- Article
Establishment and Application of a SYBR Green I qPCR Detection Method Based on the CP40 Gene of Corynebacterium pseudotuberculosis Biovar Ovi
- Jingpeng Zhang,
- Jinxiu Jiang and
- Yongliang Che
- + 3 authors
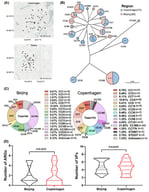
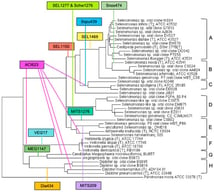
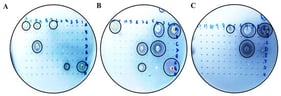
Caseous lymphadenitis (CLA), an infectious disease caused by Corynebacterium pseudotuberculosis (C. pseudotuberculosis), poses a significant economic burden to the global small ruminant industry. This study aimed to investigate genetic variations in the CP40 gene of C. pseudotuberculosis and to develop a rapid detection assay for enhanced pathogen identification. Homology analysis was performed to compare the CP40 gene sequence of the FJ-PN strain with other Corynebacterium species. Specific primers targeting CP40 were designed, and a SYBR Green I-based real-time PCR protocol was optimized. The assay’s specificity, sensitivity, and reproducibility were subsequently validated. The FJ-PN strain exhibited ≥99.65% nucleotide identity and ≥98.94% amino acid identity with C. pseudotuberculosis biovar ovi reference strains, showing 90.18–91.84% nucleotide identity and 88.63–90.77% amino acid identity with C. pseudotuberculosis biovar equi, and ≤82.71% nucleotide identity and ≤78.63% amino acid identity with other Corynebacterium species. The established qPCR assay demonstrated high specificity, the limit of detection was 52 copies/μL, and it demonstrated good reproducibility (intra- and inter-assay CV < 1.0%). Clinical sample testing revealed 18.8% positivity rates in nasal swabs, which was higher than that detected by conventional PCR (16.3%). These results indicate that the CP40 gene is evolutionarily conserved and represents a specific molecular marker for the identification of C. pseudotuberculosis biovar ovis. The developed SYBR Green I real-time PCR assay enables the efficient detection of C. pseudotuberculosis and provides technical support for CLA surveillance and control.
6 February 2026