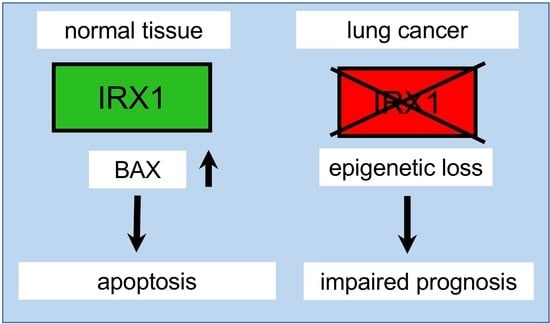
Epigenetic Inactivation of the Tumor Suppressor IRX1 Occurs Frequently in Lung Adenocarcinoma and Its Silencing Is Associated with Impaired Prognosis
Abstract
:Simple Summary
Abstract
1. Introduction
2. Results
2.1. Epigenetic Silencing of IRX1 in Cancer Cells

2.2. Epigenetic Inactivation of IRX1 in Lung Adenocarcinoma Is Correlated with Impaired Prognosis
2.3. Downregulation of IRX1 Is Associated with Increased EZH2 and DNMT3A Expression
2.4. Overexpression of IRX1 Is Associated with Fragmented Nuclei and Induction of BAX
3. Discussion
4. Materials and Methods
4.1. Cell lines, Lung Cancer Tissues, and Controls
4.2. Cell Culture and Treatment of Cell Lines
4.3. DNA Isolation and Methylation Analysis
4.4. Patient Biospecimen
4.5. RNA Expression Analysis
4.6. Plasmids
4.7. Genetic and Epigenetic Editing
4.8. Fluorescent Microscopy
4.9. Data and Statistical Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Ferlay, J.; Soerjomataram, I.; Dikshit, R.; Eser, S.; Mathers, C.; Rebelo, M.; Parkin, D.M.; Forman, D.; Bray, F. Cancer incidence and mortality worldwide: Sources, methods and major patterns in GLOBOCAN 2012. Int. J. Cancer 2015, 136, E359–E386. [Google Scholar] [CrossRef] [PubMed]
- Dammann, R.; Li, C.; Yoon, J.H.; Chin, P.L.; Bates, S.; Pfeifer, G.P. Epigenetic inactivation of a RAS association domain family protein from the lung tumour suppressor locus 3p21.3. Nat. Genet. 2000, 25, 315–319. [Google Scholar] [CrossRef] [PubMed]
- Kiehl, S.; Zimmermann, T.; Savai, R.; Pullamsetti, S.S.; Seeger, W.; Bartkuhn, M.; Dammann, R.H. Epigenetic silencing of downstream genes mediated by tandem orientation in lung cancer. Sci. Rep. 2017, 7, 3896. [Google Scholar] [CrossRef] [PubMed]
- Richter, A.M.; Kiehl, S.; Koger, N.; Breuer, J.; Stiewe, T.; Dammann, R.H. ZAR1 is a novel epigenetically inactivated tumour suppressor in lung cancer. Clin. Epigenetics 2017, 9, 60. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gomez-Skarmeta, J.L.; Modolell, J. Iroquois genes: Genomic organization and function in vertebrate neural development. Curr. Opin. Genet. Dev. 2002, 12, 403–408. [Google Scholar] [CrossRef]
- Kim, K.H.; Rosen, A.; Bruneau, B.G.; Hui, C.C.; Backx, P.H. Iroquois homeodomain transcription factors in heart development and function. Circ. Res. 2012, 110, 1513–1524. [Google Scholar] [CrossRef] [Green Version]
- Leyns, L.; Gómez-Skarmeta, J.-L.; Dambly-Chaudière, C. iroquois: A prepattern gene that controls the formation of bristles on the thorax ofDrosophila. Mech. Dev. 1996, 59, 63–72. [Google Scholar] [CrossRef]
- Hu, W.; Xin, Y.; Zhang, L.; Hu, J.; Sun, Y.; Zhao, Y. Iroquois Homeodomain transcription factors in ventricular conduction system and arrhythmia. Int. J. Med. Sci. 2018, 15, 808–815. [Google Scholar] [CrossRef] [Green Version]
- Cavodeassi, F.; Modolell, J.; Gomez-Skarmeta, J.L. The Iroquois family of genes: From body building to neural patterning. Development 2001, 128, 2847–2855. [Google Scholar]
- Yu, W.; Li, X.; Eliason, S.; Romero-Bustillos, M.; Ries, R.J.; Cao, H.; Amendt, B.A. Irx1 regulates dental outer enamel epithelial and lung alveolar type II epithelial differentiation. Dev. Biol. 2017, 429, 44–55. [Google Scholar] [CrossRef]
- Lu, Y.; Yu, Y.; Zhu, Z.; Xu, H.; Ji, J.; Bu, L.; Liu, B.; Jiang, H.; Lin, Y.; Kong, X.; et al. Identification of a new target region by loss of heterozygosity at 5p15.33 in sporadic gastric carcinomas: Genotype and phenotype related. Cancer Lett. 2005, 224, 329–337. [Google Scholar] [CrossRef] [PubMed]
- Arias-Pulido, H.; Narayan, G.; Vargas, H.; Mansukhani, M.; Murty, V.V. Mapping common deleted regions on 5p15 in cervical carcinoma and their occurrence in precancerous lesions. Mol. Cancer 2002, 1, 3. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Xu, S.F.; Peng, Z.H.; Li, D.P.; Qiu, G.Q.; Zhang, F. Refinement of heterozygosity loss on chromosome 5p15 in sporadic colorectal cancer. World J. Gastroenterol. 2003, 9, 1713–1718. [Google Scholar] [CrossRef] [PubMed]
- Wieland, I.; Bohm, M.; Arden, K.C.; Ammermuller, T.; Bogatz, S.; Viars, C.S.; Rajewsky, M.F. Allelic deletion mapping on chromosome 5 in human carcinomas. Oncogene 1996, 12, 97–102. [Google Scholar]
- Guo, X.; Liu, W.; Pan, Y.; Ni, P.; Ji, J.; Guo, L.; Zhang, J.; Wu, J.; Jiang, J.; Chen, X.; et al. Homeobox gene IRX1 is a tumor suppressor gene in gastric carcinoma. Oncogene 2010, 29, 3908–3920. [Google Scholar] [CrossRef] [Green Version]
- Bennett, K.L.; Karpenko, M.; Lin, M.T.; Claus, R.; Arab, K.; Dyckhoff, G.; Plinkert, P.; Herpel, E.; Smiraglia, D.; Plass, C. Frequently methylated tumor suppressor genes in head and neck squamous cell carcinoma. Cancer Res. 2008, 68, 4494–4499. [Google Scholar] [CrossRef] [Green Version]
- Kitchen, M.O.; Bryan, R.T.; Emes, R.D.; Glossop, J.R.; Luscombe, C.; Cheng, K.K.; Zeegers, M.P.; James, N.D.; Devall, A.J.; Mein, C.A.; et al. Quantitative genome-wide methylation analysis of high-grade non-muscle invasive bladder cancer. Epigenetics 2016, 11, 237–246. [Google Scholar] [CrossRef] [Green Version]
- Gao, C.; Zhuang, J.; Zhou, C.; Ma, K.; Zhao, M.; Liu, C.; Liu, L.; Li, H.; Feng, F.; Sun, C. Prognostic value of aberrantly expressed methylation gene profiles in lung squamous cell carcinoma: A study based on The Cancer Genome Atlas. J. Cell. Physiol. 2019, 234, 6519–6528. [Google Scholar] [CrossRef]
- Lee, J.Y.; Lee, W.K.; Park, J.Y.; Kim, D.S. Prognostic value of Iroquois homeobox 1 methylation in non-small cell lung cancers. Genes Genom. 2020, 42, 571–579. [Google Scholar] [CrossRef]
- Jung, I.H.; Jung, D.E.; Chung, Y.Y.; Kim, K.S.; Park, S.W. Iroquois Homeobox 1 Acts as a True Tumor Suppressor in Multiple Organs by Regulating Cell Cycle Progression. Neoplasia 2019, 21, 1003–1014. [Google Scholar] [CrossRef]
- Hsiung, C.A.; Lan, Q.; Hong, Y.C.; Chen, C.J.; Hosgood, H.D.; Chang, I.S.; Chatterjee, N.; Brennan, P.; Wu, C.; Zheng, W.; et al. The 5p15.33 locus is associated with risk of lung adenocarcinoma in never-smoking females in Asia. PLoS Genet. 2010, 6, e1001051. [Google Scholar] [CrossRef] [PubMed]
- McKay, J.D.; Hung, R.J.; Gaborieau, V.; Boffetta, P.; Chabrier, A.; Byrnes, G.; Zaridze, D.; Mukeria, A.; Szeszenia-Dabrowska, N.; Lissowska, J.; et al. Lung cancer susceptibility locus at 5p15.33. Nat. Genet. 2008, 40, 1404–1406. [Google Scholar] [CrossRef] [PubMed]
- Weissfeld, J.L.; Lin, Y.; Lin, H.M.; Kurland, B.F.; Wilson, D.O.; Fuhrman, C.R.; Pennathur, A.; Romkes, M.; Nukui, T.; Yuan, J.M.; et al. Lung Cancer Risk Prediction Using Common SNPs Located in GWAS-Identified Susceptibility Regions. J. Thorac. Oncol. 2015, 10, 1538–1545. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zanetti, K.A.; Wang, Z.; Aldrich, M.; Amos, C.I.; Blot, W.J.; Bowman, E.D.; Burdette, L.; Cai, Q.; Caporaso, N.; Chung, C.C.; et al. Genome-wide association study confirms lung cancer susceptibility loci on chromosomes 5p15 and 15q25 in an African-American population. Lung Cancer 2016, 98, 33–42. [Google Scholar] [CrossRef] [Green Version]
- Lonsdale, J.; Thomas, J.; Salvatore, M.; Phillips, R.; Lo, E.; Shad, S.; Hasz, R.; Walters, G.; Garcia, F.; Young, N.; et al. The Genotype-Tissue Expression (GTEx) project. Nat. Genet. 2013, 45, 580–585. [Google Scholar] [CrossRef] [PubMed]
- Marchler-Bauer, A.; Bo, Y.; Han, L.; He, J.; Lanczycki, C.J.; Lu, S.; Chitsaz, F.; Derbyshire, M.K.; Geer, R.C.; Gonzales, N.R.; et al. CDD/SPARCLE: Functional classification of proteins via subfamily domain architectures. Nucleic Acids Res. 2017, 45, D200–D203. [Google Scholar] [CrossRef]
- Wang, J.; Zhuang, J.; Iyer, S.; Lin, X.; Whitfield, T.W.; Greven, M.C.; Pierce, B.G.; Dong, X.; Kundaje, A.; Cheng, Y.; et al. Sequence features and chromatin structure around the genomic regions bound by 119 human transcription factors. Genome Res. 2012, 22, 1798–1812. [Google Scholar] [CrossRef] [Green Version]
- Dammann, G.; Teschler, S.; Haag, T.; Altmuller, F.; Tuczek, F.; Dammann, R.H. Increased DNA methylation of neuropsychiatric genes occurs in borderline personality disorder. Epigenetics 2011, 6, 1454–1462. [Google Scholar] [CrossRef] [Green Version]
- Ghandi, M.; Huang, F.W.; Jané-Valbuena, J.; Kryukov, G.V.; Lo, C.C.; McDonald, E.R.; Barretina, J.; Gelfand, E.T.; Bielski, C.M.; Li, H.; et al. Next-generation characterization of the Cancer Cell Line Encyclopedia. Nature 2019, 569, 503–508. [Google Scholar] [CrossRef]
- Molenaar, J.J.; Koster, J.; Zwijnenburg, D.A.; van Sluis, P.; Valentijn, L.J.; van der Ploeg, I.; Hamdi, M.; van Nes, J.; Westerman, B.A.; van Arkel, J.; et al. Sequencing of neuroblastoma identifies chromothripsis and defects in neuritogenesis genes. Nature 2012, 483, 589–593. [Google Scholar] [CrossRef]
- Iorio, F.; Knijnenburg, T.A.; Vis, D.J.; Bignell, G.R.; Menden, M.P.; Schubert, M.; Aben, N.; Gonçalves, E.; Barthorpe, S.; Lightfoot, H.; et al. A Landscape of Pharmacogenomic Interactions in Cancer. Cell 2016, 166, 740–754. [Google Scholar] [CrossRef] [Green Version]
- Diez-Villanueva, A.; Mallona, I.; Peinado, M.A. Wanderer, an interactive viewer to explore DNA methylation and gene expression data in human cancer. Epigenetics Chromatin 2015, 8, 22. [Google Scholar] [CrossRef] [Green Version]
- Nagy, A.; Lanczky, A.; Menyhart, O.; Gyorffy, B. Validation of miRNA prognostic power in hepatocellular carcinoma using expression data of independent datasets. Sci. Rep. 2018, 8, 9227. [Google Scholar] [CrossRef]
- Modhukur, V.; Iljasenko, T.; Metsalu, T.; Lokk, K.; Laisk-Podar, T.; Vilo, J. MethSurv: A web tool to perform multivariable survival analysis using DNA methylation data. Epigenomics 2018, 10, 277–288. [Google Scholar] [CrossRef] [Green Version]
- Fiorentino, F.P.; Giordano, A. The tumor suppressor role of CTCF. J. Cell. Physiol. 2012, 227, 479–492. [Google Scholar] [CrossRef]
- Witcher, M.; Emerson, B.M. Epigenetic silencing of the p16(INK4a) tumor suppressor is associated with loss of CTCF binding and a chromatin boundary. Mol. Cell 2009, 34, 271–284. [Google Scholar] [CrossRef] [Green Version]
- De La Rosa-Velazquez, I.A.; Rincon-Arano, H.; Benitez-Bribiesca, L.; Recillas-Targa, F. Epigenetic regulation of the human retinoblastoma tumor suppressor gene promoter by CTCF. Cancer Res. 2007, 67, 2577–2585. [Google Scholar] [CrossRef] [Green Version]
- Haag, T.; Richter, A.M.; Schneider, M.B.; Jimenez, A.P.; Dammann, R.H. The dual specificity phosphatase 2 gene is hypermethylated in human cancer and regulated by epigenetic mechanisms. BMC Cancer 2016, 16, 49. [Google Scholar] [CrossRef] [Green Version]
- Haag, T.; Herkt, C.E.; Walesch, S.K.; Richter, A.M.; Dammann, R.H. The apoptosis associated tyrosine kinase gene is frequently hypermethylated in human cancer and is regulated by epigenetic mechanisms. Genes Cancer 2014, 5, 365–374. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Victoria-Acosta, G.; Vazquez-Santillan, K.; Jimenez-Hernandez, L.; Munoz-Galindo, L.; Maldonado, V.; Martinez-Ruiz, G.U.; Melendez-Zajgla, J. Epigenetic silencing of the XAF1 gene is mediated by the loss of CTCF binding. Sci. Rep. 2015, 5, 14838. [Google Scholar] [CrossRef] [Green Version]
- Wan, L.; Li, X.; Shen, H.; Bai, X. Quantitative analysis of EZH2 expression and its correlations with lung cancer patients’ clinical pathological characteristics. Clin. Transl. Oncol. 2013, 15, 132–138. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Zhao, H.; Lv, L.; Bao, L.; Wang, X.; Han, S. Prognostic Significance of EZH2 Expression in Non-Small Cell Lung Cancer: A Meta-analysis. Sci. Rep. 2016, 6, 19239. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lu, T.P.; Tsai, M.H.; Lee, J.M.; Hsu, C.P.; Chen, P.C.; Lin, C.W.; Shih, J.Y.; Yang, P.C.; Hsiao, C.K.; Lai, L.C.; et al. Identification of a novel biomarker, SEMA5A, for non-small cell lung carcinoma in nonsmoking women. Cancer Epidemiol. Biomark. Prev. 2010, 19, 2590–2597. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Meister, M.; Belousov, A.; Xu, E.C.; Schnabel, P.; Warth, A.; Hoffmann, H.; Dienemann, H.; Riedlinger, J.; Bodenmueller, H.; Zolg, W.; et al. Intra-tumor Heterogeneity of Gene Expression Profiles in Early Stage Non-Small Cell Lung Cancer. J. Bioinform. Res. Stud. 2014, 1, 1–20. [Google Scholar]
- Huang, S.; Reitze, N.J.; Ewing, A.L.; McCreary, S.; Uihlein, A.H.; Brower, S.L.; Wang, D.; Wang, T.; Gabrin, M.J.; Keating, K.E.; et al. Analytical Performance of a 15-Gene Prognostic Assay for Early-Stage Non-Small-Cell Lung Carcinoma Using RNA-Stabilized Tissue. J. Mol. Diagn. 2015, 17, 438–445. [Google Scholar] [CrossRef] [PubMed]
- Kuhn, A.; Loscher, D.; Marschalek, R. The IRX1/HOXA connection: Insights into a novel t(4;11)- specific cancer mechanism. Oncotarget 2016, 7, 35341–35352. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Davis, S.; Meltzer, P.S. GEOquery: A bridge between the Gene Expression Omnibus (GEO) and BioConductor. Bioinformatics 2007, 23, 1846–1847. [Google Scholar] [CrossRef] [Green Version]
- Lu, J.; Song, G.; Tang, Q.; Zou, C.; Han, F.; Zhao, Z.; Yong, B.; Yin, J.; Xu, H.; Xie, X.; et al. IRX1 hypomethylation promotes osteosarcoma metastasis via induction of CXCL14/NF-kappaB signaling. J. Clin. Investig. 2015, 125, 1839–1856. [Google Scholar] [CrossRef] [Green Version]
- Wang, T.; Xu, Y.; Hou, P. Identifying novel biomarkers of gastric cancer through integration analysis of single nucleotide polymorphisms and gene expression profile. Int. J. Biol. Markers 2015, 30, 321–326. [Google Scholar] [CrossRef]
- Hark, A.T.; Schoenherr, C.J.; Katz, D.J.; Ingram, R.S.; Levorse, J.M.; Tilghman, S.M. CTCF mediates methylation-sensitive enhancer-blocking activity at the H19/Igf2 locus. Nature 2000, 405, 486–489. [Google Scholar] [CrossRef]
- Yoon, B.; Herman, H.; Hu, B.; Park, Y.J.; Lindroth, A.; Bell, A.; West, A.G.; Chang, Y.; Stablewski, A.; Piel, J.C.; et al. Rasgrf1 imprinting is regulated by a CTCF-dependent methylation-sensitive enhancer blocker. Mol. Cell. Biol. 2005, 25, 11184–11190. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Katainen, R.; Dave, K.; Pitkanen, E.; Palin, K.; Kivioja, T.; Valimaki, N.; Gylfe, A.E.; Ristolainen, H.; Hanninen, U.A.; Cajuso, T.; et al. CTCF/cohesin-binding sites are frequently mutated in cancer. Nat. Genet. 2015, 47, 818–821. [Google Scholar] [CrossRef] [PubMed]
- Vire, E.; Brenner, C.; Deplus, R.; Blanchon, L.; Fraga, M.; Didelot, C.; Morey, L.; Van Eynde, A.; Bernard, D.; Vanderwinden, J.M.; et al. The Polycomb group protein EZH2 directly controls DNA methylation. Nature 2006, 439, 871–874. [Google Scholar] [CrossRef] [PubMed]
- Strunnikova, M.; Schagdarsurengin, U.; Kehlen, A.; Garbe, J.C.; Stampfer, M.R.; Dammann, R. Chromatin inactivation precedes de novo DNA methylation during the progressive epigenetic silencing of the RASSF1A promoter. Mol. Cell. Biol. 2005, 25, 3923–3933. [Google Scholar] [CrossRef] [Green Version]
- Huston, J.H.; Ryan, J.J. The emerging role of epigenetics in pulmonary arterial hypertension: An important avenue for clinical trials (2015 Grover Conference Series). Pulm. Circ. 2016, 6, 274–284. [Google Scholar] [CrossRef] [Green Version]
- Rowbotham, S.P.; Kim, C.F. Diverse cells at the origin of lung adenocarcinoma. Proc. Natl. Acad. Sci. USA 2014, 111, 4745–4746. [Google Scholar] [CrossRef] [Green Version]
- Bennett, K.L.; Romigh, T.; Eng, C. Disruption of Transforming Growth Factor- Signaling by Five Frequently Methylated Genes Leads to Head and Neck Squamous Cell Carcinoma Pathogenesis. Cancer Res. 2009, 69, 9301–9305. [Google Scholar] [CrossRef] [Green Version]
- Dammann, R.; Strunnikova, M.; Schagdarsurengin, U.; Rastetter, M.; Papritz, M.; Hattenhorst, U.E.; Hofmann, H.S.; Silber, R.E.; Burdach, S.; Hansen, G. CpG island methylation and expression of tumour-associated genes in lung carcinoma. Eur. J. Cancer 2005, 41, 1223–1236. [Google Scholar] [CrossRef]
- Muley, T.R.; Herth, F.J.; Schnabel, P.A.; Dienemann, H.; Meister, M. From tissue to molecular phenotyping: Pre-analytical requirements heidelberg experience. Transl. Lung Cancer Res. 2012, 1, 111–121. [Google Scholar] [CrossRef]
- Helmbold, P.; Richter, A.M.; Walesch, S.; Skorokhod, A.; Marsch, W.; Enk, A.; Dammann, R.H. RASSF10 promoter hypermethylation is frequent in malignant melanoma of the skin but uncommon in nevus cell nevi. J. Investig. Dermatol. 2012, 132, 687–694. [Google Scholar] [CrossRef] [Green Version]
- McShane, L.M.; Altman, D.G.; Sauerbrei, W.; Taube, S.E.; Gion, M.; Clark, G.M.; Statistics Subcommittee of the NCIEWGoCD. Reporting recommendations for tumor marker prognostic studies. J. Clin. Oncol. 2005, 23, 9067–9072. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Schneider, M.A.; Granzow, M.; Warth, A.; Schnabel, P.A.; Thomas, M.; Herth, F.J.; Dienemann, H.; Muley, T.; Meister, M. Glycodelin: A New Biomarker with Immunomodulatory Functions in Non-Small Cell Lung Cancer. Clin. Cancer Res. 2015, 21, 3529–3540. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bustin, S.A.; Benes, V.; Garson, J.A.; Hellemans, J.; Huggett, J.; Kubista, M.; Mueller, R.; Nolan, T.; Pfaffl, M.W.; Shipley, G.L.; et al. The MIQE guidelines: Minimum information for publication of quantitative real-time PCR experiments. Clin. Chem. 2009, 55, 611–622. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Deutschmeyer, V.; Breuer, J.; Walesch, S.K.; Sokol, A.M.; Graumann, J.; Bartkuhn, M.; Boettger, T.; Rossbach, O.; Richter, A.M. Epigenetic therapy of novel tumour suppressor ZAR1 and its cancer biomarker function. Clin. Epigenetics 2019, 11, 182. [Google Scholar] [CrossRef] [PubMed]






| Hallmarks 1 | GSE33532 [44] | GSE19804 [43] | TCGALUAD | GSE63074 [45] |
|---|---|---|---|---|
| E2F_targets 2 | 1.1 × 10−6 | 1.0 × 10−15 | 1.7 × 10−19 | 6.2 × 10−9 |
| Myc_targets_V1 2 | 1.1 × 10−6 | 5.2 × 10−3 | - | - |
| mTORC1_signaling 2 | 9.2 × 10−5 | 1.6 × 10−4 | 3.6 × 10−3 | 9.0 × 10−3 |
| G2M_checkpoint 2 | 5.6 × 10−4 | 1.6 × 10−11 | 1.3 × 10−10 | 1.4 × 10−4 |
| EMT 2 | - | 6.3 × 10−4 | - | - |
| myogenesis 3 | - | 1.3 × 10−3 | - | - |
| p53_pathway 3 | - | - | 3.6 × 10−3 | - |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Küster, M.M.; Schneider, M.A.; Richter, A.M.; Richtmann, S.; Winter, H.; Kriegsmann, M.; Pullamsetti, S.S.; Stiewe, T.; Savai, R.; Muley, T.; et al. Epigenetic Inactivation of the Tumor Suppressor IRX1 Occurs Frequently in Lung Adenocarcinoma and Its Silencing Is Associated with Impaired Prognosis. Cancers 2020, 12, 3528. https://doi.org/10.3390/cancers12123528
Küster MM, Schneider MA, Richter AM, Richtmann S, Winter H, Kriegsmann M, Pullamsetti SS, Stiewe T, Savai R, Muley T, et al. Epigenetic Inactivation of the Tumor Suppressor IRX1 Occurs Frequently in Lung Adenocarcinoma and Its Silencing Is Associated with Impaired Prognosis. Cancers. 2020; 12(12):3528. https://doi.org/10.3390/cancers12123528
Chicago/Turabian StyleKüster, Miriam M., Marc A. Schneider, Antje M. Richter, Sarah Richtmann, Hauke Winter, Mark Kriegsmann, Soni S. Pullamsetti, Thorsten Stiewe, Rajkumar Savai, Thomas Muley, and et al. 2020. "Epigenetic Inactivation of the Tumor Suppressor IRX1 Occurs Frequently in Lung Adenocarcinoma and Its Silencing Is Associated with Impaired Prognosis" Cancers 12, no. 12: 3528. https://doi.org/10.3390/cancers12123528