Identification and Characterization of Dmct: A Cation Transporter in Yarrowia lipolytica Involved in Metal Tolerance
Abstract
:1. Introduction
2. Materials and Methods
2.1. In Silico Analysis
2.2. Microorganisms and Growth Conditions
2.3. Construction of Δdmct Strain
2.4. Reintegration of the DMCT Gene
2.5. Phenotypic Characterization
2.6. Intracellular Accumulation of Cations
2.7. Statistical Analysis
3. Results
3.1. In Silico Yl-Dmct Protein Analysis
3.2. In Silico Yl-Dmct Analysis: Cellular Location, Phylogenetic Analysis, and Three-Dimensional Modeling
3.3. In Silico Analysis of Putative Binding Sites for Transcription Factors in the Promoter Region from the Yl-DMCT Gene
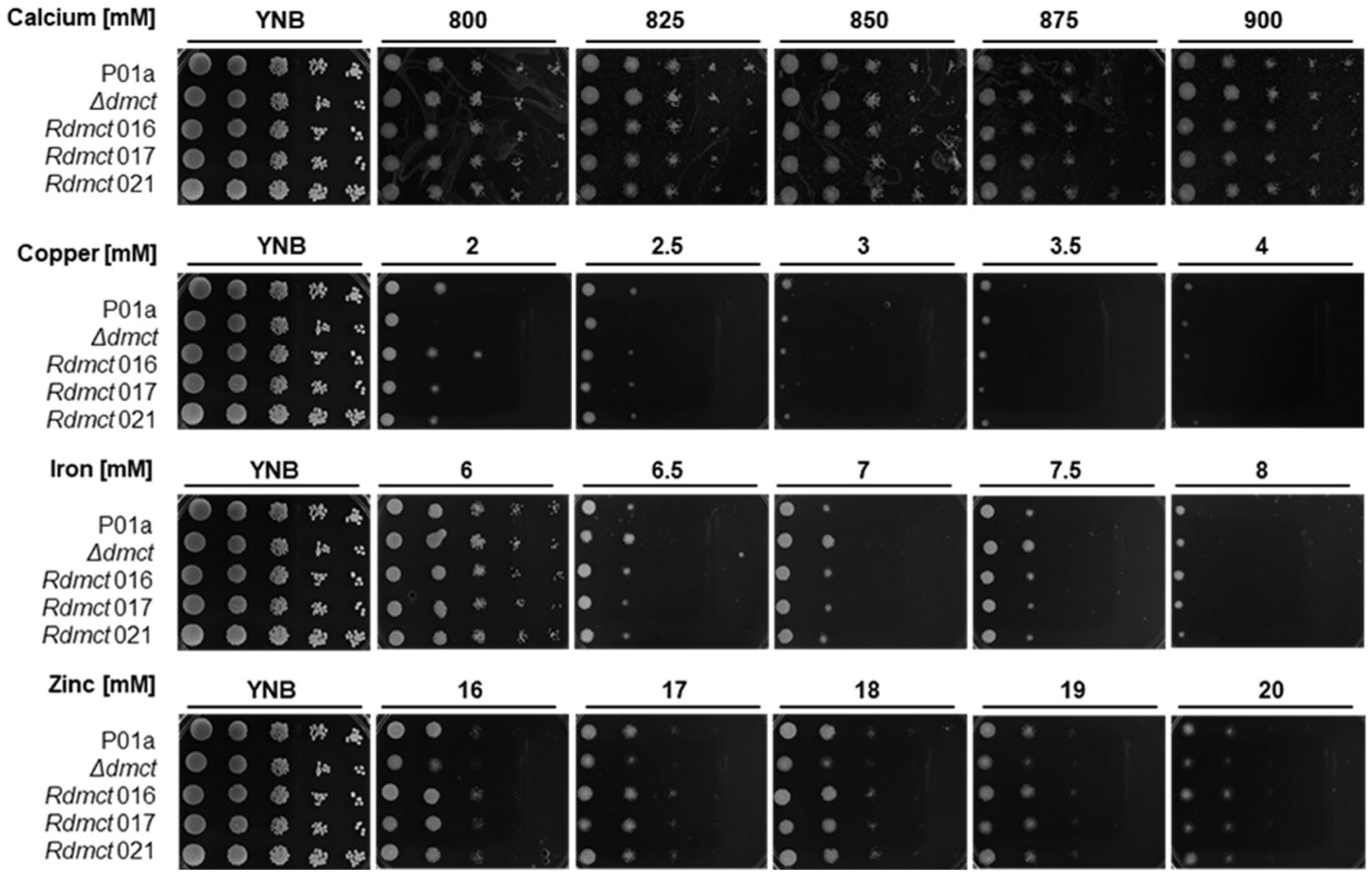
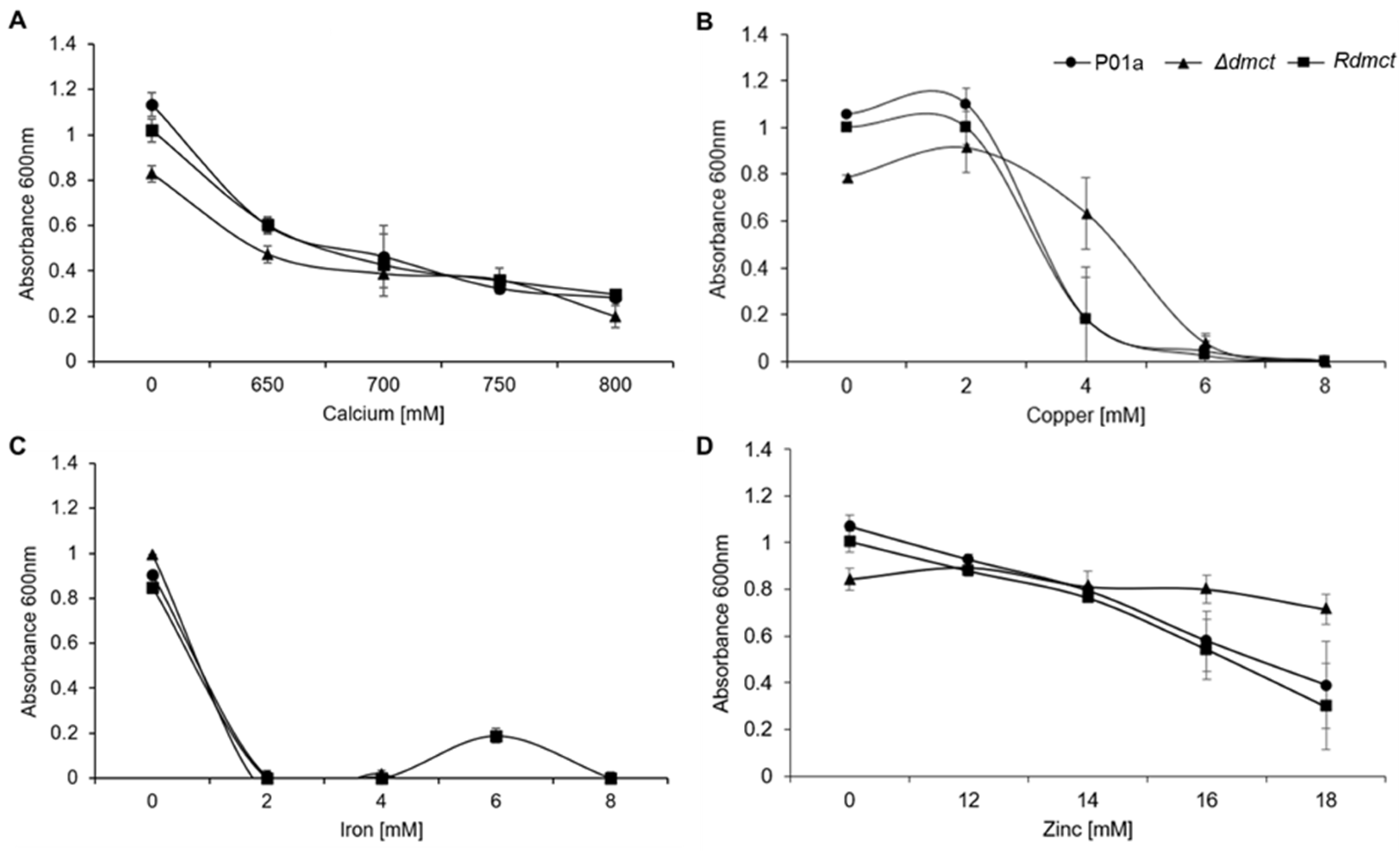
3.4. Δdmct and Rdmct Strains’ Phenotypic Characterization
3.5. Cell and Colony Morphological Characteristics
3.6. Intracellular Accumulation of Divalent Metal Cations and Cell Wall Integrity
3.7. Identification of Putative Genes That Participate in the Transport of Metals as a Possible Compensatory Response to Dmct Deletion
4. Discussion
5. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Bleackley, M.R.; MacGillivray, R.T.A. Transition metal homeostasis: From yeast to human disease. Biometals 2011, 24, 785–809. [Google Scholar] [CrossRef]
- Ito, H.; Inouhe, M.; Tohoyama, H.; Joho, M. Characteristics of copper tolerance in Yarrowia lipolytica. Biometals 2007, 20, 773–780. [Google Scholar] [CrossRef] [PubMed]
- Nzengue, Y.; Candéias, S.M.; Sauvaigo, S.; Douki, T.; Favier, A.; Rachidi, W.; Guiraud, P. The toxicity redox mechanisms of cadmium alone or together with copper and zinc homeostasis alteration: Its redox biomarkers. J. Trace Elem. Med. Biol. 2011, 25, 171–180. [Google Scholar] [CrossRef] [PubMed]
- Robinson, J.R.; Isikhuemhen, O.S.; Anike, F.N. Fungal–metal interactions: A review of toxicity and homeostasis. J. Fungi 2021, 7, 225. [Google Scholar] [CrossRef] [PubMed]
- André, B. An overview of membrane transport proteins in Saccharomyces cerevisiae. Yeast 1995, 11, 1575–1611. [Google Scholar] [CrossRef] [PubMed]
- Nies, D.H. Efflux-mediated heavy metal resistance in prokaryotes. FEMS Microbiol. Rev. 2003, 27, 313–339. [Google Scholar] [CrossRef] [PubMed]
- Singh, S.; Parihar, P.; Singh, R.; Singh, V.P.; Prasad, S.M. Heavy metal tolerance in plants: Role of transcriptomics, proteomics, metabolomics, and ionomics. Front. Plant Sci. 2016, 6, 1143. [Google Scholar] [CrossRef]
- Nies, D.H.; Silver, S. Ion efflux systems involved in bacterial metal resistances. J. Ind. Microbiol. 1995, 14, 186–199. [Google Scholar] [CrossRef]
- Wei, Y.; Fu, D. Selective metal binding to a membrane-embedded aspartate in the Escherichia coli metal transporter YiiP (FieF). J. Biol. Chem. 2005, 280, 33716–33724. [Google Scholar] [CrossRef]
- Paulsen, I.T.; Saier, M.H., Jr. A novel family of ubiquitous heavy metal ion transport proteins. J. Biol. Chem. 1997, 156, 99–103. [Google Scholar] [CrossRef]
- Montanini, B.; Blaudez, D.; Jeandroz, S.; Sanders, D.; Chalot, M. Phylogenetic and functional analysis of the cation diffusion facilitator (CDF) family: Improved signature and prediction of substrate specificity. BMC Genom. 2007, 8, 107. [Google Scholar] [CrossRef] [PubMed]
- Cubillas, C.; Vinuesa, P.; Tabche, M.L.; García-de los Santos, A. Phylogenomic analysis of cation diffusion facilitator proteins uncovers Ni2+/Co2+ transporters. Metallomics 2013, 5, 1634–1643. [Google Scholar] [CrossRef] [PubMed]
- Xu, T.; Chen, H.; Li, J.; Hong, S.; Shao, L.; Zheng, X.; Zou, Q.; Wang, Y.; Guo, S.; Jiang, J. Implications for cation selectivity and evolution by a novel cation diffusion facilitator family member from the moderate halophile Planococcus dechangensis. Front. Microbiol. 2019, 10, 607. [Google Scholar] [CrossRef] [PubMed]
- Rutherford, J.C.; Bird, A.J. Metal-responsive transcription factors that regulate iron, zinc, and copper homeostasis in eukaryotic cells. Eukaryot. Cell 2004, 3, 1–13. [Google Scholar] [CrossRef]
- Waldron, K.J.; Robinson, N.J. How do bacterial cells ensure that metalloproteins get the correct metal? Nat. Rev. Microbiol. 2009, 7, 25–35. [Google Scholar] [CrossRef]
- Martha-Paz, A.M.; Eide, D.; Mendoza-Cózatl, D.; Castro-Guerrero, N.A.; Aréchiga-Carvajal, E.T. Zinc uptake in the basidiomycota: Characterization of zinc transporters in Ustilago maydis. Mol. Membr. Biol. 2019, 35, 39–50. [Google Scholar] [CrossRef]
- Bull, P.C.; Cox, D.W. Wilson disease and Menkes disease: New handles on heavy-metal transport. Trends Genet. 1994, 10, 246–252. [Google Scholar] [CrossRef]
- Fagan, M.J.; Saier, M.H. P-Type ATPases of eukaryotes and bacteria: Sequence analyses and construction of phylogenetic trees. J. Mol. Evol. 1994, 38, 57–99. [Google Scholar] [CrossRef]
- Tam, R.; Saier, M.H. Structural, functional, and evolutionary relationships among extracellular solute-binding receptors of bacteria. Microbiol. Rev. 1993, 57, 320–346. [Google Scholar] [CrossRef]
- Kuan, G.; Dassa, E.; Saurin, W.; Hofnung, M.; Saier, M.H., Jr. Phylogenetic analyses of the ATP-binding constituents of bacterial extracytoplasmic receptor-dependent ABC-type nutrient uptake permeases. Res. Microbiol. 1995, 146, 271–278. [Google Scholar] [CrossRef]
- Dinh, T.; Paulsen, I.T.; Saier, M.H. A family of extracytoplasmic proteins that allow transport of large molecules across the outer membranes of gram-negative bacteria. J. Bacteriol. 1994, 176, 3825–3831. [Google Scholar] [CrossRef] [PubMed]
- Saier, M.H. Computer-aided analyses of transport protein sequences: Gleaning evidence concerning function, structure, biogenesis, and evolution. Microbiol. Rev. 1994, 58, 71–93. [Google Scholar] [CrossRef] [PubMed]
- MacDiarmid, C.W.; Milanick, M.A.; Eide, D.J. Biochemical properties of vacuolar zinc transport systems of Saccharomyces cerevisiae. J. Biol. Chem. 2002, 277, 39187–39194. [Google Scholar] [CrossRef] [PubMed]
- Guffanti, A.A.; Wei, Y.; Rood, S.V.; Krulwich, T.A. An antiport mechanism for a member of the cation diffusion facilitator family: Divalent cations efflux in exchange for K+ and H+. Mol. Microbiol. 2002, 45, 145–153. [Google Scholar] [CrossRef]
- Chao, Y.; Fu, D. Kinetic study of the antiport mechanism of an Escherichia coli zinc transporter, ZitB. J. Biol. Chem. 2004, 279, 12043–12050. [Google Scholar] [CrossRef]
- Grass, G.; Otto, M.; Fricke, B.; Haney, C.J.; Rensing, C.; Nies, D.H.; Munkelt, D. FieF (YiiP) from Escherichia coli mediates decreased cellular accumulation of iron and relieves iron stress. Arch. Microbiol. 2005, 183, 9–18. [Google Scholar] [CrossRef]
- Haney, C.J.; Grass, G.; Franke, S.; Rensing, C. New developments in the understanding of the cation diffusion facilitator family. J. Ind. Microbiol. Biotechnol. 2005, 32, 215–226. [Google Scholar] [CrossRef]
- Kambe, T.; Narita, H.; Yamaguchi-Iwai, Y.; Hirose, J.; Amano, T.; Sugiura, N.; Sasaki, R.; Mori, K.; Iwanaga, T.; Nagao, M. Cloning and characterization of a novel mammalian zinc transporter, zinc transporter 5, abundantly expressed in pancreatic β cells. J. Biol. Chem. 2002, 277, 19049–19055. [Google Scholar] [CrossRef]
- Cragg, R.A.; Christie, G.R.; Phillips, S.R.; Russi, R.M.; Küry, S.; Mathers, J.C.; Taylor, P.M.; Ford, D. A novel zinc-regulated human zinc transporter, HZTL1, is localized to the enterocyte apical membrane. J. Biol. Chem. 2002, 277, 22789–22797. [Google Scholar] [CrossRef] [PubMed]
- Delhaize, E.; Kataoka, T.; Hebb, D.M.; White, R.G.; Ryan, P.R. Genes encoding proteins of the cation diffusion facilitator family that confer manganese tolerance. Plant Cell 2003, 15, 1131–1142. [Google Scholar] [CrossRef]
- Mäser, P.; Thomine, S.; Schroeder, J.I.; Ward, J.M.; Hirschi, K.; Sze, H.; Talke, I.N.; Amtmann, A.; Maathuis, F.J.M.; Sanders, D.; et al. Phylogenetic relationships within Cation Transporter Families of Arabidopsis. Plant Physiol. 2001, 126, 1646–1667. [Google Scholar] [CrossRef] [PubMed]
- Munkelt, D.; Grass, G.; Nies, D.H. The chromosomally encoded Cation Diffusion Facilitator proteins DmeF and FieF from Wautersia metallidurans CH34 are transporters of broad metal specificity. J. Bacteriol. 2004, 186, 8036–8043. [Google Scholar] [CrossRef] [PubMed]
- Li, L.; Kaplan, J. Characterization of two homologous yeast genes that encode mitochondrial iron transporters. J. Biol. Chem. 1997, 272, 28485–28493. [Google Scholar] [CrossRef] [PubMed]
- Desbrosses-Fonrouge, A.-G.; Voigt, K.; Schröder, A.; Arrivault, S.; Thomine, S.; Krämer, U. Arabidopsis thaliana MTP1 is a Zn transporter in the vacuolar membrane which mediates Zn detoxification and drives leaf Zn accumulation. FEBS Lett. 2005, 579, 4165–4174. [Google Scholar] [CrossRef]
- Palmiter, R.D.; Cole, T.B.; Findley, S.D. ZnT-2, a mammalian protein that confers resistance to zinc by facilitating vesicular sequestration. EMBO J. 1996, 15, 1784–1791. [Google Scholar] [CrossRef]
- Palmiter, R.D.; Cole, T.B.; Quaife, C.J.; Findley, S.D. ZnT-3, a putative transporter of zinc into synaptic vesicles. Proc. Natl. Acad. Sci. USA 1996, 93, 14934–14939. [Google Scholar] [CrossRef]
- Huang, L.; Gitschier, J. A novel gene involved in zinc transport is deficient in the lethal milk mouse. Nat Genet. 1997, 17, 292–297. [Google Scholar] [CrossRef]
- MacDiarmid, C.W.; Gaither, L.A.; Eide, D. Zinc transporters that regulate vacuolar zinc storage in Saccharomyces cerevisiae. EMBO J. 2000, 19, 2845–2855. [Google Scholar] [CrossRef]
- Clemens, S.; Bloss, T.; Vess, C.; Neumann, D.; Nies, D.H.; Zur Nieden, U. A transporter in the endoplasmic reticulum of Schizosaccharomyces pombe cells mediates zinc storage and differentially affects transition metal tolerance. J. Biol. Chem. 2002, 277, 18215–18221. [Google Scholar] [CrossRef]
- Casaregola, S.; Neuvéglise, C.; Lépingle, A.; Bon, E.; Feynerol, C.; Artiguenave, F.; Wincker, P.; Gaillardin, C. Genomic exploration of the hemiascomycetous yeasts: 17. Yarrowia lipolytica. FEBS Lett. 2000, 487, 95–100. [Google Scholar] [CrossRef]
- Shinde, N.R.; Bankar, A.V.; Kumar, A.R.; Zinjarde, S.S. Removal of Ni (II) ions from aqueous solutions by biosorption onto two strains of Yarrowia lipolytica. J. Environ. Manag. 2012, 102, 115–124. [Google Scholar] [CrossRef] [PubMed]
- Bankar, A.V.; Kumar, A.R.; Zinjarde, S.S. Environmental and industrial applications of Yarrowia lipolytica. Appl. Microbiol. Biotechnol. 2009, 84, 847–865. [Google Scholar] [CrossRef] [PubMed]
- Wheeler, D.L.; Barrett, T.; Benson, D.A.; Bryant, S.H.; Canese, K.; Chetvernin, V.; Church, D.M.; DiCuccio, M.; Edgar, R.; Federhen, S.; et al. Database resources of the National Center for Biotechnology Information. Nucleic Acids Res. 2007, 36, D13–D21. [Google Scholar] [CrossRef] [PubMed]
- Marchler-Bauer, A. CDD: A curated entrez database of conserved domain alignments. Nucleic Acids Res. 2003, 31, 383–387. [Google Scholar] [CrossRef] [PubMed]
- Goddard, T.D.; Huang, C.C.; Meng, E.C.; Pettersen, E.F.; Couch, G.S.; Morris, J.H.; Ferrin, T.E. UCSF ChimeraX: Meeting modern challenges in visualization and analysis: UCSF ChimeraX visualization system. Protein Sci. 2018, 27, 14–25. [Google Scholar] [CrossRef]
- The UniProt Consortium. UniProt: A Hub for protein information. Nucleic Acids Res. 2015, 43, D204–D212. [Google Scholar] [CrossRef]
- Sievers, F.; Higgins, D.G. Clustal Omega, accurate alignment of very large numbers of sequences. In Multiple Sequence Alignment Methods; Methods in Molecular Biology; Russell, D.J., Ed.; Humana Press: Totowa, NJ, USA, 2014; Volume 1079, pp. 105–116. ISBN 978-1-62703-645-0. [Google Scholar]
- Thumuluri, V.; Almagro Armenteros, J.J.; Johansen, A.R.; Nielsen, H.; Winther, O. DeepLoc 2.0: Multi-label subcellular localization prediction using protein language models. Nucleic Acids Res. 2022, 50, W228–W234. [Google Scholar] [CrossRef]
- Dobson, L.; Reményi, I.; Tusnády, G.E. CCTOP: A consensus constrained TOPology Prediction web server. Nucleic Acids Res. 2015, 43, W408–W412. [Google Scholar] [CrossRef]
- Pettersen, E.F.; Goddard, T.D.; Huang, C.C.; Meng, E.C.; Couch, G.S.; Croll, T.I.; Morris, J.H.; Ferrin, T.E. UCSF ChimeraX: Structure visualization for researchers, educators, and developers. Protein Sci. 2021, 30, 70–82. [Google Scholar] [CrossRef]
- Jumper, J.; Evans, R.; Pritzel, A.; Green, T.; Figurnov, M.; Ronneberger, O.; Tunyasuvunakool, K.; Bates, R.; Žídek, A.; Potapenko, A.; et al. Highly accurate protein structure prediction with AlphaFold. Nature 2021, 596, 583–589. [Google Scholar] [CrossRef]
- Huerta-Cepas, J.; Bueno, A.; Dopazo, J.; Gabaldon, T. PhylomeDB: A database for genome-wide collections of gene phylogenies. Nucleic Acids Res. 2007, 36, D491–D496. [Google Scholar] [CrossRef] [PubMed]
- Teixeira, M.C. The YEASTRACT database: A tool for the analysis of transcription regulatory associations in Saccharomyces cerevisiae. Nucleic Acids Res. 2006, 34, D446–D451. [Google Scholar] [CrossRef] [PubMed]
- Wheeler, D.; Bhagwat, M. BLAST quickstart. In Comparative Genomics; Human Press: Totowa, NJ, USA, 2007. [Google Scholar]
- Hoffman, C.S.; Winston, F. Ten-minute DNA preparation from yeast efficiently releases autonomous plasmids for the transformation of Escherichia coli. Gene 1987, 57, 267–272. [Google Scholar] [CrossRef] [PubMed]
- Gaillardin, C.; Ribet, A.M.; Heslot, H. Integrative transformation of the yeast Yarrowia lipolytica. Curr. Genet. 1985, 10, 49–58. [Google Scholar] [CrossRef]
- Larochelle, M.; Drouin, S.; Robert, F.; Turcotte, B. Oxidative stress-activated zinc cluster protein Stb5 has dual activator/repressor functions required for pentose phosphate pathway regulation and NADPH production. Mol. Cell. Biol. 2006, 26, 6690–6701. [Google Scholar] [CrossRef] [PubMed]
- Courel, M.; Lallet, S.; Camadro, J.M.; Blaiseau, P.L. Direct activation of genes involved in intracellular iron use by the yeast iron-responsive transcription factor Aft2 without its paralog Aft1. Mol. Cell. Biol. 2005, 25, 6760–6771. [Google Scholar] [CrossRef]
- Kuge, S.; Jones, N.; Nomoto, A. Regulation of YAP-1 nuclear localization in response to oxidative stress. EMBO J. 1997, 16, 1710–1720. [Google Scholar] [CrossRef]
- Buchman, C.; Skroch, P.; Karin’, M. The CUP2 gene product, regulator of yeast metallothionein expression, is a copper-activated DNA-binding protein. Mol. Cell. Biol. 1989, 9, 4091–4095. [Google Scholar]
- Lamb, T.M.; Mitchell, A.P. The transcription factor Rim101p governs ion tolerance and cell differentiation by direct repression of the regulatory genes NRG1 and SMP1 in Saccharomyces cerevisiae. Mol. Cell. Biol. 2003, 23, 677–686. [Google Scholar] [CrossRef]
- Hanlon, S.E.; Rizzo, J.M.; Tatomer, D.C.; Lieb, J.D.; Buck, M.J. The stress response factors Yap6, Cin5, Phd1, and Skn7 direct targeting of the conserved co-repressor Tup1-Ssn6 in S. cerevisiae. PLoS ONE 2011, 6, e19060. [Google Scholar] [CrossRef]
- Dalal, C.K.; Cai, L.; Lin, Y.; Rahbar, K.; Elowitz, M.B. Pulsatile dynamics in the yeast proteome. Curr. Biol. 2014, 24, 2189–2194. [Google Scholar] [CrossRef] [PubMed]
- Liesegang, H.; Lemke, K.; Siddiqui, R.A.; Schlegel, H.G. Characterization of the inducible nickel and cobalt resistance determinant Cnr from PMOL28 of Alcaligenes eutrophus CH34. J. Bacteriol. 1993, 175, 767–778. [Google Scholar] [CrossRef] [PubMed]
- Barber-Zucker, S.; Shaanan, B.; Zarivach, R. Transition metal binding selectivity in proteins and its correlation with the Phylogenomic classification of the Cation Diffusion Facilitator protein family. Sci. Rep. 2017, 7, 16381. [Google Scholar] [CrossRef]
- Murgia, C.; Vespignani, I.; Cerase, J.; Nobili, F.; Perozzi, G. Cloning, expression, and vesicular localization of zinc transporter Dri 27/ZnT4 in intestinal tissue and cells. Am. J. Physiol. Gastrointest. Liver Physiol. 1999, 277, G1231–G1239. [Google Scholar] [CrossRef] [PubMed]
- Blaudez, D.; Kohler, A.; Martin, F.; Sanders, D.; Chalot, M. Poplar metal tolerance protein 1 confers zinc tolerance and is an oligomeric vacuolar zinc transporter with an essential leucine zipper motif. Plant Cell 2003, 15, 2911–2928. [Google Scholar] [CrossRef] [PubMed]
- Ellis, C.D.; MacDiarmid, C.W.; Eide, D.J. Heteromeric protein complexes mediate zinc transport into the secretory pathway of eukaryotic cells. J. Biol. Chem. 2005, 280, 28811–28818. [Google Scholar] [CrossRef] [PubMed]
- Lu, M.; Chai, J.; Fu, D. Structural basis for autoregulation of the zinc transporter YiiP. Nat. Struct. Mol. Biol. 2009, 16, 1063–1067. [Google Scholar] [CrossRef]
- Lin, H.; Kumánovics, A.; Nelson, J.M.; Warner, D.E.; Ward, D.M.; Kaplan, J. A single amino acid change in the yeast vacuolar metal transporters Zrc1 and Cot1 alters their substrate specificity. J. Biol. Chem. 2008, 283, 33865–33873. [Google Scholar] [CrossRef]
- Breuer, U.; Harms, H. Debaryomyces hansenii—An extremophilic yeast with biotechnological potential. Yeast 2006, 23, 415–437. [Google Scholar] [CrossRef]
- Almagro, A.; Prista, C.; Castro, S.; Quintas, C.; Madeira-Lopes, A.; Ramos, J.; Loureiro-Dias, M.C. Effects of salts on Debaryomyces hansenii and Saccharomyces cerevisiae under stress conditions. Int. J. Food Microbiol. 2000, 56, 191–197. [Google Scholar] [CrossRef]
- Sekova, V.Y.; Gessler, N.N.; Isakova, E.P.; Antipov, A.N.; Dergacheva, D.I.; Deryabina, Y.I.; Trubnikova, E.V. Redox status of extremophilic yeast Yarrowia lipolytica during adaptation to pH-stress. Appl. Biochem. Microbiol. 2015, 51, 649–654. [Google Scholar] [CrossRef]
- Sekova, V.Y.; Isakova, E.P.; Deryabina, Y.I. Biotechnological applications of the extremophilic yeast Yarrowia lipolytica (review). Appl. Biochem. Microbiol. 2015, 51, 278–291. [Google Scholar] [CrossRef]
- Blaiseau, P.L.; Lesuisse, E.; Camadro, J.M. Aft2p, a novel iron-regulated transcription activator that modulates, with Aft1p, intracellular iron use and resistance to oxidative stress in yeast. J. Biol. Chem. 2001, 276, 34221–34226. [Google Scholar] [CrossRef] [PubMed]
- Vido, K.; Spector, D.; Lagniel, G.; Lopez, S.; Toledano, M.B.; Labarre, J. A proteome analysis of the cadmium response in Saccharomyces cerevisiae. J. Biol. Chem. 2001, 276, 8469–8474. [Google Scholar] [CrossRef] [PubMed]
- Cherian, M. Metallothioneins in human tumors and potential roles in carcinogenesis. Mutat. Res. Fundam. Mol. Mech. Mutagen. 2003, 533, 201–209. [Google Scholar] [CrossRef]
- Cuizano, N.A.; Reyes, Ú.F.; Domínguez, S.; Llanos, B.P.; Navarro, A.E. Relevancia del pH en la adsorción de iones metálicos mediante algas pardas. Rev. Soc. Quím. Perú 2010, 76, 123–130. [Google Scholar]
- Beeler, T.J.; Fu, D.; Rivera, J.; Monaghan, E.; Gable, K.; Dunn, T.M. SUR1 (CSG1/BCL21), a gene necessary for growth of Saccharomyces cerevisiae in the presence of high Ca2+ concentrations at 37 °C, is required for mannosylation of inositolphosphorylceramide. Mol. Gen. Genet. 1997, 255, 570–579. [Google Scholar] [CrossRef]
- Li, C.; Xu, Y.; Jiang, W.; Dong, X.; Wang, D.; Liu, B. Effect of NaCl on the heavy metal tolerance and bioaccumulation of Zygosaccharomyces rouxii and Saccharomyces cerevisiae. Bioresour. Technol. 2013, 143, 46–52. [Google Scholar] [CrossRef]
- Lin, Z.; Li, W.-H. Expansion of hexose transporter genes was associated with the evolution of aerobic fermentation in yeasts. Mol. Biol. Evol. 2011, 28, 131–142. [Google Scholar] [CrossRef]
- Ruiz-Herrera, J.; Sentandreu, R. Different effectors of dimorphism in Yarrowia lipolytica. Arch. Microbiol. 2002, 178, 477–483. [Google Scholar] [CrossRef]
- De Silóniz, M.-I.; Balsalobre, L.; Alba, C.; Valderrama, M.-J.; Peinado, J.M. Feasibility of copper uptake by the yeast Pichia guilliermondii isolated from sewage sludge. Res. Microbiol. 2002, 153, 173–180. [Google Scholar] [CrossRef] [PubMed]
- Safarik, I.; Maderova, Z.; Pospiskova, K.; Baldikova, E.; Horska, K.; Safarikova, M. Magnetically responsive yeast cells: Methods of preparation and applications. Yeast 2015, 32, 227–237. [Google Scholar] [CrossRef] [PubMed]
- Soares, E.V.; Soares, H.M.V.M. Bioremediation of industrial effluents containing heavy metals using brewing cells of Saccharomyces cerevisiae as a green technology: A review. Environ. Sci. Pollut. Res. 2012, 19, 1066–1083. [Google Scholar] [CrossRef] [PubMed]
- Gaither, L.A.; Eide, D.J. Eukaryotic zinc transporters and their regulation. In Zinc Biochemistry, Physiology, and Homeostasis; Maret, W., Ed.; Springer: Dordrecht, The Netherlands, 2001; pp. 65–84. ISBN 978-90-481-5916-1. [Google Scholar]





| Protein * | Function | Organism | Length (Aa) | % Complete Gene Similarity | % CDF Domain Similarity |
|---|---|---|---|---|---|
| YlDmct | Hypothetical transport of metal divalent cations (This studio) | Y. lipolytica | 555 | 100 | 100 |
| Cot1 | Transport of metal cations in vacuole, resistance to cobalt and rhodium, response to stress | S. cerevisiae | 439 | 22.67 | 18.67 |
| Zcr1 | Resistance to zinc and cadmium | S. cerevisiae | 442 | 21.07 | 19.41 |
| Mmt1p | Zinc transport | S. cerevisiae | 510 | 19.28 | 23.44 |
| Zrg17 | Zinc transport in endoplasmic reticulum | Z. rouxii | 634 | 21.14 | 24.24 |
| Msc2 | Zinc homeostasis, zinc influx to the endoplasmic reticulum | S. cerevisiae | 725 | 19.12 | 19.88 |
| Mitochondrial metal transporter | Iron accumulation in mitochondria | S. cerevisiae | 510 | 19.85 | 23.44 |
| Cobalt toxicity protein | Zinc vacuolar transport and cobalt and rhodium resistance | S. cerevisiae | 439 | 19.01 | 18.67 |
| Cation diffusion facilitator family transporter | Iron, copper, zinc and cadmium transport | C. albicans | 616 | 44.30 | 56.77 |
| Metal cation transporter | Copper, zinc and cadmium ion efflux | C. albicans | 626 | 20.73 | 20.56 |
| Zinc/cadmium resistance protein | Copper, zinc and cadmium efflux | L. elongisporus | 474 | 23.20 | 20.70 |
| Mitochondrial protein with role in iron accumulation | Iron accumulation | S. stipitis | 430 | 21.73 | 22.73 |
| Zinc/cadmium resistance protein | Copper, zinc and cadmium efflux | K. phaffii | 459 | 19.93 | 19.76 |
| Homologous | Cation | Organism | Location | Accesion Number * | Aa ** | Identity (%) |
|---|---|---|---|---|---|---|
| Zrt3 | Fe2+/Zn2+ | S. c. | Vacuole/ Lysosome | P34240 | 503 | 13 |
| Fth1 | Fe2+ | S. c. | P38310 | 465 | 14 | |
| C. g. | Q6FJK8 | 435 | 15 | |||
| Fet5 | Fe2+ | S. c. | P43561 | 622 | 14 | |
| C. g. | A0A0W0DZF1 | 621 | 13 | |||
| Cot1 | Fe2+/Zn2+ | S. c. | P32798 | 439 | 16 | |
| Zrc1 | Fe2+/Zn2+ | S. c. | P20107 | 442 | 13 | |
| Smf3 | Fe2+ | S. c. | Q12078 | 473 | 15 | |
| C. a. | Q5ACZ8 | 514 | 14 | |||
| Ctr2 | Cu2+ | S. c. | P38865 | 189 | 9 | |
| Ccc2 | Fe2+/Cu2+ | S. c. | Golgi apparatus | P38995 | 1004 | 13 |
| C. a. | Q5AG51 | 1204 | 11 | |||
| Hmx1 | Fe2+ | S. c. | P32339 | 317 | 13 | |
| Fet3 | Cu2+ | S. c. | P38993 | 636 | 14 | |
| C. g. | Q96WT3 | 635 | 15 |
| Trancription Factor | Binding Site | Function | Genes That Regulate | Functional Homologous Predicted in Y. lipolytica | Reference |
|---|---|---|---|---|---|
| Stb5p | −679, −531, −407, −389, −256, −103 | Involved in the transcription of transmembrane transporters. | ZRT3 | YALI0A10637p/Q6CHB0 YALI0C15202p/Q6CBV4 YALI0F03630p/B5RSK6 YALI0F16599p/Q6C1G1 YALI0D12628p/Q6C9A9 YALI0C22990p/Q6CB01 YALI0B06853p/Q6CFH8 | [57] |
| Aft2p | −961 | In the absence of iron, which is responsible for the transcription of genes, products mobilize copper. | FTH1, FET5, FET3, SMF3 | No significant similarity found in genome | [58] |
| Yap1p | −882 | Related to cadmium tolerance, is involved in the transcription of genes that respond to ionic detoxification, besides the expression of ferredoxin and ferredoxin reductase proteins. | ZRT1, FTH1, FET5, FET3, COT1, ZRC1, SMF3. CTR2, CCC2, HMX1 | YALI0F03388p/Q6C317 YALI0D09757p/Q6C9N3 YALI0B13200p/Q6CET1 YALI0F27445p/Q6C065 | [59] |
| Cup2p | −919 | Activates the transcription of metallothionein proteins when the cell is in an environment that presents excess metals. | FET3 | CRF1/P45815 YALI0E31669p/Q6C3W4 | [60] |
| Nrg1p | −791, −773, −738, −729, −728, −718, −647, −646, −605, −604, −588, −559, −349, −348, −81 | Related to the response to stress due to pH changes | FET3 | YALI0C12364p/Q6CC55 YALI0E07942p/Q6C6N4 YALI0D23749p/Q6C809 YALI0D18678p/Q6C8L5 YALI0B21582p/F2Z5Y0 YALI0A16841p/Q6CGR7 YALI0C05995p/Q6CCW4 YALI0F21923p/Q6C0T4 YALI0F22649p/Q6C0Q3 | [61] |
| Skn7p | −531, −27 | Responsible for regulating genes involved in the antioxidant cellular response. | ZRT3, HMX1 | YALI0D14520p/Q6C937 YALI0D04785p/Q6CA95 YALI0C21340p/Q6CB75 YALI0E13948p/Q6C5Z0 | [62] |
| Msn2p | −854, −791, −773, −738, −729, −718, −646, −604, −560, −559, −348, −300, −104, −81, | Responsible for regulating genes involved in the antioxidant cellular response. | FET3, ZRC1, CTR2, CCC2, HMX1 | No significant similarity found in genome | [63] |
| Msn4p | −791, −773, −738, −729, −718, −646, −604, −559, −348, −81 | Responsible for regulating genes involved in the antioxidant cellular response. | FET3, CCC2, HMX1 | YALI0C13750p/Q6CC08 | [63] |
| Chr | Gene (Locus Tag/ID) | Similar to | Possible Function | Functional Domains |
|---|---|---|---|---|
| A | YALI_A11605g/Q6CH77 | MSF transporter | Transport of small molecules by the gradient | MSF superfamily |
| YALI_A14883g/Q6CGY6 | Ferrichrome-type siderophore transporter | Iron homeostasis | MSF superfamily | |
| B | YALI0_B06094g/Q6CFK7 | RdgB Ca2+ transporter | Metal binding and calcium transport | DDHD superfamily |
| YALI0_B17864g/Q6CE78 | High-affinity potassium absorption transporter | Potassium transport | 2a38euk superfamily | |
| YALI0_B19250g/Q6CE20 | Ferrichrome-type siderophore transporter | Iron homeostasis | MSF superfamily | |
| C | YALI0_C02541g/Q6CD98 | MSF transporter | Unknown specificity | MSF superfamily |
| YALI0_C04411g/Q6CD22 | SMF1/E protein | Divalent and trivalent metal transporter | SLC5-6-like_sbd superfamily | |
| YALI0_C06105g/Q6CCV9 | MSF transporter | Unknown specificity | MSF superfamily | |
| YALI0_C09823g/Q6CCF5 | MSF transporter | Anion-cation simporter | MSF superfamily | |
| YALI0_C10311g/Q6CCD6 | High-affinity potassium transporter | Potassium transport | K-Trans superfamily | |
| YALI0_C10670g/Q6CCC4 | MSF transporter | Unknown specificity | MSF superfamily | |
| YALI0_C12254g/Q6CC58 | Iron transporter in mitochondria MMT2 | Transport of divalent cation metals | SelP_N superfamily, FieF domain | |
| YALI0_C16225g/Q6CBR6 | MSF transporter | Unknown specificity | MSF superfamily | |
| YALI0_C17105g/Q6CBM9 | MSF transporter | Unknown specificity | MSF superfamily | |
| YALI0_C18051g/Q6CBK3 | YCFI metal resistance protein | Vacuolar transport with increased tolerance to metals | MRP_assoc_pro superfamily | |
| D | YALI0_D00319g/Q6CAS6 | Divalent cation transporter ALR1 | Divalent cation transport | Alr1p-like superfamily |
| YALI0_D20064g/Q6C8F6 | MSF transporter | Unknown specificity | MSF superfamily | |
| YALI0_D24651g/Q6C7X0 | MSF transporter | Unknown specificity | MSF superfamily | |
| YALI0_D26818g/Q6C7M8 | SMF2 carrier protein | Manganese transport | Nramp domain | |
| E | YALI0_E00748g/Q6C7H7 | Zrt2 transporter | Zinc transporter | Zip superfamily |
| YALI0_E00462g/Q6C7J0 | Alr1 transporter | Divalent cations transport | Alr1p-like superfamily | |
| YALI0_E11473g/Q6C691 | HAK1 transporter | Potassium transport | K-Trans superfamily | |
| YALI0_E14234g/Q6C5X7 | YBT1 transporter | Involved with Ca2+ and metal resistance | MRP_assoc_pro superfamily | |
| F | YALI0_F19118g/Q6C155 | Ferrichrome-type siderophore transporter | Iron homeostasis | MSF superfamily |
| YALI0_F20922g/Q6C0X7 | Ferrichrome-type siderophore transporter | Iron homeostasis | MSF superfamily | |
| YALI0_F27709g/Q6C053 | Ferrichrome-type siderophore transporter | Iron homeostasis | MSF superfamily | |
| YALI0_F29711g/Q6BZZ0 | MSF transporter | Unknown specificity | MSF superfamily |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
González-Lozano, K.J.; Aréchiga-Carvajal, E.T.; Jiménez-Salas, Z.; Valdez-Rodríguez, D.M.; León-Ramírez, C.G.; Ruiz-Herrera, J.; Adame-Rodríguez, J.M.; López-Cabanillas-Lomelí, M.; Campos-Góngora, E. Identification and Characterization of Dmct: A Cation Transporter in Yarrowia lipolytica Involved in Metal Tolerance. J. Fungi 2023, 9, 600. https://doi.org/10.3390/jof9060600
González-Lozano KJ, Aréchiga-Carvajal ET, Jiménez-Salas Z, Valdez-Rodríguez DM, León-Ramírez CG, Ruiz-Herrera J, Adame-Rodríguez JM, López-Cabanillas-Lomelí M, Campos-Góngora E. Identification and Characterization of Dmct: A Cation Transporter in Yarrowia lipolytica Involved in Metal Tolerance. Journal of Fungi. 2023; 9(6):600. https://doi.org/10.3390/jof9060600
Chicago/Turabian StyleGonzález-Lozano, Katia Jamileth, Elva Teresa Aréchiga-Carvajal, Zacarías Jiménez-Salas, Debany Marlen Valdez-Rodríguez, Claudia Geraldine León-Ramírez, José Ruiz-Herrera, Juan Manuel Adame-Rodríguez, Manuel López-Cabanillas-Lomelí, and Eduardo Campos-Góngora. 2023. "Identification and Characterization of Dmct: A Cation Transporter in Yarrowia lipolytica Involved in Metal Tolerance" Journal of Fungi 9, no. 6: 600. https://doi.org/10.3390/jof9060600





