H.; Liu, P.; Chen, W.M.; Tao, N.; Zhao, Y.C. The asymmetrical distribution of opposite mating type nuclei in single-ascospore isolates revealed Morchella importuna is a pseudohomothallic fungus. Mycosystema 2022, 1, 12. [Google Scholar] [CrossRef]
- Becker, K.; Beer, C.; Freitag, M.; Kück, U. Genome-wide identification of target genes of a mating-type α-domain transcription factor reveals functions beyond sexual development. Mol. Microbiol. 2015, 96, 1002–1022. [Google Scholar] [CrossRef]
- Keszthelyi, A.; Jeney, A.; Kerenyi, Z.; Mendes, O.; Waalwijk, C.; Hornok, L. Tagging target genes of the MAT1-2-1 transcription factor in Fusarium verticillioides (Gibberella fujikuroi MP-A). Antonie Van Leeuwenhoek 2007, 91, 373–391. [Google Scholar] [CrossRef] [PubMed]





| Name | Sequence (5′ to 3′) |
|---|---|
| mat1-1-1-QL | GCCTTCTGAGT CCGTTAT |
| mat1-1-1-QR | ATGTGAGCGTCCCTTGA |
| mat1-2-1-QL | GCCGTGACCCTCCTTCT |
| mat1-2-1-QR | TCGTTGTTCCCAATCCC |
| hyg-F | CTGTCGAGAAGTTTCTGATCG |
| hyg-R | CTGATAGAGTTGGTCAAGACC |
| mat1-1-1-RT-F | TGTCTTCGTAACGCCACT |
| mat1-1-1-RT-R | TAGCCCAACCCTTCCA |
| mat1-1-10-RT-F | TCGGAGGATGTTGGGTT |
| mat1-1-10-RT-R | ACGCTGCTTGAAGTATGGA |
| mat1-1-11-RT-F | GTGAAGCCATCAACAA |
| mat1-1-11-RT-R | GAGCCTTTCGTCAATAC |
| mat1-2-1-RT-F | CCAGGGAAAGAAAGTG |
| mat1-2-1-RT-R | CATAGGACGAGGAACAT |
| 372751-RT-F | TGGACCGATTAGGGAG |
| 372751-RT-R | TGTTGTTGCGGATGAC |
| 484741-RT-F | ACATTGTCACCACCCTC |
| 484741-RT-R | CCAAACTACTCGCCTTC |
| 504283-RT-F | TAAGGACGGCTATGTAA |
| 504283-RT-R | TCCGCAACTAGACCAC |
| 481101-RT-F | ACCTTACCCAACCTGA |
| 481101-RT-R | CTATCGTCCGCATTTA |
| 533191-RT-F | TCAACTACCACCTCACCC |
| 533191-RT-R | TCCCTCGCCAAAGAAC |
| 479298-RT-F | AACACTCTTGACCTCCAC |
| 479298-RT-R | CATCAGTAACCGCCTC |
| 543093-RT-F | GAACACTTACACGCCTAC |
| 543093-RT-R | CATCTCAGCCATCTCG |
| 533505-RT-F | ATTCCACCAAACAACG |
| 533505-RT-R | CCAACATAAACACCACC |
| 517289-RT-F | TGTCACCTCCATCTGTC |
| 517289-RT-R | GTATTCGGTCCGTCAT |
| HindIII-GPD-F1 | TGCCAAGCTTCGGGAGGTACGGGGAGAATA |
| SbfI-GPD-B1 | CAACCTGCAGGTTTGACTATTTAGTG |
| SpeI-35S-F3 | GAAACTAGTGTTTGATCGAGACCTAATACAGCC |
| SbfI-35S-B3 | ATTCACTAAATAGTCAAACCTGCAGGGGTGATGTCTGCTCAAGC |
| Part-mat1-2-1-F | TTGAGCAGACATCACCCCTGCAGGCTTCGGCCAGAACAGATG |
| Part-mat1-2-1-R | ATTCACTAAATAGTCAAACCTGCAGGGCATTCATAGGACGAG |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Liu, Q.; Qu, S.; He, G.; Wei, J.; Dong, C. Mating-Type Genes Play an Important Role in Fruiting Body Development in Morchella sextelata. J. Fungi 2022, 8, 564. https://doi.org/10.3390/jof8060564
Liu Q, Qu S, He G, Wei J, Dong C. Mating-Type Genes Play an Important Role in Fruiting Body Development in Morchella sextelata. Journal of Fungi. 2022; 8(6):564. https://doi.org/10.3390/jof8060564
Chicago/Turabian StyleLiu, Qizheng, Shan Qu, Guoqiang He, Jinkang Wei, and Caihong Dong. 2022. "Mating-Type Genes Play an Important Role in Fruiting Body Development in Morchella sextelata" Journal of Fungi 8, no. 6: 564. https://doi.org/10.3390/jof8060564







