Transcriptomics and Phenotypic Analysis of gpr56 Knockout in Zebrafish
Abstract
:1. Introduction
2. Results
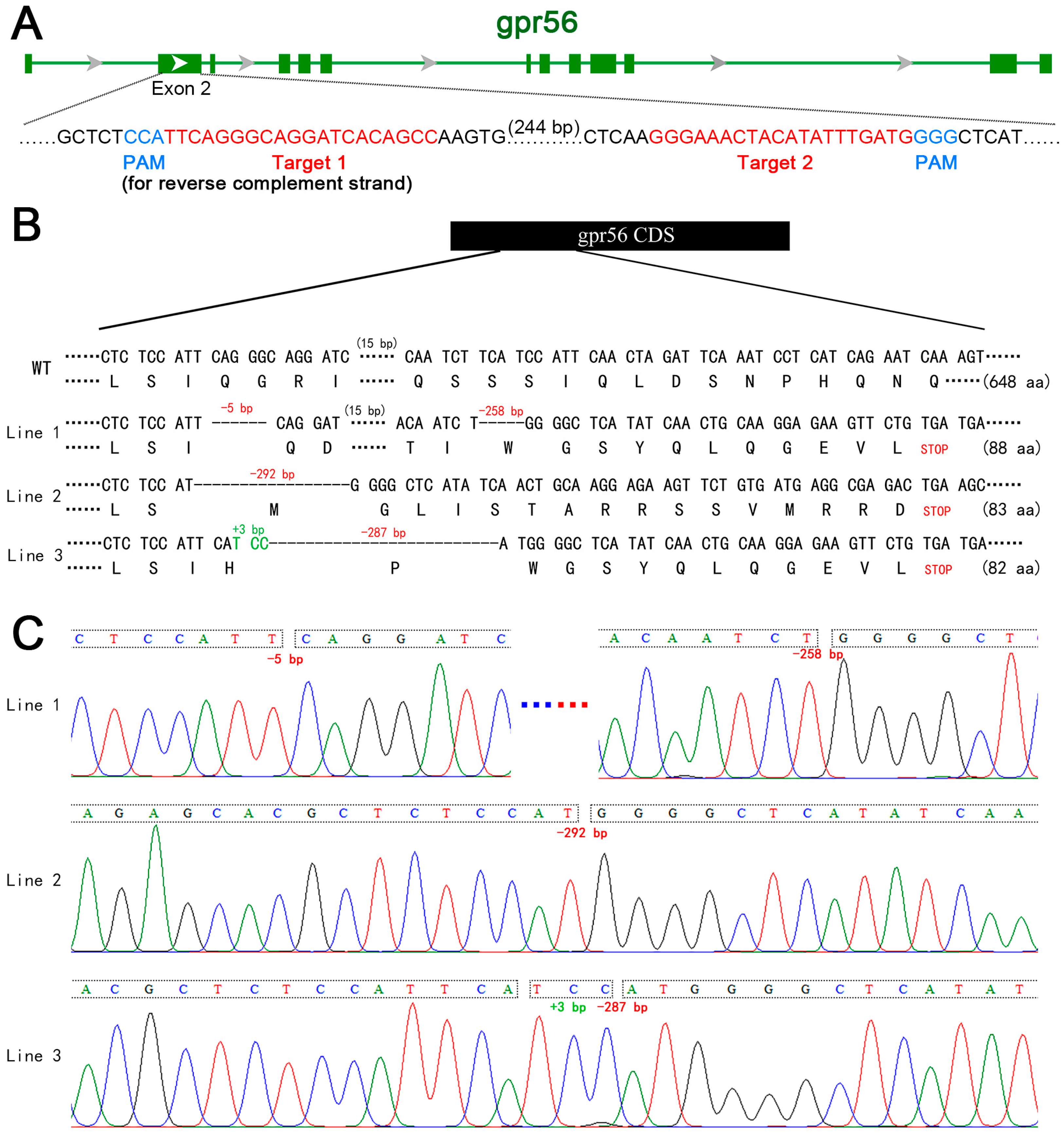
2.1. Construction of gpr56-Knockout Zebrafish Lines
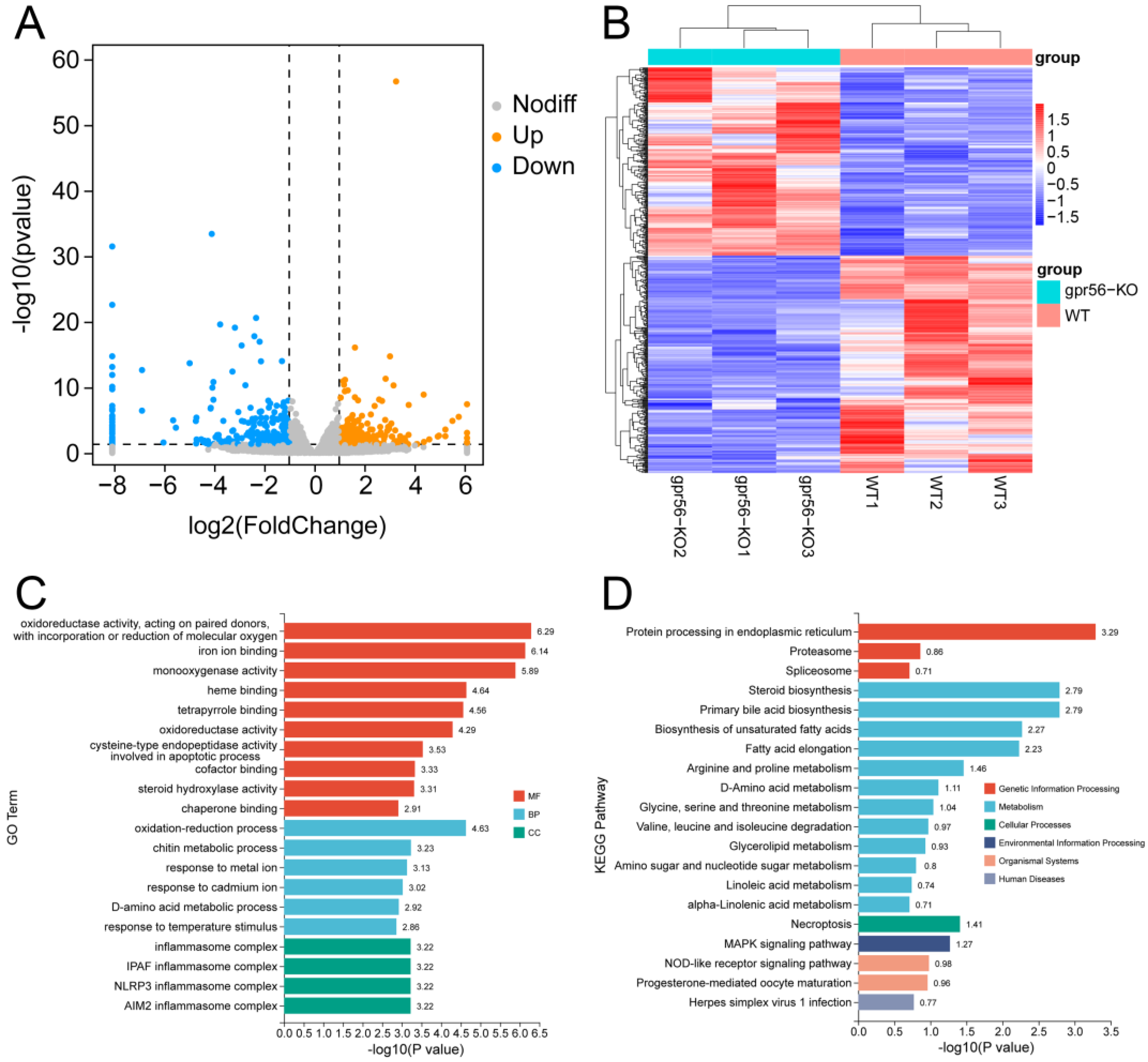
2.2. Analysis of Transcriptome Sequencing Data
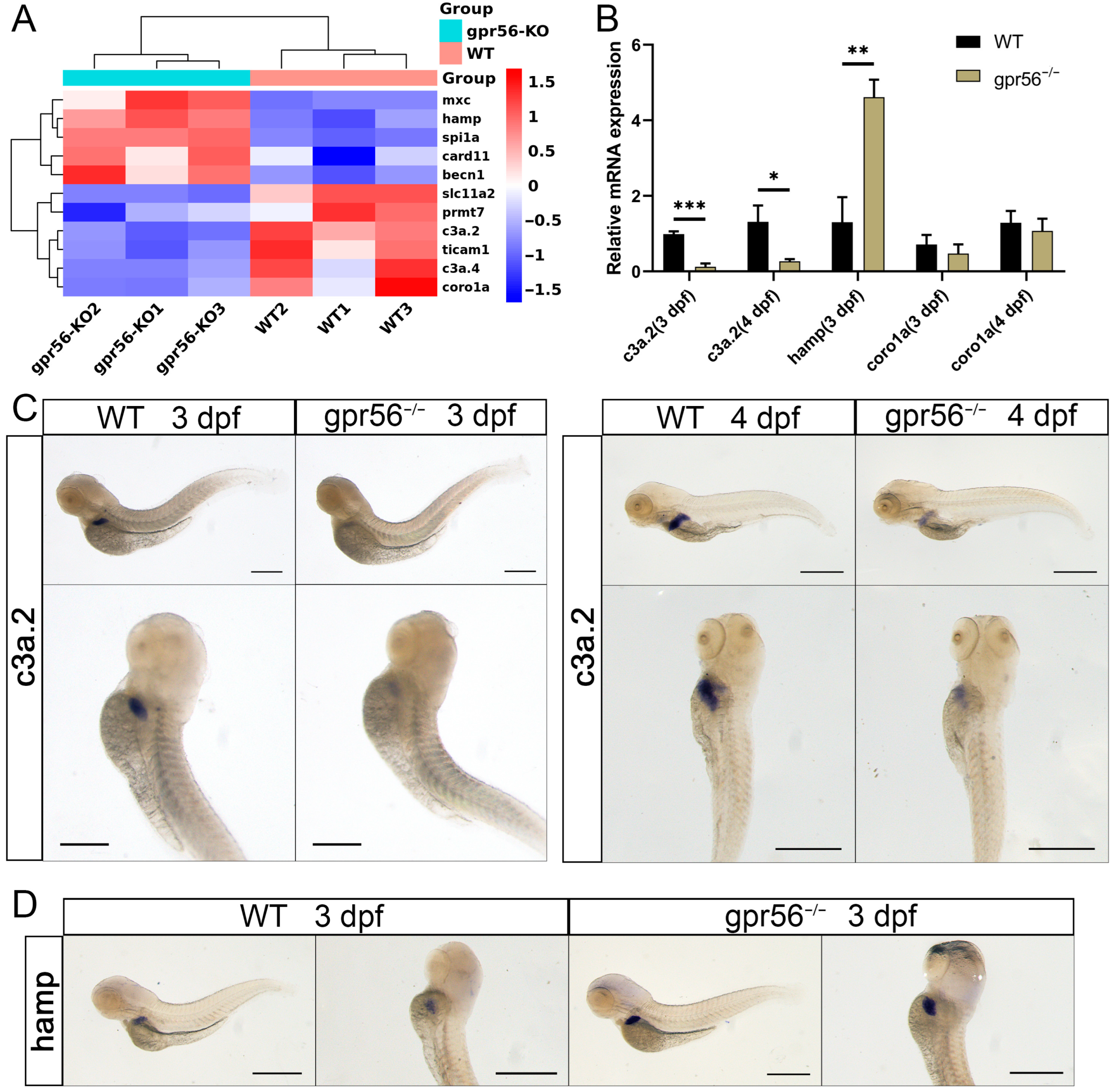
2.3. Deletion of gpr56 Leads to an Imbalance of Innate Immunity-Related Gene Expression in Zebrafish Juveniles
2.4. The Absence of gpr56 Causes Multiple Damages to Pancreatic Exocrine Secretion
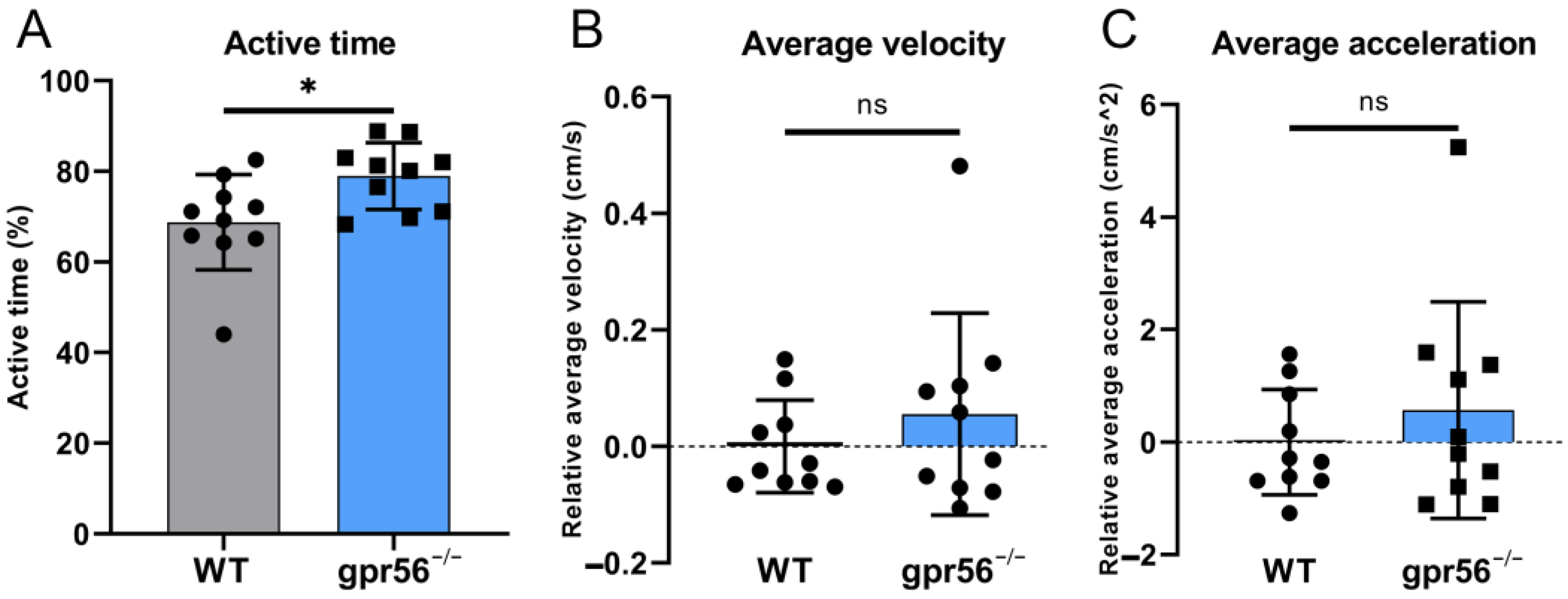
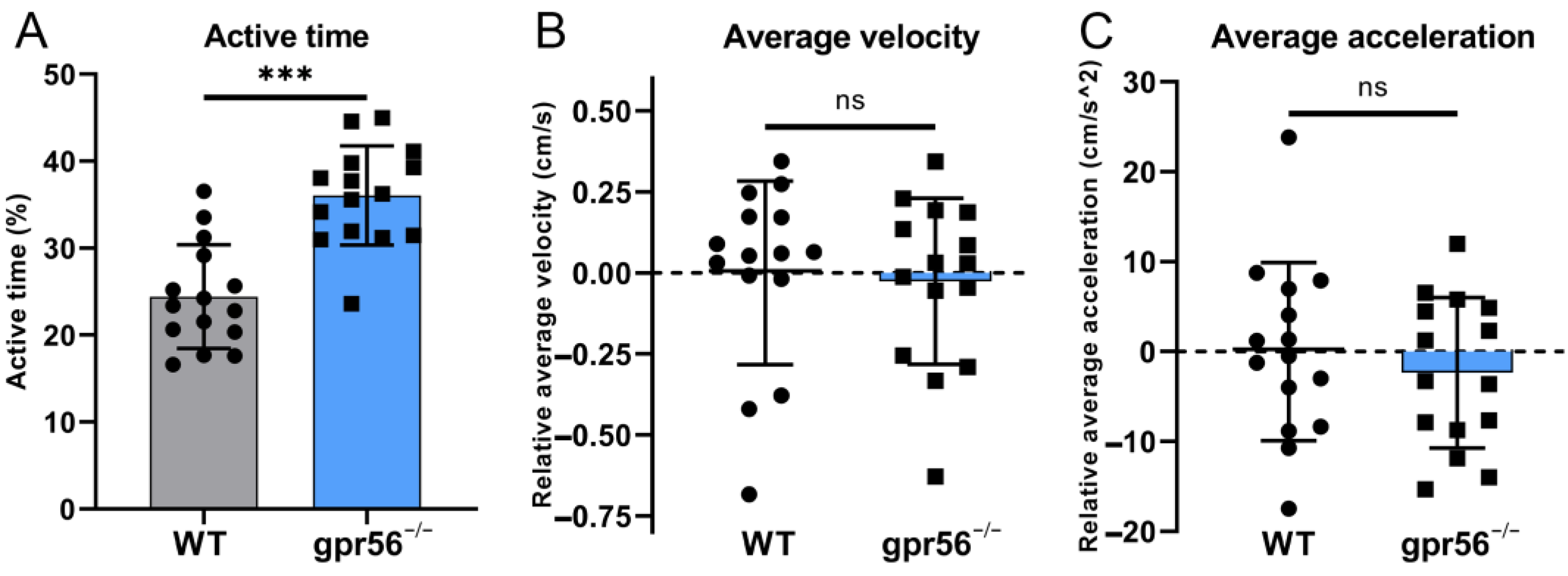
2.5. Deletion of gpr56 Makes 5 dpf Zebrafish More Active
3. Discussion
4. Materials and Methods
4.1. Experimental Zebrafish and Gene Knockout
4.2. RNA-seq Transcriptome Analysis
4.3. qPCR Detection
4.4. Synthesis of RNA Probes and Whole In Situ Hybridization (WISH)
4.5. Imaging and Image Analysis
4.6. Motor Ability Test
4.7. Statistical Analysis
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Zhou, X.E.; He, Y.; de Waal, P.W.; Gao, X.; Kang, Y.; Van Eps, N.; Yin, Y.; Pal, K.; Goswami, D.; White, T.A.; et al. Identification of Phosphorylation Codes for Arrestin Recruitment by G Protein-Coupled Receptors. Cell 2017, 170, 457–469.e413. [Google Scholar] [CrossRef] [PubMed]
- Kobilka, B.K. G protein coupled receptor structure and activation. Biochim. Biophys. Acta 2007, 1768, 794–807. [Google Scholar] [CrossRef] [PubMed]
- Kang, Y.; Kuybeda, O.; de Waal, P.W.; Mukherjee, S.; Van Eps, N.; Dutka, P.; Zhou, X.E.; Bartesaghi, A.; Erramilli, S.; Morizumi, T.; et al. Cryo-EM structure of human rhodopsin bound to an inhibitory G protein. Nature 2018, 558, 553–558. [Google Scholar] [CrossRef] [PubMed]
- Ackerman, S.D.; Garcia, C.; Piao, X.; Gutmann, D.H.; Monk, K.R. The adhesion GPCR Gpr56 regulates oligodendrocyte development via interactions with Gα12/13 and RhoA. Nat. Commun. 2015, 6, 6122. [Google Scholar] [CrossRef] [PubMed]
- Liebscher, I.; Cevheroğlu, O.; Hsiao, C.C.; Maia, A.F.; Schihada, H.; Scholz, N.; Soave, M.; Spiess, K.; Trajković, K.; Kosloff, M.; et al. A guide to adhesion GPCR research. FEBS J. 2021, 289, 7610–7630. [Google Scholar] [CrossRef]
- Strokes, N.; Piao, X. Adhesion-GPCRs in the CNS. Adv. Exp. Med. Biol. 2010, 706, 87–97. [Google Scholar] [CrossRef]
- Singh, A.K.; Lin, H.H. The role of GPR56/ADGRG1 in health and disease. Biomed. J. 2021, 44, 534–547. [Google Scholar] [CrossRef]
- Yang, T.Y.; Chiang, N.Y.; Tseng, W.Y.; Pan, H.L.; Peng, Y.M.; Shen, J.J.; Wu, K.A.; Kuo, M.L.; Chang, G.W.; Lin, H.H. Expression and immunoaffinity purification of recombinant soluble human GPR56 protein for the analysis of GPR56 receptor shedding by ELISA. Protein Expr. Purif. 2015, 109, 85–92. [Google Scholar] [CrossRef]
- Kaiser, F.; Morawski, M.; Krohn, K.; Rayes, N.; Hsiao, C.C.; Quaas, M.; Aust, G. Adhesion GPCR GPR56 Expression Profiling in Human Tissues. Cells 2021, 10, 3557. [Google Scholar] [CrossRef]
- Piao, X.; Hill, R.S.; Bodell, A.; Chang, B.S.; Basel-Vanagaite, L.; Straussberg, R.; Dobyns, W.B.; Qasrawi, B.; Winter, R.M.; Innes, A.M.; et al. G protein-coupled receptor-dependent development of human frontal cortex. Science 2004, 303, 2033–2036. [Google Scholar] [CrossRef]
- Öncü-Öner, T.; Ünalp, A.; Porsuk-Doru, İ.; Ağılkaya, S.; Güleryüz, H.; Saraç, A.; Ergüner, B.; Yüksel, B.; Hız-Kurul, S.; Cingöz, S. GPR56 homozygous nonsense mutation p.R271* associated with phenotypic variability in bilateral frontoparietal polymicrogyria. Turk. J. Pediatr. 2018, 60, 229–237. [Google Scholar] [CrossRef]
- Fujii, Y.; Ishikawa, N.; Kobayashi, Y.; Kobayashi, M.; Kato, M. Compound heterozygosity in GPR56 with bilateral frontoparietal polymicrogyria. Brain Dev. 2014, 36, 528–531. [Google Scholar] [CrossRef]
- Parrini, E.; Ferrari, A.R.; Dorn, T.; Walsh, C.A.; Guerrini, R. Bilateral frontoparietal polymicrogyria, Lennox-Gastaut syndrome, and GPR56 gene mutations. Epilepsia 2009, 50, 1344–1353. [Google Scholar] [CrossRef]
- Sawal, H.A.; Harripaul, R.; Mikhailov, A.; Vleuten, K.; Naeem, F.; Nasr, T.; Hassan, M.J.; Vincent, J.B.; Ayub, M.; Rafiq, M.A. Three Mutations in the Bilateral Frontoparietal Polymicrogyria Gene GPR56 in Pakistani Intellectual Disability Families. J. Pediatr. Genet. 2018, 7, 60–66. [Google Scholar] [CrossRef]
- Koirala, S.; Jin, Z.; Piao, X.; Corfas, G. GPR56-regulated granule cell adhesion is essential for rostral cerebellar development. J. Neurosci. Off. J. Soc. Neurosci. 2009, 29, 7439–7449. [Google Scholar] [CrossRef]
- Thisse, B.; Thisse, C. Fast Release Clones: A High Throughput Expression Analysis; University of Virginia: Charlottesville, VA, USA, 2004. [Google Scholar]
- Harty, B.L.; Krishnan, A.; Sanchez, N.E.; Schiöth, H.B.; Monk, K.R. Defining the gene repertoire and spatiotemporal expression profiles of adhesion G protein-coupled receptors in zebrafish. BMC Genom. 2015, 16, 62. [Google Scholar] [CrossRef]
- Ackerman, S.D.; Luo, R.; Poitelon, Y.; Mogha, A.; Harty, B.L.; D’Rozario, M.; Sanchez, N.E.; Lakkaraju, A.K.K.; Gamble, P.; Li, J.; et al. GPR56/ADGRG1 regulates development and maintenance of peripheral myelin. J. Exp. Med. 2018, 215, 941–961. [Google Scholar] [CrossRef]
- Horzmann, K.A.; Freeman, J.L. Making Waves: New Developments in Toxicology With the Zebrafish. Toxicol. Sci. Off. J. Soc. Toxicol. 2018, 163, 5–12. [Google Scholar] [CrossRef]
- Berg, E.M.; Björnfors, E.R.; Pallucchi, I.; Picton, L.D.; El Manira, A. Principles Governing Locomotion in Vertebrates: Lessons From Zebrafish. Front. Neural Circuits 2018, 12, 73. [Google Scholar] [CrossRef]
- Ran, F.A.; Hsu, P.D.; Wright, J.; Agarwala, V.; Scott, D.A.; Zhang, F. Genome engineering using the CRISPR-Cas9 system. Nat. Protoc. 2013, 8, 2281–2308. [Google Scholar] [CrossRef]
- Xia, B.; Amador, G.; Viswanatha, R.; Zirin, J.; Mohr, S.E.; Perrimon, N. CRISPR-based engineering of gene knockout cells by homology-directed insertion in polyploid Drosophila S2R+ cells. Nat. Protoc. 2020, 15, 3478–3498. [Google Scholar] [CrossRef] [PubMed]
- Jiang, F.; Doudna, J.A. CRISPR-Cas9 Structures and Mechanisms. Annu. Rev. Biophys. 2017, 46, 505–529. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.; Zhou, Q.; Kesinger, J.W.; Norris, C.; Valdez, C. Heme regulates exocrine peptidase precursor genes in zebrafish. Exp. Biol. Med. 2007, 232, 1170–1180. [Google Scholar] [CrossRef] [PubMed]
- Yee, N.S.; Lorent, K.; Pack, M. Exocrine pancreas development in zebrafish. Dev. Biol. 2005, 284, 84–101. [Google Scholar] [CrossRef]
- Qu, H.Q.; Fisher-Hoch, S.P.; McCormick, J.B. Molecular immunity to mycobacteria: Knowledge from the mutation and phenotype spectrum analysis of Mendelian susceptibility to mycobacterial diseases. Int. J. Infect. Dis. IJID Off. Publ. Int. Soc. Infect. Dis. 2011, 15, e305–e313. [Google Scholar] [CrossRef]
- Majumder, M.; Huang, C.; Snider, M.D.; Komar, A.A.; Tanaka, J.; Kaufman, R.J.; Krokowski, D.; Hatzoglou, M. A novel feedback loop regulates the response to endoplasmic reticulum stress via the cooperation of cytoplasmic splicing and mRNA translation. Mol. Cell. Biol. 2012, 32, 992–1003. [Google Scholar] [CrossRef]
- Hato, T.; Dagher, P.C. How the Innate Immune System Senses Trouble and Causes Trouble. Clin. J. Am. Soc. Nephrol. CJASN 2015, 10, 1459–1469. [Google Scholar] [CrossRef]
- Rothlin, C.V.; Rathinam, V.A.; Ghosh, S. Editorial overview: Innate immunity, from host defense and beyond. Curr. Opin. Immunol. 2021, 68, iii–v. [Google Scholar] [CrossRef]
- Chang, G.W.; Hsiao, C.C.; Peng, Y.M.; Vieira Braga, F.A.; Kragten, N.A.; Remmerswaal, E.B.; van de Garde, M.D.; Straussberg, R.; König, G.M.; Kostenis, E.; et al. The Adhesion G Protein-Coupled Receptor GPR56/ADGRG1 Is an Inhibitory Receptor on Human NK Cells. Cell Rep. 2016, 15, 1757–1770. [Google Scholar] [CrossRef]
- Hamann, J.; Hsiao, C.C.; Lee, C.S.; Ravichandran, K.S.; Lin, H.H. Adhesion GPCRs as Modulators of Immune Cell Function. Handb. Exp. Pharmacol. 2016, 234, 329–350. [Google Scholar] [CrossRef]
- Su, T.; Guan, Q.; Cheng, H.; Zhu, Z.; Jiang, C.; Guo, P.; Tai, Y.; Sun, H.; Wang, M.; Wei, W.; et al. Functions of G protein-coupled receptor 56 in health and disease. Acta Physiol. 2022, 236, e13866. [Google Scholar] [CrossRef]
- Yona, S.; Lin, H.H.; Stacey, M. Immunity and adhesion-GPCRs. Adv. Exp. Med. Biol. 2010, 706, 121–127. [Google Scholar] [CrossRef]
- Locati, M.; Curtale, G.; Mantovani, A. Diversity, Mechanisms, and Significance of Macrophage Plasticity. Annu. Rev. Pathol. 2020, 15, 123–147. [Google Scholar] [CrossRef]
- Weiss, G.; Schaible, U.E. Macrophage defense mechanisms against intracellular bacteria. Immunol. Rev. 2015, 264, 182–203. [Google Scholar] [CrossRef]
- Ferrari, G.; Langen, H.; Naito, M.; Pieters, J. A coat protein on phagosomes involved in the intracellular survival of mycobacteria. Cell 1999, 97, 435–447. [Google Scholar] [CrossRef]
- Dufner, A.; Mak, T.W. CARD tricks: Controlling the interactions of CARD6 with RICK and microtubules. Cell Cycle Georget. Tex. 2006, 5, 797–800. [Google Scholar] [CrossRef]
- Hara, H.; Wada, T.; Bakal, C.; Kozieradzki, I.; Suzuki, S.; Suzuki, N.; Nghiem, M.; Griffiths, E.K.; Krawczyk, C.; Bauer, B.; et al. The MAGUK family protein CARD11 is essential for lymphocyte activation. Immunity 2003, 18, 763–775. [Google Scholar] [CrossRef]
- Zhu, J.; Li, X.; Cai, X.; Zha, H.; Zhou, Z.; Sun, X.; Rong, F.; Tang, J.; Zhu, C.; Liu, X.; et al. Arginine monomethylation by PRMT7 controls MAVS-mediated antiviral innate immunity. Mol. Cell 2021, 81, 3171–3186.e3178. [Google Scholar] [CrossRef]
- Zhu, J.; Liu, X.; Cai, X.; Ouyang, G.; Fan, S.; Wang, J.; Xiao, W. Zebrafish prmt7 negatively regulates antiviral responses by suppressing the retinoic acid-inducible gene-I-like receptor signaling. FASEB J. Off. Publ. Fed. Am. Soc. Exp. Biol. 2020, 34, 988–1000. [Google Scholar] [CrossRef]
- Luo, L.; Lu, J.; Wang, Q.; Chen, S.; Xu, A.; Yuan, S. Autophagy participates in innate immune defense in lamprey. Fish Shellfish Immunol. 2018, 83, 416–424. [Google Scholar] [CrossRef]
- Cui, J.; Jin, S.; Wang, R.F. The BECN1-USP19 axis plays a role in the crosstalk between autophagy and antiviral immune responses. Autophagy 2016, 12, 1210–1211. [Google Scholar] [CrossRef] [PubMed]
- Mei, Y.; Glover, K.; Su, M.; Sinha, S.C. Conformational flexibility of BECN1: Essential to its key role in autophagy and beyond. Protein Sci. A Publ. Protein Soc. 2016, 25, 1767–1785. [Google Scholar] [CrossRef] [PubMed]
- Kuo, C.J.; Hansen, M.; Troemel, E. Autophagy and innate immunity: Insights from invertebrate model organisms. Autophagy 2018, 14, 233–242. [Google Scholar] [CrossRef] [PubMed]
- Lo, M.W.; Woodruff, T.M. Complement: Bridging the innate and adaptive immune systems in sterile inflammation. J. Leukoc. Biol. 2020, 108, 339–351. [Google Scholar] [CrossRef]
- Ricklin, D.; Reis, E.S.; Mastellos, D.C.; Gros, P.; Lambris, J.D. Complement component C3—The “Swiss Army Knife” of innate immunity and host defense. Immunol. Rev. 2016, 274, 33–58. [Google Scholar] [CrossRef]
- Krause, A.; Neitz, S.; Mägert, H.J.; Schulz, A.; Forssmann, W.G.; Schulz-Knappe, P.; Adermann, K. LEAP-1, a novel highly disulfide-bonded human peptide, exhibits antimicrobial activity. FEBS Lett. 2000, 480, 147–150. [Google Scholar] [CrossRef]
- Park, C.H.; Valore, E.V.; Waring, A.J.; Ganz, T. Hepcidin, a urinary antimicrobial peptide synthesized in the liver. J. Biol. Chem. 2001, 276, 7806–7810. [Google Scholar] [CrossRef]
- Jiang, X.F.; Liu, Z.F.; Lin, A.F.; Xiang, L.X.; Shao, J.Z. Coordination of Bactericidal and Iron Regulatory Functions of Hepcidin in Innate Antimicrobial Immunity in a Zebrafish Model. Sci. Rep. 2017, 7, 4265. [Google Scholar] [CrossRef]
- Biemar, F.; Argenton, F.; Schmidtke, R.; Epperlein, S.; Peers, B.; Driever, W. Pancreas Development in Zebrafish: Early Dispersed Appearance of Endocrine Hormone Expressing Cells and Their Convergence to Form the Definitive Islet. Dev. Biol. 2001, 230, 189–203. [Google Scholar] [CrossRef]
- Field, H.A.; Dong, P.D.; Beis, D.; Stainier, D.Y. Formation of the digestive system in zebrafish. II. Pancreas morphogenesis. Dev. Biol. 2003, 261, 197–208. [Google Scholar] [CrossRef]
- Hui, D.Y.; Howles, P.N. Carboxyl ester lipase. J. Lipid Res. 2002, 43, 2017–2030. [Google Scholar] [CrossRef]
- Qiu, Y.; Sun, S.; Yu, X.; Zhou, J.; Cai, W.; Qian, L. Carboxyl ester lipase is highly conserved in utilizing maternal supplied lipids during early development of zebrafish and human. Biochim. Biophys. Acta Mol. Cell Biol. Lipids 2020, 1865, 158663. [Google Scholar] [CrossRef]
- Xie, H.; Ye, D.; Sepich, D.; Lin, F. S1pr2/Gα13 signaling regulates the migration of endocardial precursors by controlling endoderm convergence. Dev. Biol. 2016, 414, 228–243. [Google Scholar] [CrossRef]
- Lu, X.; Tian, J.; Wen, H.; Jiang, M.; Liu, W.; Wu, F.; Yu, L.; Zhong, S. Microcystin-LR-regulated transcriptome dynamics in ZFL cells. Aquat. Toxicol. 2019, 212, 222–232. [Google Scholar] [CrossRef]
- Wang, L.; Feng, Z.; Wang, X.; Wang, X.; Zhang, X. DEGseq: An R package for identifying differentially expressed genes from RNA-seq data. Bioinformatics 2010, 26, 136–138. [Google Scholar] [CrossRef]
- Team, R. R: A Language and Environment for Statistical Computing. R Foundation for Statistical Computing:Vienna, Austria. Computing 2009, 14, 12–21. [Google Scholar]
- Alexa, A.; Rahnenführer, J.; Lengauer, T. Improved scoring of functional groups from gene expression data by decorrelating GO graph structure. Bioinformatics 2006, 22, 1600–1607. [Google Scholar] [CrossRef]
- Yu, G.; Wang, L.G.; Han, Y.; He, Q.Y. clusterProfiler: An R package for comparing biological themes among gene clusters. OMICS A J. Integr. Biol. 2012, 16, 284–287. [Google Scholar] [CrossRef] [PubMed]
- Schmittgen, T.D.; Livak, K.J. Analyzing real-time PCR data by the comparative CT method. Nat. Protoc. 2008, 3, 1101–1108. [Google Scholar] [CrossRef]
- Thisse, C.; Thisse, B. High-resolution in situ hybridization to whole-mount zebrafish embryos. Nat. Protoc. 2008, 3, 59–69. [Google Scholar] [CrossRef]

Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Sun, L.; Yang, B.; Peng, Z.; Yang, T.; Qin, B.; Ao, J.; Yang, Y.; Wang, J.; Zheng, L.; Xie, H. Transcriptomics and Phenotypic Analysis of gpr56 Knockout in Zebrafish. Int. J. Mol. Sci. 2023, 24, 7740. https://doi.org/10.3390/ijms24097740
Sun L, Yang B, Peng Z, Yang T, Qin B, Ao J, Yang Y, Wang J, Zheng L, Xie H. Transcriptomics and Phenotypic Analysis of gpr56 Knockout in Zebrafish. International Journal of Molecular Sciences. 2023; 24(9):7740. https://doi.org/10.3390/ijms24097740
Chicago/Turabian StyleSun, Luning, Boyu Yang, Zheng Peng, Tianle Yang, Bin Qin, Jieyu Ao, Yanqun Yang, Jingling Wang, Lan Zheng, and Huaping Xie. 2023. "Transcriptomics and Phenotypic Analysis of gpr56 Knockout in Zebrafish" International Journal of Molecular Sciences 24, no. 9: 7740. https://doi.org/10.3390/ijms24097740





