MicroRNAs in Brain Metastases: Potential Role as Diagnostics and Therapeutics
Abstract
:1. Introduction

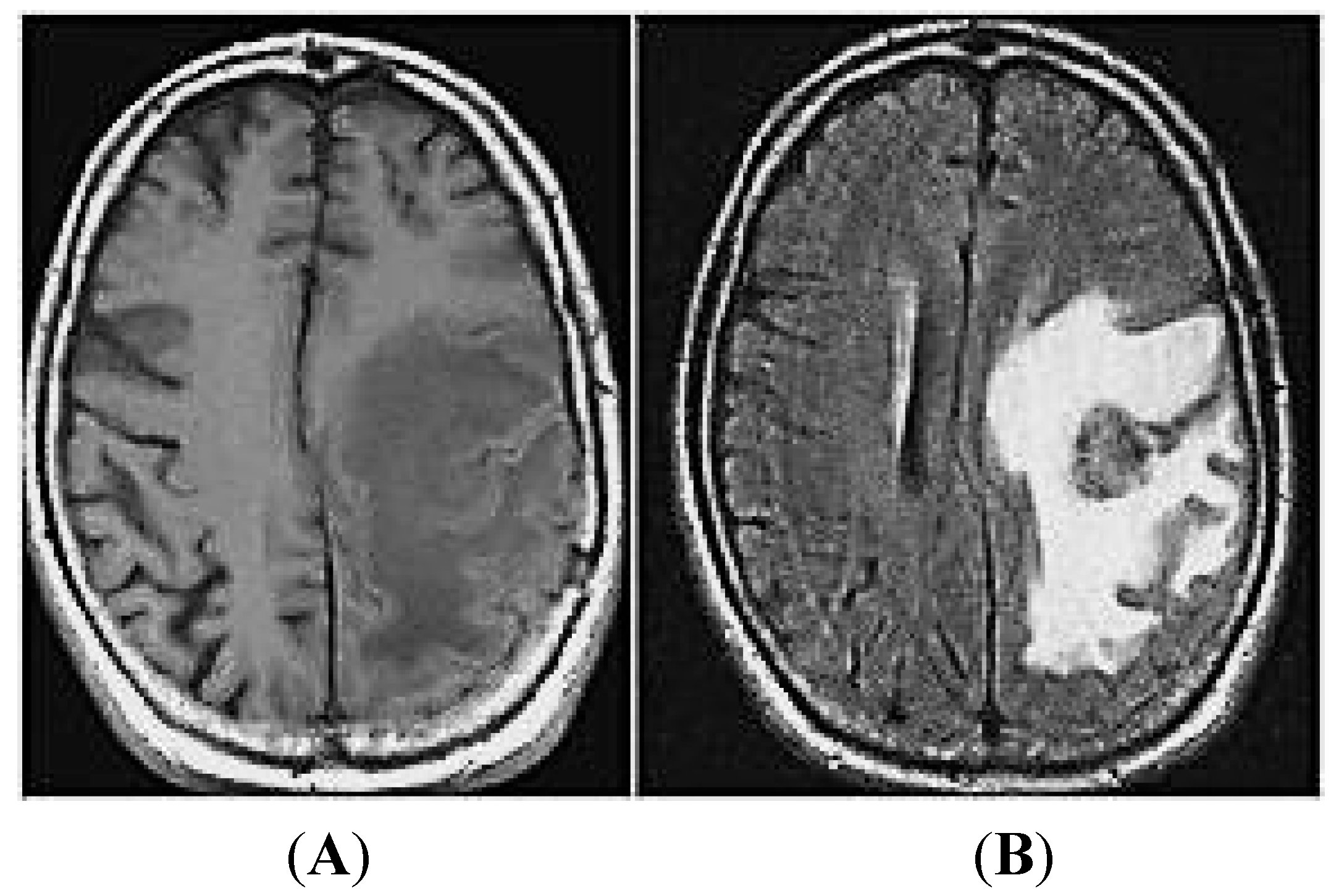
2. MiRNAs and Brain Metastases
| Deregulated MiRNA | Direction of Expression in Brain Metastases | Primary Tumor | Putative Target |
|---|---|---|---|
| miR-1258 [41] | Down-regulated | Breast | Heparanase |
| miR-7 [43] | Down-regulated | Breast | KLF4 gene |
| miR-145 [44] | Down-regulated | Lung adenocarcinoma | 3'-UTR of the JAM-A and fascin |
| miR-146-a [48] | Down-regulated | Breast | B-catenin and hnRNPC |
| miR-768-3p [49] | Down-regulated | Lung and breast | K-RAS |
| miR-19a [50] | Down-regulated | Breast | 3'-UTR of tissue factor transcript [36] |
| miR-29c [50] | Down-regulated | Breast and melanoma | Induced myeloid leukemia cell differentiation protein MCL1 [37] |
| miR-31 [51] | Down-regulated | Colon | p53 [38] |
| miR-328 [45] | Up-regulated | NSCLC | PRKCA gene |
| miR-378 [46] | Up-regulated | NSCLC | MMP-7, MMP-9 and VEGF |
| miR-200 [47] | Up-regulated | Breast and lung | E-cadherin transcriptional repressors ZEB1 and ZEB2 [39] |
| miR-210 [50] | Up-regulated | Breast and melanoma | PTP1b and HIF-1α [40] |
| miR-1, miR-145, miR-146a, miR-143, miR-10b, miR-22 [51] | Up-regulated | Colon | Multiple genes related to apoptosis and oncogenesis |
3. MiRNA Diagnostics
4. MiRNA Therapeutics
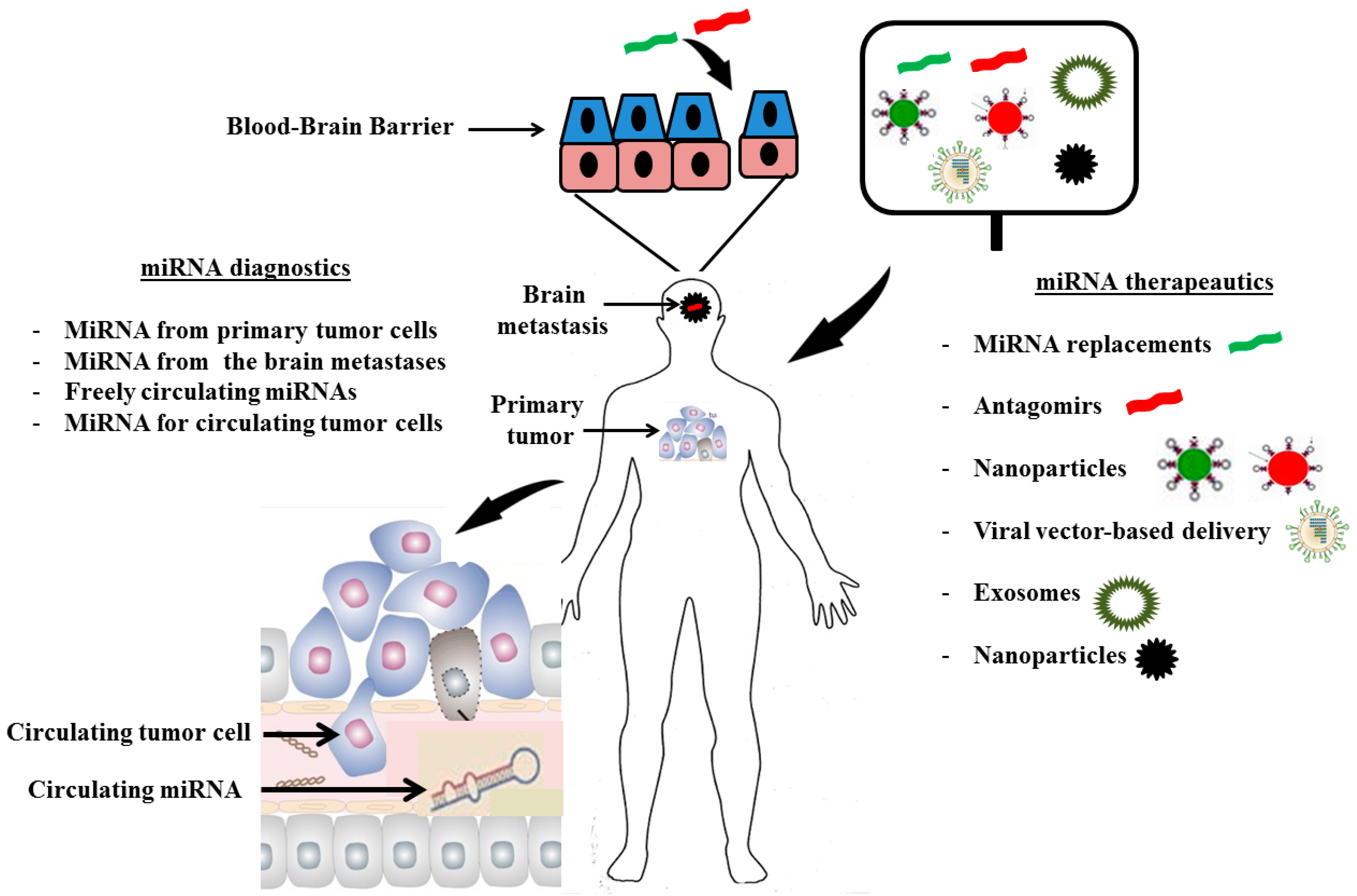
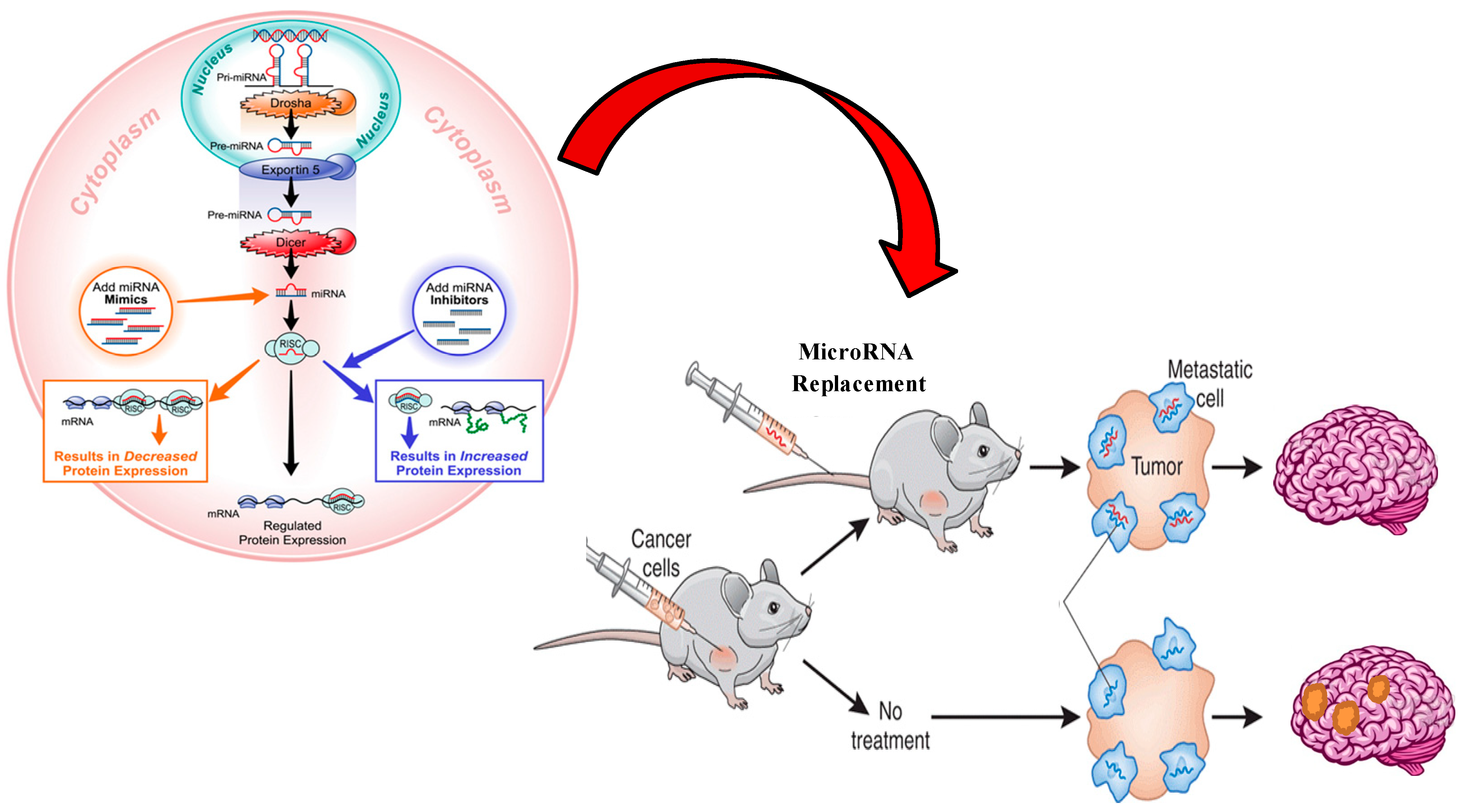
5. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Jemal, A.; Siegel, R.; Ward, E.; Hao, Y.; Xu, J.; Murray, T.; Thun, M.J. Cancer statistics, 2008. CA Cancer J. Clin. 2008, 58, 71–96. [Google Scholar] [CrossRef]
- Nayak, L.; Lee, E.Q.; Wen, P.Y. Epidemiology of brain metastases. Curr. Oncol. Rep. 2012, 14, 48–54. [Google Scholar] [CrossRef]
- Disibio, G.; French, S.W. Metastatic patterns of cancers: Results from a large autopsy study. Arch. Pathol. Lab. Med. 2008, 132, 931–939. [Google Scholar]
- Barnholtz-Sloan, J.S.; Sloan, A.E.; Davis, F.G.; Vigneau, F.D.; Lai, P.; Sawaya, R.E. Incidence proportions of brain metastases in patients diagnosed (1973 to 2001) in the Metropolitan Detroit Cancer Surveillance System. J. Clin. Oncol. 2004, 22, 2865–2872. [Google Scholar] [CrossRef]
- Schouten, L.J.; Rutten, J.; Huveneers, H.A.; Twijnstra, A. Incidence of brain metastases in a cohort of patients with carcinoma of the breast, colon, kidney, and lung and melanoma. Cancer 2002, 94, 2698–2705. [Google Scholar] [CrossRef]
- Eichler, A.F.; Loeffler, J.S. Multidisciplinary management of brain metastases. Oncologist 2007, 12, 884–898. [Google Scholar] [CrossRef]
- Rizzi, A.; Tondini, M.; Rocco, G.; Rossi, G.; Robustellini, M.; Radaelli, F.; Della Pona, C. Lung cancer with a single brain metastasis: Therapeutic options. Tumori 1990, 76, 579–581. [Google Scholar]
- Nugent, J.L.; Bunn, P.A., Jr.; Matthews, M.J.; Ihde, D.C.; Cohen, M.H.; Gazdar, A.; Minna, J.D. CNS metastases in small cell bronchogenic carcinoma: Increasing frequency and changing pattern with lengthening survival. Cancer 1979, 44, 1885–1893. [Google Scholar] [CrossRef]
- Knights, E.M., Jr. Metastatic tumors of the brain and their relation to primary and secondary pulmonary cancer. Cancer 1954, 7, 259–265. [Google Scholar] [CrossRef]
- Langley, R.R.; Fidler, I.J. The seed and soil hypothesis revisited—The role of tumor–stroma interactions in metastasis to different organs. Int. J. Cancer 2011, 128, 2527–2535. [Google Scholar] [CrossRef]
- Chishima, T.; Miyagi, Y.; Wang, X.; Yamaoka, H.; Shimada, H.; Moossa, A.; Hoffman, R.M. Cancer invasion and micrometastasis visualized in live tissue by green fluorescent protein expression. Cancer Res. 1997, 57, 2042–2047. [Google Scholar]
- Yang, M.; Baranov, E.; Jiang, P.; Sun, F.-X.; Li, X.-M.; Li, L.; Hasegawa, S.; Bouvet, M.; Al-Tuwaijri, M.; Chishima, T. Whole-body optical imaging of green fluorescent protein-expressing tumors and metastases. Proc. Natl. Acad. Sci. USA 2000, 97, 1206–1211. [Google Scholar] [CrossRef]
- Hoffman, R.M. Orthotopic metastatic mouse models for anticancer drug discovery and evaluation: A bridge to the clinic. Investig. New Drugs 1999, 17, 343–360. [Google Scholar] [CrossRef]
- Hoffman, R. In vivo imaging of metastatic cancer with fluorescent proteins. Cell Death Differ. 2002, 9, 786–789. [Google Scholar] [CrossRef]
- Kim, S.J.; Kim, J.S.; Park, E.S.; Lee, J.S.; Lin, Q.; Langley, R.R.; Maya, M.; He, J.; Kim, S.W.; Weihua, Z.; et al. Astrocytes up-regulate survival genes in tumor cells and induce protection from chemotherapy. Neoplasia 2011, 13, 286–298. [Google Scholar]
- Langley, R.R.; Fan, D.; Guo, L.; Zhang, C.; Lin, Q.; Brantley, E.C.; McCarty, J.H.; Fidler, I.J. Generation of an immortalized astrocyte cell line from H-2Kb-tsA58 mice to study the role of astrocytes in brain metastasis. Int. J. Oncol. 2009, 35, 665–672. [Google Scholar]
- JuanYin, J.; Tracy, K.; Zhang, L.; Munasinghe, J.; Shapiro, E.; Koretsky, A.; Kelly, K. Noninvasive imaging of the functional effects of anti-VEGF therapy on tumor cell extravasation and regional blood volume in an experimental brain metastasis model. Clin. Exp. Metastasis 2009, 26, 403–414. [Google Scholar] [CrossRef]
- Lorger, M.; Felding-Habermann, B. Capturing changes in the brain microenvironment during initial steps of breast cancer brain metastasis. Am. J. Pathol. 2010, 176, 2958–2971. [Google Scholar] [CrossRef]
- Deeken, J.F.; Loscher, W. The blood–brain barrier and cancer: Transporters, treatment, and Trojan horses. Clin. Cancer Res. 2007, 13, 1663–1674. [Google Scholar] [CrossRef]
- Fukumura, D.; Xu, L.; Chen, Y.; Gohongi, T.; Seed, B.; Jain, R.K. Hypoxia and acidosis independently up-regulate vascular endothelial growth factor transcription in brain tumors in vivo. Cancer Res. 2001, 61, 6020–6024. [Google Scholar]
- Hobbs, S.K.; Monsky, W.L.; Yuan, F.; Roberts, W.G.; Griffith, L.; Torchilin, V.P.; Jain, R.K. Regulation of transport pathways in tumor vessels: Role of tumor type and microenvironment. Proc. Natl. Acad. Sci. USA 1998, 95, 4607–4612. [Google Scholar]
- Sharom, F.J. Complex interplay between the p-glycoprotein multidrug efflux pump and the membrane: Its role in modulating protein function. Front. Oncol. 2014, 4, 41. [Google Scholar]
- Hida, K.; Akiyama, K.; Ohga, N.; Maishi, N.; Hida, Y. Tumour endothelial cells acquire drug resistance in a tumour microenvironment. J. Biochem. 2013, 153, 243–249. [Google Scholar] [CrossRef]
- Hanahan, D.; Weinberg, R.A. Hallmarks of cancer: The next generation. Cell 2011, 144, 646–674. [Google Scholar] [CrossRef]
- Melo, S.A.; Esteller, M. Dysregulation of microRNAs in cancer: Playing with fire. FEBS Lett. 2011, 585, 2087–2099. [Google Scholar] [CrossRef]
- Calin, G.A.; Croce, C.M. MicroRNA signatures in human cancers. Nat. Rev. Cancer 2006, 6, 857–866. [Google Scholar] [CrossRef]
- Aigner, A. MicroRNAs (miRNAs) in cancer invasion and metastasis: Therapeutic approaches based on metastasis-related miRNAs. J. Mol. Med. 2011, 89, 445–457. [Google Scholar] [CrossRef]
- Calin, G.A.; Sevignani, C.; Dumitru, C.D.; Hyslop, T.; Noch, E.; Yendamuri, S.; Shimizu, M.; Rattan, S.; Bullrich, F.; Negrini, M. Human microRNA genes are frequently located at fragile sites and genomic regions involved in cancers. Proc. Natl. Acad. Sci. USA 2004, 101, 2999–3004. [Google Scholar] [CrossRef]
- Janssen, E.A.; Slewa, A.; Gudlaugsson, E.; Jonsdottir, K.; Skaland, I.; Soiland, H.; Baak, J.P. Biologic profiling of lymph node negative breast cancers by means of microRNA expression. Mod. Pathol. 2010, 23, 1567–1576. [Google Scholar] [CrossRef]
- Cheng, H.; Zhang, L.; Cogdell, D.E.; Zheng, H.; Schetter, A.J.; Nykter, M.; Harris, C.C.; Chen, K.; Hamilton, S.R.; Zhang, W. Circulating plasma miR-141 is a novel biomarker for metastatic colon cancer and predicts poor prognosis. PLoS One 2011, 6, e17745. [Google Scholar]
- Zhu, L.; Chen, H.; Zhou, D.; Li, D.; Bai, R.; Zheng, S.; Ge, W. MicroRNA-9 up-regulation is involved in colorectal cancer metastasis via promoting cell motility. Med. Oncol. 2012, 29, 1037–1043. [Google Scholar] [CrossRef]
- Ji, Q.; Hao, X.; Zhang, M.; Tang, W.; Yang, M.; Li, L.; Xiang, D.; Desano, J.T.; Bommer, G.T.; Fan, D.; et al. MicroRNA miR-34 inhibits human pancreatic cancer tumor-initiating cells. PLoS One 2009, 4, e6816. [Google Scholar] [CrossRef]
- Zimmerman, A.L.; Wu, S. MicroRNAs, cancer and cancer stem cells. Cancer Lett. 2011, 300, 10–19. [Google Scholar] [CrossRef]
- Mueller, W.C.; Spector, Y.; Edmonston, T.B.; Cyr, B.S.; Jaeger, D.; Lass, U.; Aharonov, R.; Rosenwald, S.; Chajut, A. Accurate classification of metastatic brain tumors using a novel microRNA-based test. Oncologist 2011, 16, 165–174. [Google Scholar] [CrossRef]
- Nass, D.; Rosenwald, S.; Meiri, E.; Gilad, S.; Tabibian-Keissar, H.; Schlosberg, A.; Kuker, H.; Sion-Vardy, N.; Tobar, A.; Kharenko, O.; et al. MiR-92b and miR-9/9* are specifically expressed in brain primary tumors and can be used to differentiate primary from metastatic brain tumors. Brain Pathol. 2009, 19, 375–383. [Google Scholar] [CrossRef]
- Zhang, X.; Yu, H.; Lou, J.R.; Zheng, J.; Zhu, H.; Popescu, N.I.; Lupu, F.; Lind, S.E.; Ding, W.Q. MicroRNA-19 (miR-19) regulates tissue factor expression in breast cancer cells. J. Biol. Chem. 2011, 286, 1429–1435. [Google Scholar]
- Mott, J.L.; Kobayashi, S.; Bronk, S.F.; Gores, G.J. Mir-29 regulates Mcl-1 protein expression and apoptosis. Oncogene 2007, 26, 6133–6140. [Google Scholar] [CrossRef]
- Creighton, C.J.; Fountain, M.D.; Yu, Z.; Nagaraja, A.K.; Zhu, H.; Khan, M.; Olokpa, E.; Zariff, A.; Gunaratne, P.H.; Matzuk, M.M.; et al. Molecular profiling uncovers a p53-associated role for microRNA-31 in inhibiting the proliferation of serous ovarian carcinomas and other cancers. Cancer Res. 2010, 70, 1906–1915. [Google Scholar] [CrossRef]
- Gregory, P.A.; Bert, A.G.; Paterson, E.L.; Barry, S.C.; Tsykin, A.; Farshid, G.; Vadas, M.A.; Khew-Goodall, Y.; Goodall, G.J. The miR-200 family and miR-205 regulate epithelial to mesenchymal transition by targeting ZEB1 and SIP1. Nat. Cell Biol. 2008, 10, 593–601. [Google Scholar] [CrossRef]
- Li, L.; Huang, K.; You, Y.; Fu, X.; Hu, L.; Song, L.; Meng, Y. Hypoxia-induced miR-210 in epithelial ovarian cancer enhances cancer cell viability via promoting proliferation and inhibiting apoptosis. Int. J. Oncol. 2014, 44, 2111–2120. [Google Scholar]
- Zhang, L.; Sullivan, P.S.; Goodman, J.C.; Gunaratne, P.H.; Marchetti, D. MicroRNA-1258 suppresses breast cancer brain metastasis by targeting heparanase. Cancer Res. 2011, 71, 645–654. [Google Scholar]
- Ridgway, L.D.; Wetzel, M.D.; Ngo, J.A.; Erdreich-Epstein, A.; Marchetti, D. Heparanase-induced GEF-H1 signaling regulates the cytoskeletal dynamics of brain metastatic breast cancer cells. Mol. Cancer Res. 2012, 10, 689–702. [Google Scholar] [CrossRef]
- Okuda, H.; Xing, F.; Pandey, P.R.; Sharma, S.; Watabe, M.; Pai, S.K.; Mo, Y.Y.; Iiizumi-Gairani, M.; Hirota, S.; Liu, Y.; et al. miR-7 suppresses brain metastasis of breast cancer stem-like cells by modulating KLF4. Cancer Res. 2013, 73, 1434–1444. [Google Scholar] [CrossRef]
- Zhao, C.; Xu, Y.; Zhang, Y.; Tan, W.; Xue, J.; Yang, Z.; Zhang, Y.; Lu, Y.; Hu, X. Down-regulationof miR-145 contributes to lung adenocarcinoma cell growth to form brain metastases. Oncol. Rep. 2013, 30, 2027–2034. [Google Scholar]
- Arora, S.; Ranade, A.R.; Tran, N.L.; Nasser, S.; Sridhar, S.; Korn, R.L.; Ross, J.T.; Dhruv, H.; Foss, K.M.; Sibenaller, Z.; et al. MicroRNA-328 is associated with (non-small) cell lung cancer (NSCLC) brain metastasis and mediates NSCLC migration. Int. J. Cancer 2011, 129, 2621–2631. [Google Scholar] [CrossRef]
- Chen, L.T.; Xu, S.D.; Xu, H.; Zhang, J.F.; Ning, J.F.; Wang, S.F. MicroRNA-378 is associated with non-small cell lung cancer brain metastasis by promoting cell migration, invasion and tumor angiogenesis. Med. Oncol. 2012, 29, 1673–1680. [Google Scholar] [CrossRef]
- Teplyuk, N.M.; Mollenhauer, B.; Gabriely, G.; Giese, A.; Kim, E.; Smolsky, M.; Kim, R.Y.; Saria, M.G.; Pastorino, S.; Kesari, S.; et al. MicroRNAs in cerebrospinal fluid identify glioblastoma and metastatic brain cancers and reflect disease activity. Neuro-Oncol. 2012, 14, 689–700. [Google Scholar] [CrossRef]
- Hwang, S.J.; Seol, H.J.; Park, Y.M.; Kim, K.H.; Gorospe, M.; Nam, D.H.; Kim, H.H. MicroRNA-146a suppresses metastatic activity in brain metastasis. Mol. Cells 2012, 34, 329–334. [Google Scholar] [CrossRef]
- Subramani, A.; Alsidawi, S.; Jagannathan, S.; Sumita, K.; Sasaki, A.T.; Aronow, B.; Warnick, R.E.; Lawler, S.; Driscoll, J.J. The brain microenvironment negatively regulates miRNA-768-3p to promote K-ras expression and lung cancer metastasis. Sci. Rep. 2013, 3, 2392. [Google Scholar]
- Camacho, L.; Guerrero, P.; Marchetti, D. MicroRNA and protein profiling of brain metastasis competent cell-derived exosomes. PLoS One 2013, 8, e73790. [Google Scholar] [CrossRef]
- Li, Z.; Gu, X.; Fang, Y.; Xiang, J.; Chen, Z. MicroRNA expression profiles in human colorectal cancers with brain metastases. Oncol. Lett. 2012, 3, 346–350. [Google Scholar]
- Grupenmacher, A.T.; Halpern, A.L.; Bonaldo Mde, F.; Huang, C.C.; Hamm, C.A.; de Andrade, A.; Tomita, T.; Sredni, S.T. Study of the gene expression and microRNA expression profiles of malignant rhabdoid tumors originated in the brain (AT/RT) and in the kidney (RTK). Childs Nerv. Syst. 2013, 29, 1977–1983. [Google Scholar] [CrossRef]
- Lu, Y.; Govindan, R.; Wang, L.; Liu, P.Y.; Goodgame, B.; Wen, W.; Sezhiyan, A.; Pfeifer, J.; Li, Y.F.; Hua, X.; et al. MicroRNA profiling and prediction of recurrence/relapse-free survival in stage I lung cancer. Carcinogenesis 2012, 33, 1046–1054. [Google Scholar] [CrossRef]
- Ahmad, N.; Haider, S.; Jagannathan, S.; Anaissie, E.; Driscoll, J. MicroRNA theragnostics for the clinical management of multiple myeloma. Leukemia 2014, 28, 732–738. [Google Scholar] [CrossRef]
- Mitchell, P.S.; Parkin, R.K.; Kroh, E.M.; Fritz, B.R.; Wyman, S.K.; Pogosova-Agadjanyan, E.L.; Peterson, A.; Noteboom, J.; O’Briant, K.C.; Allen, A.; et al. Circulating microRNAs as stable blood-based markers for cancer detection. Proc. Natl. Acad. Sci. USA 2008, 105, 10513–10518. [Google Scholar] [CrossRef]
- Jones, C.I.; Zabolotskaya, M.V.; King, A.J.; Stewart, H.J.; Horne, G.A.; Chevassut, T.J.; Newbury, S.F. Identification of circulating microRNAs as diagnostic biomarkers for use in multiple myeloma. Br. J. Cancer 2012, 107, 1987–1996. [Google Scholar] [CrossRef]
- Vlassov, A.V.; Magdaleno, S.; Setterquist, R.; Conrad, R. Exosomes: Current knowledge of their composition, biological functions, and diagnostic and therapeutic potentials. Biochim. Biophys. Acta 2012, 1820, 940–948. [Google Scholar] [CrossRef]
- Cazzoli, R.; Buttitta, F.; di Nicola, M.; Malatesta, S.; Marchetti, A.; Rom, W.N.; Pass, H.I. MicroRNAs derived from circulating exosomes as noninvasive biomarkers for screening and diagnosing lung cancer. J. Thorac. Oncol. 2013, 8, 1156–1162. [Google Scholar] [CrossRef]
- Manterola, L.; Guruceaga, E.; Pérez-Larraya, J.G.; González-Huarriz, M.; Jauregui, P.; Tejada, S.; Diez-Valle, R.; Segura, V.; Samprón, N.; Barrena, C. A small noncoding RNA signature found in exosomes of GBM patient serum as a diagnostic tool. Neuro-Oncol. 2014. [Google Scholar] [CrossRef]
- Zomer, A.; Vendrig, T.; Hopmans, E.S.; van Eijndhoven, M.; Middeldorp, J.M.; Pegtel, D.M. Exosomes: Fit to deliver small RNA. Commun. Integr. Biol. 2010, 3, 447–450. [Google Scholar] [CrossRef]
- Lawrie, C.H.; Gal, S.; Dunlop, H.M.; Pushkaran, B.; Liggins, A.P.; Pulford, K.; Banham, A.H.; Pezzella, F.; Boultwood, J.; Wainscoat, J.S.; et al. Detection of elevated levels of tumour-associated microRNAs in serum of patients with diffuse large B-cell lymphoma. Br. J. Haematol. 2008, 141, 672–675. [Google Scholar]
- Tanaka, M.; Oikawa, K.; Takanashi, M.; Kudo, M.; Ohyashiki, J.; Ohyashiki, K.; Kuroda, M. Down-regulation of miR-92 in human plasma is a novel marker for acute leukemia patients. PLoS One 2009, 4, e5532. [Google Scholar]
- Fayyad-Kazan, H.; Bitar, N.; Najar, M.; Lewalle, P.; Fayyad-Kazan, M.; Badran, R.; Hamade, E.; Daher, A.; Hussein, N.; ElDirani, R.; et al. Circulating miR-150 and miR-342 in plasma are novel potential biomarkers for acute myeloid leukemia. J. Transl. Med. 2013, 11, 31. [Google Scholar] [CrossRef]
- Huang, Z.; Huang, D.; Ni, S.; Peng, Z.; Sheng, W.; Du, X. Plasma microRNAs are promising novel biomarkers for early detection of colorectal cancer. Int. J. Cancer 2010, 127, 118–126. [Google Scholar] [CrossRef]
- Heneghan, H.M.; Miller, N.; Lowery, A.J.; Sweeney, K.J.; Newell, J.; Kerin, M.J. Circulating microRNAs as novel minimally invasive biomarkers for breast cancer. Ann. Surg. 2010, 251, 499–505. [Google Scholar] [CrossRef]
- Si, H.; Sun, X.; Chen, Y.; Cao, Y.; Chen, S.; Wang, H.; Hu, C. Circulating microRNA-92a and microRNA-21 as novel minimally invasive biomarkers for primary breast cancer. J. Cancer Res. Clin. Oncol. 2013, 139, 223–229. [Google Scholar] [CrossRef]
- Brase, J.C.; Johannes, M.; Schlomm, T.; Falth, M.; Haese, A.; Steuber, T.; Beissbarth, T.; Kuner, R.; Sultmann, H. Circulating miRNAs are correlated with tumor progression in prostate cancer. Int. J. Cancer 2011, 128, 608–616. [Google Scholar] [CrossRef]
- Resnick, K.E.; Alder, H.; Hagan, J.P.; Richardson, D.L.; Croce, C.M.; Cohn, D.E. The detection of differentially expressed microRNAs from the serum of ovarian cancer patients using a novel real-time PCR platform. Gynecol. Oncol. 2009, 112, 55–59. [Google Scholar]
- Wang, J.; Chen, J.; Chang, P.; LeBlanc, A.; Li, D.; Abbruzzesse, J.L.; Frazier, M.L.; Killary, A.M.; Sen, S. MicroRNAs in plasma of pancreatic ductal adenocarcinoma patients as novel blood-based biomarkers of disease. Cancer Prev. Res. 2009, 2, 807–813. [Google Scholar] [CrossRef]
- Ganepola, G.A.; Rutledge, J.R.; Suman, P.; Yiengpruksawan, A.; Chang, D.H. Novel blood-basedmicroRNA biomarker panel for early diagnosis of pancreatic cancer. World J. Gastrointest. Oncol. 2014, 6, 22–33. [Google Scholar] [CrossRef]
- Gao, L.; He, S.B.; Li, D.C. Effects of miR-16 plus CA19-9 detections on pancreatic cancer diagnostic performance. Clin. Lab. 2014, 60, 73–77. [Google Scholar]
- Schultz, N.A.; Dehlendorff, C.; Jensen, B.V.; Bjerregaard, J.K.; Nielsen, K.R.; Bojesen, S.E.; Calatayud, D.; Nielsen, S.E.; Yilmaz, M.; Hollander, N.H.; et al. MicroRNA biomarkers in whole blood for detection of pancreatic cancer. JAMA 2014, 311, 392–404. [Google Scholar] [CrossRef]
- Tsujiura, M.; Ichikawa, D.; Komatsu, S.; Shiozaki, A.; Takeshita, H.; Kosuga, T.; Konishi, H.; Morimura, R.; Deguchi, K.; Fujiwara, H.; et al. Circulating microRNAs in plasma of patients with gastric cancers. Br. J. Cancer 2010, 102, 1174–1179. [Google Scholar] [CrossRef]
- Ma, G.J.; Gu, R.M.; Zhu, M.; Wen, X.; Li, J.T.; Zhang, Y.Y.; Zhang, X.M.; Chen, S.Q. Plasma post-operative miR-21 expression in the prognosis of gastric cancers. Asian Pac. J. Cancer Prev. 2013, 14, 7551–7554. [Google Scholar] [CrossRef]
- Tsujiura, M.; Komatsu, S.; Ichikawa, D.; Shiozaki, A.; Konishi, H.; Takeshita, H.; Moriumura, R.; Nagata, H.; Kawaguchi, T.; Hirajima, S.; et al. Circulating miR-18a in plasma contributes to cancer detection and monitoring in patients with gastric cancer. Gastric Cancer 2014. [Google Scholar] [CrossRef]
- Hu, Z.; Chen, X.; Zhao, Y.; Tian, T.; Jin, G.; Shu, Y.; Chen, Y.; Xu, L.; Zen, K.; Zhang, C.; et al. Serum microRNA signatures identified in a genome-wide serum microRNA expression profiling predict survival of non-small-cell lung cancer. J. Clin. Oncol. 2010, 28, 1721–1726. [Google Scholar] [CrossRef]
- Yu, H.; Jiang, L.; Sun, C.; Guo, L.; Lin, M.; Huang, J.; Zhu, L. Decreased circulating miR-375: A potential biomarker for patients with non-small-cell lung cancer. Gene 2014, 534, 60–65. [Google Scholar] [CrossRef]
- Pacifici, M.; Delbue, S.; Kadri, F.; Peruzzi, F. Cerebrospinal fluid microRNA profiling using quantitative real time PCR. J. Vis. Exp. 2014, 83, e51172. [Google Scholar]
- Sheinerman, K.S.; Umansky, S.R. Circulating cell-free microRNA as biomarkers for screening, diagnosis and monitoring of neurodegenerative diseases and other neurologic pathologies. Front. Cell. Neurosci. 2013, 7, 150. [Google Scholar]
- D’Asti, E.; Garnier, D.; Lee, T.H.; Montermini, L.; Meehan, B.; Rak, J. Oncogenic extracellular vesicles in brain tumor progression. Front. Physiol. 2012, 3, 294. [Google Scholar]
- Zhang, Y.; Wang, Z.; Gemeinhart, R.A. Progress in microRNA delivery. J. Control. Release 2013, 172, 962–974. [Google Scholar] [CrossRef]
- Garzon, R.; Marcucci, G.; Croce, C.M. Targeting microRNAs in cancer: Rationale, strategies and challenges. Nat. Rev. Drug Discov. 2010, 9, 775–789. [Google Scholar] [CrossRef]
- Corsten, M.F.; Miranda, R.; Kasmieh, R.; Krichevsky, A.M.; Weissleder, R.; Shah, K. MicroRNA-21 knockdown disrupts glioma growth in vivo and displays synergistic cytotoxicity with neural precursor cell delivered S-TRAIL in human gliomas. Cancer Res. 2007, 67, 8994–9000. [Google Scholar] [CrossRef]
- Cook, P.D. Medicinal chemistry of antisense oligonucleotides—Future opportunities. Anti-Cancer Drug Des. 1991, 6, 585–607. [Google Scholar]
- Crooke, S.T.; Graham, M.J.; Zuckerman, J.E.; Brooks, D.; Conklin, B.S.; Cummins, L.L.; Greig, M.J.; Guinosso, C.J.; Kornbrust, D.; Manoharan, M.; et al. Pharmacokinetic properties of several novel oligonucleotide analogs in mice. J. Pharmacol. Exp. Ther. 1996, 277, 923–937. [Google Scholar]
- Yoo, B.H.; Bochkareva, E.; Bochkarev, A.; Mou, T.C.; Gray, D.M. 2'-O-Methyl-modified phosphorothioate antisense oligonucleotides have reduced non-specific effects in vitro. Nucleic Acids Res. 2004, 32, 2008–2016. [Google Scholar] [CrossRef]
- Wahlestedt, C.; Salmi, P.; Good, L.; Kela, J.; Johnsson, T.; Hokfelt, T.; Broberger, C.; Porreca, F.; Lai, J.; Ren, K.; et al. Potent and nontoxic antisense oligonucleotides containing locked nucleic acids. Proc. Natl. Acad. Sci. USA 2000, 97, 5633–5638. [Google Scholar] [CrossRef]
- Vester, B.; Wengel, J. LNA (locked nucleic acid): High-affinity targeting of complementary RNA and DNA. Biochemistry 2004, 43, 13233–13241. [Google Scholar] [CrossRef]
- Zhilina, Z.V.; Ziemba, A.J.; Ebbinghaus, S.W. Peptide nucleic acid conjugates: Synthesis, properties and applications. Curr. Top. Med. Chem. 2005, 5, 1119–1131. [Google Scholar] [CrossRef]
- Pallan, P.S.; Greene, E.M.; Jicman, P.A.; Pandey, R.K.; Manoharan, M.; Rozners, E.; Egli, M. Unexpected origins of the enhanced pairing affinity of 2'-fluoro-modified RNA. Nucleic Acids Res. 2011, 39, 3482–3495. [Google Scholar]
- Ebert, M.S.; Neilson, J.R.; Sharp, P.A. MicroRNA sponges: Competitive inhibitors of small RNAs in mammalian cells. Nat. Methods 2007, 4, 721–726. [Google Scholar] [CrossRef]
- Choi, W.Y.; Giraldez, A.J.; Schier, A.F. Target protectors reveal dampening and balancing of Nodal agonist and antagonist by miR-430. Science 2007, 318, 271–274. [Google Scholar] [CrossRef]
- Terasawa, K.; Shimizu, K.; Tsujimoto, G. Synthetic pre-miRNA-based shRNA as potent RNAi triggers. J. Nucleic Acids 2011, 2011, 131579. [Google Scholar]
- Bonci, D.; Coppola, V.; Musumeci, M.; Addario, A.; Giuffrida, R.; Memeo, L.; D’Urso, L.; Pagliuca, A.; Biffoni, M.; Labbaye, C. The miR-15a–miR-16-1 cluster controls prostate cancer by targeting multiple oncogenic activities. Nat. Med. 2008, 14, 1271–1277. [Google Scholar] [CrossRef]
- Xiong, Y.; Fang, J.H.; Yun, J.P.; Yang, J.; Zhang, Y.; Jia, W.H.; Zhuang, S.M. Effects of microRNA-29 on apoptosis, tumorigenicity, and prognosis of hepatocellular carcinoma. Hepatology 2010, 51, 836–845. [Google Scholar]
- Garzon, R.; Heaphy, C.E.; Havelange, V.; Fabbri, M.; Volinia, S.; Tsao, T.; Zanesi, N.; Kornblau, S.M.; Marcucci, G.; Calin, G.A. MicroRNA 29b functions in acute myeloid leukemia. Blood 2009, 114, 5331–5341. [Google Scholar] [CrossRef]
- Michelfelder, S.; Trepel, M. Adeno-associated viral vectors and their redirection to cell-type specific receptors. Adv. Genet. 2009, 67, 29–60. [Google Scholar] [CrossRef]
- Kota, J.; Chivukula, R.R.; O’Donnell, K.A.; Wentzel, E.A.; Montgomery, C.L.; Hwang, H.-W.; Chang, T.-C.; Vivekanandan, P.; Torbenson, M.; Clark, K.R. Therapeutic microRNA delivery suppresses tumorigenesis in a murine liver cancer model. Cell 2009, 137, 1005–1017. [Google Scholar] [CrossRef]
- Wu, Y.; Crawford, M.; Yu, B.; Mao, Y.; Nana-Sinkam, S.P.; Lee, L.J. MicroRNA delivery by cationic lipoplexes for lung cancer therapy. Mol. Pharm. 2011, 8, 1381–1389. [Google Scholar] [CrossRef]
- Lv, H.; Zhang, S.; Wang, B.; Cui, S.; Yan, J. Toxicity of cationic lipids and cationic polymers in gene delivery. J. Control. Release 2006, 114, 100–109. [Google Scholar] [CrossRef]
- Chen, Y.; Zhu, X.; Zhang, X.; Liu, B.; Huang, L. Nanoparticles modified with tumor-targeting scFv deliver siRNA and miRNA for cancer therapy. Mol. Ther. 2010, 18, 1650–1656. [Google Scholar] [CrossRef]
- Trang, P.; Wiggins, J.F.; Daige, C.L.; Cho, C.; Omotola, M.; Brown, D.; Weidhaas, J.B.; Bader, A.G.; Slack, F.J. Systemic delivery of tumor suppressor microRNA mimics using a neutral lipid emulsion inhibits lung tumors in mice. Mol. Ther. 2011, 19, 1116–1122. [Google Scholar] [CrossRef]
- Chirila, T.V.; Rakoczy, P.E.; Garrett, K.L.; Lou, X.; Constable, I.J. The use of synthetic polymers for delivery of therapeutic antisense oligodeoxynucleotides. Biomaterials 2002, 23, 321–342. [Google Scholar] [CrossRef]
- Ren, Y.; Kang, C.S.; Yuan, X.B.; Zhou, X.; Xu, P.; Han, L.; Wang, G.X.; Jia, Z.; Zhong, Y.; Yu, S.; et al. Co-delivery of as-miR-21 and 5-FU by poly(amidoamine) dendrimer attenuates human glioma cell growth in vitro. J. Biomater. Sci. 2010, 21, 303–314. [Google Scholar] [CrossRef]
- Zhao, X.; Pan, F.; Holt, C.M.; Lewis, A.L.; Lu, J.R. Controlled delivery of antisense oligonucleotides: A brief review of current strategies. Expert Opin. Drug Deliv. 2009, 6, 673–686. [Google Scholar] [CrossRef]
- Babar, I.A.; Cheng, C.J.; Booth, C.J.; Liang, X.; Weidhaas, J.B.; Saltzman, W.M.; Slack, F.J. Nanoparticle-based therapy in an in vivo microRNA-155 (miR-155)-dependent mouse model of lymphoma. Proc. Natl. Acad. Sci. USA 2012, 109, E1695–E1704. [Google Scholar]
- Pardridge, W.M. Drug and gene delivery to the brain: The vascular route. Neuron 2002, 36, 555–558. [Google Scholar] [CrossRef]
- Boado, R.J. Blood–brain barrier transport of non-viral gene and RNAi therapeutics. Pharm. Res. 2007, 24, 1772–1787. [Google Scholar] [CrossRef]
- Zhang, Y.; Zhang, Y.F.; Bryant, J.; Charles, A.; Boado, R.J.; Pardridge, W.M. Intravenous RNA interference gene therapy targeting the human epidermal growth factor receptor prolongs survival in intracranial brain cancer. Clin. Cancer Res. 2004, 10, 3667–3677. [Google Scholar] [CrossRef]
- Boussif, O.; Lezoualc’h, F.; Zanta, M.A.; Mergny, M.D.; Scherman, D.; Demeneix, B.; Behr, J.-P. A versatile vector for gene and oligonucleotide transfer into cells in culture and in vivo: polyethylenimine. Proc. Natl. Acad. Sci. USA 1995, 92, 7297–7301. [Google Scholar]
- Hornung, V.; Guenthner-Biller, M.; Bourquin, C.; Ablasser, A.; Schlee, M.; Uematsu, S.; Noronha, A.; Manoharan, M.; Akira, S.; de Fougerolles, A. Sequence-specific potent induction of IFN-α by short interfering RNA in plasmacytoid dendritic cells through TLR7. Nat. Med. 2005, 11, 263–270. [Google Scholar] [CrossRef]
- Hwang, D.W.; Son, S.; Jang, J.; Youn, H.; Lee, S.; Lee, D.; Lee, Y.-S.; Jeong, J.M.; Kim, W.J.; Lee, D.S. A brain-targeted rabies virus glycoprotein-disulfide linked PEI nanocarrier for delivery of neurogenic microRNA. Biomaterials 2011, 32, 4968–4975. [Google Scholar] [CrossRef]
- Grimm, D.; Streetz, K.L.; Jopling, C.L.; Storm, T.A.; Pandey, K.; Davis, C.R.; Marion, P.; Salazar, F.; Kay, M.A. Fatality in mice due to oversaturation of cellular microRNA/short hairpin RNA pathways. Nature 2006, 441, 537–541. [Google Scholar] [CrossRef]
© 2014 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Alsidawi, S.; Malek, E.; Driscoll, J.J. MicroRNAs in Brain Metastases: Potential Role as Diagnostics and Therapeutics. Int. J. Mol. Sci. 2014, 15, 10508-10526. https://doi.org/10.3390/ijms150610508
Alsidawi S, Malek E, Driscoll JJ. MicroRNAs in Brain Metastases: Potential Role as Diagnostics and Therapeutics. International Journal of Molecular Sciences. 2014; 15(6):10508-10526. https://doi.org/10.3390/ijms150610508
Chicago/Turabian StyleAlsidawi, Samer, Ehsan Malek, and James J. Driscoll. 2014. "MicroRNAs in Brain Metastases: Potential Role as Diagnostics and Therapeutics" International Journal of Molecular Sciences 15, no. 6: 10508-10526. https://doi.org/10.3390/ijms150610508





