1. Introduction
Heart rate variability (HRV) signals are extricated from an electrocardiogram (ECG), a technique utilized in many disciplines including the study of cardiac and non-cardiac diseases, such as myocardial infarction (
MI) [
1], sudden cardiac death (SCD) and ventricular arrhythmias [
2], diabetes mellitus (DM) [
3], and hypertension [
4]. Patients suffering from congestive heart failure (CHF) present with a depressed or low HRV. Moreover, minute variations in HRV signals are difficult to identify, as these signals contain baseline shifts in ECG signals, along with noise. It is difficult and challenging to analyze these signals with traditional methods. The HRV signal parameters are affected by instantaneous variation [
5], respiration [
6], and motion artifacts [
7]. To minimize the obstacles of manual and visual interpretation, computer aided diagnostic (CAD) techniques are used to analyze the HRV signals.
Around the world, there are about 26 million people suffering from CHF [
8]. This pathophysiological condition prevents the heart from circulating enough blood around the body. These different types of health pathologies reduce the ventricle’s ability to pump blood [
9]. The common indications of CHF include edema, dyspnea, and fatigue [
8,
9], myocardial infarction (
MI), heart valve disease, etc. [
10]. Patients suffering from CHF are prone to cardiac death [
11]. Thus, we need to develop automated tools which can help us to investigate the underlying hidden dynamics at the earliest stages, so that concerned cardiologists can take the relevant measures to treat these diseases.
Soni et al. (2011) [
12] employed methods from data mining to detect heart diseases. The methods from data mining, including Bayesian networks (BNs) and decision trees (DTs) yielded the higher performance than other predictive models such as neural networks and KNN. Moreover, the genetic algorithms applied to DT and BN further enhanced the detection performance for heart disease prediction [
13].
Probabilistic propagation networks using Bayesian networks (BNs) have recently been used to investigate the parametric information from the data using associations and the degree of uncertainty of the variables, and to make the expert opinions, etc. [
14]. Recently, BNs have successfully been used in many applications, as detailed in Kocian et al. [
15], Amaral et al. [
16], Laurila-Pant et al. [
17], and Zhang et al. [
18]. When other variables in the models are known, the Bayes approach helps to determine causal relationships. BNs have also been utilized in other applications, including the prediction of coffee rust disease using Bayesian networks [
19], predicting energy crop yields [
20], sustainable planning and management decisions, [
21] etc. Moreover, BNs are used to visualize and determine the complex interrelationships between interdisciplinary variables resulting from the impacts of climate change in agricultural scenarios [
22]. Most recently, BNs have been used to determine the complex causal interaction between the environment (i.e., climate, weather, and their causes and impact severity) and plant disease in Canada.
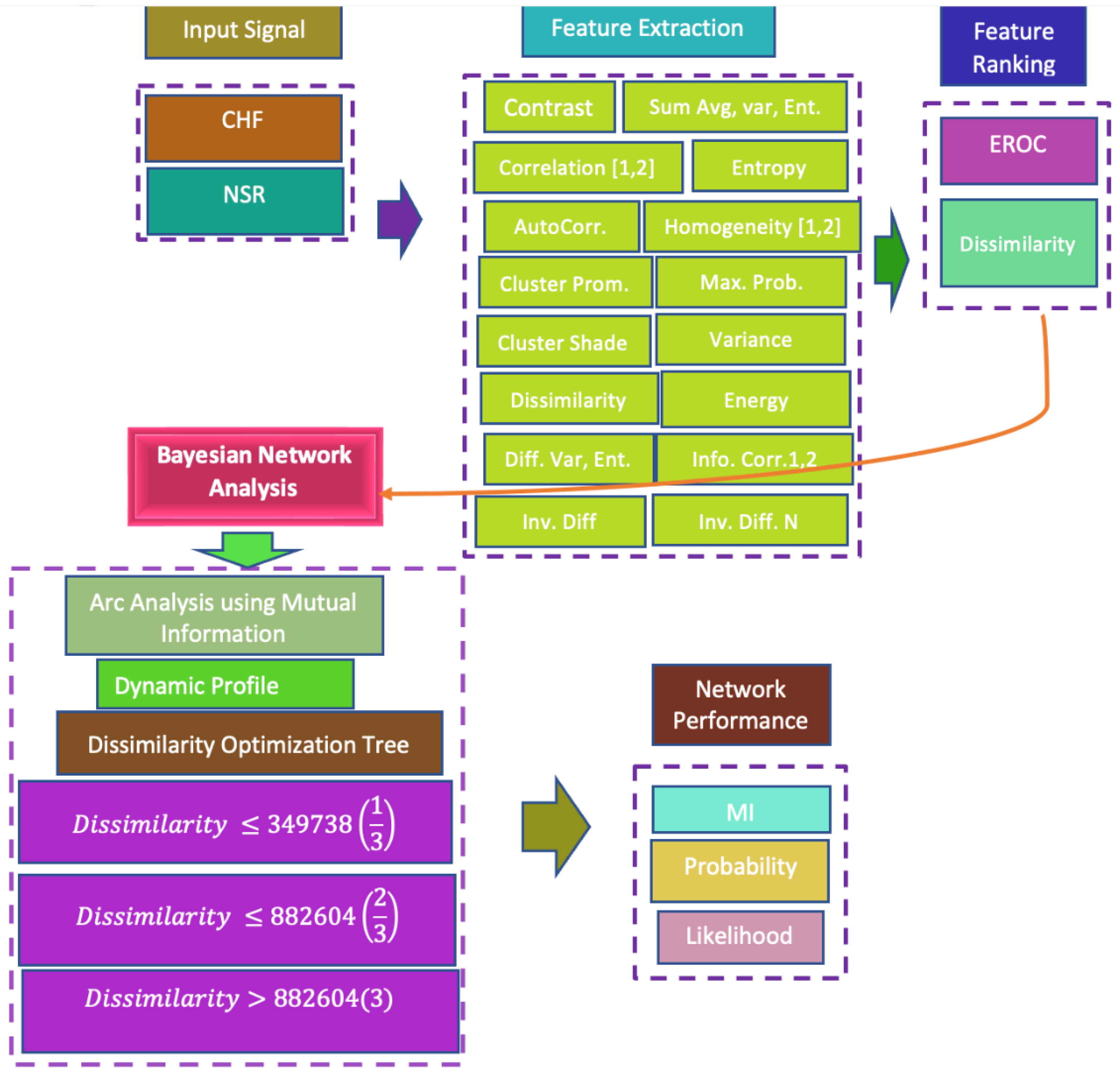
In the present era, the most common and sudden deaths occur due to cardiac and congestive heart failure (CHF). The proper and timely treatment of CHF is highly desired and demanded; researchers are therefore devising automated tools for automatic diagnosis. The dynamics produced from these signals are of a highly complex, nonlinear, and nonstationary nature. Recently, researchers have mostly utilized classification methods to distinguish the CHF from NSR. However, this study was specifically designed to first extract the GLCM features from CHF and NSR subjects, in order to capture these dynamics. We then ranked the features to determine the features’ importance, based on an empirical receiver operating characteristics (EROC) curve and random classifier slope. Finally, we utilized the robust Bayesian inference approach to determine the associations between the extracted GLCM features by computing arc analysis using mutual information, the significance of a cluster’s prominence, and overall analysis of highly ranked dissimilarity features among other extracted features. The proposed approach will further elucidate the underlying dynamics of highly complex heart variability signals, and can be used for better diagnosis and prognosis by health professionals and cardiologists for timely treatment. The associations and strengths of these relationships will be useful for further micro-level analysis and determining the significance of these features among all computed features.
The first section of the paper describes the background of the problem, the existing methods utilized and their limitations, and the proposed innovative methods. The second section explains the dataset used, feature extraction and ranking methods, the Bayesian inference methods used, along with exploratory analysis of unsupervised network analysis. The next section describes and interprets the results, and the last section discusses the main findings and limitations of this study, and future research directions.
3. Results and Discussion
In this study, we first computed the GLCM features from heart failure signals. We determined these features based on ROC values. The highly important dissimilarity value was selected as our target node, and its association with other extracted target variables was computed. The detailed analysis will help to elucidate further dynamics of our complex analysis for a deeper understanding of the extracted features.
We first extracted the GLCM features from pituitary and meningioma brain MRI images. We then ranked the features based on entropy values. The ranked values, based on entropy, are shown in
Figure 2. The importance of the features based on entropy value are energy (3.069), hoomogenity1 (2.7317), homogenity2 (2.6927), maximum probability (2.6818), and so on. We then chose energy as the target variable to compute its association with other computed GLCM features using a Bayesian inference approach.
Figure 3 shows the arc analysis using the
MI method. The arcs between the nodes show the strength of the relationship between the nodes. The bolder arc with greater values shows the highest strength between the nodes; the arc line decreases accordingly as the value and strength of the relationship decreases. There are also other methods for computing arc analysis, including Pearson’s correlation (PC), etc. However, in the present study, we only utilized the
MI for our arc analysis. Moreover, node size can be computed using different methods, including mean, normalized mean value, node force, etc., to reflect the node size. However, in this study, the node size was computed using the normalized mean value.
In this study, we computed the GLCM features and computed the associations among them using mutual information, as reflected in
Figure 3. The arcs show the strengths of relation, and the nodes show the normalized mean values. The relationship of the highest strength was yielded by
followed by
,
, and so on.
Table 1 shows the outgoing, incoming, and total force of extracted GLCM features from congestive heart failure signals. The highest incoming force was yielded at node dissimilarity (1.0146), outgoing force (0.9330), and total force (1.9477), followed by cluster shade with incoming force (0.9330), outgoing force (0.000), and total force (0.9330); this contrasts with incoming force (0.6281), outgoing force (0.5446), and total force (1.1727), and so on.
The target node of dissimilarity had a probability of 58.62% and a joint probability of 100%. The first level of the tree contains child nodes with probabilities such as cluster prominence (100%), contrast (100%), cluster shade (91.78%), energy (73.53%), entropy (68.23%), homogenity1 (68.23%), autocorrelation (67.79%), correlation (63.95%), and correlation1 (62.51%). The red colors show the 1/3rd cluster, green shows the 3/3rd cluster ranges. Navy blue color denote the probabilities at each node. Light purple denotes the joint probability and white denote the scores as reflected in
Figure 4.
At the second level, cluster shades produce child nodes with probabilities such as energy (95.38%), entropy (94.21%), homogenity1 (94.21%), autocorrelation (93.56%), correlation (92.91%), and correlation1 (92.64%). Likewise, energy has child nodes of autocorrelation, correlation, and correlation2. Moreover, homogeneity1 has child nodes of autocorrelation, correlation, and correlation2. At the third level, energy, entropy, and homogenity1 have the child nodes autocorrelation, correlation, and correlation2.
Figure 5 shows the unsupervised clustering when dissimilarity was the selected target node. The arrows indicate the direction of relationship. Further dynamics are computed based on the target node’s associations with other nodes. The right part of the figure indicates the different cluster states with their occurrence out of the total subjects.
Figure 6 shows the significance of the selected top ranked dissimilarity node with other nodes such as cluster prominence, cluster shade, contrast, autocorrelation, energy, entropy, correlation, correlation2, and homogeinity1. The highest association was yielded in the cluster ≤349,738 (58.62%) with the red lines showing cluster prominence, contrast, and cluster shade with probability in the range of 0.0 to 1.0, and with other nodes in the range 0.3 to 0.75. The second highest occurrence of dissimilarity was yielded in the cluster ≤882,604 (34.48%), indicated in the green color, with cluster prominence and cluster shades occurring in the highest probability range of 0.0 to 0.90.
Table 2 shows the overall analysis of the high-ranked feature of dissimilarity as the target node, alongside other nodes. The highest performance was yielded using cluster prominence with mutual information (
MI) as 1.0146, normalized
MI (64.01%), relative
MI (81.33%), and relative significance (1.000), followed by cluster shade, contrast, autocorrelation, energy, entropy, correlation, correlation2, and homogeneity.
Table 3 shows the local analysis for cluster 1 of 3. The highest significance was yielded with cluster prominence with binary
MI (0.7847), relative binary significance, (80.19%), binary relative significance (1.000), and max. Bayes factor (92.64%), followed by cluster shade, contrast, and so on.
Table 4 shows the local analysis for cluster 2 of 3. The highest significance was yielded with cluster prominence with binary
MI (0.6966), relative binary significance (74.95%), binary relative significance (1.000), and max. Bayes factor (97.50%), followed by cluster shade, contrast, and so on.
Table 5 shows the local analysis for cluster 3 of 3. The highest significance was yielded with cluster prominence with binary
MI (0.3621), relative binary significance (100%), binary relative significance (1.000), and max. Bayes factor (100%), followed by cluster shade, contrast, and so on.
Recently, the need has arisen for a comprehensive analysis to compute the associations among the computed features, in order to understand the strength of relationships, associations, the incoming and outgoing forces between parent and child nodes, and the significance of target nodes and target nodes trees; this analysis can be carried out using the robust Bayesian inference approach. This approach will further help us to determine the underlying dynamics and relationships among the extracted nodes, which will help the relevant healthcare professionals to further improve their decision-making capabilities and diagnostic procedures. Hussain et al. [
28] extracted the morphological features from prostate cancer data, in order to compute the associations between these features for deeper analysis.