1. Introduction
Colorectal cancer (CRC) is a leading cause of morbidity and cancer death worldwide. In Malaysia, CRC is identified as the most common cancer in men and the second most common cancer in women [
1]. This disease is highly heterogeneous, with varying responses to cancer therapy and prognosis. The heterogeneity of CRC evolved from multiple pathways, including Chromosomal Instability (CIN), Microsatellite Instability (MSI), and CpG Island Methylator Phenotype (CIMP) [
2]. Unlike other pathways, CIMP is an epigenetic mechanism that is more dynamic [
3] and often reversible in the presence of inducing factors such as demethylating agents. The concept of CIMP was first introduced by Toyota et al. in 1999 and defined as widespread methylation in the CpG island of the genes [
4]. The methylation process was described by the addition of the methyl group (CH
3−) at the carbon 5 of the cytosine ring to form 5-methylcytosine, and the process is catalysed by the enzyme DNA methyltransferase (DNMT) in which S-adenosyl-methionine (SAM) acts as a methyl donor [
5,
6]. Aberrant methylation in CRC has correlated with the inactivation of tumour suppressor genes [
7] and the activation of oncogenes [
8] that function to control a variety of cellular processes, including apoptosis, proliferation, invasion, and migration [
9]. Aberrant DNA methylation is a hallmark of cancer that occurs early in cancer development [
10], increases with the progression of the disease, and is involved in the treatment response [
11,
12]. Therefore, DNA methylation may serve as a potential biomarker for cancer diagnosis, predicting patient prognosis, and monitoring response towards therapy. Several studies have been conducted to identify a methylation biomarker with high specificity and sensitivity to be used in the diagnosis of CRC. For instance, Freitas and his colleagues discovered promoter methylation of the three-genes panel (
MGMT,
RASSF1A, and
SEPT9) in accurately diagnosed CRC with 96.6% sensitivity and 74% specificity of detection [
13]. Furthermore, a meta-analysis from 38 studies has offered the potential diagnostic markers of hypermethylation
SFRP1,
SFRP2,
NDRG2, and
VIM genes in CRC patients [
14].
As previously stated, the methylation status of specific genes may also predict the prognosis of the patients. An interesting study by Maija et al. 2013, discovered that the activation of oncogene
KRAS along with promoter methylation of
CDKN2A leads to more aggressive rectal cancers [
15]. In addition,
CHFR is another promising prognostic marker whereby promoter methylation of this gene indicates poor prognosis in stage II microsatellite stable CRC [
16]. Numerous investigations have identified a therapeutic response mediated by DNA methylation [
17,
18,
19]. In our previous research, we identified the aberrant methylation of five potential therapeutic targets,
CCNE1,
CCNDBP1,
PON3,
CHL1, and
DDX43, involved in CRC chemoresistance [
20]. Moreover, CRC patients with the hypermethylation of
NKX6.1 [
21],
TFAP2E-DKK4 [
22], and
IGFBP3 genes [
23] were unresponsive to 5-FU chemotherapy treatment. Patients with unmethylated
RASSF1A [
24] and
SRBC [
18] were highly responsive to oxaliplatin chemotherapy drugs than patients who exhibited aberrant methylation of these genes. However, the methylation of
GPX3 was associated with oxaliplatin sensitivity [
25]. The hypermethylation of
BNIP3 reduces the sensitivity of CRC towards irinotecan chemotherapeutic drugs [
26].
DNA methylation has been recognised as a potential biomarker in CRC; however, only a few methylation markers are currently used in cancer diagnostics. DNA methylation-based biomarkers are still relatively new, hence, careful assessments of the potential biomarkers are required to further validate them prior to being used in clinical diagnostics. Therefore, in this study, we aim to profile the DNA methylation in CRC and explore cancer-specific methylation biomarkers to provide evidence that can support their use in clinical practice.
4. Discussion
We analysed the global methylation status of 54 paired CRC and the corresponding normal tissue samples. Our demographic data showed that most patients were diagnosed over 50 years of age. Many studies have reported an increase in the incidence rate of CRC among individuals aged more than 50 years old [
1,
38]. To the best of our knowledge, this study provides the largest epigenome-wide DNA methylation profiles in CRC–adjacent normal colon tissue pairs using the 450K BeadChip. The Cancer Genome Atlas [
39] has data on the DNA methylation status in 308 CRCs; however, only 38 matched tumour–adjacent normal samples were included. In 2013, Naumov and colleagues performed genome-wide methylation profiling in 22 paired CRC and adjacent normal tissues in addition to 19 colon tissue samples from cancer-free donors [
40]. Recently, Gu et al. analysed 12 pairs of CRC and adjacent normal tissues using the newest version of methylation chip; the MethylationEpic Beadchip [
41].
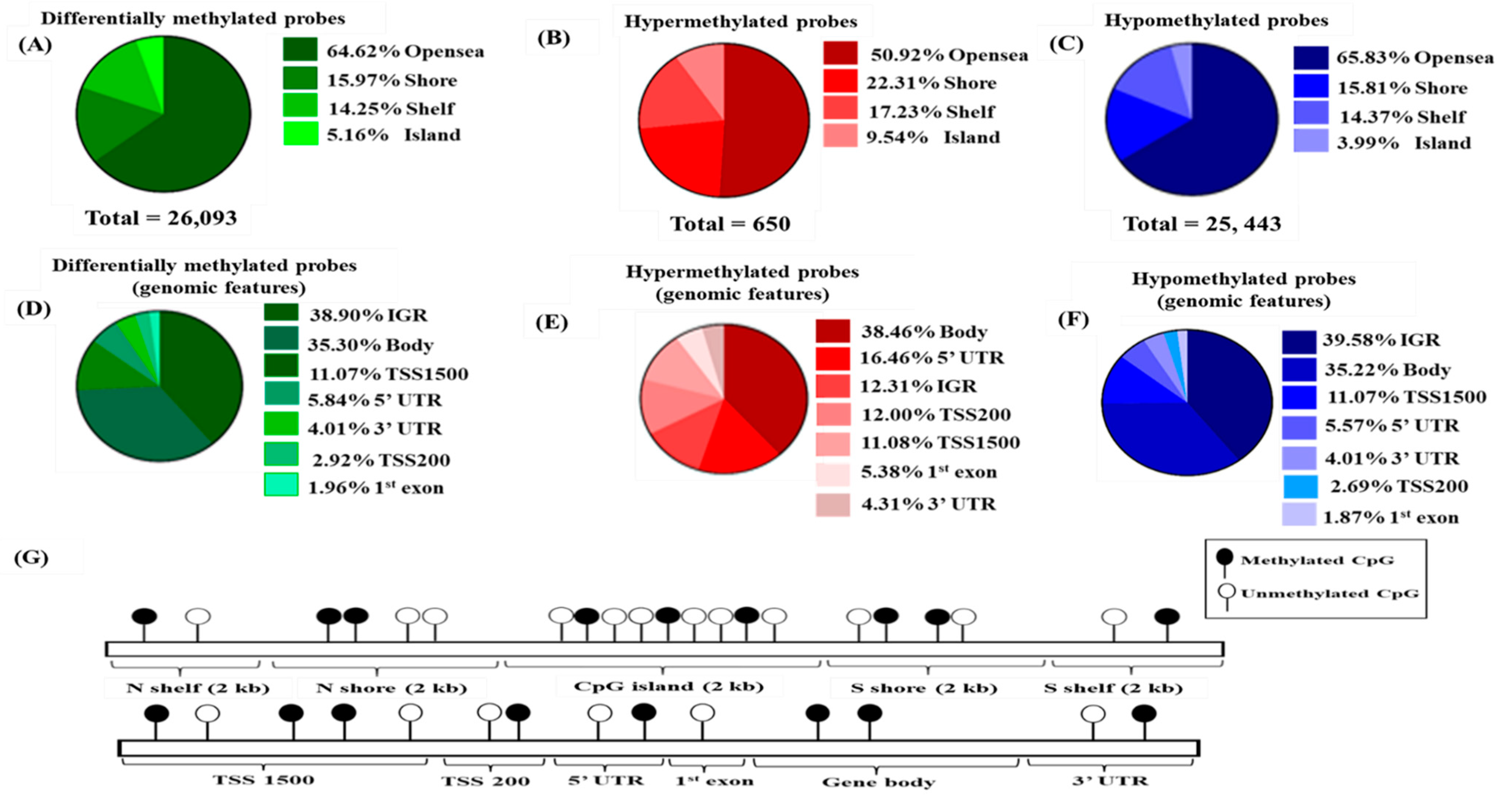
Our study revealed 26,093 differentially methylated probes that were distributed over the CpG sites of the genome. The CpG sites were comprised of several regions such as CpG island, shores (2 kb upstream from the island), shelves (2 kb upstream from the shores), and open sea, which make up for the remaining genomic region [
42]. Notably, we discovered that most of the identified CpG sites were hypomethylated rather than hypermethylated, contrasting with previous findings in other cancers [
43,
44]. Our findings corroborated recent research by Gu and colleagues, which revealed that approximately 87% of differentially methylated CpG sites were hypomethylated, whereas just 13% were hypermethylated [
41]. This may be explained by the genomic distribution of the HumanMethylation 450K microarray, which identified more probes in the open sea region (36.3%) than in CpG islands (30.9%), shores (23%), and shelves (9.7%) [
45,
46].
The CpG island is a region that is rich in CG sequences and often associated with the transcription start site. Our results showed that half of the hypermethylated probes were in the CpG island which is associated with promoter regions. This finding is supported by Sproul et al. where they showed that most of the cancers frequently exhibit hypermethylation at the CpG rich regions [
47]. Conversely, hypomethylation often occurs in the open sea area of the genome. A similar pattern has been observed in another study where hypomethylation of CpG sites was enriched at the open sea area and intergenic region [
48,
49,
50].
Various studies have reported septin 9 gene (
SEPT9) methylation in CRC, highlighting the relevance of
SEPT9 methylation in cancer [
51,
52,
53,
54,
55,
56].
SEPT9 is one of the widely studied hypermethylated genes in CRC, and our finding also supported its role in this cancer. We also uncovered many potential genes with interesting profiles. For instance, a group of Homeobox A (
HOXA) cluster genes, the members of the HOX family, and an important gene in normal organ development was found to be significantly hypermethylated in CRC compared to the normal adjacent tissues. On top of that, four genes in this family, namely
HOXA5, HOXA3, HOXA2, and
HOXA6, were hypermethylated at multiple loci. Our discovery is supported by a recent study by Li and colleagues, who reported the hypermethylation of
HOXA5,
HOXA2, and
HOXA6 [
57]. Furthermore,
HOXA5 methylation was shown to be associated with age, stage, and tumour status, while
HOXA6 methylation was linked to age and
KRAS mutation [
57]. The
HOXA family has been the subject of substantial research in cancer. Numerous clinical trials have been conducted on the
HOXA genes, but none have focused exclusively on the
HOXA2 gene. Recently, the methylation level of
HOXA1 was used to accurately differentiate between cholangiocarcinoma and benign biliary stricture from brushed biliary samples in clinical trial NCT04568512 [
58].
HOXA9 is among the biomarkers studied in the myeloid leukaemia clinical trial NCT03701295. The expression level of
HOXA9 was measured in myeloid leukaemia after treatment with chemotherapeutic drugs, including Pinometostat and Azacitidine [
59]. Despite the fact that several
HOXA family genes have been translated into clinical trials, none have focused entirely on the
HOXA2 gene. The relationship of
HOXA2 with cancer progression is limited, and the role of
HOXA2 in cancer prognosis and response to treatment is unknown. In 2019, Li and colleagues established a link between
HOXA2 and age, cancer staging, lymphovascular invasion, and lymph node involvement in CRC [
57]. However, this is the only study within the last decade that revealed the significance of
HOXA2 in the clinicopathological characteristics of CRC. The identification of hypermethylation of the
HOXA2 gene as a biomarker for CRC in our study adds to the evidence of
HOXA2′s association with cancer. Hence, our research may aid in the clinical development of
HOXA2. In addition, it will be interesting to assess the clinicopathological correlation with the methylation status of
HOXA2 genes in our patients, which will be a subject for future research.
Human 5-oxoprolinase,
OPLAH, was shown to be hypermethylated in our study, which had a significant impact on the gene’s downregulation, suggesting a possible contribution to CRC through the dysregulation of gene expression. Numerous investigations have also revealed that
OPLAH is frequently hypermethylated in CRC versus normal tissues [
40,
60]. Despite a paucity of information on
OPLAH methylation in cancer, several patents have been filed for its applicability in cancer detection. Recently,
OPLAH was identified as one of the biomarkers that have been patented for the diagnosis of lung cancer (patent number: US 11028447 B2) [
61] and CRC (patent number: US 11078539 B2) [
62]. Taken together, our findings suggest that hypermethylated
OPLAH has a role in the identification of CRC in the Malaysian population. In the future, the methylation status of
OPLAH can be determined in blood, urine, and saliva, thereby establishing
OPLAH as a non-invasive biomarker and accelerating the translation of molecular evidence to clinical practice.
Next, we discovered that
TRIM31 displayed a global loss of DNA methylation in CRC tissues. DNA hypomethylation is a common epigenetic alteration observed in human oncogenes.
TRIM31 is an oncogene that has been shown to be overexpressed in different types of cancer, including pancreatic [
63], acute myeloid leukaemia [
64], hepatocellular carcinoma [
65], breast [
66], and CRC [
67]. Whilst
TRIM31 is commonly overexpressed in cancers, the mechanism by which
TRIM31 is overexpressed remains unknown. Our study is the first to demonstrate that
TRIM31 is hypomethylated in CRC. DNA hypomethylation may enhance gene expression by allowing transcription factors to bind to the promoter region of the gene. Therefore, we postulated that the overexpression of
TRIM31 in CRC is mediated by a global loss of DNA methylation. Nonetheless, additional research is necessary to confirm the association between DNA hypomethylation and
TRIM31 overexpression. In addition, increased
TRIM31 expression was associated with an aggressive phenotype and poor prognosis in pancreatic cancer. Moreover, this gene lowered the chemosensitivity of pancreatic cancer to gemcitabine, a commonly used chemotherapy treatment for pancreatic cancer [
63]. In CRC, the high expression of
TRIM31 promotes cancer invasion and metastasis [
67]. According to the findings presented, the overexpression of
TRIM31 had a role in the development, invasion, and metastasis of cancer, as well as resistance to chemotherapy.
Barrow and colleagues [
68] performed an epigenome-wide analysis of DNA methylation in CRC patients with different smoking statuses, and among the significant hypomethylated genes were the
SDK1, PTPRN2, and
TNXB genes. In our study, we identified 116 hypomethylated
SDK1 loci and 83 hypomethylated loci in each of the
PTPRN2 and
TNXB genes. These three genes also contained the highest number of hypomethylated loci. Sidekick cell adhesion molecule 1 (
SDK1) hypomethylation was also reported in sporadic colorectal cancer [
69] and is concordant with our findings. Receptor-type tyrosine-protein phosphatase N2 (
PTPRN2) hypomethylation, on the other hand, has been rarely reported. The
TNXB (tenascin XB) gene was first implicated in Ehlers–Danlos syndrome [
70], but its role in several human cancers have been established, including nasopharyngeal [
71] and mesothelioma [
72]. Recent evidence further supports its role in malignancy, whereby
TNXB is indicated as one of the triple-evidenced genes, which exhibit aberrant methylation, differentially expressed and associated with somatic mutation, hence, displaying the superior predictive ability in cancer diagnosis and prognosis [
73].
The relationship between DNA methylation and gene expression is highly complex, and traditionally, DNA methylation-silenced gene expression was primarily affected in the CpG island promoter region [
74,
75,
76] since the regulation of expression is controlled by transcription factors that bind to the promoter [
77]. To gain further insight into the role of promoter methylation in silencing the gene expression, we performed an in silico analysis of gene expression profiles of normal and CRC samples from the TCGA datasets. We focused on the methylation profile of the promoter methylated genes and the association with the gene expression. Our analysis showed supporting evidence that genes associated with hypermethylated promoters display reduced gene expressions in CRC patients [
78,
79]. The addition of a methyl group at the promoter of the genes inhibits the binding of the transcription factor to the promoter region, hence, initiating the activation of genes [
80].
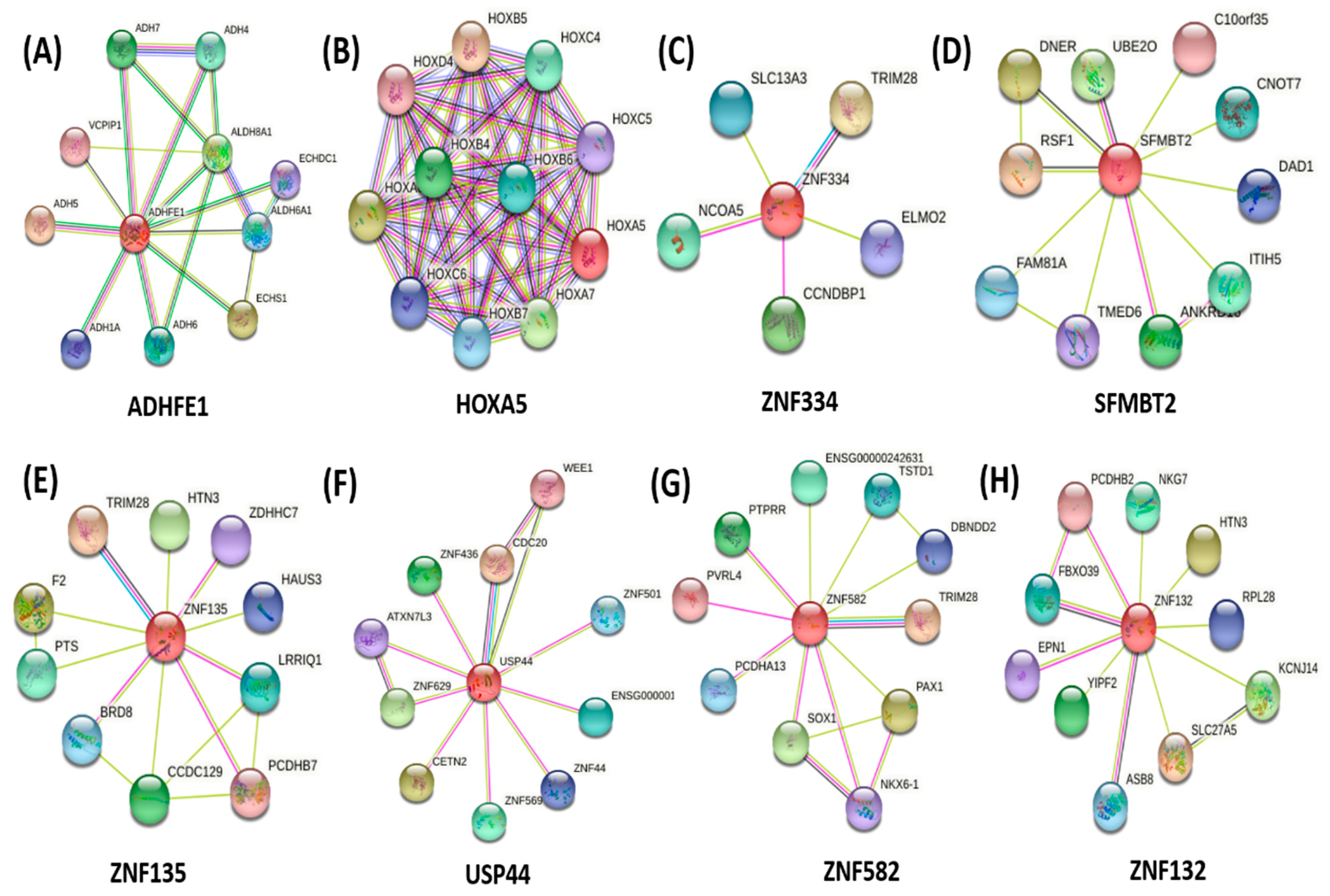
A protein-protein interaction network was constructed to identify the regulator protein among the selected promoter hypermethylated.
HOXA5 and
ADHFE1 had strong protein-protein interactions with neighbouring proteins to form a regulatory network. The strong regulatory protein may influence the expression of neighbouring proteins, which contribute to the pathogenesis of CRC. Furthermore, the strong connection among the proteins is likely to form a protein complex and often direct the biological processes [
81,
82]. Our findings discovered the interaction of
HOXA5 with
HOXB6 and
HOXB7. The downregulation of
HOXA5 increased the expression of
HOXB6 and
HOXB7, which were associated with poor clinical outcomes in cancer patients [
83,
84,
85]. On the other hand, the hypermethylation of
ADHFE1 further reduced the expression of neighbouring proteins,
ADH6,
ADH7, as well as
ADH1A, in which the genes were associated with the patient’s prognosis and cancer pathogenesis [
86,
87,
88].
Taken together, this summarises the potential of
HOXA5 and
ADHFE1 as promising biomarkers in CRC. However, when we performed ROC analysis for the top ten hypermethylated promoters mediating gene silencing,
ADHFE1 showed the highest discriminative values that significantly differentiated the CRCs from the normal colonic tissues. Compared with
HOXA5, the better discrimination of normal and cancer tissues by
ADHFE1 signifies the great potential for this gene as a methylation marker to indicate pathological changes.
ADHFE1, known as Alcohol Dehydrogenase Ion Containing 1, is a member of the iron-activated alcohol dehydrogenase family [
89]. Consistent with previous studies, we observed that the CpG island methylation status of the
ADHFE1 promoter was higher in CRC tissues in contrast to their adjacent normal mucosa, and the loss expression of
ADHFE1 in CRC was associated with promoter methylation [
90,
91].
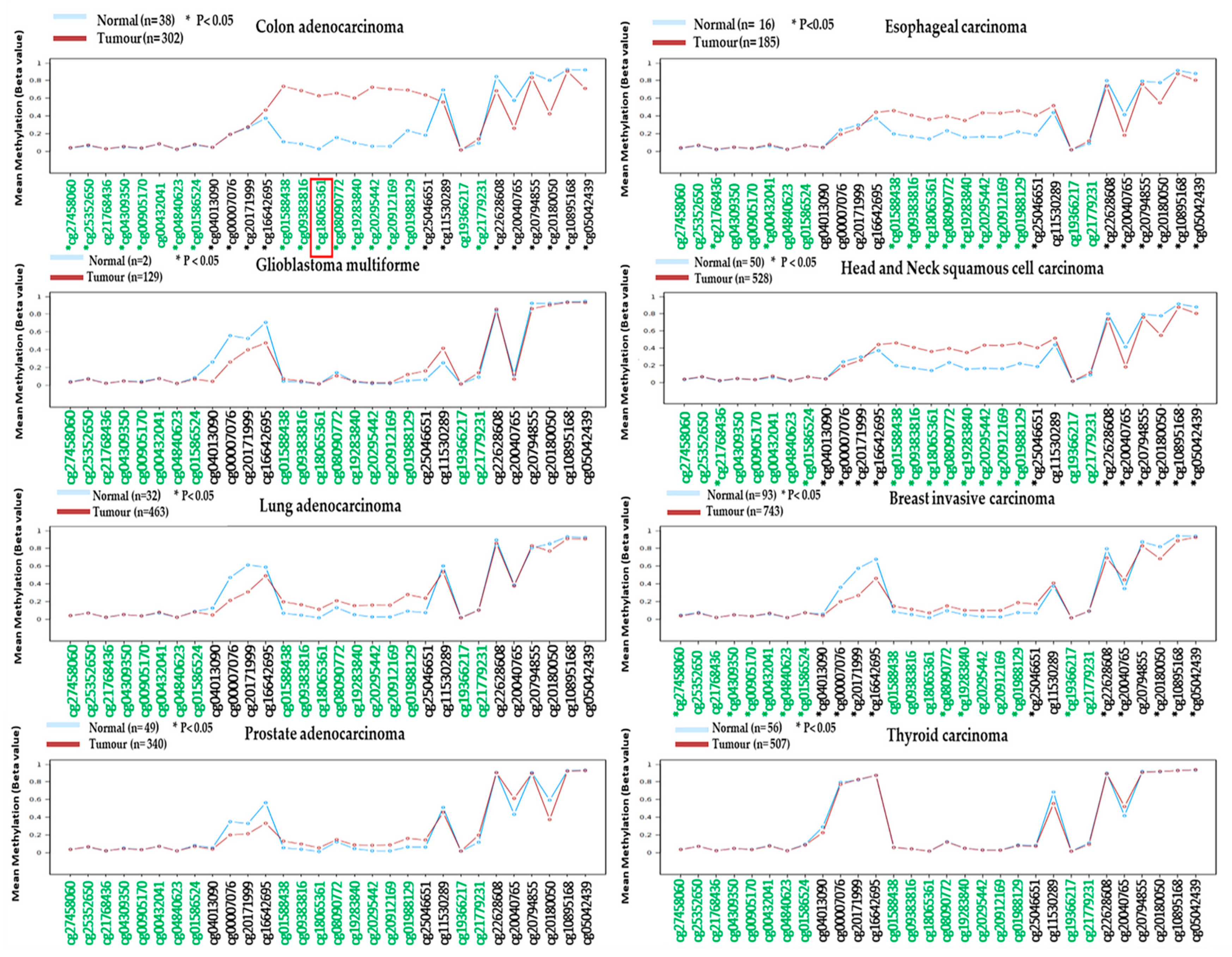
Our additional methylation analysis using the Wanderer database further strengthens the diagnostic potential of hypermethylated
ADHFE1 in detecting CRC as we observed that the methylation of
ADHFE1 displayed significant differences in CRCs versus normal tissues as compared to other cancers, for instance, glioblastoma multiforme (GBM), breast-invasive carcinoma, esophageal carcinoma, lung adenocarcinoma, head and neck squamous cell carcinoma, prostate adenocarcinoma, and thyroid carcinoma. The potential of
ADHFE1 as early detection of CRC was also discovered by Fan and his colleagues, whereby they observed hypermethylated
ADHFE1 in colorectal adenoma [
92]. According to Moon et al., the hypermethylation of the
ADHFE1 gene promotes cell proliferation in CRC [
91]. This finding was supported by Hu and colleagues, who demonstrated that the hypermethylation of the
ADHFE1 gene enhances CRC proliferation via altering cell cycle progression [
89].
ADHFE1 acted as a tumour suppressor gene in esophageal squamous cell carcinoma and was reported to be hypermethylated in a Chinese Han population [
93]. More recently, the downregulation of the
ADHFE1 gene has been linked to decreased cancer survival [
94]. Additionally, this study discovered that
ADHFE1 might contribute to cancer progression through its interactions with signalling pathways such as energy metabolism, DNA replication, and the cell cycle. With the evidence provided, we believed the promoter methylation-mediated downregulated gene of
ADHFE1 could be one of the potential DNA methylation biomarkers in detecting CRC. However, the diagnostic roles of
ADHFE1 will be subjected to further validation in a larger patient cohort in our country. It will be interesting if the methylation of
ADHFE1 can be detected in the blood for an early, accurate, and non-invasive blood-based biomarker.