Comprehensive Analysis of Copy Number Variations on Glycoside Hydrolase 45 Genes among Different Bursaphelenchus xylophilus Strains
Abstract
:1. Introduction
2. Results
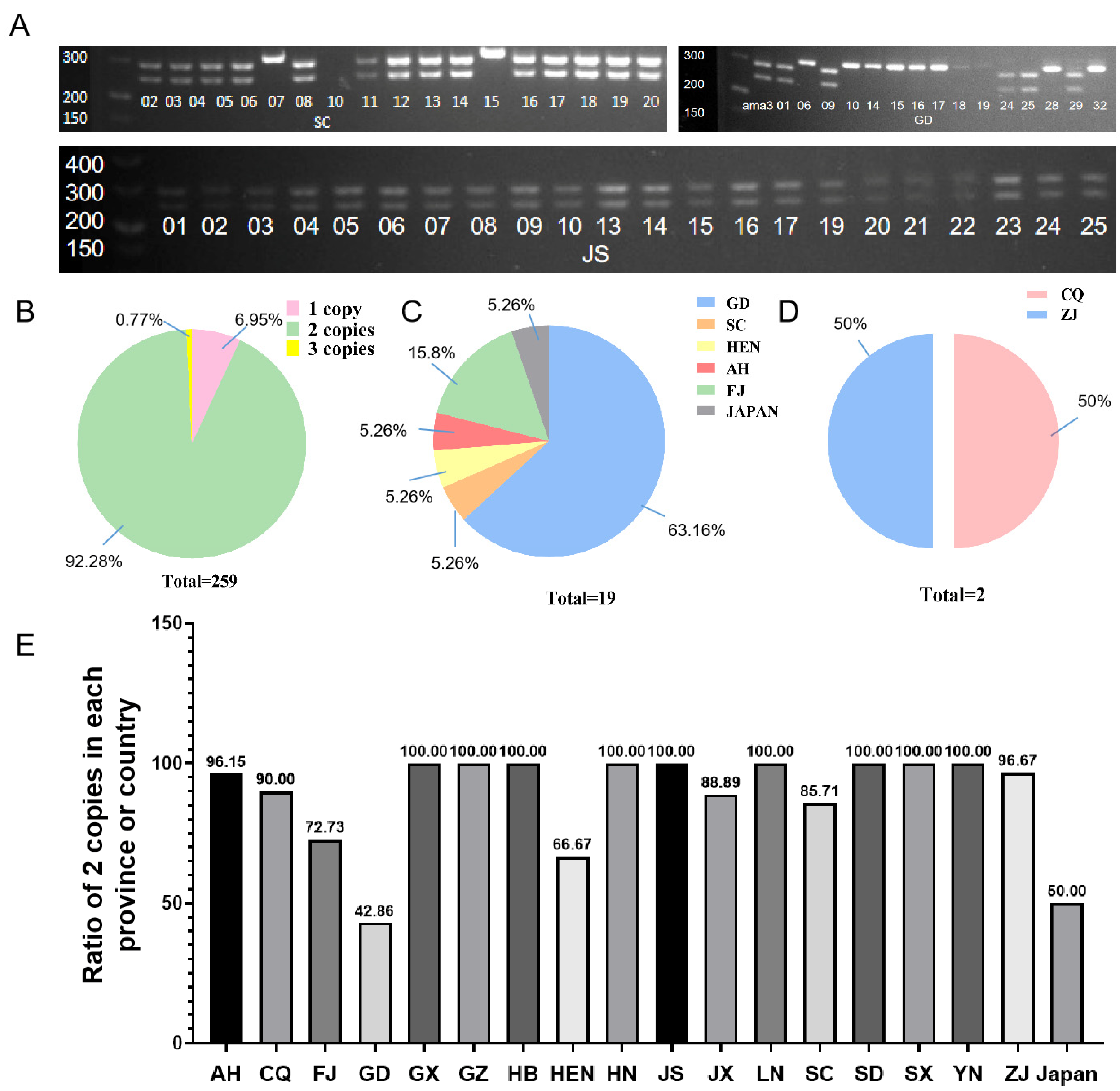
2.1. CNV of GH45 Gene Identified among Different B. xylophilus Strains
2.2. In Vitro Quantification of CNVs in GH45
2.3. Post-Inoculation Analysis of GH45 CNVs among Different B. xylophilus Strains
3. Discussion
4. Materials and Methods
4.1. Collection of Bursaphelenchus xylophilus Strains
4.2. Identification of CNV on Glycoside Hydrolase 45
4.3. Inoculation Test and Disease Severity Analysis
4.4. Quantification of GH45 Gene with Different Copies
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Mamiya, Y. Pathology of the pine wilt disease caused by Bursaphelenchus xylophilus. Ann. Rev. Phytopathol. 1983, 21, 201–220. [Google Scholar] [CrossRef]
- Soliman, T.; Mourits, M.C.M.; van der Werf, W.; Hengeveld, G.M.; Robinet, C.; Lansink, A.G.J.M.O. Framework for modelling economic impacts of invasive species, applied to pine wood nematode in Europe. PLoS ONE 2012, 7, e45505. [Google Scholar] [CrossRef] [Green Version]
- Kikuchi, T.; Shibuya, H.; Aikawa, T.; Jones, J.T. Cloning and characterization of pectate lyases expressed in the esophageal gland of the pine wood nematode Bursaphelenchus xylophilus. Mol. Plant-Microbe Interact. 2006, 19, 280–287. [Google Scholar] [CrossRef] [Green Version]
- Kang, J.S.; Lee, H.; Moon, I.S.; Lee, Y.; Koh, Y.H.; Je, Y.H.; Lim, K.J.; Lee, S.H. Construction and characterization of subtractive stage-specific expressed sequence tag (EST) libraries of the pinewood nematode Bursaphelenchus xylophilus. Genomics 2009, 94, 70–77. [Google Scholar] [CrossRef] [Green Version]
- Hirao, T.; Fukatsu, E.; Watanabe, A.J.B.P.B. Characterization of resistance to pine wood nematode infection in Pinus thunbergii using suppression subtractive hybridization. BMC Plant Biol. 2012, 12, 13. [Google Scholar] [CrossRef] [Green Version]
- Kikuchi, T.; Cotton, J.A.; Dalzell, J.J.; Hasegawa, K.; Kanzaki, N.; McVeigh, P.; Takanashi, T.; Tsai, I.J.; Assefa, S.A.; Cock, P.J.A.; et al. Genomic insights into the origin of parasitism in the emerging plant pathogen Bursaphelenchus xylophilus. PLoS Pathog. 2011, 7, e1002219. [Google Scholar] [CrossRef] [Green Version]
- Ding, X.L.; Guo, Y.F.; Ye, J.R.; Wu, X.Q.; Lin, S.X.; Chen, F.M.; Zhu, L.H.; Huang, L.; Song, X.F.; Zhang, Y.; et al. Population differentiation and epidemic tracking of Bursaphelenchus xylophilus in China based on chromosome-level assembly and whole-genome sequencing data. Pest. Manag. Sci. 2022, 78, 1213–1226. [Google Scholar] [CrossRef]
- Figueiredo, J.; Simoes, M.J.; Gomes, P.; Barroso, C.; Pinho, D.; Conceicao, L.; Fonseca, L.; Abrantes, I.; Pinheiro, M.; Egas, C. Assessment of the geographic origins of pinewood nematode isolates via single nucleotide polymorphism in effector genes. PLoS ONE 2013, 8, e83542. [Google Scholar] [CrossRef] [Green Version]
- Palomares-Rius, J.E.; Tsai, I.J.; Karim, N.; Akiba, M.; Kato, T.; Maruyama, H.; Takeuchi, Y.; Kikuchi, T. Genome-wide variation in the pinewood nematode Bursaphelenchus xylophilus and its relationship with pathogenic traits. BMC Genom. 2015, 16, 845. [Google Scholar] [CrossRef] [Green Version]
- Tsai, I.J.; Tanaka, R.; Kanzaki, N.; Akiba, M.; Yokoi, T.; Espada, M.; Jones, J.T.; Kikuchi, T. Transcriptional and morphological changes in the transition from mycetophagous to phytophagous phase in the plant-parasitic nematode Bursaphelenchus xylophilus. Mol. Plant Pathol. 2016, 17, 77–83. [Google Scholar] [CrossRef]
- Li, Y.X.; Meng, F.L.; Deng, X.; Wang, X.; Feng, Y.Q.; Zhang, W.; Pan, L.; Zhang, X.Y. Comparative transcriptome analysis of the pinewood nematode Bursaphelenchus xylophilus reveals the molecular mechanism underlying its defense response to host-derived -pinene. Int. J. Mol. Sci. 2019, 20, 911. [Google Scholar] [CrossRef] [Green Version]
- Ding, X.L.; Ye, J.R.; Wu, X.Q.; Huang, L.; Zhu, L.H.; Lin, S.X. Deep sequencing analyses of pine wood nematode Bursaphelenchus xylophilus microRNAs reveal distinct miRNA expression patterns during the pathological process of pine wilt disease. Gene 2015, 555, 346–356. [Google Scholar] [CrossRef]
- Ding, X.; Ye, J.; Lin, S.; Wu, X.; Li, D.; Nian, B. Deciphering the molecular variations of pine wood nematode Bursaphelenchus xylophilus with different virulence. PLoS ONE 2016, 11, e0156040. [Google Scholar] [CrossRef] [Green Version]
- Kikuchi, T.; Jones, J.T.; Aikawa, T.; Kosaka, H.; Ogura, N. A family of glycosyl hydrolase family 45 cellulases from the pine wood nematode Bursaphelenchus xylophilus. FEBS Lett. 2004, 572, 201–205. [Google Scholar] [CrossRef] [Green Version]
- Palomares-Rius, J.E.; Hirooka, Y.; Tsai, I.J.; Masuya, H.; Hino, A.; Kanzaki, N.; Jones, J.T.; Kikuchi, T. Distribution and evolution of glycoside hydrolase family 45 cellulases in nematodes and fungi. BMC Evol. Biol. 2014, 14, 69. [Google Scholar] [CrossRef] [Green Version]
- Ding, X.; Wang, Q.; Guo, Y.; Li, Y.; Lin, S.; Zeng, Q.; Sun, F.; Li, D.-W.; Ye, J. Copy Number Variations of Glycoside Hydrolase 45 Genes in Bursaphelenchus xylophilus and Their Impact on the Pathogenesis of Pine Wilt Disease. Forests 2021, 12, 275. [Google Scholar] [CrossRef]
- Brahmachary, M.; Guilmatre, A.; Quilez, J.; Hasson, D.; Borel, C.; Warburton, P.; Sharp, A.J. Digital genotyping of macrosatellites and multicopy genes reveals novel biological functions associated with copy number variation of large tandem repeats. PLoS Genet. 2014, 10, e1004418. [Google Scholar] [CrossRef] [Green Version]
- Redon, R.; Ishikawa, S.; Fitch, K.R.; Feuk, L.; Perry, G.H.; Andrews, T.D.; Fiegler, H.; Shapero, M.H.; Carson, A.R.; Chen, W.W.; et al. Global variation in copy number in the human genome. Nature 2006, 444, 444–454. [Google Scholar] [CrossRef] [Green Version]
- Veerappa, A.M.; Suresh, R.V.; Vishweswaraiah, S.; Lingaiah, K.; Murthy, M.; Manjegowda, D.S.; Padakannaya, P.; Ramachandra, N.B. Global patterns of large copy number variations in the human genome reveal complexity in chromosome organization. Genet. Res. 2015, 97, e18. [Google Scholar] [CrossRef]
- Stranger, B.E.; Forrest, M.S.; Dunning, M.; Ingle, C.E.; Beazley, C.; Thorne, N.; Redon, R.; Bird, C.P.; de Grassi, A.; Lee, C.; et al. Relative impact of nucleotide and copy number variation on gene expression phenotypes. Science 2007, 315, 848–853. [Google Scholar] [CrossRef]
- Verbitsky, M.; Westland, R.; Perez, A.; Kiryluk, K.; Liu, Q.X.; Krithivasan, P.; Mitrotti, A.; Fasel, D.A.; Batourina, E.; Sampson, M.G.; et al. The copy number variation landscape of congenital anomalies of the kidney and urinary tract. Nat. Genet. 2019, 51, 117–127. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Fadista, J.; Nygaard, M.; Holm, L.E.; Thomsen, B.; Bendixen, C. A Snapshot of CNVs in the Pig Genome. PLoS ONE 2008, 3, e3916. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Fiston-Lavier, A.S.; Anxolabehere, D.; Quesneville, H. A model of segmental duplication formation in Drosophila melanogaster. Genome Res. 2007, 17, 1458–1470. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Cantsilieris, S.; Baird, P.N.; White, S.J. Molecular methods for genotyping complex copy number polymorphisms. Genomics 2013, 101, 86–93. [Google Scholar] [CrossRef] [Green Version]
- Yu, L.-Z.; Wu, X.-Q.; Ye, J.-R.; Zhang, S.-N.; Wang, C. NOS-like-mediated nitric oxide is involved in Pinus thunbergii response to the invasion of Bursaphelenchus xylophilus. Plant Cell Rep. 2012, 31, 1813–1821. [Google Scholar] [CrossRef]
- Lupski, J.R. Genomic disorders: Structural features of the genome can lead to DNA rearrangements and human disease traits. Trends Genet. 1998, 14, 417–422. [Google Scholar] [CrossRef]
- Cahan, P.; Li, Y.; Izumi, M.; Graubert, T.A. The impact of copy number variation on local gene expression in mouse hematopoietic stem and progenitor cells. Nat. Genet. 2009, 41, 430–437. [Google Scholar] [CrossRef] [Green Version]
- Gallet, R.; Di Mattia, J.; Ravel, S.; Zeddam, J.L.; Vitalis, R.; Michalakis, Y.; Blanc, S. Gene copy number variations at the within-host population level modulate gene expression in a multipartite virus. Virus Evol. 2022, 8, 11. [Google Scholar] [CrossRef]
- Duda, T.F., Jr.; Palumbi, S.R. Molecular genetics of ecological diversification: Duplication and rapid evolution of toxin genes. Proc. Natl. Acad. Sci. USA 1999, 96, 6820–6823. [Google Scholar] [CrossRef] [Green Version]
- Orozco, L.D.; Cokus, S.J.; Ghazalpour, A.; Ingram-Drake, L.; Wang, S.; van Nas, A.; Che, N.; Araujo, J.A.; Pellegrini, M.; Lusis, A.J. Copy number variation influences gene expression and metabolic traits in mice. Hum. Mol. Genet. 2009, 18, 4118–4129. [Google Scholar] [CrossRef]
- Seo, B.Y.; Park, E.W.; Ahn, S.J.; Lee, S.H.; Kim, J.H.; Im, H.T.; Lee, J.H.; Cho, I.C.; Kong, I.K.; Jeon, J.T. An accurate method for quantifying and analyzing copy number variation in porcine KIT by an oligonucleotide ligation assay. BMC Genet. 2007, 8, 81. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wright, D.; Boije, H.; Meadows, J.; Bed’Hom, B.; Gourichon, D.; Vieaud, A.; Tixier-Boichard, M.; Rubin, C.-J.; Imsland, F.; Hallböök, F.; et al. Copy number variation in intron 1 of SOX5 causes the pea-comb phenotype in chickens. PLoS Genet. 2009, 5, e1000512. [Google Scholar] [CrossRef] [PubMed]
- Feuk, L.; Carson, A.R.; Scherer, S.W. Structural variation in the human genome. Nat. Rev. Genet. 2006, 7, 85–97. [Google Scholar] [CrossRef] [PubMed]
- Goidts, V.; Cooper, D.N.; Armengol, L.; Schempp, W.; Conroy, J.; Estivill, X.; Nowak, N.; Hameister, H.; Kehrer-Sawatzki, H. Complex patterns of copy number variation at sites of segmental duplications: An important category of structural variation in the human genome. Hum. Genet. 2006, 120, 270–284. [Google Scholar] [CrossRef] [PubMed]
- Hou, S.; Qi, J.; Liao, D.; Zhang, Q.; Fang, J.; Zhou, Y.; Liu, Y.; Bai, L.; Zhang, M.; Kijlstra, A.; et al. Copy Number Variations of Complement Component C4 Are Associated With Behçet’s Disease but Not With Ankylosing Spondylitis Associated With Acute Anterior Uveitis. Arthritis Rheum. 2013, 65, 2963–2970. [Google Scholar] [CrossRef] [PubMed]
- Lin, S.; Jian, H.; Zhao, H.; Yang, D.; Liu, Q. Cloning and characterization of a venom allergen-like protein gene cluster from the pinewood nematode Bursaphelenchus xylophilus. Exp. Parasitol. 2011, 127, 440–447. [Google Scholar] [CrossRef]
- Volker, M.; Backstrom, N.; Skinner, B.M.; Langley, E.J.; Bunzey, S.K.; Ellegren, H.; Griffin, D.K. Copy number variation, chromosome rearrangement, and their association with recombination during avian evolution. Genome Res. 2010, 20, 503–511. [Google Scholar] [CrossRef] [Green Version]
- Rice, A.M.; McLysaght, A. Dosage sensitivity is a major determinant of human copy number variant pathogenicity. Nat. Commun. 2017, 8, 14366. [Google Scholar] [CrossRef] [Green Version]
- Riggs, E.; Church, D.; Hanson, K.; Horner, V.; Kaminsky, E.; Kuhn, R.; Wain, K.; Williams, E.; Aradhya, S.; Kearney, H.; et al. Towards an evidence-based process for the clinical interpretation of copy number variation. Clin. Genet. 2012, 81, 403–412. [Google Scholar] [CrossRef]
- Nickle, W.R.; Golden, A.M.; Mamiya, Y.; Wergin, W.P. On the Taxonomy and Morphology of the Pine Wood Nematode, Bursaphelenchus xylophilus (Steiner & Buhrer 1934) Nickle 1970. J. Nematol. 1981, 13, 385–392. [Google Scholar]
- Braasch, H. Morphology of Bursaphelenchus xylophilus compared with other Bursaphelenchus species. In Proceedings of the 2001 International Workshop, Évora, Portugal, 20–22 August 2001; Volume 1, pp. 127–143. [Google Scholar]
- Wang, Y.; Chen, F.; Wang, L.; Li, M. Investigation of beetle species that carry the pine wood nematode, Bursaphelenchus xylophilus (Steiner and Buhrer) Nickle, in China. J. For. Res. 2021, 32, 1745–1751. [Google Scholar] [CrossRef]
- Haymes, K.M. Mini-prep method suitable for a plant breeding program. Plant Mol. Biol. Rep. 1996, 14, 280–284. [Google Scholar] [CrossRef]
- Meng, F.; Li, Y.; Wang, X.; Feng, Y.; Liu, Z.; Zhang, W.; Zhang, X. Thaumatin-Like Protein-1 Gene (Bx-tlp-1) Is Associated with the Pathogenicity of Bursaphelenchus xylophilus. Phytopathology 2019, 109, 1949–1956. [Google Scholar] [CrossRef]
- Xue, Q.; Xiang, Y.; Wu, X.-Q.; Li, M.-J. Bacterial Communities and Virulence Associated with Pine Wood Nematode Bursaphelenchus xylophilus from Different Pinus spp. Int. J. Mol. Sci. 2019, 20, 3342. [Google Scholar] [CrossRef] [Green Version]
- Hu, L.-J.; Wu, X.-Q.; Wen, T.-Y.; Ye, J.; Qiu, Y.-J.; Rui, L.; Zhang, Y. The key molecular pattern BxCDP1 of Bursaphelenchus xylophilus induces plant immunity and enhances plant defense response via two small peptide regions. Front. Plant Sci. 2022, 13, 937473. [Google Scholar] [CrossRef]
- Deng, L.-N.; Wu, X.-Q.; Ye, J.-R.; Xue, Q. Identification of Autophagy in the Pine Wood Nematode Bursaphelenchus xylophilus and the Molecular Characterization and Functional Analysis of Two Novel Autophagy-Related Genes, BxATG1 and BxATG8. Int. J. Mol. Sci. 2016, 17, 279. [Google Scholar] [CrossRef]

Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ding, X.; Zhao, R.; Dai, Y.; Zhang, Y.; Lin, S.; Ye, J. Comprehensive Analysis of Copy Number Variations on Glycoside Hydrolase 45 Genes among Different Bursaphelenchus xylophilus Strains. Int. J. Mol. Sci. 2022, 23, 15323. https://doi.org/10.3390/ijms232315323
Ding X, Zhao R, Dai Y, Zhang Y, Lin S, Ye J. Comprehensive Analysis of Copy Number Variations on Glycoside Hydrolase 45 Genes among Different Bursaphelenchus xylophilus Strains. International Journal of Molecular Sciences. 2022; 23(23):15323. https://doi.org/10.3390/ijms232315323
Chicago/Turabian StyleDing, Xiaolei, Ruiwen Zhao, Yonglin Dai, Yue Zhang, Sixi Lin, and Jianren Ye. 2022. "Comprehensive Analysis of Copy Number Variations on Glycoside Hydrolase 45 Genes among Different Bursaphelenchus xylophilus Strains" International Journal of Molecular Sciences 23, no. 23: 15323. https://doi.org/10.3390/ijms232315323



