-
 Insights into Missense SNPs on Amyloidogenic Proteins
Insights into Missense SNPs on Amyloidogenic Proteins -
 Cross-Species Analysis of ABA-Induced Phosphosignaling Landscapes in Rice, Soybean, and Arabidopsis
Cross-Species Analysis of ABA-Induced Phosphosignaling Landscapes in Rice, Soybean, and Arabidopsis -
 Exploring Mitoxantrone Sensitivity in BRCA1-Deficient Ovarian Cancer Using Quantitative Proteomics
Exploring Mitoxantrone Sensitivity in BRCA1-Deficient Ovarian Cancer Using Quantitative Proteomics -
 Stage-Specific Proteomic Signatures in Non-Muscle Invasive Bladder Cancer
Stage-Specific Proteomic Signatures in Non-Muscle Invasive Bladder Cancer
Journal Description
Proteomes
Proteomes
is an international, peer-reviewed, open access journal on all aspects of proteomics published quarterly online by MDPI.
- Open Access— free for readers, with article processing charges (APC) paid by authors or their institutions.
- High Visibility: indexed within Scopus, ESCI (Web of Science), PubMed, PMC, CAPlus / SciFinder, and other databases.
- Journal Rank: JCR - Q2 (Biochemistry and Molecular Biology) / CiteScore - Q1 (Structural Biology)
- Rapid Publication: manuscripts are peer-reviewed and a first decision is provided to authors approximately 28.6 days after submission; acceptance to publication is undertaken in 5.6 days (median values for papers published in this journal in the second half of 2025).
- Recognition of Reviewers: reviewers who provide timely, thorough peer-review reports receive vouchers entitling them to a discount on the APC of their next publication in any MDPI journal, in appreciation of the work done.
Impact Factor:
3.6 (2024);
5-Year Impact Factor:
3.9 (2024)
Latest Articles
Beyond Reanalysis: Critical Issues in Data Reuse for Solid Tumor Proteomics
Proteomes 2026, 14(2), 16; https://doi.org/10.3390/proteomes14020016 - 7 Apr 2026
Abstract
►
Show Figures
Proteomics represents a fundamental layer for understanding the molecular complexity of solid tumors by quantifying protein abundance and capturing proteoforms and post-translational modifications undetected in genomics or transcriptomics analyses. As mass spectrometry-based technologies and public proteomics repositories have expanded, opportunities for large-scale data
[...] Read more.
Proteomics represents a fundamental layer for understanding the molecular complexity of solid tumors by quantifying protein abundance and capturing proteoforms and post-translational modifications undetected in genomics or transcriptomics analyses. As mass spectrometry-based technologies and public proteomics repositories have expanded, opportunities for large-scale data reuse have grown accordingly. Nevertheless, data availability has not been translated into straightforward reuse: differences in experimental design, acquisition strategies, quantification workflows and metadata quality still limit the reproducibility and cross-study comparability. In this review, proteomics data reuse is defined as the systematic reanalysis and integration of publicly available datasets to support precision oncology applications such as biomarker assessment and antibody–drug conjugate target prioritization. We discuss reuse as an end-to-end analytical process, focusing on data analysis workflows, harmonization strategies, and the impact of heterogeneous experimental and analytical choices on interoperability. The increased application of artificial intelligence in proteomics data integration and reuse is also addressed, highlighting its analytical potential while underscoring the risks of overinterpretation when biological context and data structure are not adequately considered. Using colorectal and prostate cancer as representative examples, we illustrate how proteomics data reuse can support biological discovery and translational research, while critically examining the factors that limit robustness and clinical relevance.
Full article
Open AccessArticle
Proteomic Insights into the Immune and Sex-Specific Proteins in the Skin Mucus of Barramundi (Lates calcarifer)
by
Varsha V. Balu, Dean R. Jerry and Andreas L. Lopata
Proteomes 2026, 14(1), 15; https://doi.org/10.3390/proteomes14010015 - 20 Mar 2026
Abstract
►▼
Show Figures
Background: Fish skin mucus contains proteins involved in diverse biological pathways, representing a valuable non-invasive diagnostic of fish health. Methods: Skin mucus from three male and three female barramundi was analysed using liquid chromatography-tandem mass spectrometry (LC-MS/MS) following protein extraction and S-Trap digestion.
[...] Read more.
Background: Fish skin mucus contains proteins involved in diverse biological pathways, representing a valuable non-invasive diagnostic of fish health. Methods: Skin mucus from three male and three female barramundi was analysed using liquid chromatography-tandem mass spectrometry (LC-MS/MS) following protein extraction and S-Trap digestion. Results and Discussion: A total of 1801 protein groups were matched to the L. calcarifer reference proteome and functionally annotated using Gene Ontology (GO) terms via UniProt ID mapping, with representation across Biological Process, Cellular Component, and Molecular Function categories. Functional classification using eggNOG-mapper further associated leading protein group sequences with Clusters of Orthologous Groups (COGs) and Kyoto Encyclopaedia of Genes and Genomes (KEGG) pathways. GO-based screening prioritised 352 putatively immune-relevant protein groups and 24 protein groups associated with sex- and reproduction-related processes, highlighting the functional complexity of the skin mucus proteome. Comparative analysis revealed sex-associated patterns in protein group detection and relative abundance, with differential abundance analysis identifying 244 protein groups exhibiting statistically significant differences between male and female samples. Conclusions: This study provides the first comprehensive discovery-based characterisation of the barramundi skin mucus proteome and establishes a baseline reference dataset for this aquaculture-relevant species. The findings support the utility of skin mucus proteomics for exploring immune and sex-associated molecular patterns and provide a baseline dataset for future validation studies investigating non-invasive health and reproductive monitoring.
Full article

Graphical abstract
Open AccessReview
Emergence of Catalytic Activity in VRK3: Phosphoproteomic Insights into the Regulatory Network of a Former Pseudokinase
by
Ayadathil Sujina, Amal Fahma, Suhail Subair, Rajesh Raju and Poornima Ramesh
Proteomes 2026, 14(1), 14; https://doi.org/10.3390/proteomes14010014 - 18 Mar 2026
Abstract
Vaccinia-Related Kinase 3 (VRK3) is increasingly recognized as a crucial signaling modulator in both normal and pathological processes. This kinase was long thought of as a catalytically inactive pseudokinase, until recently it was established to phosphorylate Barrier to Autointegration Factor (BAF) proteins through
[...] Read more.
Vaccinia-Related Kinase 3 (VRK3) is increasingly recognized as a crucial signaling modulator in both normal and pathological processes. This kinase was long thought of as a catalytically inactive pseudokinase, until recently it was established to phosphorylate Barrier to Autointegration Factor (BAF) proteins through its extracatalytic domain. VRK3 regulates diverse cellular pathways through scaffold interactions and context-dependent phosphorylation. This review is centered around the phosphoregulatory network that modulates VRK3 phosphorylation with implications in its abundance and function. A large-scale phosphoproteomic data integration was performed by combining phosphoproteomics profiling and differential phosphorylation from 115 mass spectrometry studies, identifying 32 high-confidence phosphorylation sites on VRK3. Notably, VRK3 (S59), (S82), and (S83) were predominantly observed highlighting plausible functional significance. These phosphorylation sites share 33 potential upstream kinases, and multiple interactor proteins, which in combination are known to regulate ERK, Hippo, and GPCR pathways. These insights advance the understanding of phosphorylation control by kinases and highlight opportunities to target VRK3-associated networks for therapeutic intervention in diseases such as glioma and liver cancer.
Full article
(This article belongs to the Section Proteome Bioinformatics)
►▼
Show Figures

Graphical abstract
Open AccessArticle
Dimethyl Sulfoxide Enhances HLA Peptide Identification
by
Terry C. C. Lim Kam Sian, Yue Ding, Scott A. Blundell, Ralf B. Schittenhelm and Pouya Faridi
Proteomes 2026, 14(1), 13; https://doi.org/10.3390/proteomes14010013 - 13 Mar 2026
Abstract
Background: Mass spectrometry (MS)-based immunopeptidomics has emerged as the gold standard for profiling HLA-bound peptides, yet detection remains challenging due to their non-tryptic nature, variable lengths, and lack of basic residues, which limit ionisation and fragmentation efficiency. Methods: To address these limitations, we
[...] Read more.
Background: Mass spectrometry (MS)-based immunopeptidomics has emerged as the gold standard for profiling HLA-bound peptides, yet detection remains challenging due to their non-tryptic nature, variable lengths, and lack of basic residues, which limit ionisation and fragmentation efficiency. Methods: To address these limitations, we investigated the impact of incorporating 5% dimethyl sulfoxide (DMSO) into LC-MS/MS mobile-phase buffers on immunopeptidomic workflows. Using B-lymphoblastoid cell lines expressing HLA class I and II alleles and elastase-digested HeLa lysates as a surrogate for non-tryptic peptides, we assessed peptide identification, ionisation efficiency, charge state distribution, and fragmentation quality. Results: DMSO significantly increased peptide identifications across all sample types, with gains of ~1.33 folds for HLA class I, ~1.55 folds for HLA class II, and ~1.24 folds for elastase digests. Improvements were systematic and reproducible, driven by enhanced electrospray ionisation, higher charge states, and superior MS2 spectral quality, evidenced by ~2-fold increase in b- and y-ion intensities. Importantly, DMSO did not introduce major sequence bias, preserving motif integrity and predicted binding characteristics. Conclusions: Overall, these findings establish DMSO as a robust additive for improving sensitivity and reliability in immunopeptidomics, particularly for low-input or clinically derived samples.
Full article
(This article belongs to the Section Proteomics Technology and Methodology Development)
►▼
Show Figures

Graphical abstract
Open AccessReview
Proteomic Analysis in Search of New Biomarkers of Immune Thrombocytopenia (ITP)—A Review of Current Data
by
Anastasia Boura-Theodorou, Konstantina Psatha, Stefania Maniatsi, Areti Kourti, Georgia Kaiafa, Michalis Aivaliotis and Kali Makedou
Proteomes 2026, 14(1), 12; https://doi.org/10.3390/proteomes14010012 - 12 Mar 2026
Abstract
►▼
Show Figures
Immune thrombocytopenia (ITP) is a hematological disorder commonly found in individuals of any gender, race, or age. Patients with ITP will present with thrombocytopenia either in a primary form or because of an infection or a dysfunction in the immune system. The severity
[...] Read more.
Immune thrombocytopenia (ITP) is a hematological disorder commonly found in individuals of any gender, race, or age. Patients with ITP will present with thrombocytopenia either in a primary form or because of an infection or a dysfunction in the immune system. The severity of ITP is linked to diminished production of platelets due to the blockage of production in the bone marrow niche and increased destruction of platelets, which confirms the diagnosis of the disorder. The investigation of the pathogenesis of ITP is of critical importance as it can give an important indication of the state of the patient, guiding us through risk assessment and treatment. Proteomics can provide tools to explore the protein profile of ITP. In this review, we aimed to uncover different biomarkers, both diagnostic and prognostic, that have been investigated with proteomic methodologies and that might help in understanding the pathogenesis of ITP and providing personalized treatment to patients. Several differentially abundant proteins were identified, including haptoglobin isoforms, heat shock proteins (HSPA6, HSPA8), integrin β3 (ITGB3), 14-3-3 protein eta (YWHAH), vitamin D-binding protein, fibrinogen chains, MYH9, and FETUB, which are involved in key signaling pathways, such as PI3K/akt, TNF-a, and mTOR, and they demonstrate potential as diagnostic and prognostic biomarkers. Collectively, current data support the value of proteomics for uncovering the molecular landscape of ITP and guiding the development of precision diagnostics and personalized therapeutic strategies.
Full article

Figure 1
Open AccessArticle
Cellular Responses of Maize Roots to Long-Term Cadmium Exposure: Adjustments of Class III Peroxidases, Plasma Membrane and Tonoplast Sub-Proteomes
by
Sabine Lüthje, Ayse Gül Yilmaz, Kalaivani Ramanathan, Waldemar Gräfenstein, Jenny M. Tabbert, Stefanie Wienkoop, Katrin Heino, François Clement Perrineau and Sönke Harder
Proteomes 2026, 14(1), 11; https://doi.org/10.3390/proteomes14010011 - 25 Feb 2026
Abstract
Background: Crop plants have to deal with long-term cadmium exposure to farmlands contaminated by intensive use of fertilizers and pesticides. For uptake and sequestration, Cd2+ has to pass the plasma membrane and tonoplast. Class III peroxidases, plasma membrane, and tonoplast sub-proteomes were
[...] Read more.
Background: Crop plants have to deal with long-term cadmium exposure to farmlands contaminated by intensive use of fertilizers and pesticides. For uptake and sequestration, Cd2+ has to pass the plasma membrane and tonoplast. Class III peroxidases, plasma membrane, and tonoplast sub-proteomes were studied. Methods: Control and Cd2+-treated maize (Zea mays L.) were grown in hydroponics for 18 days. Soluble peroxidases were partially purified by chromatofocusing and characterized by substrate specificity. Membrane-bound peroxidases were analyzed spectrophotometrically and by non-reducing SDS-PAGE. Soluble and plasma membrane-bound peroxidases were identified by mass spectrometry. Shotgun proteomics was used to identify membrane proteins of differential abundance. Results: Guaiacol peroxidase activities increased in soluble fractions of Cd2+ samples. A Cd2+-specific soluble peroxidase (ZmPrx101) was identified, and ZmPrx85 abundance increased significantly in the plasma membrane. Substrate specificity of peroxidases revealed a preference for ferulic acid and esculetin, which was confirmed by docking analyses. Primary active transporters increased auxin efflux (brachytic2, ABCB9, and ABCB21), Cd2+ exclusion (ABCG34), and sequestration into the vacuole (HMA2, ABCB27). Evaluation of sub-proteome fractions demonstrated significant changes for proteins involved in disease resistance responses and cell wall modification. Conclusions: Molecular adjustments of maize root proteome to long-term Cd2+ exposure revealed relevance of low-abundant proteins for Cd2+ tolerance and putative stress markers.
Full article
(This article belongs to the Special Issue Plant Genomics and Proteomics)
►▼
Show Figures

Graphical abstract
Open AccessArticle
Comparison of the Trapping Efficiency for Tryptic Peptides on Particle-Packed and Micro-Pillar Trap Columns for Proteomics Analyses
by
Jadranka Miletić Vukajlović, Bojana Ilić, Bella Bruszel, Tanja Panić-Janković and Goran Mitulović
Proteomes 2026, 14(1), 10; https://doi.org/10.3390/proteomes14010010 - 18 Feb 2026
Abstract
Background: Low-volume trapping columns are essential for sample enrichment, desalting, and injection profile focusing on nano-LC–MS-based proteomics. They enable higher sample loading, improve chromatographic performance, and protect the analytical column by removing salts and contaminants. Recently, monolithic trap columns with micropillar architecture have
[...] Read more.
Background: Low-volume trapping columns are essential for sample enrichment, desalting, and injection profile focusing on nano-LC–MS-based proteomics. They enable higher sample loading, improve chromatographic performance, and protect the analytical column by removing salts and contaminants. Recently, monolithic trap columns with micropillar architecture have emerged as alternatives to conventionally packed traps. This study compares the performance of a packed and a micropillar monolithic trap column for the analysis of tryptic peptides. Methods: A tryptic digest of HeLa cell lysate was analyzed under identical LC–MS conditions using both trap types. Peptides were detected at 214 nm and analyzed by nano-ESI on a Q Exactive Plus Orbitrap. Data were searched against the human UniProt database (February 2023) using FragPipe v20.0, and statistical evaluation of MaxLFQ intensities was performed in Perseus using Welch’s t-test and clustering analysis. Results: Over 2500 proteins were identified with both setups. The packed trap column yielded more total peptides, particularly those with post-translational modifications and higher hydrophilicity, whereas the monolithic column favored peptides of intermediate hydrophobicity. Chromatographic profiles confirmed a slight reduction in the trapping efficiency of hydrophilic peptides by the monolithic trap. Conclusions: Trap column design significantly influences peptide recovery and proteome coverage.
Full article
(This article belongs to the Section Proteomics Technology and Methodology Development)
►▼
Show Figures

Figure 1
Open AccessArticle
Scout-Triggered Multiple Reaction Monitoring Enables Robust Quantification of Host Cell Proteins Across Bioprocess Matrices
by
Julie Flecheux, Chloé Bardet, Laura Herment, Tanguy Fortin and Jérôme Lemoine
Proteomes 2026, 14(1), 9; https://doi.org/10.3390/proteomes14010009 - 17 Feb 2026
Abstract
Background: Host cell proteins (HCPs) are process-related impurities that must be monitored in biopharmaceutical products due to their potential impact on product quality and patient safety. Targeted LC–MS/MS approaches such as multiple reaction monitoring (MRM) enable protein-specific HCP quantification but are difficult to
[...] Read more.
Background: Host cell proteins (HCPs) are process-related impurities that must be monitored in biopharmaceutical products due to their potential impact on product quality and patient safety. Targeted LC–MS/MS approaches such as multiple reaction monitoring (MRM) enable protein-specific HCP quantification but are difficult to apply in highly multiplexed assays because of retention time (RT) variability across complex bioprocess matrices. Methods: Here, we show that conventional RT-scheduled MRM workflows lack transferability when applied to heterogeneous drug substances and process intermediates. Using a targeted assay comprising 240 peptides corresponding to 97 CHO-derived HCPs, RT shifts of several minutes resulted in truncated chromatographic peaks and peptide signal loss, even when wide scheduling windows were used. To overcome this limitation, a scout-triggered MRM (st-MRM) acquisition strategy based on event-driven monitoring was implemented. Results: This approach enabled robust peptide detection across diverse matrices within a single injection, without method re-optimization. Absolute quantification using stable isotope-labeled peptides spanned six orders of magnitude, with HCPs quantified down to 2.9 ppm in purified drug substances. Conclusion: Overall, st-MRM improves the robustness and transferability of highly multiplexed targeted proteomics workflows for HCP analysis.
Full article
(This article belongs to the Section Proteomics Technology and Methodology Development)
►▼
Show Figures

Graphical abstract
Open AccessArticle
Proteome-Wide Analysis of Functional Phosphosites in the FGFR Family of Proteins: Insights from Large-Scale Phosphoproteomic Analysis
by
Akhina Palollathil, Althaf Mahin, Athira Perunelly Gopalakrishnan, Tejaswini R Poojari, Alimath Sambreena, Prathik Basthikoppa Shivamurthy and Rajesh Raju
Proteomes 2026, 14(1), 8; https://doi.org/10.3390/proteomes14010008 - 13 Feb 2026
Abstract
Background: Fibroblast growth factor receptors (FGFRs) play a crucial role in tissue homeostasis and organ development by regulating cellular processes, including proliferation, differentiation, and survival. Dysregulation of FGFRs contributes to developmental disorders and carcinogenesis. As membrane-bound receptors, they represent promising targets for therapeutic
[...] Read more.
Background: Fibroblast growth factor receptors (FGFRs) play a crucial role in tissue homeostasis and organ development by regulating cellular processes, including proliferation, differentiation, and survival. Dysregulation of FGFRs contributes to developmental disorders and carcinogenesis. As membrane-bound receptors, they represent promising targets for therapeutic intervention and drug development. Methods: This study employed a systematic in silico analysis of publicly available phosphoproteomics datasets to provide a comprehensive overview of the phosphorylation regulatory network of the FGFR family. Results: We identified predominant phosphosites in FGFR1-4 that exhibited differential abundance across diverse experimental conditions, specifically, Y653 in FGFR1; S453, Y586, Y656, and Y657 in FGFR2; S444 and S445 in FGFR3; and S573 in FGFR4. Our analysis identified 32 and 89 significantly co-modulated phosphosites on other proteins with FGFR3 and FGFR4, respectively. Beyond the upstream kinases from the FGFR family, we also identified MAPK1 as a potential upstream kinase of FGFR4. Furthermore, disease enrichment analysis revealed that proteins co-modulated with FGFR3 were primarily involved in skeletal developmental disorders, such as brachydactyly, short toe, and syndactyly of fingers, whereas those associated with FGFR4 were linked to various cancers. Conclusions: Our findings highlight key disease-associated phosphosites within the FGFRs and offer a foundation for advancing phosphosite-focused therapeutic research.
Full article
(This article belongs to the Section Proteome Bioinformatics)
►▼
Show Figures

Graphical abstract
Open AccessArticle
PTMs_Closed_Search: Multiple Post-Translational Modification Closed Search Using Reduced Search Space and Transferred FDR
by
Yury Yu. Strogov, Sergey A. Spirin, Mark V. Ivanov, Maria A. Kulebyakina, Anastasia Yu. Efimenko and Oleg I. Klychnikov
Proteomes 2026, 14(1), 7; https://doi.org/10.3390/proteomes14010007 - 2 Feb 2026
Abstract
Background: Currently, post-translational modification (PTM) search in MS/MS data is performed using either open modification search (OMS) or closed search (CS) algorithms. The OMS method allows for the determination of many PTMs and unknown mass-shifts in one run. In contrast, closed search
[...] Read more.
Background: Currently, post-translational modification (PTM) search in MS/MS data is performed using either open modification search (OMS) or closed search (CS) algorithms. The OMS method allows for the determination of many PTMs and unknown mass-shifts in one run. In contrast, closed search algorithms are more sensitive but limited in the number of PTMs that can be specified in one search. Methods: In this manuscript, we propose an optimized Python algorithm based on the IdentiPy search engine that performs an automated sequential search for each PTM based on previous annotations from public databases and customized protein lists. We also determined the sufficient size of the search space to increase the significance of false discovery rate (FDR) estimation. We modified the FDR calculation algorithm by implementing a spline approximation of the ratio of the modified decoys, and by calculating error propagation to filter out unstable data and determine the cutoff value. Results: The results of this pipeline for a test dataset were comparable to previously published data in terms of the number of unmodified peptides and proteins. Additionally, we identified 13 different types of peptide PTMs and achieved an increase in relative protein coverage. Our filtration method based on spline transferred FDR showed a superior number of identified peptides compared to separate FDR. Conclusions: Our developed pipeline can be used as a standalone application or as a module of multiple PTM search in data analysis platforms.
Full article
(This article belongs to the Section Proteome Bioinformatics)
►▼
Show Figures

Figure 1
Open AccessArticle
Evaluating the Impact of Two Different Diets on the Protein Profile of the Brain, Liver, and Intestine of the Barramundi
by
Mohadeseh Montazeri Shatouri, Igor Pirozzi, Pinar Demir Soker, Zeshan Ali, Ardeshir Amirkhani and Paul A. Haynes
Proteomes 2026, 14(1), 6; https://doi.org/10.3390/proteomes14010006 - 29 Jan 2026
Abstract
Background: Commercial feed formulations are increasingly being evaluated for their nutritional impacts on aquaculture species, yet the molecular consequences of commonly used commercial diets remain underexplored. Methods: This study investigated the effects of two commercial diets, diet A (higher land animal protein) and
[...] Read more.
Background: Commercial feed formulations are increasingly being evaluated for their nutritional impacts on aquaculture species, yet the molecular consequences of commonly used commercial diets remain underexplored. Methods: This study investigated the effects of two commercial diets, diet A (higher land animal protein) and diet B (higher fish meal content), on the protein profile in the brain, liver, and intestine of barramundi (Lates calcarifer). A 12-week feeding trial was conducted with controlled water quality, and proteomic profiling was performed using data-independent acquisition. Results: Differential analysis revealed consistent changes between diets across all tissues, with a higher percentage of differentially abundant proteins observed in between-diet comparisons (12.99% in brain, 12.73% in liver, and 16.59% in intestine) than within-diet controls (<8%), confirming a measurable dietary effect size. In total, 3901 proteins in the brain, 3660 in the liver, and 5025 in the intestine were quantified. Functional enrichment highlighted upregulation of ferroptosis pathways, downregulation of apelin signaling in the brain, and increased digestive proteases in the liver. ICP-MS confirmed elevated iron concentrations in the brain, liver, and intestine of fish fed on diet B. Conclusions: These findings demonstrate that molecular pathways linked to iron metabolism, digestion, and growth regulation are very sensitive to dietary composition, highlighting how proteomics can help identify subtle impacts of compositional differences in aquaculture feeding. Although physiological parameters did not differ significantly, the proteomic alterations observed across tissues likely indicate organ-specific metabolic adaptations to the differing nutrient availability between diets.
Full article
(This article belongs to the Section Animal Proteomics)
►▼
Show Figures

Figure 1
Open AccessArticle
Integrated Phosphoproteomics Identifies TGFβ-Dependent Phosphorylation Events Linking Kinase Signaling to Autophagy in Palatogenesis
by
Xia Peng, Jing Chen, Xiaoyu Zheng, Xige Zhao, Yijia Wang, Xiaotong Wang and Juan Du
Proteomes 2026, 14(1), 5; https://doi.org/10.3390/proteomes14010005 - 23 Jan 2026
Abstract
Background: Cleft palate (CP) is a prevalent craniofacial malformation, with the TGFβ pathway playing a critical role. Recent evidence links autophagy to regulating mouse embryonic palatal mesenchyme (MEPM) cells, but its interaction with TGFβ-activated phosphorylation cascades remains largely unknown. Here, we investigated the
[...] Read more.
Background: Cleft palate (CP) is a prevalent craniofacial malformation, with the TGFβ pathway playing a critical role. Recent evidence links autophagy to regulating mouse embryonic palatal mesenchyme (MEPM) cells, but its interaction with TGFβ-activated phosphorylation cascades remains largely unknown. Here, we investigated the interplay between these pathways during palatogenesis. Methods: H&E and IHC analyses revealed increased expression of Beclin 1 and LC3 during the critical period of palatal shelf elevation and fusion (E13.5–E15.5). Bulk RNA sequencing (Bulk RNA-seq) further revealed enrichment of autophagy-related pathways and their interaction with TGFβ signaling. TMT-based phosphoproteomics was performed on TGFβ2-treated MEPM cells. Results: We identified 23,471 peptides and 3952 proteins, including 6339 phosphopeptides corresponding to 2195 phosphoproteins. Differential analysis found 477 phosphopeptides with increased abundance and 53 with decreased abundance, revealing the enrichment of seven serine (p-Ser) motifs (RxxS, SDxD, SDxE, SP, SxDE, SxEE, SxxxxD) and one threonine (p-Thr) motif (TP). Notably, kinase-substrate enrichment analysis identified CSNK2A as a previously unrecognized phosphorylation regulator, together with MAPKs and CDKs. Functional enrichment showed significant involvement of mTOR, MAPK, and autophagy/mitophagy pathways. Conclusions: Our findings revealed that TGFβ2 reshapes the MEPM phosphoproteome through Smad-independent pathway, expanding the palate-specific phospho-signaling atlas beyond the canonical Smad cascade.
Full article
(This article belongs to the Section Animal Proteomics)
►▼
Show Figures

Graphical abstract
Open AccessArticle
Cross-Species Analysis of ABA-Induced Phosphosignaling Landscapes in Rice, Soybean, and Arabidopsis
by
Hinano Takase, Sotaro Katagiri, Takuma Ide, Aina Nagano, Haruki Sakurai, Hana Kokubo, Taiki Yanagisawa, Masanori Okamoto and Taishi Umezawa
Proteomes 2026, 14(1), 4; https://doi.org/10.3390/proteomes14010004 - 20 Jan 2026
Abstract
Background: Abscisic acid (ABA) is a key phytohormone that regulates plant growth and stress responses through protein phosphorylation. While ABA-induced phosphosignaling has been extensively studied in Arabidopsis thaliana, its conservation and divergence across plant species remain unclear. Methods: Here, we performed phosphoproteomic
[...] Read more.
Background: Abscisic acid (ABA) is a key phytohormone that regulates plant growth and stress responses through protein phosphorylation. While ABA-induced phosphosignaling has been extensively studied in Arabidopsis thaliana, its conservation and divergence across plant species remain unclear. Methods: Here, we performed phosphoproteomic analysis using LC-MS/MS in Arabidopsis, rice (Oryza sativa), and soybean (Glycine max) to compare ABA-responsive phosphorylation profiles among monocots, dicots, and legumes. Results: ABA treatments on Arabidopsis, rice, and soybean seedlings yielded approximately 24,604, 18,865, and 24,930 phosphopeptides, respectively. Comparative analyses revealed both conserved and species-specific ABA-responsive phosphoproteins. Conclusions: This work provides insights into the evolutionary diversification of ABA signaling and its potential applications in improving crop stress tolerance.
Full article
(This article belongs to the Special Issue Plant Genomics and Proteomics)
►▼
Show Figures

Figure 1
Open AccessArticle
The Proteome of Dictyostelium discoideum Across Its Entire Life Cycle Reveals Sharp Transitions Between Developmental Stages
by
Sarena Banu, P. V. Anusha, Pedro Beltran-Alvarez, Mohammed M. Idris, Katharina C. Wollenberg Valero and Francisco Rivero
Proteomes 2026, 14(1), 3; https://doi.org/10.3390/proteomes14010003 - 8 Jan 2026
Abstract
►▼
Show Figures
Background: Dictyostelium discoideum is widely used in developmental and evolutionary biology due to its ability to transition from a single cell to a multicellular organism in response to starvation. While transcriptome information across its life cycle is widely available, only early-stage data exist
[...] Read more.
Background: Dictyostelium discoideum is widely used in developmental and evolutionary biology due to its ability to transition from a single cell to a multicellular organism in response to starvation. While transcriptome information across its life cycle is widely available, only early-stage data exist at the proteome level. This study characterizes and compares the proteomes of D. discoideum cells at the vegetative, aggregation, mound, culmination and fruiting body stages. Methods: Samples were collected from cells developing synchronously on nitrocellulose filters. Proteins were extracted and digested with trypsin, and peptides were analyzed by liquid chromatography–tandem mass spectrometry. Data were processed using Proteome Discoverer™ for protein identification and label-free quantification. Results: A total of 4502 proteins were identified, of which 1848 (41%) were present across all stages. Pairwise comparisons between adjacent stages revealed clear transitions, the largest ones occurring between the culmination and fruiting body and between the fruiting body and vegetative stage, involving 29% and 52% of proteins, respectively. Hierarchical clustering assigned proteins to one of nine clusters, each displaying a distinct pattern of abundances across the life cycle. Conclusions: This study presents the first complete developmental proteomic time series for D. discoideum, revealing changes that contribute to multicellularity, cellular differentiation and morphogenesis.
Full article

Figure 1
Open AccessArticle
Proteomics and Bioinformatics Profiles of Human Mesothelial Cell Line MeT-5A
by
Rachel L. Watkin, Avedis A. Kazanjian, Jennifer R. Damicis and Elizabeth Yohannes
Proteomes 2026, 14(1), 2; https://doi.org/10.3390/proteomes14010002 - 4 Jan 2026
Abstract
►▼
Show Figures
Background: Despite existing proteomics studies of other cell types, a comprehensive proteome of mesothelial cells has not been characterized. This study establishes a crucial baseline proteome for mesothelial cells to better understand their fundamental bioprocesses in healthy and injured states. Methods:
[...] Read more.
Background: Despite existing proteomics studies of other cell types, a comprehensive proteome of mesothelial cells has not been characterized. This study establishes a crucial baseline proteome for mesothelial cells to better understand their fundamental bioprocesses in healthy and injured states. Methods: Using mass spectrometry-based shotgun proteomics, we characterized the cellular fraction (CF) and conditioned medium (CM) proteomes of mesothelial cell line MeT-5A. The datasets were analyzed for Gene Ontology (GO) terms and canonical pathway enrichments to identify biological themes. Results: Our analysis identified 5087 protein groups, including 1532 shared proteins, 3122 unique to the CF and 433 exclusive to the CM. GO annotation revealed distinct functional enrichment profiles, reflecting the differing roles of intracellular and secreted proteins. While intracellular proteins were linked to core cellular functions, the extracellular proteome was enriched for signaling and cell-to-cell interaction pathways. The proteins shared by both compartments provided an integrated view of the molecular coordination between the cellular and extracellular environments. Conclusions: This study provides the first comprehensive baseline proteome for mesothelial cells and their secreted medium, offering a vital resource for future investigations into the mesothelium, particularly in the context of disease or injury.
Full article

Figure 1
Open AccessArticle
Beyond Repression: ArsR Functions as a Global Activator of Metabolic and Redox Responses in Escherichia coli
by
Brett Sather, James Larson, Kian Hutt Vater, Jade Westrum, Timothy R. McDermott and Brian Bothner
Proteomes 2026, 14(1), 1; https://doi.org/10.3390/proteomes14010001 - 4 Jan 2026
Abstract
►▼
Show Figures
Background: The arsenic-responsive repressor, ArsR, has long been understood as a canonical regulator of the arsRBC operon, which confers resistance to arsenic stress. However, recent studies suggest a broader regulatory scope for ArsR. Here, we investigated the proteomic landscape of Escherichia coli strains
[...] Read more.
Background: The arsenic-responsive repressor, ArsR, has long been understood as a canonical regulator of the arsRBC operon, which confers resistance to arsenic stress. However, recent studies suggest a broader regulatory scope for ArsR. Here, we investigated the proteomic landscape of Escherichia coli strains with and without ArsR to elucidate ArsR as an activator in both non-stressing and arsenic-stressing conditions. Methods: Using mass-spectrometry-based shotgun proteomics and statistical analyses, we characterized the differential abundance of proteins across AW3110 (ΔarsRBC), AW3110 complemented with arsR, and wild-type K-12 strains under control and arsenite-stressed conditions. Results: Our study shows that ArsR influences proteomic networks beyond the ars operon, integrating metabolic and redox responses crucial for cellular adaptation and survival. This suggests that ArsR has a significant role in gut microbiome metabolomic profiles in response to arsenite. Proteins involved in alanine, lactaldehyde, arginine, thioredoxin, and proline pathways were significantly elevated in strains where ArsR was detected, both with and without arsenite. We identified proteins exhibiting an “ArsR-dependent” activation pattern, highlighting ArsR’s potential role in redox balance and energy metabolism. Conclusions: These findings challenge the classical view of ArsR as a repressor and position it as a pleiotropic regulator, including broad activation.
Full article

Figure 1
Open AccessCommunication
A Proteomic View of Butterfly Metamorphosis
by
Andrew Hesketh, Juned Kadiwala, Vaishnavi Ravikumar, Ana Rita Garizo, Patrícia Beldade, Marjorie Fournier and Rameen Shakur
Proteomes 2025, 13(4), 68; https://doi.org/10.3390/proteomes13040068 - 18 Dec 2025
Cited by 1
Abstract
►▼
Show Figures
Background: Insect metamorphosis is one of the most fascinating developmental processes in the natural world. Complete metamorphosis requires the breakdown and reorganisation of larval tissues and the coordinated construction and development of adult structures. The molecular events that achieve this transformation are, however,
[...] Read more.
Background: Insect metamorphosis is one of the most fascinating developmental processes in the natural world. Complete metamorphosis requires the breakdown and reorganisation of larval tissues and the coordinated construction and development of adult structures. The molecular events that achieve this transformation are, however, incompletely understood, and there is a particular shortage of data describing changes in protein abundance that occur during the process. Methods: Here, using a label-free quantitative bottom-up approach, we perform a novel whole-organism proteomic analysis of consecutive developmental stages of male Bicyclus anynana butterflies as they develop from caterpillars into adults via pupation. Results: Our analysis generated a dynamic reference dataset representing 2749 detected proteins. Statistical analysis identified 90 proteins changing significantly in abundance during metamorphosis, and functional interpretation highlights cuticle formation, apoptosis and autophagy during the pupal stages, and the up-regulation of respiration and energy metabolism upon completion of the fully formed adult. A preliminary search for potential peptide phosphorylation modifications identified 15 candidates, including three proteins with roles in muscle function. Conclusions: The study provides a basis for future protein-level analysis of butterfly metamorphosis and suggests the importance of dissecting the post-translational regulation associated with this fascinating developmental transformation.
Full article

Figure 1
Open AccessArticle
Comprehensive Insights into Obesity and Type 2 Diabetes from Protein Network, Canonical Pathway, Phosphorylation and Antimicrobial Peptide Signatures of Human Serum
by
Petra Magdolna Bertalan, Erdenetsetseg Nokhoijav, Ádám Pap, George C. Neagu, Miklós Káplár, Zsuzsanna Darula, Gergő Kalló, Laszlo Prokai and Éva Csősz
Proteomes 2025, 13(4), 67; https://doi.org/10.3390/proteomes13040067 - 17 Dec 2025
Abstract
Background: Obesity is a major risk factor for type 2 diabetes (T2D); however, the molecular links between these conditions are not fully understood. Methods: We performed an integrative serum proteomics study on samples from 134 individuals (healthy controls, patients with obesity and/or T2D)
[...] Read more.
Background: Obesity is a major risk factor for type 2 diabetes (T2D); however, the molecular links between these conditions are not fully understood. Methods: We performed an integrative serum proteomics study on samples from 134 individuals (healthy controls, patients with obesity and/or T2D) using both data-independent (DIA) and data-dependent (DDA) liquid chromatography-mass spectrometry approaches, complemented by phosphopeptide enrichment, kinase activity prediction, network and pathway analyses to get more information on the different proteoforms involved in the pathophysiology of the diseases. Results: We identified 235 serum proteins, including 13 differentially abundant proteins (DAPs) between groups. Both obesity and T2D were characterized by activation of complement and coagulation cascades, as well as alterations in lipid metabolism. Ingenuity Pathway Analysis® (IPA) revealed shared canonical pathways, while phosphorylation-based regulation differentiated the two conditions. Elevated hemopexin (HPX), vitronectin (VTN), kininogen-1 (KNG1) and pigment epithelium-derived factor (SERPINF1), along with decreased adiponectin (ADIPOQ) and apolipoprotein D (APOD), indicated a pro-inflammatory, pro-coagulant serum profile. Network analyses of antimicrobial and immunomodulatory peptides (AMPs) revealed strong overlaps between immune regulation and lipid metabolism. Phosphoproteomics and kinase prediction highlighted altered CK2 and AGC kinase activities in obesity, suggesting signaling-level modulation. Conclusions: Our comprehensive proteomic and phosphoproteomic profiling reveals overlapping yet distinct molecular signatures in obesity and T2D, emphasizing inflammation, complement activation and phosphorylation-driven signaling as central mechanisms that potentially contribute to disease progression and therapeutic targeting.
Full article
(This article belongs to the Special Issue Proteomics in Diabetes: From Mechanisms to Biomarkers)
►▼
Show Figures

Figure 1
Open AccessArticle
Proteome Profiling of Rabies-Infected and Uninfected Dog Brain Tissues, Cerebrospinal Fluids and Serum Samples
by
Ukamaka U. Eze, Rethabile Mokoena, Kenneth I. Ogbu, Sinegugu Dubazana, Ernest C. Ngoepe, Mparamoto Munangatire, Romanus C. Ezeokonkwo, Boniface M. Anene, Sindisiwe G. Buthelezi and Claude T. Sabeta
Proteomes 2025, 13(4), 66; https://doi.org/10.3390/proteomes13040066 - 15 Dec 2025
Abstract
Background: Rabies is among the oldest known zoonotic viral diseases and is caused by members of the Lyssavirus genus. The prototype species, Lyssavirus rabies, effectively evades the host immune response, allowing the infection to progress unnoticed until the onset of clinical signs.
[...] Read more.
Background: Rabies is among the oldest known zoonotic viral diseases and is caused by members of the Lyssavirus genus. The prototype species, Lyssavirus rabies, effectively evades the host immune response, allowing the infection to progress unnoticed until the onset of clinical signs. At this stage, the disease is irreversible and invariably fatal, with definitive diagnosis possible only post-mortem. Given the advances in modern proteomics, this study aimed to identify potential protein biomarkers for antemortem diagnosis of rabies in dogs, which are the principal reservoir hosts of the rabies virus. Methods: Two hundred and thirty-one samples (brain tissues (BT), cerebrospinal fluids (CSF), and serum (SR) samples) were collected from apparently healthy dogs brought for slaughter for human consumption in South-East and North-Central Nigeria. All the BT were subjected to a direct fluorescent antibody test to confirm the presence of lyssavirus antigen, and 8.7% (n = 20) were positive. Protein extraction, quantification, reduction, and alkylation were followed by on-bead (HILIC) cleanup and tryptic digestion. The resulting peptides from each sample were injected into the Evosep One LC system, coupled to the timsTOF HT MS, using the standard dia-PASEF short gradient data acquisition method. Data was processed using SpectronautTM (v19). An unpaired t-test was performed to compare identified protein groups (proteins and their isoforms) between the rabies-infected and uninfected BT, CSF, and SR samples. Results: The study yielded 54 significantly differentially abundant proteins for the BT group, 299 for the CSF group, and 280 for the SR group. Forty-five overlapping differentially abundant proteins were identified between CSF and SR, one between BT and CSF, and two between BT and SR; none were found that overlapped all three groups. Within the BT group, 33 proteins showed increased abundance, while 21 showed decreased abundance in the rabies-positive samples. In the CSF group, 159 proteins had increased abundance and 140 had decreased abundance in the rabies-positive samples. For the SR group, 215 proteins showed increased abundance, and 65 showed decreased abundance in the rabies-positive samples. Functional enrichment analysis revealed that pathways associated with CSF, spinocerebellar ataxia, and neurodegeneration were among the significant findings. Conclusion: This study identified canonical proteins in CSF and SR that serve as candidate biomarkers for rabies infection, offering insights into neuronal dysfunction and potential tools for early diagnosis.
Full article
(This article belongs to the Section Animal Proteomics)
►▼
Show Figures

Graphical abstract
Open AccessArticle
Proteomic Profiling of Non-Muscle Invasive Bladder Cancer Reveals Stage-Specific Molecular Signatures and Prognostic Biomarkers
by
Lorenza Vantaggiato, Marco Frisenda, Enxhi Shaba, Chiara Splendore, Beatrice Sciarra, Luca Bini, Alessandro Sciarra and Claudia Landi
Proteomes 2025, 13(4), 65; https://doi.org/10.3390/proteomes13040065 - 10 Dec 2025
Abstract
Background: Non-muscle invasive bladder cancer (NMIBC) comprises high-grade (HG) and low-grade (LG) variants, classified by aggressiveness, recurrence risk, and stage—either non-invasive (pTa) or invading the lamina propria (pT1). Cystoscopy remains the diagnostic gold standard, with no less-invasive alternatives, while molecular mechanisms driving
[...] Read more.
Background: Non-muscle invasive bladder cancer (NMIBC) comprises high-grade (HG) and low-grade (LG) variants, classified by aggressiveness, recurrence risk, and stage—either non-invasive (pTa) or invading the lamina propria (pT1). Cystoscopy remains the diagnostic gold standard, with no less-invasive alternatives, while molecular mechanisms driving tumorigenesis and treatment response are poorly understood. Methods: To address this gap, we conducted a preliminary top-down proteomic study on fresh biopsies from pTa-LG and pT1-HG NMIBC at initial diagnosis to identify molecular differences and potential prognostic biomarkers. Results: Distinct protein profiles were observed between stages. Highly abundant proteins in pT1-HG were associated with nitric oxide biosynthesis, signal transduction, inhibition of apoptosis, protein folding, and immune response. Proteins of low abundance were related to cellular localization, cytoskeleton organization, cell adhesion, phagocytosis, and tissue development. Notably, multiple proteoforms of PDC6I/ALIX, a protein implicated in the regulation of apoptosis, proliferation, and PD-L1 surface presentation, were significantly downregulated in pT1-HG tumors. Furthermore, the abundance of proteins such as GANAB, GALE, THIC, SEPT8, and MYDGF/C19orf10 correlated with tumor size, suggesting their potential as prognostic biomarkers. Conclusions: These proteins, taken together, indicate that they may serve as valuable prognostic markers, offering a path toward more personalized management of NMIBC beyond the traditional one-size-fits-all approach.
Full article
(This article belongs to the Section Identification of Potential Biomarkers and Potential Therapeutic Targets)
►▼
Show Figures
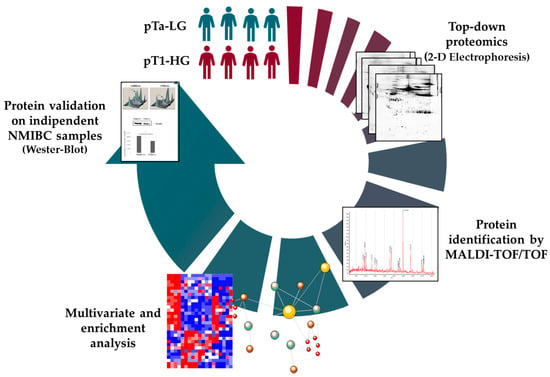
Figure 1
Highly Accessed Articles
Latest Books
E-Mail Alert
News
Topics
Topic in
Biomolecules, IJMS, Cancers, Proteomes
Extracellular Vesicles: Isolation, Characterization, Function, Application and Utility
Topic Editors: Suresh Mathivanan, Sai Vara Prasad ChittiDeadline: 30 October 2026
Topic in
Biomedicines, Metabolites, Proteomes, Genes, J
Multi-Omics in Precision Medicine
Topic Editors: Michele Costanzo, Armando CeveniniDeadline: 31 December 2026

Special Issues
Special Issue in
Proteomes
Proteomics in Diabetes: From Mechanisms to Biomarkers
Guest Editor: Sindre Lee-ØdegårdDeadline: 31 May 2026
Special Issue in
Proteomes
Plant Genomics and Proteomics
Guest Editors: Setsuko Komatsu, Pingfang YangDeadline: 30 June 2026
Special Issue in
Proteomes
Clinical Proteomics: Unravelling Drug and Disease Mechanisms (Fifth Edition)
Guest Editor: Vikram SharmaDeadline: 30 June 2026









