Plant Disease Prognosis Using Spatial-Exploitation-Based Deep-Learning Models †
Abstract
:1. Introduction
2. Literature Review
3. Methodology
4. Proposed Approach
4.1. Infrastructure and Tools
4.2. Predictive Analytics Process
- Defining a Project: Identification and definition of research objectives, scope, and datasets used for experimentation.
- Data Gathering: Preparation and formulation of data through data-mining techniques from multiple sources.
- Data Analysis: Preprocessing stages, such as resizing, normalizing, and modeling data, to extract usable information and draw conclusions.
- Statistics: Validation of hypotheses and assumptions through statistical analysis using appropriate models.
- Modelling: Creation of precise predictive models automatically, allowing for multiple evaluations to select the optimal solution.
- Deployment: Automating decisions based on the models to integrate analytical results into routine decision-making processes, generating results, reports, and output.

4.3. Knowledge-Based Expert Systems for Crop Disease Diagnosis
4.4. Plant Disease Diagnosis
- Accurate Plant Identification: Identifying the infected plants, including scientific and generic names.
- Distinguishing Characteristics: Recognizing the distinctive traits of healthy and diseased parts, accounting for variations in patterns, coloration, and growth rates.
- Symptom and Sign Analysis: Identifying specific symptoms, such as stunted growth, tissue overgrowth, tissue death, and variations in appearance. Differentiating between symptoms and analyzing ecological causative agents.
- Affected-Plant-Part Detection: Noting which plant parts are affected, such as roots, leaves, or stems.
- Symptom Distribution: Observing the spread of affected plants in the area, noting patterns and distributions.
- Host Specificity: Determining if the issue affects specific plant species or multiple species, aiding in understanding potential causes.
4.5. Plant Disease Management
- Exclusion: Preventing disease spread through geographical barriers and local prevention methods.
- Eradication: Eliminating the disease after introduction but before widespread dissemination.
- Protection: Implementing barriers, either mechanical, temporal, or economic, to prevent infection.
- Resistance: Using disease-resistant plants as a primary prevention method.
- Integrated Disease Management (IDM): Employing a combination of tactics, methods, disease diagnosis, and environmental monitoring to manage diseases effectively.
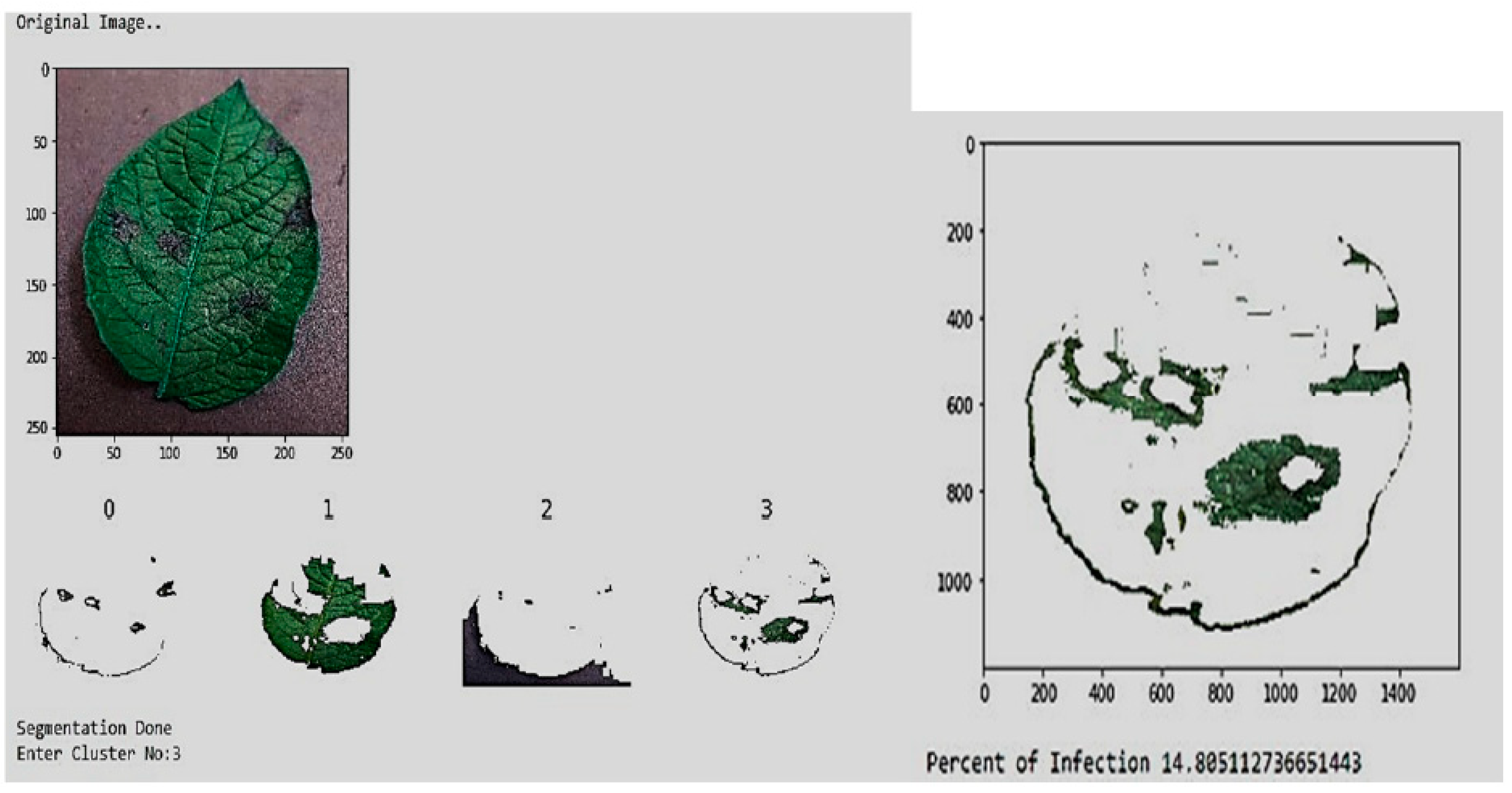
4.6. Methodology: Deep CNN and Otsu-Based Image Segmentation
- Data Acquisition: Utilizing real-time field images and the “PlantVillage” dataset, dividing the data into training, validation, and testing sets.
- Model Construction: Creating a multiclass multilayer CNN architecture suited for processing various images independent of size or orientation.
- Training and Validation: Scaling, normalizing, and training the model iteratively on the dataset to adapt to different images.
- Classification: Employing the trained deep CNN to categorize images into predefined classes, assessing its real-time performance on unseen images.

4.7. Algorithm
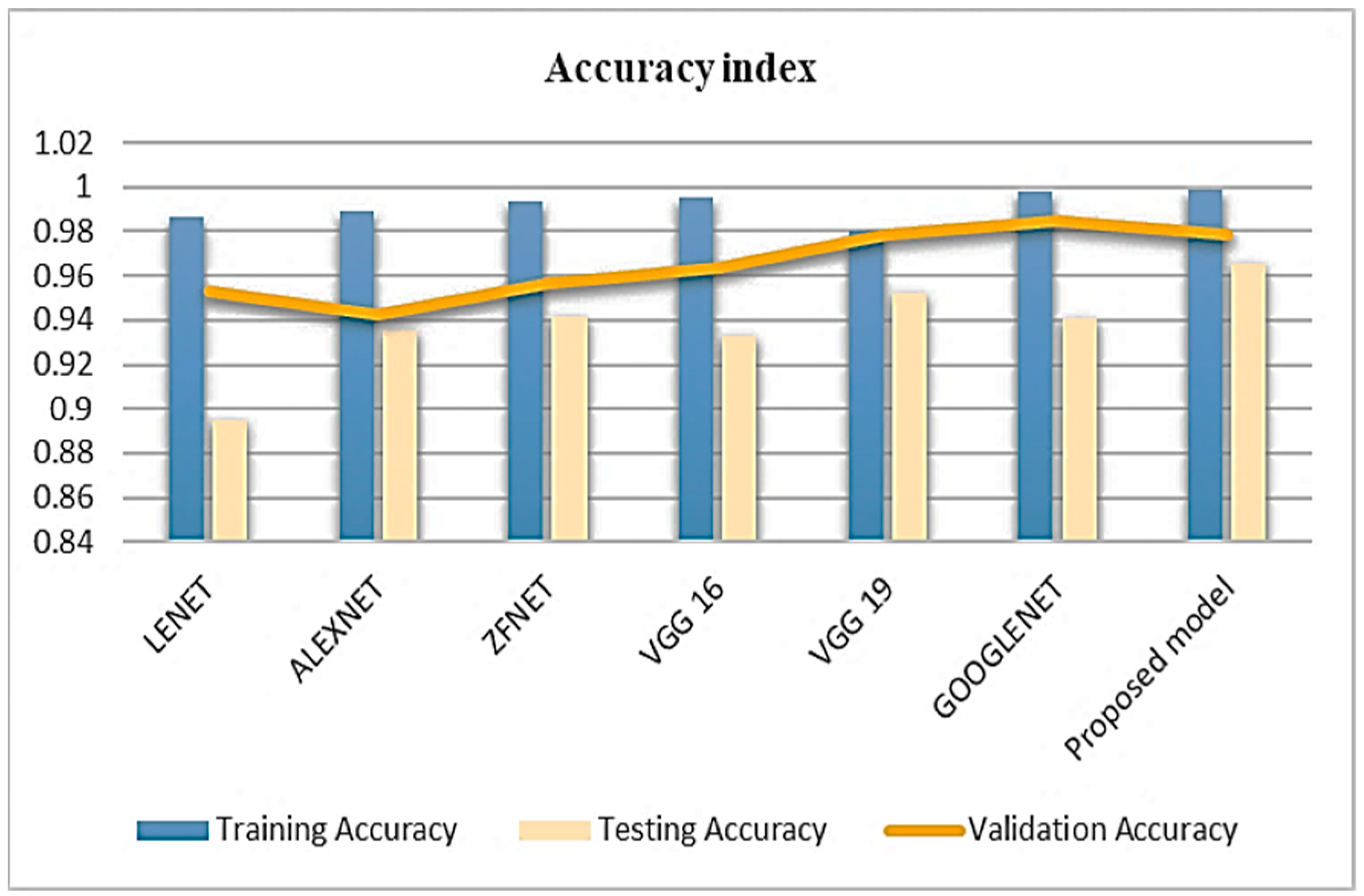
5. Results and Discussion
6. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Picon, A.; Seitz, M.; Alvarez-Gila, A.; Mohnke, P.; Ortiz-Barredo, A.; Echazarra, J. Crop conditional Convolutional Neural Networks for massive multi-crop plant disease classification over cell phone acquired images taken on real field conditions. Comput. Electron. Agric. 2019, 167, 105093. [Google Scholar] [CrossRef]
- Chaudhury, A.; Barron, J.L. Plant species identification from occluded leaf images. IEEE/ACM Trans. Comput. Biol. Bioinform. 2018, 17, 1042–1055. [Google Scholar] [CrossRef] [PubMed]
- Richey, B.; Majumder, S.; Shirvaikar, M.; Kehtarnavaz, N. Real-time detection of maize crop disease via a deep learning-based smartphone app. In Real-Time Image Processing and Deep Learning; SPIE: Bellingham, DC, USA, 2020; Volume 11401, pp. 23–29. [Google Scholar]
- Yang, C.K.; Lee, C.Y.; Wang, H.S.; Huang, S.C.; Liang, P.I.; Chen, J.S.; Kuo, C.F.; Tu, K.H.; Yeh, C.Y.; Chen, T.D. Glomerular disease classification and lesion identification by machine learning. Biomed. J. 2022, 45, 675–685. [Google Scholar] [CrossRef] [PubMed]
- Shrestha, Y.R.; Krishna, V.; von Krogh, G. Augmenting organizational decision-making with deep learning algorithms: Principles, promises, and challenges. J. Bus. Res. 2021, 123, 588–603. [Google Scholar] [CrossRef]
- Cevallos, C.; Ponce, H.; Moya-Albor, E.; Brieva, J. Vision-based analysis on leaves of tomato crops for classifying nutrient deficiency using convolutional neural networks. In Proceedings of the 2020 International Joint Conference on Neural Networks (IJCNN), Glasgow, UK, 19–24 July 2020; pp. 1–7. [Google Scholar]
- Veys, C.; Chatziavgerinos, F.; AlSuwaidi, A.; Hibbert, J.; Hansen, M.; Bernotas, G.; Smith, M.; Yin, H.; Rolfe, S.; Grieve, B. Multispectral imaging for presymptomatic analysis of light leaf spot in oilseed rape. Plant Methods 2019, 15, 4. [Google Scholar] [CrossRef] [PubMed]
- Argüeso, D.; Picon, A.; Irusta, U.; Medela, A.; San-Emeterio, M.G.; Bereciartua, A.; Alvarez-Gila, A. Few-Shot Learning approach for plant disease classification using images taken in the field. Comput. Electron. Agric. 2020, 175, 105542. [Google Scholar] [CrossRef]
- Ashourloo, D.; Aghighi, H.; Matkan, A.A.; Mobasheri, M.R.; Rad, A.M. An investigation into machine learning regression techniques for the leaf rust disease detection using hyperspectral measurement. IEEE J. Sel. Top. Appl. Earth Obs. Remote Sens. 2016, 9, 4344–4351. [Google Scholar] [CrossRef]
- Nazki, H.; Yoon, S.; Fuentes, A.; Park, D.S. Unsupervised image translation using adversarial networks for improved plant disease recognition. Comput. Electron. Agric. 2020, 168, 105117. [Google Scholar] [CrossRef]
- Chen, J.; Zhang, D.; Nanehkaran, Y.A. Identifying plant diseases using deep transfer learning and enhanced lightweight network. Multimed. Tools Appl. 2020, 31, 497–515. [Google Scholar] [CrossRef]
- Golhani, K.; Balasundram, S.K.; Vadamalai, G.; Pradhan, B. A review of neural networks in plant disease detection using hyperspectral data. Inf. Process. Agric. 2018, 5, 354–371. [Google Scholar] [CrossRef]
- Krishnan, V.G.; Deepa, J.R.; Rao, P.V.; Divya, V.; Kaviarasan, S. An automated segmentation and classification model for banana leaf disease detection. J. Appl. Biol. Biotechnol. 2022, 10, 213–220. [Google Scholar]
- Ngugi, L.C.; Abelwahab, M.; Abo-Zahhad, M. Recent advances in image processing techniques for automated leaf pest and disease recognition–A review. Inf. Process. Agric. 2021, 8, 27–51. [Google Scholar] [CrossRef]
- Brahimi, M.; Arsenovic, M.; Laraba, S.; Sladojevic, S.; Boukhalfa, K.; Moussaoui, A. Deep learning for plant diseases: Detection and saliency map visualisation. In Human and Machine Learning: Visible, Explainable, Trustworthy and Transparent; Springer: Cham, Switzerland, 2018; pp. 93–117. [Google Scholar]
- Lamba, M.; Gigras, Y.; Dhull, A. Classification of plant diseases using machine and deep learning. Open Comput. Sci. 2021, 11, 491–508. [Google Scholar] [CrossRef]
- Harakannanavar, S.S.; Rudagi, J.M.; Puranikmath, V.I.; Siddiqua, A.; Pramodhini, R. Plant leaf disease detection using computer vision and machine learning algorithms. Glob. Transit. Proc. 2022, 3, 305–310. [Google Scholar] [CrossRef]
- Vadivel, T.; Suguna, R. Automatic recognition of tomato leaf disease using fast enhanced learning with image processing. Acta Agric. Scand. Sect. B—Soil Plant Sci. 2022, 72, 312–324. [Google Scholar] [CrossRef]
- Sujatha, R.; Chatterjee, J.M.; Jhanjhi, N.Z.; Brohi, S.N. Performance of deep learning vs machine learning in plant leaf disease detection. Microprocess. Microsyst. 2021, 80, 103615. [Google Scholar] [CrossRef]
- Essah, R.; Anand, D.; Singh, S. An intelligent cocoa quality testing framework based on deep learning techniques. Meas. Sens. 2022, 24, 100466. [Google Scholar] [CrossRef]


| Reference | Methodology | Finding | Limitation | Advantage |
|---|---|---|---|---|
| [4] | Machine learning for disease detection | Enhanced accuracy in diagnosis | Dependent on data quality and quantity | Rapid, noninvasive diagnosis |
| [5] | Automated diagnosis challenges and opportunities | Integration of technology in agriculture | Limited access to advanced technology | Potential for early intervention |
| [6] | Convolutional neural networks | Improved early detection | Model complexity and training time | High accuracy and speed |
| [7] | Multi spectral image processing | Training and testing process | Model complexity | Proof of concept |
| [8] | Few shot learning approach | Fast processing | Complex model | Improved Speed |
| [9] | Regression technique | Hyperspectral images | Less accuracy | Improved speed |
| [10] | Deep learning for disease detection | Accurate multiclass classification | Requires large labeled datasets | Robust and scalable detection |
| [11] | Deep transfer learning | Enhanced network | Requires less data | Improved Accuracy |
| [12] | Hyperspectral images and machine learning | Enhanced spectral disease detection | Hardware and cost limitations | Improved spectral resolution |
| [13] | Deep-learning model for citrus diseases | High accuracy in citrus disease identification | Limited to specific diseases | Accurate and quick diagnosis |
| [14] | New image processing Techniques | Improved accuracy | Time delay in processing | Accuracy improved |
| [15] | Review of deep-learning techniques | Comprehensive overview of deep-learning applications | Lack of standardization | Wide applicability and effectiveness |
| [16] | AI and ML applications | Diverse AI and ML applications | Lack of interpretability | Broad coverage of AI techniques |
| [17] | Computer vision with ML algorithms | Integration of technique | Improved interpretability | Improved accuracy |
| [18] | Machine learning in smart agriculture | Integration of AI in agriculture | Infrastructure constraints | Enhanced efficiency and productivity |
| [19] | Deep learning for disease classification | High accuracy in classification | Data imbalance issues | Effective for large datasets |
| Models | Transfer Learning | Training from Scratch | ||||||
|---|---|---|---|---|---|---|---|---|
| A | P | R | F1 | A | P | R | F1 | |
| LENET | 0.97 | 0.96 | 0.97 | 0.94 | 0.92 | 0.94 | 0.90 | 0.90 |
| ALEXNET | 0.98 | 0.97 | 0.98 | 0.96 | 0.94 | 0.92 | 0.92 | 0.90 |
| ZFNET | 0.97 | 0.99 | 0.99 | 0.98 | 0.94 | 0.91 | 0.91 | 0.91 |
| VGG-16 | 0.99 | 0.99 | 0.99 | 0.98 | 0.94 | 0.93 | 0.91 | 0.91 |
| VGG-19 | 0.99 | 0.99 | 0.99 | 0.99 | 0.95 | 0.95 | 0.93 | 0.94 |
| GOOGLENET | 0.99 | 0.99 | 0.99 | 0.99 | 0.97 | 0.96 | 0.94 | 0.95 |
| Number of Epochs | Training Accuracy | Training Loss | Validation Accuracy | Validation Loss |
|---|---|---|---|---|
| 25 | 0.8965 | 0.3865 | 0.9025 | 0.2145 |
| 40 | 0.9251 | 0.2456 | 0.9365 | 0.156 |
| 50 | 0.9564 | 0.0952 | 0.9657 | 0.123 |
| 75 | 0.9765 | 0.1365 | 0.9898 | 0.1021 |
| 100 | 0.8678 | 0.5862 | 0.8742 | 0.4658 |
| Number of Epochs | Learning Rate | Dropout Rate | Training Accuracy | Validation Accuracy | Training Loss | Validation Loss |
|---|---|---|---|---|---|---|
| 25 | 0.001 | 0.25 | 0.9354 | 0.8624 | 0.4362 | 0.4521 |
| 50 | 0.0001 | 0.25 | 0.9264 | 0.8951 | 0.3561 | 0.3125 |
| 75 | 0.1 | 0.15 | 0.9021 | 0.8999 | 0.2531 | 0.2001 |
| 75 | 0.001 | 0.25 | 0.9985 | 0.9854 | 0.1254 | 0.1564 |
| 75 | 0.0001 | 0.40 | 0.8694 | 0.832 | 0.3214 | 0.2154 |
| 75 | 0.00001 | 0.50 | 0.8216 | 0.8021 | 0.2145 | 0.5641 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Vankara, J.; Nandini, S.S.; Muddada, M.K.; Kuppili, N.S.C.; Naidu, K.S. Plant Disease Prognosis Using Spatial-Exploitation-Based Deep-Learning Models. Eng. Proc. 2023, 59, 137. https://doi.org/10.3390/engproc2023059137
Vankara J, Nandini SS, Muddada MK, Kuppili NSC, Naidu KS. Plant Disease Prognosis Using Spatial-Exploitation-Based Deep-Learning Models. Engineering Proceedings. 2023; 59(1):137. https://doi.org/10.3390/engproc2023059137
Chicago/Turabian StyleVankara, Jayavani, Sekharamahanti S. Nandini, Murali Krishna Muddada, N. Satya Chitra Kuppili, and K Sowjanya Naidu. 2023. "Plant Disease Prognosis Using Spatial-Exploitation-Based Deep-Learning Models" Engineering Proceedings 59, no. 1: 137. https://doi.org/10.3390/engproc2023059137






