Assessment of Nuclear Gem Quantity for Evaluating the Efficacy of Antisense Oligonucleotides in Spinal Muscular Atrophy Cells
Abstract
:1. Introduction
2. Experimental Design
2.1. Materials
- Single-channel pipettes (0.5–10 µL, 5–50 µL, 20–200 µL, 100–1000 µL) (Lenpipet, Saint-Petersburg, Russia; Cat. No.: 4027482, 40270282, 40270292, 40270302);
- Culture flask 75 cm2, for work with adhesive cell cultures (TC treated), lid with filter, sterile (Guangzhou JET Bio-Filtration Co. Ltd., Guangzhou, China Cat. No.: TCF012250);
- Plastic sterile containers, volume 120 mL (Medpolymer, Saint-Petersburg, Russia; Cat. No.: 2620304);
- Micro centrifuge tubes (2.0 mL) (BIOplastics BV, Landgraaf, The Netherlands Cat. No.: B71420);
- Microtubes (1.5 mL) (Axygen, Union City, CA, USA; Cat. No.: MCT-150-C);
- PCR tubes, 0.5 µL (SSI, Lodi, CA, USA, Cat. No.: SSI-3320-00);
- 8-well Permanox chamber slide (Thermo Fisher Scientific, Grand Island, NY, USA; Cat. No.: 177445);
- 24-well plates (TC treated) (Sarstedt AG & Co., Nümbrecht, Germany; Cat. No.: 83.3922);
- Hemocytometer counting chamber;
- 200 μL beveled pipette tips (SSI, Lodi, CA, USA; graduated, yellow, SKU: 4220-06);
- Pipette micro tips (0.1–10 μL) (Jet Biofil, Guangzhou, China; Sterile, Cat. no: PPT151010);
- Filtered pipette tips, sterile (1000 μL) (SSI, Lodi, CA, USA; SKU: 4337NSF);
- Pipette tips (1000 μL) (Jiangsu Huida medical instruments Co., Ltd., Yancheng city, China; Cat. no: HP2036-1);
- Cryotubes, sterile (Deltalab S.L., Barcelona, Spain; Cat. No.: 409106.1);
- Nutrient medium DMEM liquid, with stable glutamine (1 L) (alanyl-glutamine) glucose 1 g/L, sterile (Bilot, Russia; Cat. No.: 1.3.6.3.);
- Fetal bovine serum, Gibco (100 mL) (Thermo Fisher Scientific, Waltham, MA, USA; Cat. No.: 10270-098);
- Penicillin-Streptomycin, for cell culture, dry powder, 50,000 U/vial penicillin G and 50 mg/vial streptomycin, sterile (BIOFIL, Saint-Petersburg, Russia; Cat. No.: 1.3.18.);
- Gibco® Trypsin-EDTA (0.25%), phenol red (Thermo Fisher Scientific, Waltham, MA, USA; Cat. No.: 25200056);
- 3UP8 RNA oligonucleotide 5′-GCUGGCAG-3′ with phosphorothioate and 2′-O-methyl modifications [25] (Syntol JSC, Moscow, Russia);
- X-tremeGENE™ siRNA Transfection Reagent (Roche, Mannheim, Germany; SKU: 4476093001);
- Triton X-100 (Helicon, Moscow, Russia; vendor code: SB-G1204-100ML);
- Bovine serum albumin, BSA, 500g (Sigma-Aldrich, Steinheim, Germany; SKU: A3059-500G);
- SMN antibody (2B1) NB100-1936 Unit Size: 0.1 mL (Novus Biologicals, Littleton, CO, USA; Cat. No.: NB100-1936);
- Donkey Anti-Mouse IgG NorthernLights™ NL493-conjugated Antibody (BioTech R&D systems, Minneapolis, MN, USA; Cat. No.: NL009);
- VECTASHIELD® Antifade Mounting Medium with DAPI (H-1200-10) (Vector laboratories, Burlingame, CA, USA; Cat. No.: H-12000);
- Acrylamide 2K Standard grade, extra-pure, 500 g (PanReac AppliChem, Darmstadt, Germany; product code: A1089).;
- Boric acid, chemical grade, 500 g (Helicon, Moscow, Russia; vendor code: H-0202-0.5);
- EDTA, disodium salt, dihydrate 500 g (Helicon, Moscow, Russia; vendor code: H-E5134-0.5);
- Ethidium bromide (Helicon, Moscow, Russia; vendor code: SRL-17220-5G);
- N,N,N′,N′-Tetramethylethylenediamine TEMED (Helicon, Moscow, Russia; vendor code: SB-GC203001);
- Ammonium persulfate (APS) (Merck; Darmstadt, Germany; Cat. No.: h-248614-0.1);
- 1X 0.25% trypsin solution (BioloT, Saint-Petersburg, Russia; Cat. No.: 1.2.2.5.);
- 0.3% Versen’s solution (BioloT, Saint-Petersburg, Russia; Cat. No.: 1.2.3.2.);
- DPBS without Ca2+ and Mg2+ (Sigma-Aldrich, Steinheim, Germany; Cat. No.: 59331C-1000ML);
- 96° ethyl alcohol P.O.A. (Merck, Darmstadt, Germany; Cat. No.: 8.18760.1000);
- TRIzol reagent (Invitrogen, Carlsbad, CA, USA; Cat. No.: 15596026);
- Chloroform (Vekton, Saint-Petersburg, Russia);
- Isopropanol (Vekton, Saint-Petersburg, Russia);
- Dimethylsulfoxide (DMSO) (VWR (Amresco), Montreal, QC, Canada; Cat. No.: Am-0231-0.1);
- M-MLV reverse transcriptase (Sileks, Moscow, Russia; Cat. No.: E1211);
- Random hexa primer, 15 o.u./mL (Sileks, Moscow, Russia; Cat. No.: D0310);
- Mixture of dNTP, 25 mM each (Syntol, Moscow, Russia; Cat. No.: dNTP-100-010);
- Taq polymerase, buffer without Mg2+ and 25 mM MgCl2 (Syntol, Moscow, Russia; Cat. No.: E0120).
2.2. Equipment
- Biosafety Cabinet Class II (Laminar systems, Miass, Russia; Cat. No.: 1R-D.001-12ada);
- DNA/RNA UV-cleaner box UVT-S (BIOSAN, Riga, Latvia);
- Ultraviolet germicidal irradiator (recirculator) Desar (Himmed, Moscow, Russia; Cat. No.: av345);
- Centrifuge for 15 mL tubes up to 2300 g (ELMI Ltd., Riga, Latvia; Cat. No.: Elmi CM-6M);
- Centrifuge MiniSpin (Eppendorf, Hamburg, Germany; Cat. No.: 00000030762);
- CO2 incubator MCO-19AIC (UV) (SANYO Electr.Co., Ltd., Osaka, Japan; Cat. No.: SA-MCO19);
- Incubator +37 °C (Memmert, Schwabach, Germany; Cat. No.: 9537930);
- Two-compartment refrigerator: +4 °C and −20 °C (POZIS, Zelenodolsk, Russia; Cat. No.: 00000031036);
- Refrigerator −80 °C (SANYO Electr.Co., Ltd., Osaka, Japan; Cat. No.: MDF-U32V);
- Inverted microscope MIBR with a digital camera (LOMO, Saint-Petersburg, Russia; Cat. No.: 00000074356);
- Centrifuge mini-vortex microspin (BIOSAN, Riga, Latvia; Cat. No.: 00000026197);
- Refrigerated centrifuge 5417R (Eppendorf, Hamburg, Germany; Cat. No.: 5407000317);
- UV transilluminator (Vilber-Lourmart, Marné La Vallée, France);
- Camera for vertical electrophoresis (Helicon, Moscow, Russia; Cat. No.: VE-20);
- Microscope Leica, DM 2500 (Leica, Wetzlar, Germany).
3. Procedure
3.1. Fibroblast Transfection
 CRITICAL STEP: Twenty minutes before handling the material, turn on the UV lamp and Dezar in the Biosafety Cabinet Class II. Ensure that all the procedures are performed under sterile conditions.
CRITICAL STEP: Twenty minutes before handling the material, turn on the UV lamp and Dezar in the Biosafety Cabinet Class II. Ensure that all the procedures are performed under sterile conditions.- Twenty-four hours prior to transfection: Culture the healthy fibroblast cells as well as those obtained from the patient with SMA II into an 8-well Permanox chamber slide (for protein analysis) or 24-well plate (for transcripts analysis) in DMEM medium with L-glutamine and 10% fetal bovine serum to reach ~50% confluency per well on the day of transfection (volume 250 μL for each well of 8-well Permanox chamber slide and volume 500 μL for 24-well plate). Incubate the plate with cells in a CO2 incubator at 37° C and 5% CO2.
- After 24 h, prepare the transfection complexes of the 3UP8 antisense oligonucleotide with the carrier according to the following: the ratio of X-treme GENE (µL) to RNA (µg) should be 10:1; the molar concentration of 400 nM of the oligonucleotide per well is recommended.
- Add the complexes of the RNA and the carrier to the cells.
- Incubate in the CO2 incubator at 37° C in 5% CO2 for 4 h.
- After that, change the medium in all wells to full DMEM with 10% FBS and antibiotic (penicillin 100 U/mL, streptomycin 100 μg/mL). Add 250 μL of the full medium in each well of the 8-well Permanox chamber slide and 500 μL in each well of the 24-well plate.
- Incubate the plates or slides with cells in a CO2 incubator at 37 °C and 5% CO2 for 48 h.
3.2. Immunocytochemical Staining
- Fixation, permeabilization, and blocking:
- Remove all the medium from the slide wells, and wash the cells with Dulbecco’s PBS without Ca2+ and Mg2+ (1x) (about 100 µL volume for each well). After each wash, all the drops should be removed with pipetting. Then, add 50 µL of 4% paraformaldehyde–PBS in each well and incubate for 10 min at room temperature (RT) to allow cells to be fixed into the bottom of the slide.
- Remove all the liquid, and wash the cells with PBS. Repeat this procedure one more time.
- Add 50 µL of 0.1% Triton X-100 in PBS in each well, and incubate at RT for 5 min for permeabilization.
- Remove all the liquid, and wash the cells again with PBS;
- Add 50 µL of 1% BSA (Albumin, bovine serum) freshly prepared in PBS, and incubate for 1 h at RT to reduce the background fluorescence.
- Antibody Incubation:
- 6.
- Prepare primary mouse anti-SMN (2B1) antibody dilution (5 µg/mL) in 1% BSA.
- 7.
- Remove all the liquid from the wells, and add 50 µL of primary antibody solution.
- 8.
- Incubate the cells overnight at 4 °C with the primary antibody.
- 9.
- Next day: remove all the liquid from the wells, and wash the cells with PBS three times for 5 min each.
- 10.
- Prepare a dilution of secondary antibody Anti-Mouse IgG NL493-conjugated Donkey antibody (1:200 in PBS).
- 11.
- Add 50 µL of secondary antibody solution in each well.
- 12.
- Incubate for 1 h at RT with secondary antibody in a dark environment.
- Mounting and Imaging:
- 13.
- Remove all the liquid, and wash the cells in PBS three times, each time for 5 min.
- 14.
- Take away the upper part of the chamber to obtain flat slides for microscope analysis.
- 15.
- Mount the cells in Vectashield with DAPI to stain the nuclei (about 10 µL for each well).
- 16.
- Put on the coverslip, and seal with nail polish.
3.3. RNA Isolation and cDNA Synthesis
 CRITICAL STEP: The following protocol is to be performed in a DNA/RNA UV-cleaner box.
CRITICAL STEP: The following protocol is to be performed in a DNA/RNA UV-cleaner box.- Remove the medium from the plate, then wash with 200 μL of 1x PBS (without Ca2+, Mg2+).
- Add 200 μL trypsin-Versene solution (1:3) to each well to detach the cells.
- Incubate for 10 min in a CO2 incubator at +37 °C.
- Inactivate the trypsin by adding 300 μL of PBS to each well.
- Resuspend the cells, then transfer them into 1.5 mL tubes and centrifuge at 2200 r.p.m. for 10 min at +4 °C.
- Remove the supernatant, add 125 μL of TRIzol reagent to each tube, resuspend, and incubate for 5 min at RT.
- After that, add 25 μL of chloroform; mix and incubate for 3 min at RT.
- Centrifuge the resulting mixture at 12,000 r.p.m. for 15 min at +4 °C.
- Place the upper transparent phase in 1.5 mL tubes, then add 63 μL of isopropanol.
- Stir the tubes, and place them overnight at −70 °C.
- Next day: Centrifuge the mixture at 10,000 r.p.m. for 20 min at +4 °C.
- Remove the supernatant, and wash the residue with 125 μL of cooled 70% ethanol.
- Centrifuge the tubes at 14,000 r.p.m. for 4 min at +4 °C, and then dry the residue for an hour.
- Add 20 μL of water treated with diethyl pyrocarbonate (DEPC) to the residue, and dissolve within 40 min by periodically stirring on a vortex.
- Using a first strand cDNA synthesis kit and random primers, 700 ng of total RNA should be reverse-transcribed following the manufacturer’s protocol instructions.
3.4. Semiquantitative RT-PCR
- Prepare the PCR mix: 1 µL of 10x PCR buffer with MgCl2, 0.31 mM of each dNTP, 1 µM of each primer, and 5 U of Taq DNA polymerase.
- Add 1 µL of cDNA from the total volume of 28 µL to the PCR mix.
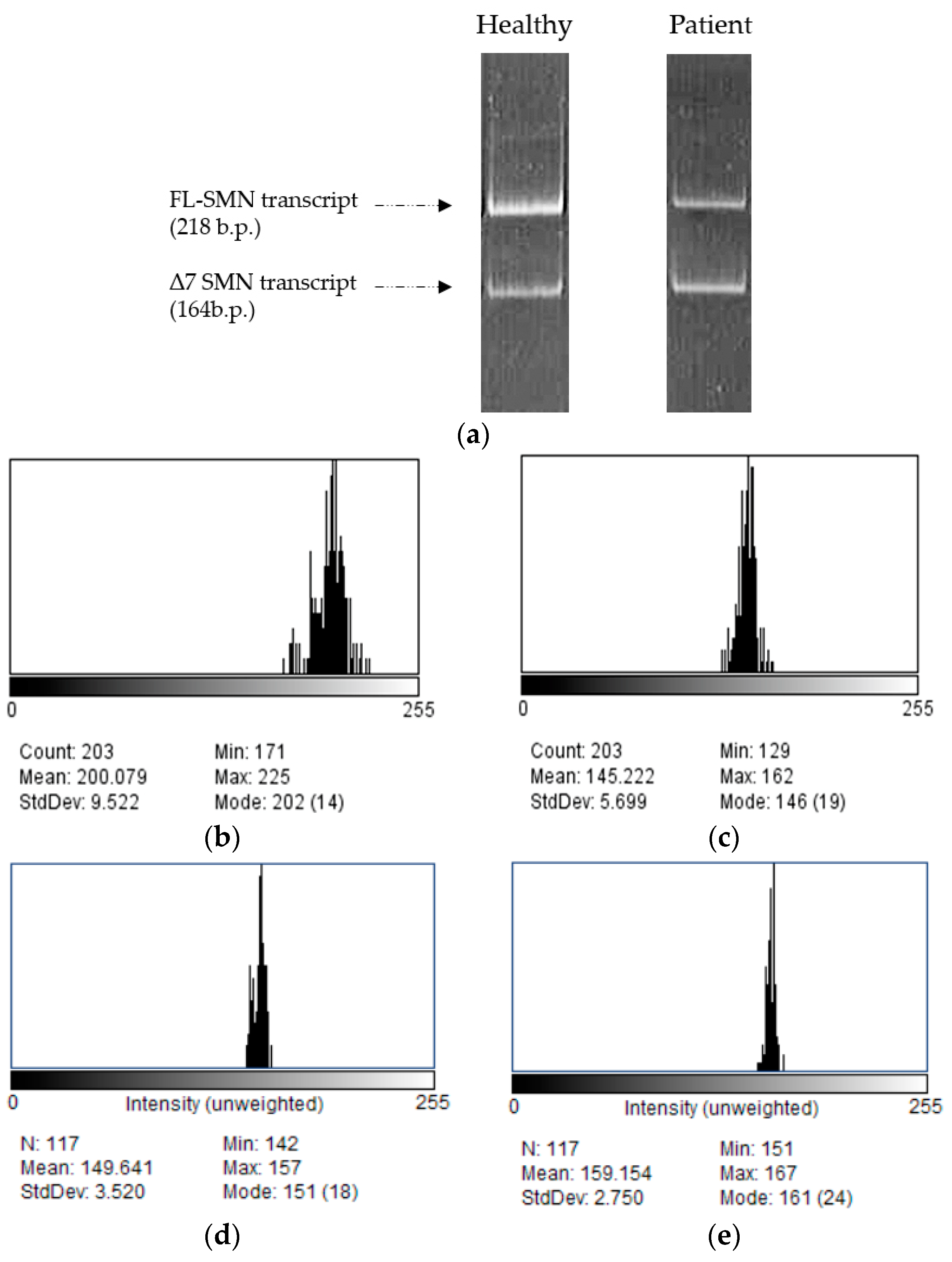
- We recommend using the following primers for full-length and Δ7 SMN transcript amplification: SMN F 5′-GTCCAGATTCTCTTGATGAT-3′, complementary to SMN exon 6 region, and SMN R 5′-CTATAACGCTTCACATTCCA-3′, complementary to SMN exon 8 region [26].
- The amplification reaction is to be conducted at 94 °C for 4 min, with n cycles of 94 °C for 45 s, 50 °C for 45 s, 72 °C for 45 s, and final synthesis at 72 °C for 8 min.
- The number of cycles (n) should not exceed 26–28 to stop the reaction on the exponential phase.
- Amplification of each cDNA sample should be performed at least 2 times.
3.5. Polyacrylamide Gel Electrophoresis
- Perform 6% polyacrylamide gel electrophoresis for 80 min at voltage 360 V and current 80 mA.
- Stain the gel in a solution of ethidium bromide (0.5 μg/mL).
- The results of polyacrylamide gel electrophoresis are to be recorded on a UV transilluminator.
- Electrophoresis is processed using densitometric analysis in the software ImageJ v. 1.54d (NIH, Bethesda, MD, USA).
- The proportion of full-length SMN transcripts as well as the correlation factor are calculated using the program Microsoft® Office Excel® 2007.
3.6. Statistical Analysis
4. Expected Results
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- D’Amico, A.; Mercuri, E.; Tiziano, F.D.; Bertini, E. Spinal muscular atrophy. Orphanet J. Rare Dis. 2011, 6, 1–10. [Google Scholar] [CrossRef]
- Wirth, B.; Brichta, L.; Hahnen, E. Spinal Muscular Atrophy: From Gene to Therapy. Semin. Pediatr. Neurol. 2006, 13, 121–131. [Google Scholar] [CrossRef] [PubMed]
- Zerres, K.; Wirth, B.; Rudnik-Schöneborn, S. Spinal muscular atrophy—Clinical and genetic correlations. Neuromuscul. Disord. 1997, 7, 202–207. [Google Scholar] [CrossRef] [PubMed]
- Russman, B.S. Spinal muscular atrophy: Clinical classification and disease heterogeneity. J. Child Neurol. 2007, 22, 946–951. [Google Scholar] [CrossRef] [PubMed]
- Mercuri, E.; Sumner, C.J.; Muntoni, F.; Darras, B.T.; Finkel, R.S. Spinal muscular atrophy. Nat. Rev. Dis. Prim. 2022, 8, 52. [Google Scholar] [CrossRef]
- Lefebvre, S.; Bürglen, L.; Reboullet, S.; Clermont, O.; Burlet, P.; Viollet, L.; Benichou, B.; Cruaud, C.; Millasseau, P.; Zeviani, M.; et al. Identification and characterization of a spinal muscular atrophy-determining gene. Cell 1995, 80, 155–165. [Google Scholar] [CrossRef] [PubMed]
- Pellizzoni, L.; Yong, J.; Dreyfuss, G. Essential role for the SMN complex in the specificity of snRNP assembly. Science 2002, 298, 1775–1779. [Google Scholar] [CrossRef]
- Monani, U.R.; Lorson, C.L.; Parsons, D.W.; Prior, T.W.; Androphy, E.J.; Burghes, A.H.M.; McPherson, J.D. A single nucleotide difference that alters splicing patterns distinguishes the SMA gene SMN1 from the copy gene SMN2. Hum. Mol. Genet. 1999, 8, 1177–1183. [Google Scholar] [CrossRef]
- Wirth, B.; Brichta, L.; Schrank, B.; Lochmüller, H.; Blick, S.; Baasner, A.; Heller, R. Mildly affected patients with spinal muscular atrophy are partially protected by an increased SMN2 copy number. Hum. Genet. 2006, 119, 422–428. [Google Scholar] [CrossRef]
- Patrizi, A.L.; Tiziano, F.; Zappata, S.; Donati, M.A.; Neri, G.; Brahe, C. SMN protein analysis in fibroblast, amniocyte and CVS cultures from spinal muscular atrophy patients and its relevance for diagnosis. Eur. J. Hum. Genet. 1999, 7, 301–309. [Google Scholar] [CrossRef]
- Liu, Q.; Dreyfuss, G. A novel nuclear structure containing the survival of motor neurons protein. EMBO J. 1996, 15, 3555–3565. [Google Scholar] [CrossRef] [PubMed]
- Staněk, D.; Neugebauer, K.M. The Cajal body: A meeting place for spliceosomal snRNPs in the nuclear maze. Chromosoma 2006, 115, 343–354. [Google Scholar] [CrossRef] [PubMed]
- Cacciottolo, R.; Ciantar, J.; Lanfranco, M.; Borg, R.M.; Vassallo, N.; Bordonné, R.; Cauchi, R.J. SMN complex member Gemin3 self-interacts and has a functional relationship with ALS-linked proteins TDP-43, FUS and Sod1. Sci. Rep. 2019, 9, 18666. [Google Scholar] [CrossRef] [PubMed]
- Coovert, D.D.; Le, T.T.; McAndrew, P.E.; Strasswimmer, J.; Crawford, T.O.; Mendell, J.R.; Coulson, S.E.; Androphy, E.J.; Prior, T.W.; Burghes, A.H.M. The survival motor neuron protein in spinal muscular atrophy. Hum. Mol. Genet. 1997, 6, 1205–1214. [Google Scholar] [CrossRef] [PubMed]
- Lefebvre, S.; Burlet, P.; Liu, Q.; Bertrandy, S.; Clermont, O.; Munnich, A.; Dreyfuss, G.; Melki, J. Correlation between severity and SMN protein level in spinal muscular atrophy. Nat. Genet. 1997, 16, 265–269. [Google Scholar] [CrossRef] [PubMed]
- Ebert, A.D.; Yu, J.; Rose, F.F.; Mattis, V.B.; Lorson, C.L.; Thomson, J.A.; Svendsen, C.N. Induced pluripotent stem cells from a spinal muscular atrophy patient. Nature 2009, 457, 277–280. [Google Scholar] [CrossRef] [PubMed]
- Mattis, V.B.; Rai, R.; Wang, J.; Chang, C.W.T.; Coady, T.; Lorson, C.L. Novel aminoglycosides increase SMN levels in spinal muscular atrophy fibroblasts. Hum. Genet. 2006, 120, 589–601. [Google Scholar] [CrossRef] [PubMed]
- Andreassi, C.; Angelozzi, C.; Tiziano, F.D.; Vitali, T.; De Vincenzi, E.; Boninsegna, A.; Villanova, M.; Bertini, E.; Pini, A.; Neri, G.; et al. Phenylbutyrate increases SMN expression in vitro: Relevance for treatment of spinal muscular atrophy. Eur. J. Hum. Genet. 2004, 12, 59–65. [Google Scholar] [CrossRef]
- Wolstencroft, E.C. A non-sequence-specific requirement for SMN protein activity: The role of aminoglycosides in inducing elevated SMN protein levels. Hum. Mol. Genet. 2005, 14, 1199–1210. [Google Scholar] [CrossRef]
- Riessland, M.; Brichta, L.; Hahnen, E.; Wirth, B. The benzamide M344, a novel histone deacetylase inhibitor, significantly increases SMN2 RNA/protein levels in spinal muscular atrophy cells. Hum. Genet. 2006, 120, 101–110. [Google Scholar] [CrossRef]
- Lumpkin, C.J.; Harris, A.W.; Connell, A.J.; Kirk, R.W.; Whiting, J.A.; Saieva, L.; Pellizzoni, L.; Burghes, A.H.M.; Butchbach, M.E.R. Evaluation of the orally bioavailable 4-phenylbutyrate-tethered trichostatin A analogue AR42 in models of spinal muscular atrophy. Sci. Rep. 2023, 13, 10374. [Google Scholar] [CrossRef]
- Coady, T.H.; Baughan, T.D.; Shababi, M.; Passini, M.A.; Lorson, C.L. Development of a single vector system that enhances Trans-splicing of SMN2 transcripts. PLoS ONE 2008, 3, e3468. [Google Scholar] [CrossRef]
- Grigor’eva, E.V.; Valetdinova, K.R.; Ustyantseva, E.I.; Shevchenko, A.I.; Medvedev, S.P.; Mazurok, N.A.; Maretina, M.A.; Kuranova, M.L.; Kiselev, A.V.; Baranov, V.S.; et al. Neural differentiation of patient-specific induced pluripotent stem cells from patients with a hereditary form of spinal muscular atrophy. Genes Cells 2016, 11, 70–79. [Google Scholar]
- Nozdracheva, A.; Pleskach, N.; Kuranova, M. Features of DNA repair in dermal fibroblasts in patients with breast cancer and persons with medical history of cancer. RAP Conf. Proc. 2020, 9973, 38–43. [Google Scholar]
- Singh, N.N.; Shishimorova, M.; Cao, L.C.; Gangwani, L.; Singh, R.N. A short antisense oligonucleotide masking a unique intronic motif prevents skipping of a critical exon in spinal muscular atrophy. RNA Biol. 2009, 6, 341–350. [Google Scholar] [CrossRef]
- Maretina, M.; Egorova, A.; Lanko, K.; Baranov, V.; Kiselev, A. Evaluation of Mean Percentage of Full-Length SMN Transcripts as a Molecular Biomarker of Spinal Muscular Atrophy. Genes 2022, 13, 1911. [Google Scholar] [CrossRef]

Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Al-Hilal, H.; Maretina, M.; Egorova, A.; Glotov, A.; Kiselev, A. Assessment of Nuclear Gem Quantity for Evaluating the Efficacy of Antisense Oligonucleotides in Spinal Muscular Atrophy Cells. Methods Protoc. 2024, 7, 9. https://doi.org/10.3390/mps7010009
Al-Hilal H, Maretina M, Egorova A, Glotov A, Kiselev A. Assessment of Nuclear Gem Quantity for Evaluating the Efficacy of Antisense Oligonucleotides in Spinal Muscular Atrophy Cells. Methods and Protocols. 2024; 7(1):9. https://doi.org/10.3390/mps7010009
Chicago/Turabian StyleAl-Hilal, Haya, Marianna Maretina, Anna Egorova, Andrey Glotov, and Anton Kiselev. 2024. "Assessment of Nuclear Gem Quantity for Evaluating the Efficacy of Antisense Oligonucleotides in Spinal Muscular Atrophy Cells" Methods and Protocols 7, no. 1: 9. https://doi.org/10.3390/mps7010009






