Abstract
A multi-channel microfluidic system was designed and fabricated with an integrated electrode array to be capable of trapping and analysing single cells or populations in the individual channels in a controlled chemical environment. The analytical system was interfaced with a dedicated printed circuit board designed for parallel EIS and optical screening. The continuous impedance measurement also monitors the adequate filling of the channels and the cell trapping events. Yeast cells were characterised by recording the EIS spectra of individual channels, allowing the differentiation between the populations considering their size, viability, or proliferation. The EIS analysis was supported by fluorescent optical microscopy, also using cell staining.
1. Introduction
The application of organ-on-chip devices, designed for the in vitro study of cell populations or even single cells in a chemical environment, can be a significant step forward in drug discovery and screening and developing of therapeutic strategies [1,2]. Application-specific devices integrating microfluidic and sensing systems are capable of simultaneously maintaining artificial cell populations and tissues, treating them with chemical agents, and monitoring their physiological behaviour in real time. This multifunctional behaviour is crucial in personalised medicine and drug development, with further prospects for antibiotic susceptibility testing.
Electrochemical impedance spectroscopy (EIS) can be used to characterise different properties of cells and cell cultures, allowing non-invasive and real-time measurements. By integrating microelectrodes, the microfluidic structure can be adapted to perform impedance-based cell analysis, including cell trapping and growth monitoring. In biological applications, a wide range of frequencies from 1 Hz to 10 GHz is used, with the distinct dispersions regions α, β, and γ [3]. The combined range of α and β dispersions (10 Hz–1 MHz) refers to both cellular and electrode responses.
In this work, a compact multi-channel microfluidic system was created with an integrated electrode array that enables fast, precise, and multi-parallel EIS measurements with single-cell resolution and additional optical feedback.
2. Materials and Methods
The microfluidic layout was inspired by Taylor et al., although with significant modifications for cell trapping and electrode integration [4]. It consists of two main channels connected by 15 smaller cross-channels that narrow vertically. The electrodes were designed to monitor each channel individually (Figure 1a). A dedicated interfacing PCB was designed and manufactured to perform measurements on individual single to four parallel channels connected to a PalmSens4 (PalmSens BV, Houten, The Netherlands) potentiostat (Figure 1b). Data evaluation and circuit fitting can be achieved with the associated controlling and analysing software. A hole in the bottom centre of the PCB provides insight into the channels and allows optical screening in parallel with the EIS measurements.
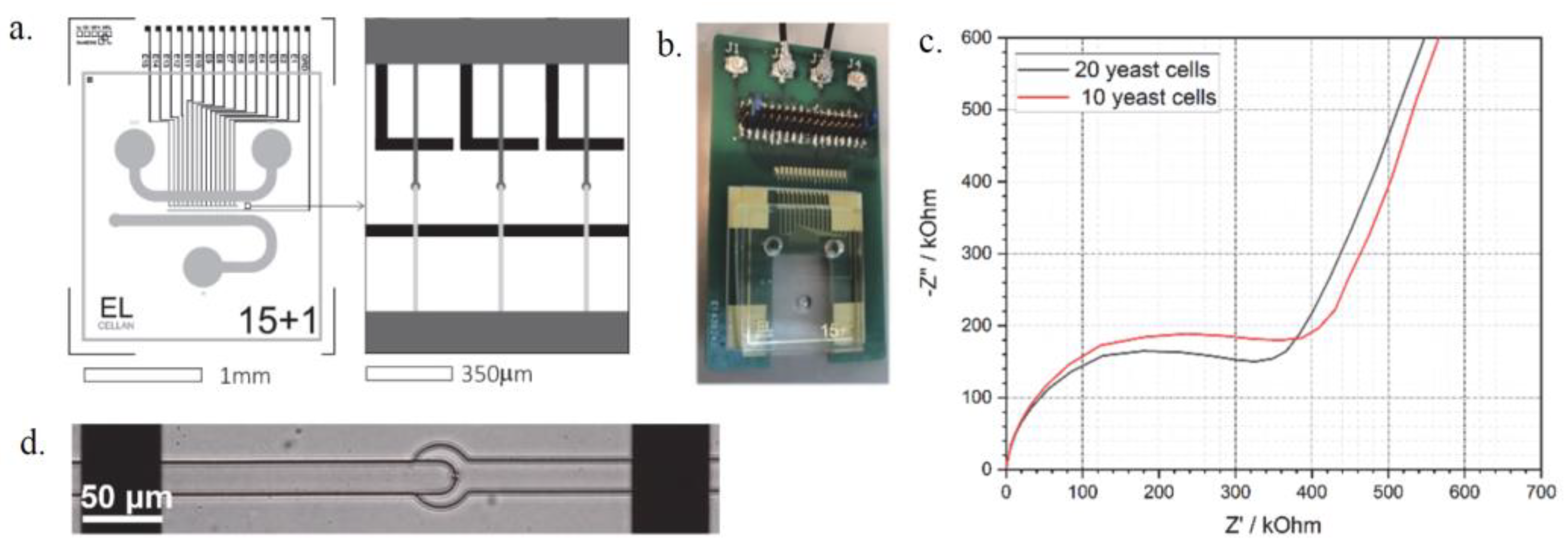
Figure 1.
The layout and electrode design of the microfluidic chip (a) and the assembled device (b) the EIS spectra of trapped yeast cells (c) and a single cell trapped between the electrodes (d).
The electrodes were prepared by vacuum evaporation of Au layer onto BorofloatTM glass substrate, applying Ti adhesive layer, and patterned via lift-off lithography. The electrodes are coated with SU-8 photoresist insulating layer with contact windows opened. Their distance is 300 µm, which enables manual aligning with the channels under a microscope. The microfluidic chip was fabricated by standard soft lithography and replica moulding of Polydimethylsiloxane (PDMS) on 4” wafers. Two-step photolithography was applied to form the cell trapping structures. The cross-channels narrow vertically from 10 µm to 5 µm in this case, which is optimal for trapping yeast cells. Raw PDMS material was modified by PDMS-b-PEO molecules to render the channels hydrophilic. The PDMS was cured at 65 °C for 2 h, and every chip was individually cleaned in an ultrasonic shaking bath in DI water and IPA and aligned to the electrode array.
3. Discussion
A microfluidic system with underlying electrodes and a PCB was designed to simultaneously trap cells and monitor their characteristics. Yeast cells (S. cerevisiae) were trapped in the narrowing channels between the electrodes and were analysed via EIS spectroscopy. The device can measure the difference in cell number (Figure 1c) and can even trap single cells (Figure 1d) that can be studied individually. The proposed microfluidic system offers possibilities for cell viability testing or growth monitoring in critical chemical environments combined with EIS measurements and optical screening with potential applications in drug efficiency tests or in antibiotic susceptibility testing.
Author Contributions
Chip design, methodology, L.B. and P.F.; manufacturing, analysis, L.B; writing, L.B. and P.F. All authors have read and agreed to the published version of the manuscript.
Funding
The work was supported by ECSEL JU and the Ministry of Innovation and Technology of Hungary from the NKFIA Fund via Moore4Medical H2020-ECSEL-2019-IA-876190 and INBIOM TKP2021-EGA-04 grants.
Institutional Review Board Statement
Not applicable.
Informed Consent Statement
Not applicable.
Data Availability Statement
Data are available in this manuscript.
Conflicts of Interest
The authors declare no conflicts of interest.
References
- Low, L.A.; Mummery, C.; Berridge, B.R.; Austin, C.P.; Tagle, D.A. Organs-on-chips: Into the next decade. Nat. Rev. Drug Discov. 2020, 20, 345–361. [Google Scholar] [CrossRef] [PubMed]
- Li, Z.; Hui, J.; Yang, P.; Mao, H. Microfluidic organ-on-a-chip system for disease modeling and drug development. Biosensors 2022, 12, 370. [Google Scholar] [CrossRef] [PubMed]
- Asami, K. Characterization of biological cells by dielectric spectroscopy. J. Non-Cryst. Solids 2002, 305, 268–277. [Google Scholar] [CrossRef]
- Taylor, A.M.; Blurton-Jones, M.; Rhee, S.W.; Cribbs, D.H.; Cotman, C.W.; Jeon, N.L. A microfluidic culture platform for CNS axonal injury, regeneration and transport. Nat. Methods 2005, 2, 599–605. [Google Scholar] [CrossRef] [PubMed]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).