Abstract
The most critical issues in computational biology are characterizing and predicting uncharacterized proteins’ secondary and tertiary structures from their uploaded amino acid sequences in databases. Aedes albopictus (A. albopictus), sometimes referred to as the Asian tiger mosquito or forest mosquito and the carrier of dengue-like diseases, has many proteins, many of which are still poorly understood. The current work aims at elucidating the physicochemical properties and structures of the as-yet-uncharacterized A. albopictus protein AEW48448.1. ExPASy Protaram, CD Search, SOPMA, PSIPRED, and other advanced computerized tools were used following the standard flowchart for characterizing a hypothetical protein to ascertain the roles and structures of AEW48448.1. After identifying the protein’s secondary and tertiary structures, the structures were evaluated for quality using tools like PROCHECK and the ProSA-web. Later, the active site was also discovered using CASTp v3.0. The protein is more stable because it has a higher aliphatic index value and more negatively charged than positively charged residues. The modeling of the proteins’ 2D and 3D structures using multiple bioinformatics tools confirmed that they had domains, indicating that they were functional proteins involved in the host’s antiviral, cytokine, and interferon production pathways. Additionally, the protein was revealed to have active regions where ligands may bind. This work aims at elucidating the characteristics and structures of an uncharacterized A. albopictus protein that may serve as a therapeutic target for the creation of antiviral candidates and vaccines.
1. Introduction
A. albopictus spreads the arboviruses Dengue (DENV), Zika virus (ZIKV), and Chikungunya (CHIKV) [1]. In temperate and tropical parts of the planet, this mosquito is a strong and very invasive species [2]. Understanding the basic biology of A. albopictus is crucial for preventing its aggressive spread, as its exceptional adaptability has led to its rapid expansion and global public concern [3]. Studies estimate that nearly 3.9 billion people worldwide are at risk of contracting DENV, putting about half of the world’s population at risk. Despite the lack of a specific cure for severe dengue, early discoveries and access to quality medical care significantly reduce its fatality rates [4]. The structure of cells and organisms is one of the many roles that proteins carry out within living things. They also participate in several crucial processes in vivo through interactions with other molecules [5]. Uncovering the biological characteristics and activities of these uncharacterized proteins from various animals is currently a frequent activity in the domain of bioinformatics because there are millions of proteins that are still uncharacterized [6,7]. A. albopictus contains several functional proteins, many of which are still unknown or are poorly understood [8,9]. Developments in computational biology have led to the development of numerous platforms and techniques for predicting protein structure, binding sites, and biological activity [10,11]. Protein studies use bioinformatics techniques to classify novel domains, evaluate 3D structural conformations, and determine function [12]. Furthermore, this comprehensive understanding can provide effective pharmacological tactics for developing potential treatments for a variety of disorders [13].
2. Materials and Methods
2.1. Selection of the Hypothetical Protein and Sequence Retrieval
We obtained the amino acid (aa) sequence for cytochrome c oxidase subunit I, partial mitochondrion protein (A. albopictus), in FASTA format from the NCBI database [14].
2.2. Physicochemical Properties Analysis
Using the ExPASy server ProtParam tool, we found the theoretical isoelectric point (pI), the instability index, the aliphatic index, the GRAVY (which measures how hydrophobic or hydrophilic a protein is), the extinction coefficients, and the amino acid sequence composition [15].
2.3. Identification and Validation of the Secondary Structure
We applied the self-optimized prediction method with alignment (SOPMA) to anticipate secondary structural elements [16].
2.4. Three-Dimensional Structure Prediction and Validation
We predicted the three-dimensional structure of the chosen protein using the SWISS-MODEL [17] and AlphaFold [18]. Furthermore, we used the PROCHECK software of the SAVES program (v.6.1) to structurally validate the modeled 3D protein structure [19].
2.5. Ligand Binding Site or Active Site Determination
We used the DeepSite tool [20], PrankWeb server [21], and CASTp v.3.0 server [22] to predict the active sites of the modeled protein.
2.6. Pathogenecity Prediction
We used CDPred to predict the celiac disease-causing peptides from the protein’s FASTA sequences [23]. We also utilized the FASTA sequence to construct the protein’s 3D structure and detect disease-causing protein mutations. We utilized the FASTA sequence to forecast and scrutinize the location of a protein sequence mutation. Additionally, we made use of a variety of services, including Polyphen-2, SIFT, predictSNP, MAPP, PhD-SNP, and SNAP [24]. The Align-GVGD program also used substitution, predicted by predictSNP, and the FASTA format of the amino acid (AA) sequence to calculate GV and GD [25].
3. Results and Discussion
3.1. Protein Sequence Retrieval
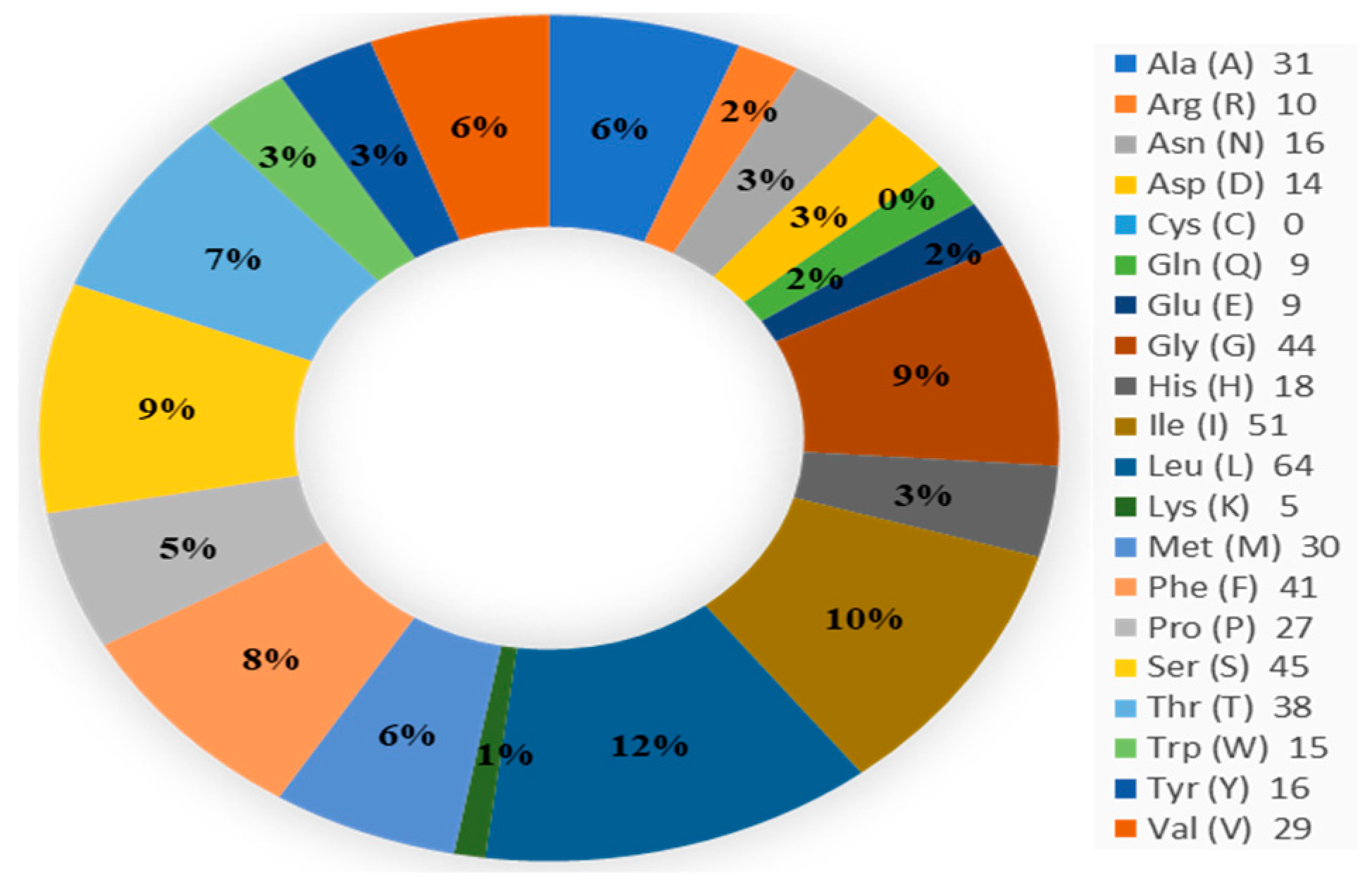
The amino acid (aa) sequence of the cytochrome c oxidase subunit I, partial mitochondrial protein of A. albopictus (AEW48448.1), was obtained from the NCBI database. We modeled the tertiary structure of the protein using the 152 amino acid-long protein sequence (Figure 1).
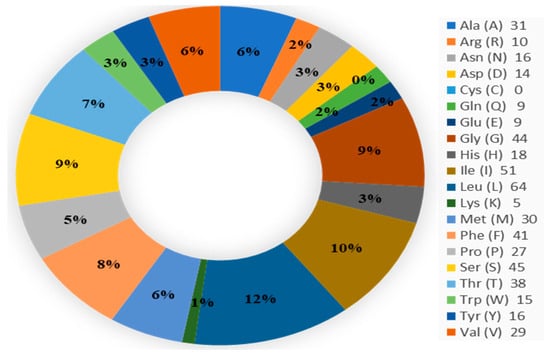
Figure 1.
Amino acid composition.
3.2. Identification of the Physicochemical Properties
We used ProtParam ExPASy to measure physicochemical parameters using the amino acid sequence of the protein AEW48448.1 found in A. albopictus. We retrieved the sequence in FASTA format. The instability index, at 30.29, indicates the protein’s stability with an index smaller than 40 [26]. The protein’s hypothetical pI (pH 6.10) indicates that it is in a weak acidic form [5]. There is 56,800.02 Dalton in the molecular weight, 110.08 in the aliphatic index, 30.29 in the instability index, and 0.731 in the GRAVY, respectively.
The protein has a higher aliphatic index value of 110.08, which is a positive indicator of greater thermostability over a broad temperature range [27]. A GRAVY index value of −0.731 revealed the protein’s hydrophilic nature and its potential for increased water interaction [19]. In vitro, Escherichia coli had an estimated half-life of >10 h; yeast had an estimated half-life of >20 h; and mammalian reticulocytes had an estimated half-life of 1.9 h.
3.3. Identification and Validation of the Secondary Structure
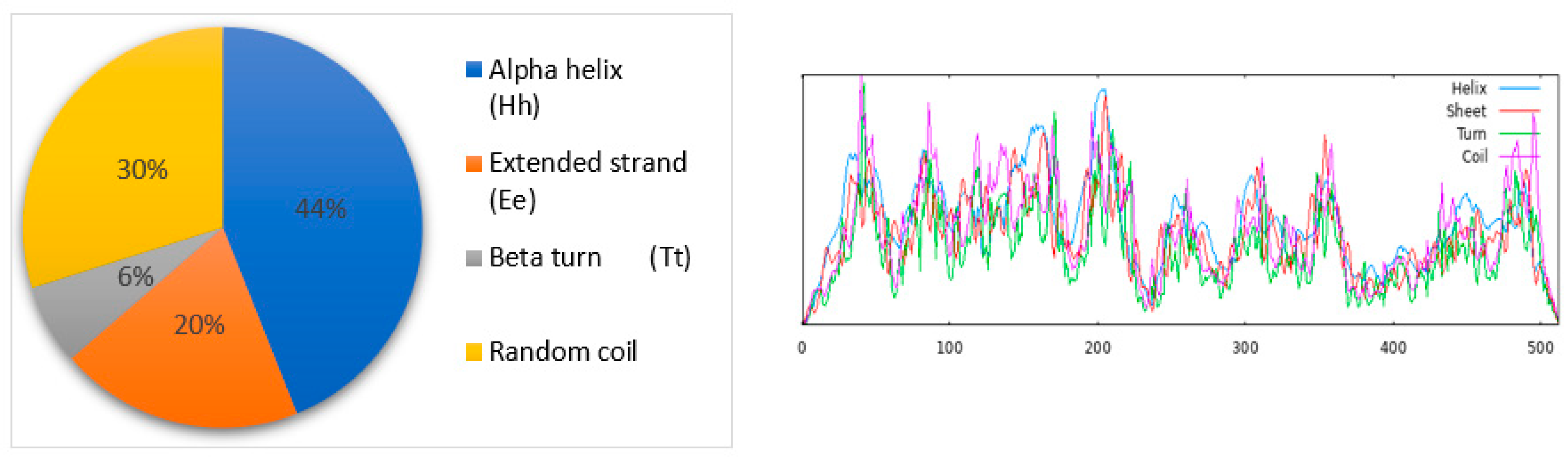
The cytochrome c oxidase subunit I partial (mitochondrion) protein (A. albopictus) predicted secondary structural components using the SOPMA program’s default parameters (window width of 17, number of states of 4, and similarity threshold of 8) (Figure 2).
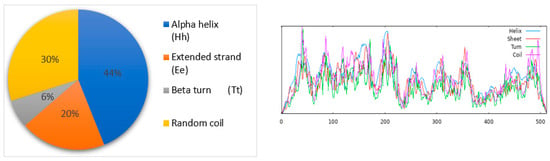
Figure 2.
The SOPMA tool identified the alpha helix (blue color), extended strand (red color), beta-turn (light-green color), and random coil (purple color) as the secondary structural elements of the selected protein.
3.4. The Three-Dimensional Protein Structure Anticipation and Assessment
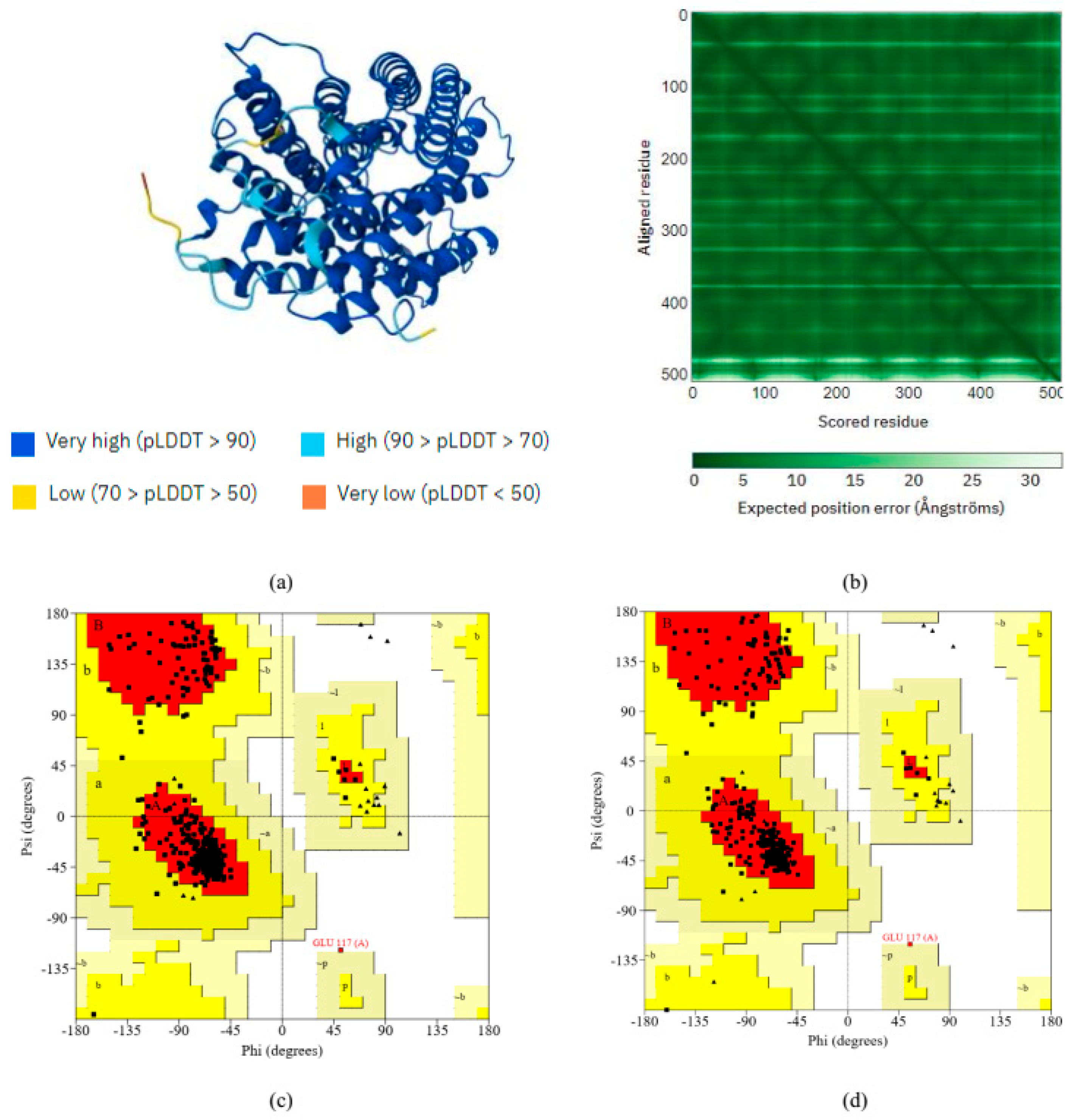
We used the SAVESv6.1 program’s PROCHECK tool to assess the modeled structures from AlphaFold and SWISS-MODEL servers [5]. We used the Ramachandran plot to assess the quality of the predicted model (by SWISS-MODEL), finding 404 (92.7%) of the total 510 residues in the core, 6.9% in the additional allowed regions, and 0.2% in the generously allowed regions (a, b, l, and p). There was a total of 436 residues that were not glycine or proline. There were three end-residues that were not Gly or Pro, and 44 and 27 glycine and proline residues out of a total of 510 residues (Figure 3). We also examined the protein residues (structure predicted by AlphaFold) in the most favored regions (94.5%), additional allowed regions (5.2%), generously allowed regions (0.0%), and disallowed regions (0.2%) (Figure 3). The quality assessment analysis showed that AlphaFold’s predicted structure outperformed the SWISS-MODEL in terms of quality.
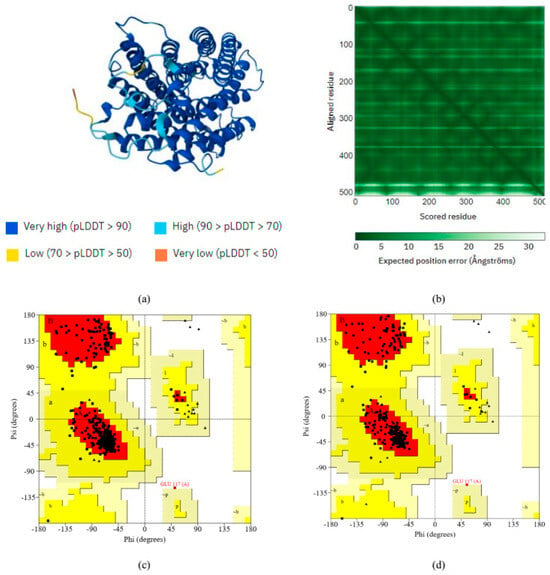
Figure 3.
(a) AlphaFold predicted the selected protein’s tertiary structure. (b) AlphaFold’s per-residue model confidence score. We used the Ramachandran plot analyses by (c) AlphaFold and (d) SWISS-MODEL to assess the quality of the selected protein. When modeled using the SWISS-MODEL method, 92.7% of the selected proteins fell within the most preferred region. However, AlphaFold’s prediction of the selected protein indicates its superior quality (94.5%).
3.5. Ligand Binding Site or Active Site Determination
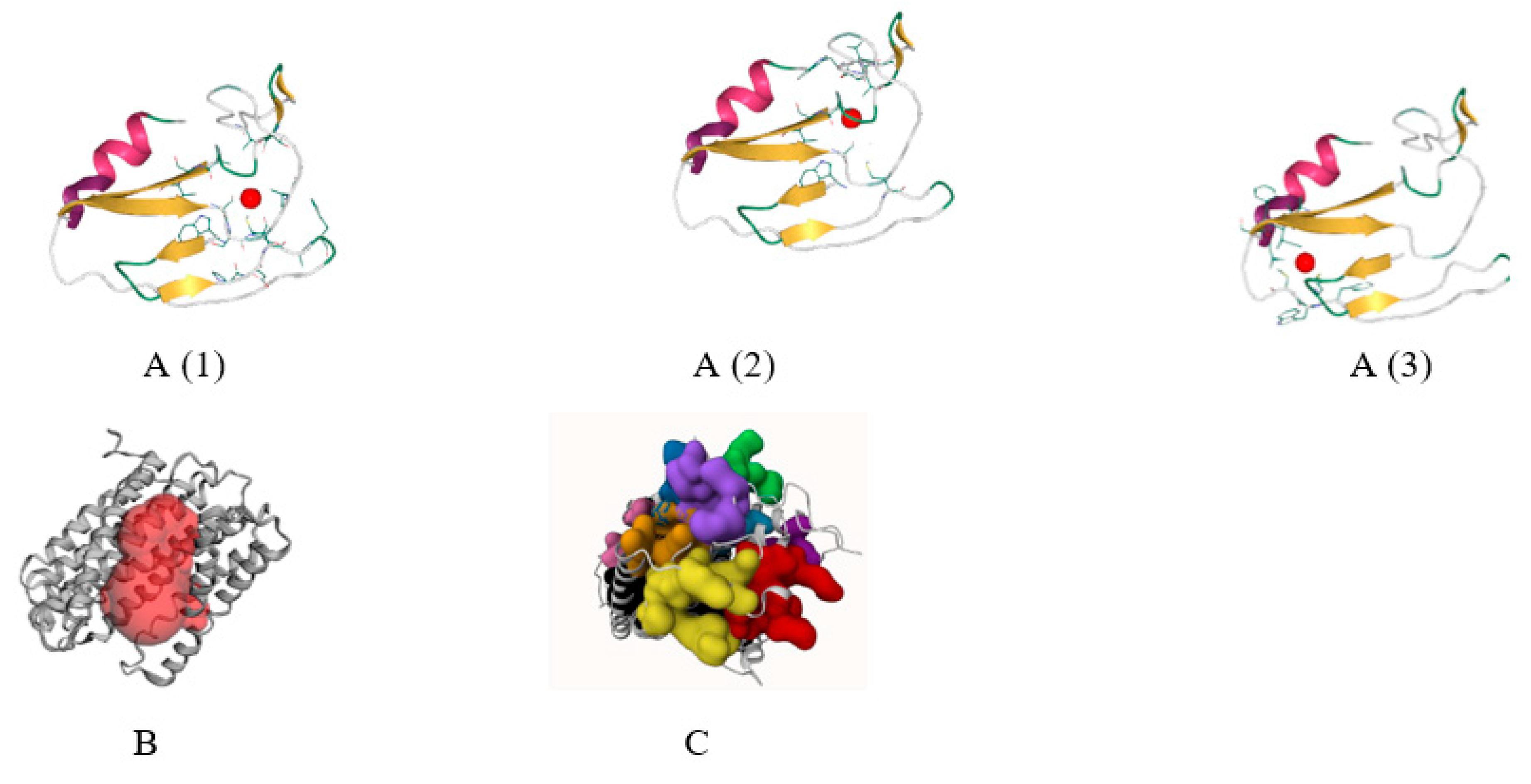
Ligand binding cavity prediction is particularly significant for drug design because it cleverly utilizes the protein’s geometric, chemical, and evolutionary properties. One prediction tool, DeepSite, can predict the protein’s active site where a drug or protein can bind to reduce protein toxicity [28]. DeepSite predicted that the AEW48448.1 protein contained three cavities (Figure 4A). Another ligand-binding tool, PrankWeb, predicted 12 pockets in the given protein (Figure 4C). CASTp could detect regions on proteins, outline them, identify their dimensions, and compute their areas [5]. The highest active sites were located between the modeled protein’s 800.210 and 1784.976 areas (Figure 4B).
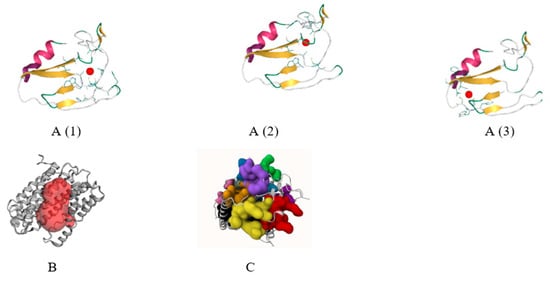
Figure 4.
DeepSite tool (A(1)–A(3)), the CASTp server (B), and the PrankWeb server (C) all use the active site determination method.
3.6. Pathogenecity Prediction
CDPred generates a tabular output of the peptides from the proteins that cause celiac disease, making it possible to predict disease-associated peptides. The table shows the amino acid sequence, the start position, the end position, and the physicochemical properties of the celiac disease-causing peptide from the protein sequence [23]. Table 1 summarizes the protein’s results (sequence id: AEW48448.1).

Table 1.
Prediction of disease associated peptides and its properties.
Predicting SNP, MAPP, PhD-SNP, Polyphen-2, SIFT, and SNAP makes it simple to identify pathogenic SNVs. Compared to other mutation prediction servers, Mutation Assessor has the highest sensitivity for all substitutions evaluated by the predict SNP server (except H41Q). Table 2 lists the values for the measured parameters. All parameters indicate that M63P, F65M, M69P, M95S, N96M, N97V, M98E, S99N, F100G, and F416P have high predictive accuracy, while F92L is likely to have high variable apparent accuracy when compared to T479P. We ranked all parameters from highly deleterious to less predicted, as shown in Table 2.

Table 2.
Prediction of disease-related mutation.
The Align-GVGD program’s GD and GV values are crucial as they indicate the distance between the mutant amino acid and the permitted variation, as indicated by GV [29]. We predicted all missense mutants in the protein to be neutral, harmful, or unclassified using the following Align-GVGD criteria: If the substitute falls into the box, then GD = 0. If not, then GD is higher than 0. Therefore, according to the MSA, GD is a measure of the biochemical difference between the mutant and the observed variation at that location [30]. Out of the 12 mutants we know of, M63P, M69P, M95S, N96M, N97V, M98E, F100G, and F416P are very likely to go through missense substitutions, while S99N, T479P, F65M, and F92L have very little chance (Table 3).

Table 3.
Prediction of missense substitutions.
4. Conclusions
Using advanced bioinformatics methods to characterize a protein is another novel challenge, similar to various system biology activities. Our research aims to shed light on the physicochemical properties, chemical structures, and biological roles of the putative protein AEW48448.1 from A. albopictus. The protein contains amino acids that have more negatively charged residues, and it is more temperature-stable due to its high aliphatic index and low instability index. Using a variety of bioinformatics techniques, we modeled the protein’s secondary structure and confirmed that it included a domain, indicating its functional nature. Numerous servers verified the protein’s precise 3D structure through tertiary structure prediction. Furthermore, the study discovered an active site in the protein where ligands could bind. More research is needed on the protein to discover innovative treatment options for dengue, which is a result of A. albopictus’s targeting of the protein.
Author Contributions
Conceptualization, methodology, software, and validation, M.A.A., S.A.S. and A.S.M.S.; formal analysis, M.A.A., S.A.S., M.M., R.D.N., A.S.M.S. and M.E.U.; investigation, A.S.M.S. and M.A.A.; resources, and data curation A.S.M.S., M.A.A. and S.A.S.; writing—original draft preparation, M.A.A. and S.A.S.; writing—review and editing, A.S.M.S.; visualization, M.A.A., S.A.S., M.M., R.D.N., A.S.M.S. and M.E.U. supervision, A.S.M.S. and M.E.U.; project administration, A.S.M.S. and M.E.U. All authors have read and agreed to the published version of the manuscript.
Funding
This research received no external funding.
Data Availability Statement
The original contributions presented in the study are included in the article, further inquiries can be directed to the corresponding author.
Conflicts of Interest
The authors declare no conflicts of interest.
References
- Gloria-Soria, A.; Payne, A.F.; Bialosuknia, S.M.; Stout, J.; Mathias, N.; Eastwood, G.; Ciota, A.T.; Kramer, L.D.; Armstrong, P.M. Vector competence of Aedes albopictus populations from the Northeastern United States for chikungunya, dengue, and Zika Viruses. Am. J. Trop. Med. Hyg. 2021, 104, 1123. [Google Scholar] [CrossRef]
- Ryan, S.J.; Carlson, C.J.; Mordecai, E.A.; Johnson, L.R. Global expansion and redistribution of Aedes-borne virus transmission risk with climate change. PLoS Neglected Trop. Dis. 2019, 13, e0007213. [Google Scholar] [CrossRef] [PubMed]
- Xie, L.M.; Yin, X.; Bi, J.; Luo, H.M.; Cao, X.J.; Ma, Y.W.; Liu, Y.L.; Su, J.W.; Lin, G.L.; Guo, X.G. Identification of potential biomarkers in dengue via integrated bioinformatic analysis. PLoS Neglected Trop. Dis. 2021, 15, e0009633. [Google Scholar] [CrossRef] [PubMed]
- Murray, N.E.A.; Quam, M.B.; Wilder-Smith, A. Epidemiology of dengue: Past, present and future prospects. Clin. Epidemiol. 2013, 5, 299–309. [Google Scholar] [PubMed]
- Al Asad, M.; Shorna, S.A.; Saikat, A.S.; Uddin, E. Structure-Based Functional Annotation of an Uncharacterized Conserved Protein of Acinetobacter baumannii: An in-silico Approach. Eng. Proc. 2023, 37, 25. [Google Scholar] [CrossRef]
- Zhao, J.; Cao, Y.; Zhang, L. Exploring the computational methods for protein-ligand binding site prediction. Comput. Struct. Biotechnol. J. 2020, 18, 417–426. [Google Scholar] [CrossRef]
- Dukka, B.K. Structure-based Methods for Computational Protein Functional Site Prediction. Comput. Struct. Biotechnol. J. 2013, 8, e201308005. [Google Scholar] [CrossRef] [PubMed]
- Martin-Martin, I.; Smith, L.B. Aedes albopictus D7 Salivary Protein Prevents Host Hemostasis and Inflammation. Biomolecules 2020, 10, 1372. [Google Scholar] [CrossRef] [PubMed]
- Saboia-Vahia, L.; Borges-Veloso, A.; Cuervo, P.; Junqueira, M.; Mesquita-Rodrigues, C.; Britto, C.; Domont, G.B.; De Jesus, J.B. Protein expression in the midgut of sugar-fed Aedes albopictus females. Parasites Vectors 2012, 5, 1–25. [Google Scholar] [CrossRef]
- Gong, J.; Chen, Y.; Pu, F.; Sun, P.; He, F.; Zhang, L.; Li, Y.; Ma, Z.; Wang, H. Understanding membrane protein drug targets in computational perspective. Curr. Drug Targets 2019, 20, 551–564. [Google Scholar] [CrossRef]
- Rahman, M.F.; Rahman, M.R.; Islam, T.; Zaman, T.; Shuvo, M.A.; Hossain, M.T.; Islam, M.R.; Karim, M.R.; Moni, M.A. A bioinformatics approach to decode core genes and molecular pathways shared by breast cancer and endometrial cancer. Inform. Med. Unlocked 2019, 17, 100274. [Google Scholar] [CrossRef]
- Mills, C.L.; Beuning, P.J.; Ondrechen, M.J. Biochemical functional predictions for protein structures of unknown or uncertain function. Comput. Struct. Biotechnol. J. 2015, 13, 182–191. [Google Scholar] [CrossRef] [PubMed]
- F de Azevedo, W. Molecular dynamics simulations of protein targets identified in Mycobacterium tuberculosis. Curr. Med. Chem. 2011, 18, 1353–1366. [Google Scholar] [CrossRef] [PubMed]
- Sayers, E.W.; Bolton, E.E.; Brister, J.R.; Canese, K.; Chan, J.; Comeau, D.C.; Connor, R.; Funk, K.; Kelly, C.; Kim, S.; et al. Database resources of the National Center for Biotechnology Information. Nucleic Acids Res. 2021, 49, D10–D17. [Google Scholar] [CrossRef]
- Gasteiger, E.; Gattiker, A.; Hoogland, C.; Ivanyi, I.; Appel, R.D.; Bairoch, A. ExPASy: The proteomics server for in-depth protein knowledge and analysis. Nucleic Acids Res. 2003, 31, 3784–3788. [Google Scholar] [CrossRef] [PubMed]
- Geourjon, C.; Deleage, G. SOPMA: Significant improvements in protein secondary structure prediction by consensus prediction from multiple alignments. Bioinformatics 1995, 11, 681–684. [Google Scholar] [CrossRef]
- Waterhouse, A.; Bertoni, M.; Bienert, S.; Studer, G.; Tauriello, G.; Gumienny, R.; Heer, F.T.; de Beer, T.A.; Rempfer, C.; Bordoli, L.; et al. SWISS-MODEL: Homology modelling of protein structures and complexes. Nucleic Acids Res. 2018, 46, W296–W303. [Google Scholar] [CrossRef] [PubMed]
- Varadi, M.; Bertoni, D.; Magana, P.; Paramval, U.; Pidruchna, I.; Radhakrishnan, M.; Tsenkov, M.; Nair, S.; Mirdita, M.; Yeo, J.; et al. AlphaFold Protein Structure Database in 2024: Providing structure coverage for over 214 million protein sequences. Nucleic Acids Res. 2024, 52, D368–D375. [Google Scholar] [CrossRef]
- Hollingsworth, S.A.; Karplus, P.A. A fresh look at the Ramachandran plot and the occurrence of standard structures in proteins. Biomol. Concepts 2010, 1, 271–283. [Google Scholar] [CrossRef]
- Jiménez, J.; Doerr, S.; Martínez-Rosell, G.; Rose, A.S.; De Fabritiis, G. DeepSite: Protein-binding site predictor using 3D-convolutional neural networks. Bioinformatics 2017, 33, 3036–3042. [Google Scholar] [CrossRef] [PubMed]
- Jendele, L.; Krivak, R.; Skoda, P.; Novotny, M.; Hoksza, D. PrankWeb: A web server for ligand binding site prediction and visualization. Nucleic Acids Res. 2019, 47, W345–W349. [Google Scholar] [CrossRef] [PubMed]
- Binkowski, T.A.; Naghibzadeh, S.; Liang, J. CASTp: Computed atlas of surface topography of proteins. Nucleic Acids Res. 2003, 31, 3352–3355. [Google Scholar] [CrossRef]
- Tomer, R.; Patiyal, S.; Dhall, A.; Raghava, G.P. Prediction of celiac disease associated epitopes and motifs in a protein. Front. Immunol. 2023, 14, 1056101. [Google Scholar] [CrossRef] [PubMed]
- Bendl, J.; Stourac, J.; Salanda, O.; Pavelka, A.; Wieben, E.D.; Zendulka, J.; Brezovsky, J.; Damborsky, J. PredictSNP: Robust and accurate consensus classifier for prediction of disease-related mutations. PLoS Comput. Biol. 2014, 10, e1003440. [Google Scholar] [CrossRef] [PubMed]
- Tavtigian, S.V.; Deffenbaugh, A.M.; Yin, L.; Judkins, T.; Scholl, T.; Samollow, P.B.; de Silva, D.; Zharkikh, A.; Thomas, A. Comprehensive statistical study of 452 BRCA1 missense substitutions with classification of eight recurrent substitutions as neutral. J. Med. Genet. 2006, 43, 295–305. [Google Scholar] [CrossRef]
- Sahay, A.; Piprodhe, A.; Pise, M. In silico analysis and homology modeling of strictosidine synthase involved in alkaloid biosynthesis in catharanthus roseus. J. Genet. Eng. Biotechnol. 2020, 18, 44. [Google Scholar] [CrossRef] [PubMed]
- Ikai, A. Thermostability and aliphatic index of globular proteins. J. Biochem. 1980, 88, 1895–1898. [Google Scholar] [PubMed]
- Bhowmick, S.; Saha, A.; AlFaris, N.A.; ALTamimi, J.Z.; ALOthman, Z.A.; Aldayel, T.S.; Wabaidur, S.M.; Islam, M.A. Structure-based identification of galectin-1 selective modulators in dietary food polyphenols: A pharmacoinformatics approach. Mol. Divers. 2022, 26, 1697–1714. [Google Scholar] [CrossRef]
- Mathe, E.; Olivier, M.; Kato, S.; Ishioka, C.; Hainaut, P.; Tavtigian, S.V. Computational approaches for predicting the biological effect of p53 missense mutations: A comparison of three sequence analysis based methods. Nucleic Acids Res. 2006, 34, 1317–1325. [Google Scholar] [CrossRef]
- Montenegro, L.R.; Lerário, A.M.; Nishi, M.Y.; Jorge, A.A.; Mendonca, B.B. Performance of mutation pathogenicity prediction tools on missense variants associated with 46, XY differences of sex development. Clinics 2021, 76, e2052. [Google Scholar] [CrossRef] [PubMed]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).