Killer Knots: Molecular Evolution of Inhibitor Cystine Knot Toxins in Wandering Spiders (Araneae: Ctenidae)
Abstract
1. Introduction
2. Results
2.1. Venom Gland Transcriptome and Proteome
2.2. Phylogenetic Results
2.3. Inhibitor Cystine Knot Annotation
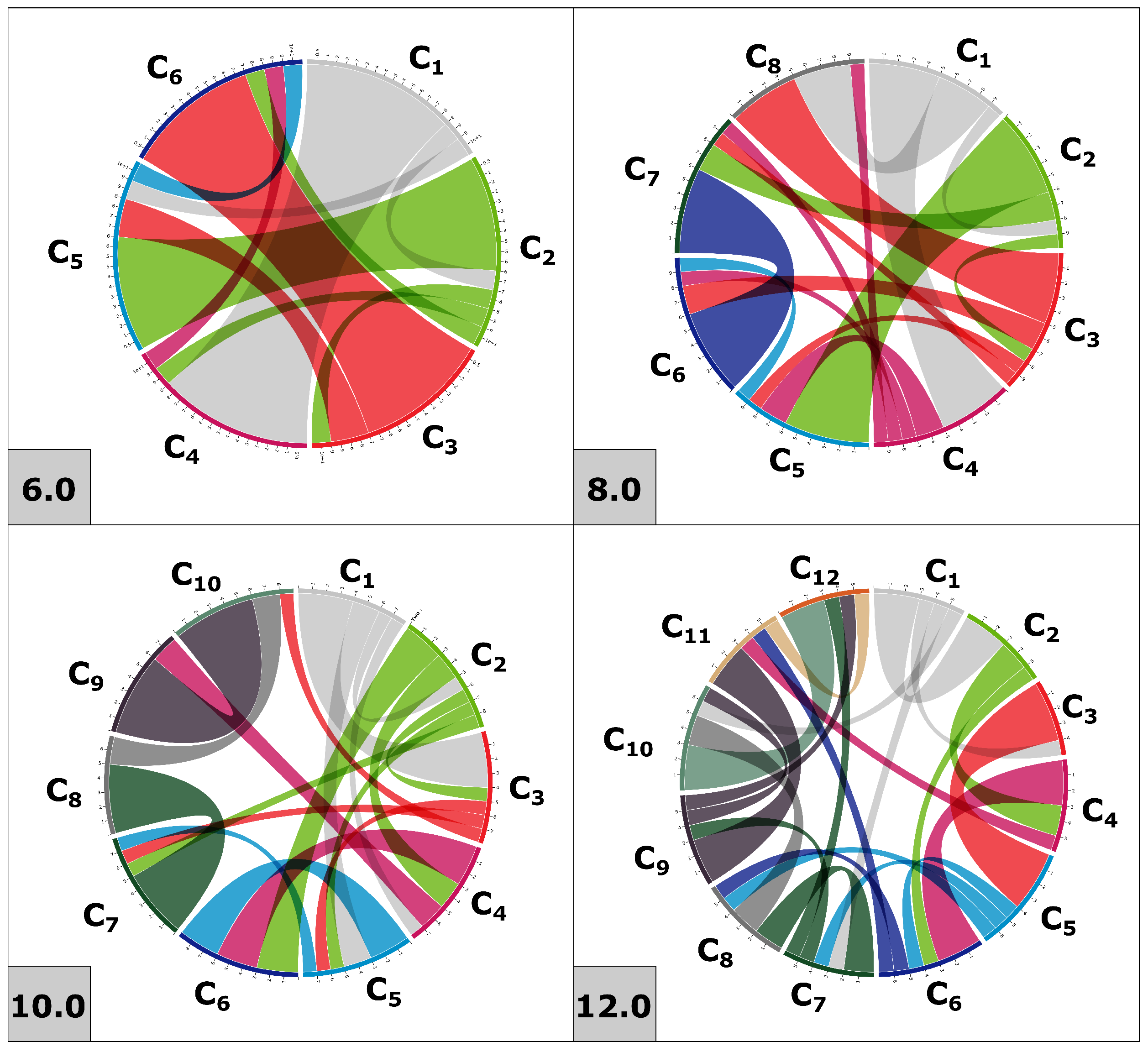
2.4. Disulfide Connectivity Predictions
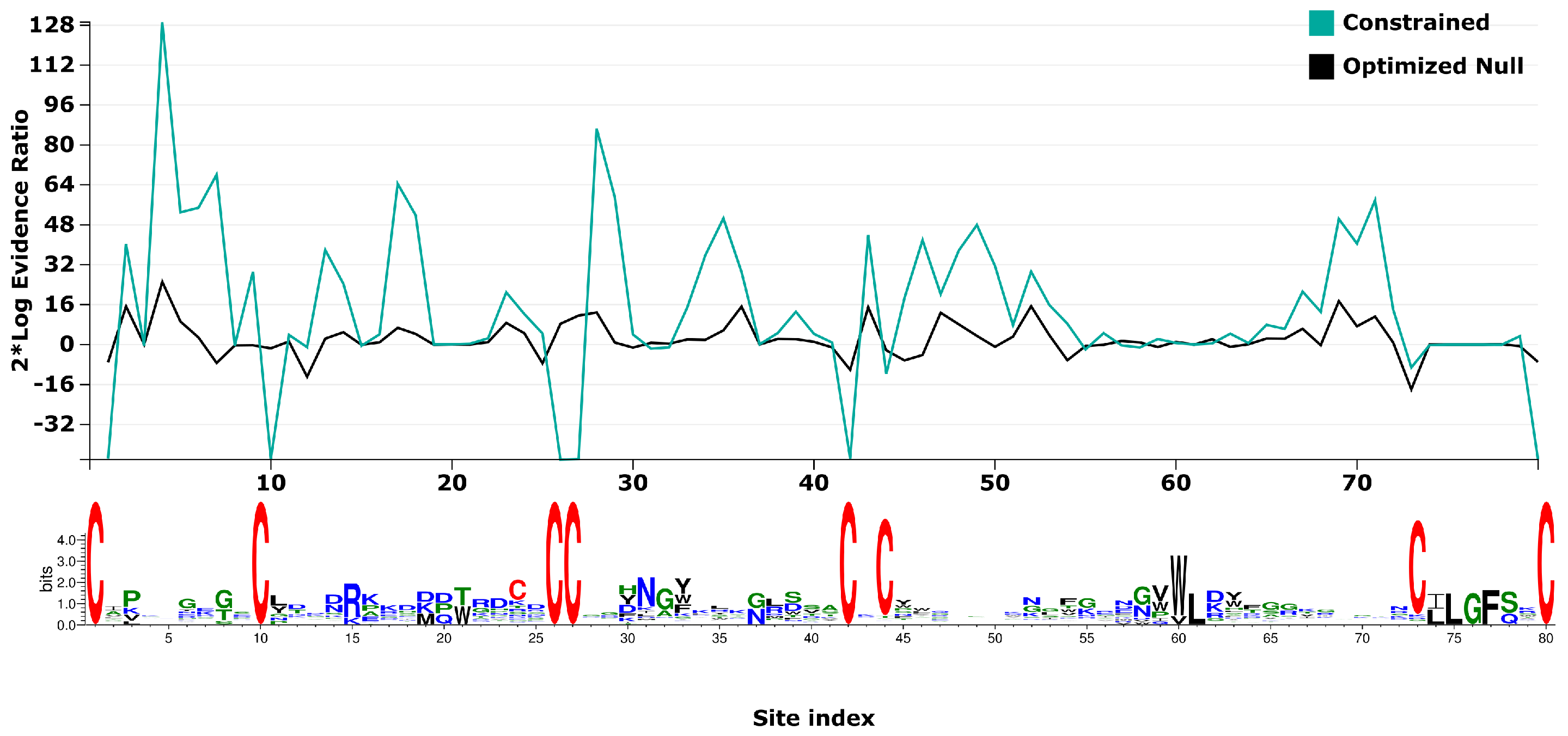
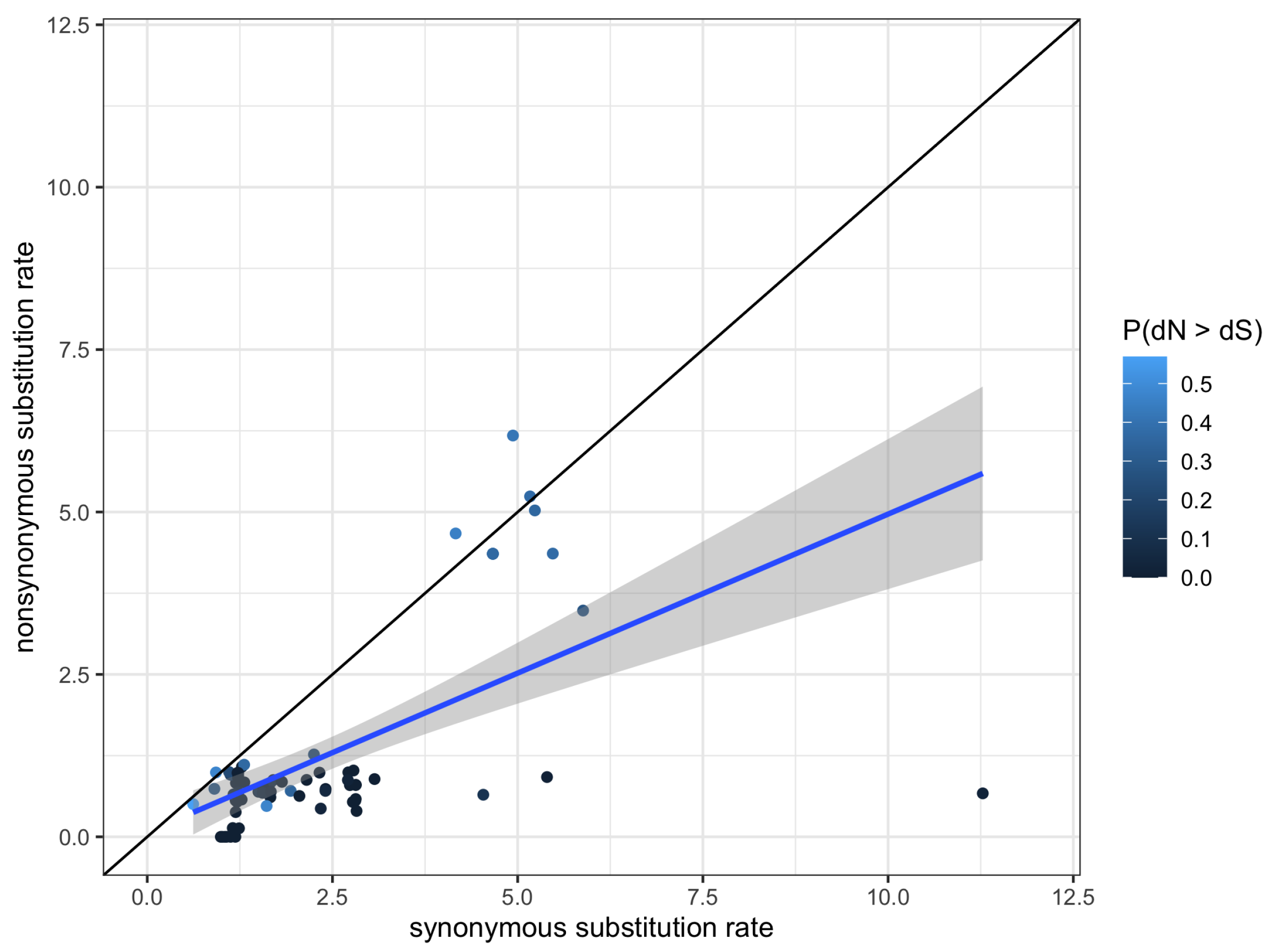
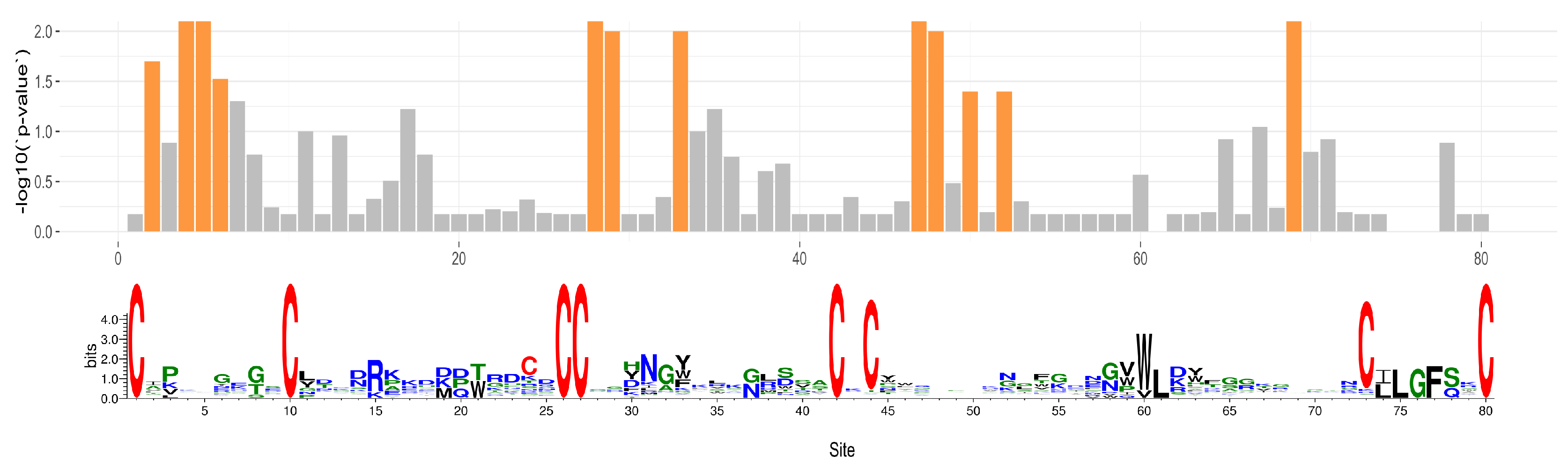
2.5. Phylogenetic Tests for Selection
3. Discussion
4. Conclusions
5. Materials and Methods
5.1. Taxon Sampling
5.2. Venom and RNA Isolation
5.3. Sequencing and Processing
5.4. Transcript Reconstruction and Expression Quantification
5.5. Venom Proteomics
5.6. Locus Sampling
5.7. Phylogenetic Reconstruction
5.8. Inhibitor Cystine Knot Annotation
5.9. Disulfide Connectivity Predictions
- 1.
- DISULFIND collectively decides the bonding state assignment of the entire chain using a Support Vector Machine binary classifier followed by a refinement stage [25]. DISULFIND v1.1 was used to generate a total of three alternative disulfide connection predictions.
- 2.
- CYSCON uses a hierarchical order reduction protocol to identify the most confident disulfide bonds and then evaluate what remains using Support Vector Regression [26]. CYSCON v2015.09.27 was used to generate a single disulfide prediction per ICK representative per unique cysteine framework.
- 3.
- CRISP v1.0 not only predicts disulfide bonds, but also the entire structure of a cysteine rich peptide by searching a customized template database with cysteine-specific sequence alignment with three separate machine learning models to filter templates, rank models, and estimate model quality [27]. CRISP was used to generate five structural models for each ICK representative per unique cysteine framework.
5.10. Phylogenetic Tests for Selection
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Walker, A.A.; Robinson, S.D.; Yeates, D.K.; Jin, J.; Baumann, K.; Dobson, J.; Fry, B.G.; King, G.F. Entomo-venomics: The evolution, biology and biochemistry of insect venoms. Toxicon 2018, 154, 15–27. [Google Scholar] [CrossRef] [PubMed]
- Sunagar, K.; Moran, Y. The rise and fall of an evolutionary innovation: Contrasting strategies of venom evolution in ancient and young animals. PLoS Genet. 2015, 11, e1005596. [Google Scholar] [CrossRef]
- Casewell, N.R.; Wüster, W.; Vonk, F.J.; Harrison, R.A.; Fry, B.G. Complex cocktails: The evolutionary novelty of venoms. Trends Ecol. Evol. 2013, 28, 219–229. [Google Scholar] [CrossRef]
- Holford, M.; Daly, M.; King, G.F.; Norton, R.S. Venoms to the rescue. Science 2018, 361, 842–844. [Google Scholar] [CrossRef]
- World Spider Catalog. World Spider Catalog, Version 19.5; Natural History Museum: Bern, Switzerland, 2019. [Google Scholar]
- Pineda, S.S.; Chin, Y.K.Y.; Undheim, E.A.; Senff, S.; Mobli, M.; Dauly, C.; Escoubas, P.; Nicholson, G.M.; Kaas, Q.; Guo, S.; et al. Structural venomics reveals evolution of a complex venom by duplication and diversification of an ancient peptide-encoding gene. Proc. Natl. Acad. Sci. USA 2020, 117, 11399–11408. [Google Scholar] [CrossRef]
- Wong, E.S.; Belov, K. Venom evolution through gene duplications. Gene 2012, 496, 1–7. [Google Scholar] [CrossRef] [PubMed]
- Schwager, E.E.; Sharma, P.P.; Clarke, T.; Leite, D.J.; Wierschin, T.; Pechmann, M.; Akiyama-Oda, Y.; Esposito, L.; Bechsgaard, J.; Bilde, T.; et al. The house spider genome reveals an ancient whole-genome duplication during arachnid evolution. BMC Biol. 2017, 15, 62. [Google Scholar] [CrossRef] [PubMed]
- Escoubas, P.; Rash, L. Tarantulas: Eight-legged pharmacists and combinatorial chemists. Toxicon 2004, 43, 555–574. [Google Scholar] [CrossRef] [PubMed]
- Arújo, D.A.; Cordeiro, M.N.; Diniz, C.R.; Beirão, P.S. Effects of a toxic fraction, PhTx 2, from the spider Phoneutria nikriventer on the sodium current. Naunyn-Schmiedeberg’s Arch. Pharmacol. 1993, 347, 205–208. [Google Scholar] [CrossRef]
- Gomez, M.V.; Kalapothakis, E.; Guatimosim, C.; Prado, M.A. Phoneutria nigriventer venom: A cocktail of toxins that affect ion channels. Cell. Mol. Neurobiol. 2002, 22, 579–588. [Google Scholar] [CrossRef]
- Nunes, K.P.; Costa-Gonçalves, A.; Lanza, L.F.; Côrtes, S.d.F.; Cordeiro, M.d.N.; Richardson, M.; Pimenta, A.M.d.C.; Webb, R.C.; Leite, R.; De Lima, M. Tx2-6 toxin of the Phoneutria nigriventer spider potentiates rat erectile function. Toxicon 2008, 51, 1197–1206. [Google Scholar] [CrossRef]
- Richardson, M.; Pimenta, A.; Bemquerer, M.; Santoro, M.; Beirao, P.; Lima, M.; Figueiredo, S.; Bloch, C., Jr.; Vasconcelos, E.; Campos, F.; et al. Comparison of the partial proteomes of the venoms of Brazilian spiders of the genus Phoneutria. Comp. Biochem. Physiol. Part C Toxicol. Pharmacol. 2006, 142, 173–187. [Google Scholar] [CrossRef]
- Inns, R.; Tuckwell, N.; Bright, J.; Marrs, T. Histochemical demonstration of calcium accumulation in muscle fibres after experimental organophosphate poisoning. Hum. Exp. Toxicol. 1990, 9, 245–250. [Google Scholar] [CrossRef] [PubMed]
- Mesilaakso, M. Chemical Weapons Convention Chemicals Analysis: Sample Collection, Preparation and Analytical Methods; John Wiley & Sons: Hoboken, NJ, USA, 2005. [Google Scholar]
- Sollod, B.L.; Wilson, D.; Zhaxybayeva, O.; Gogarten, J.P.; Drinkwater, R.; King, G.F. Were arachnids the first to use combinatorial peptide libraries? Peptides 2005, 26, 131–139. [Google Scholar] [CrossRef]
- Narasimhan, L.; Singh, J.; Humblet, C.; Guruprasad, K.; Blundell, T. Snail and spider toxins share a similar tertiary structure and ‘cystine motif’. Nat. Struct. Biol. 1994, 1, 850–852. [Google Scholar] [CrossRef] [PubMed]
- Pallaghy, P.K.; Norton, R.S.; Nielsen, K.J.; Craik, D.J. A common structural motif incorporating a cystine knot and a triple-stranded β-sheet in toxic and inhibitory polypeptides. Protein Sci. 1994, 3, 1833–1839. [Google Scholar] [CrossRef]
- Diniz, M.R.; Paiva, A.L.; Guerra-Duarte, C.; Nishiyama, M.Y., Jr.; Mudadu, M.A.; De Oliveira, U.; Borges, M.H.; Yates, J.R.; Junqueira-de Azevedo, I.d.L. An overview of Phoneutria nigriventer spider venom using combined transcriptomic and proteomic approaches. PLoS ONE 2018, 13, e0200628. [Google Scholar] [CrossRef] [PubMed]
- Cheng, D.Q.; Piel, W.H. The origins of the Psechridae: Web-building lycosoid spiders. Mol. Phylogenet. Evol. 2018, 125, 213–219. [Google Scholar] [CrossRef]
- Simó, M.; Brescovit, A.D. Revision and cladistic analysis of the Neotropical spider genus Phoneutria Perty, 1833 (Araneae, Ctenidae), with notes on related Cteninae. Bull.-Br. Arachnol. Soc. 2001, 12, 67–82. [Google Scholar]
- Davila, D.S. Higher-level relationships of the spider family Ctenidae (Araneae: Ctenoidea). Bull. Am. Mus. Nat. Hist. 2003, 2003, 1–86. [Google Scholar] [CrossRef]
- Brescovit, A.D.; Simó, M. On the Brazilian Atlantic Forest species of the spider genus Ctenus Walckenaer, with the description of a neotype for C. dubius Walckenaer (Araneae, Ctenidae, Cteninae). Arachnology 2007, 14, 1–17. [Google Scholar] [CrossRef]
- Polotow, D.; Brescovit, A.D. Revision of the neotropical spider genus Gephyroctenus (Araneae: Ctenidae: Calocteninae). Rev. Bras. Zool. 2008, 25, 705–715. [Google Scholar] [CrossRef]
- Ceroni, A.; Passerini, A.; Vullo, A.; Frasconi, P. DISULFIND: A disulfide bonding state and cysteine connectivity prediction server. Nucleic Acids Res. 2006, 34, W177–W181. [Google Scholar] [CrossRef] [PubMed]
- Yang, J.; He, B.J.; Jang, R.; Zhang, Y.; Shen, H.B. Accurate disulfide-bonding network predictions improve ab initio structure prediction of cysteine-rich proteins. Bioinformatics 2015, 31, 3773–3781. [Google Scholar] [PubMed]
- Liu, Z.L.; Hu, J.H.; Jiang, F.; Wu, Y.D. CRiSP: Accurate structure prediction of disulfide-rich peptides with cystine-specific sequence alignment and machine learning. Bioinformatics 2020, 36, 3385–3392. [Google Scholar] [CrossRef] [PubMed]
- Van, V.; Van Valen, L. A new evolutionary law. Evol. Theroy 1973, 1, 1–30. [Google Scholar]
- Dawkins, R.; Krebs, J.R. Arms races between and within species. Proc. R. Soc. Lond. Ser. B Biol. Sci. 1979, 205, 489–511. [Google Scholar]
- Endler, J. Defence against predators. In Predator-Prey Relationships; University of Chicago Press: Chicago, IL USA, 1986. [Google Scholar]
- Daltry, J.C.; Wüster, W.; Thorpe, R.S. Diet and snake venom evolution. Nature 1996, 379, 537–540. [Google Scholar] [CrossRef] [PubMed]
- Juárez, P.; Comas, I.; González-Candelas, F.; Calvete, J.J. Evolution of snake venom disintegrins by positive Darwinian selection. Mol. Biol. Evol. 2008, 25, 2391–2407. [Google Scholar] [CrossRef]
- Sunagar, K.; Jackson, T.N.; Undheim, E.A.; Ali, S.; Antunes, A.; Fry, B.G. Three-fingered RAVERs: Rapid Accumulation of Variations in Exposed Residues of snake venom toxins. Toxins 2013, 5, 2172–2208. [Google Scholar] [CrossRef]
- Haller, B.C.; Hendry, A.P. Solving the paradox of stasis: Squashed stabilizing selection and the limits of detection. Evolution 2014, 68, 483–500. [Google Scholar] [CrossRef] [PubMed]
- Barrio, A.; Brazil, O.V. Ein neues verfahren der Giftentnahme bei spinnen. Experientia 1950, 6, 112–113. [Google Scholar] [CrossRef]
- Munekiyo, S.M.; Mackessy, S.P. Effects of temperature and storage conditions on the electrophoretic, toxic and enzymatic stability of venom components. Comp. Biochem. Physiol. Part B Biochem. Mol. Biol. 1998, 119, 119–127. [Google Scholar] [CrossRef]
- Binford, G.J.; Wells, M.A. The phylogenetic distribution of sphingomyelinase D activity in venoms of Haplogyne spiders. Comp. Biochem. Physiol. Part B Biochem. Mol. Biol. 2003, 135, 25–33. [Google Scholar] [CrossRef]
- Clarke, T.H.; Garb, J.E.; Hayashi, C.Y.; Haney, R.A.; Lancaster, A.K.; Corbett, S.; Ayoub, N.A. Multi-tissue transcriptomics of the black widow spider reveals expansions, co-options, and functional processes of the silk gland gene toolkit. BMC Genomics 2014, 15, 365. [Google Scholar] [CrossRef]
- Chen, S.; Zhou, Y.; Chen, Y.; Gu, J. fastp: An ultra-fast all-in-one FASTQ preprocessor. Bioinformatics 2018, 34, i884–i890. [Google Scholar] [CrossRef]
- MacManes, M.D. The Oyster River Protocol: A multi-assembler and kmer approach for de novo transcriptome assembly. PeerJ 2018, 6, e5428. [Google Scholar] [CrossRef]
- Hart, T.; Komori, H.K.; LaMere, S.; Podshivalova, K.; Salomon, D.R. Finding the active genes in deep RNA-seq gene expression studies. BMC Genomics 2013, 14, 778. [Google Scholar] [CrossRef]
- Longo, M.S.; O’Neill, M.J.; O’Neill, R.J. Abundant human DNA contamination identified in non-primate genome databases. PLoS ONE 2011, 6, e16410. [Google Scholar] [CrossRef] [PubMed]
- Lusk, R.W. Diverse and widespread contamination evident in the unmapped depths of high throughput sequencing data. PLoS ONE 2014, 9, e110808. [Google Scholar] [CrossRef] [PubMed]
- Merchant, S.; Wood, D.E.; Salzberg, S.L. Unexpected cross-species contamination in genome sequencing projects. PeerJ 2014, 2, e675. [Google Scholar] [CrossRef] [PubMed]
- Bergmann, E.A.; Chen, B.J.; Arora, K.; Vacic, V.; Zody, M.C. Conpair: Concordance and contamination estimator for matched tumor–normal pairs. Bioinformatics 2016, 32, 3196–3198. [Google Scholar] [CrossRef] [PubMed]
- Edgar, R.C. UNCROSS: Filtering of high-frequency cross-talk in 16S amplicon reads. bioRxiv 2016, 088666. [Google Scholar] [CrossRef]
- Borner, J.; Burmester, T. Parasite infection of public databases: A data mining approach to identify apicomplexan contaminations in animal genome and transcriptome assemblies. BMC Genomics 2017, 18, 100. [Google Scholar] [CrossRef]
- Lafond-Lapalme, J.; Duceppe, M.O.; Wang, S.; Moffett, P.; Mimee, B. A new method for decontamination of de novo transcriptomes using a hierarchical clustering algorithm. Bioinformatics 2017, 33, 1293–1300. [Google Scholar] [CrossRef]
- Ballenghien, M.; Faivre, N.; Galtier, N. Patterns of cross-contamination in a multispecies population genomic project: Detection, quantification, impact, and solutions. BMC Biol. 2017, 15, 25. [Google Scholar] [CrossRef]
- Patro, R.; Duggal, G.; Love, M.I.; Irizarry, R.A.; Kingsford, C. Salmon provides fast and bias-aware quantification of transcript expression. Nat. Methods 2017, 14, 417. [Google Scholar] [CrossRef]
- Haas, B.J.; Papanicolaou, A.; Yassour, M.; Grabherr, M.; Blood, P.D.; Bowden, J.; Couger, M.B.; Eccles, D.; Li, B.; Lieber, M.; et al. De novo transcript sequence reconstruction from RNA-seq using the Trinity platform for reference generation and analysis. Nature Protoc. 2013, 8, 1494. [Google Scholar] [CrossRef]
- Davidson, N.M.; Hawkins, A.D.; Oshlack, A. SuperTranscripts: A data driven reference for analysis and visualisation of transcriptomes. Genome Biol. 2017, 18, 148. [Google Scholar] [CrossRef]
- McIlwain, S.; Tamura, K.; Kertesz-Farkas, A.; Grant, C.E.; Diament, B.; Frewen, B.; Howbert, J.J.; Hoopmann, M.R.; Kall, L.; Eng, J.K.; et al. Crux: Rapid open source protein tandem mass spectrometry analysis. J. Proteome Res. 2014, 13, 4488–4491. [Google Scholar] [CrossRef]
- Haas, B.; Papanicolaou, A. TransDecoder (Find Coding Regions within Transcripts). 2015. Available online: https://github.com/TransDecoder/TransDecoder (accessed on 17 May 2018).
- Simão, F.A.; Waterhouse, R.M.; Ioannidis, P.; Kriventseva, E.V.; Zdobnov, E.M. BUSCO: Assessing genome assembly and annotation completeness with single-copy orthologs. Bioinformatics 2015, 31, 3210–3212. [Google Scholar] [CrossRef] [PubMed]
- Katoh, K.; Standley, D.M. MAFFT multiple sequence alignment software version 7: Improvements in performance and usability. Mol. Biol. Evol. 2013, 30, 772–780. [Google Scholar] [CrossRef]
- Capella-Gutiérrez, S.; Silla-Martínez, J.M.; Gabaldón, T. trimAl: A tool for automated alignment trimming in large-scale phylogenetic analyses. Bioinformatics 2009, 25, 1972–1973. [Google Scholar] [CrossRef] [PubMed]
- Nguyen, L.T.; Schmidt, H.A.; Von Haeseler, A.; Minh, B.Q. IQ-TREE: A fast and effective stochastic algorithm for estimating maximum-likelihood phylogenies. Mol. Biol. Evol. 2015, 32, 268–274. [Google Scholar] [CrossRef]
- Mirarab, S.; Reaz, R.; Bayzid, M.S.; Zimmermann, T.; Swenson, M.S.; Warnow, T. ASTRAL: Genome-scale coalescent-based species tree estimation. Bioinformatics 2014, 30, i541–i548. [Google Scholar] [CrossRef] [PubMed]
- Gelly, J.C.; Gracy, J.; Kaas, Q.; Le-Nguyen, D.; Heitz, A.; Chiche, L. The KNOTTIN website and database: A new information system dedicated to the knottin scaffold. Nucleic Acids Res. 2004, 32, D156–D159. [Google Scholar] [CrossRef]
- Altschul, S.F.; Madden, T.L.; Schäffer, A.A.; Zhang, J.; Zhang, Z.; Miller, W.; Lipman, D.J. Gapped BLAST and PSI-BLAST: A new generation of protein database search programs. Nucleic Acids Res. 1997, 25, 3389–3402. [Google Scholar] [CrossRef]
- Finn, R.D.; Clements, J.; Eddy, S.R. HMMER web server: Interactive sequence similarity searching. Nucleic Acids Res. 2011, 39, W29–W37. [Google Scholar] [CrossRef] [PubMed]
- Armenteros, J.J.A.; Tsirigos, K.D.; Sønderby, C.K.; Petersen, T.N.; Winther, O.; Brunak, S.; von Heijne, G.; Nielsen, H. SignalP 5.0 improves signal peptide predictions using deep neural networks. Nat. Biotechnol. 2019, 37, 420–423. [Google Scholar] [CrossRef]
- Miele, V.; Penel, S.; Duret, L. Ultra-fast sequence clustering from similarity networks with SiLiX. BMC Bioinform. 2011, 12, 116. [Google Scholar] [CrossRef]
- Rubinstein, R.; Fiser, A. Predicting disulfide bond connectivity in proteins by correlated mutations analysis. Bioinformatics 2008, 24, 498–504. [Google Scholar] [CrossRef] [PubMed]
- Bostock, M.; Ogievetsky, V.; Heer, J. D3 data-driven documents. IEEE Trans. Vis. Comput. Graph. 2011, 17, 2301–2309. [Google Scholar] [CrossRef] [PubMed]
- Herzig, V.; Wood, D.L.; Newell, F.; Chaumeil, P.A.; Kaas, Q.; Binford, G.J.; Nicholson, G.M.; Gorse, D.; King, G.F. ArachnoServer 2.0, an updated online resource for spider toxin sequences and structures. Nucleic Acids Res. 2010, 39, D653–D657. [Google Scholar] [CrossRef] [PubMed]
- Shafee, T.M.; Robinson, A.J.; van der Weerden, N.; Anderson, M.A. Structural homology guided alignment of cysteine rich proteins. SpringerPlus 2016, 5, 27. [Google Scholar] [CrossRef]
- Löytynoja, A. Phylogeny-aware alignment with PRANK. In Multiple Sequence Alignment Methods; Springer: Berlin/Heidelberg, Germany, 2014; pp. 155–170. [Google Scholar]
- Pond, S.L.K.; Muse, S.V. HyPhy: Hypothesis testing using phylogenies. In Statistical Methods in Molecular Evolution; Springer: Berlin/Heidelberg, Germany, 2005; pp. 125–181. [Google Scholar]
- Murrell, B.; Wertheim, J.O.; Moola, S.; Weighill, T.; Scheffler, K.; Pond, S.L.K. Detecting individual sites subject to episodic diversifying selection. PLoS Genet. 2012, 8, e1002764. [Google Scholar] [CrossRef]
- Smith, M.D.; Wertheim, J.O.; Weaver, S.; Murrell, B.; Scheffler, K.; Kosakovsky Pond, S.L. Less is more: An adaptive branch-site random effects model for efficient detection of episodic diversifying selection. Mol. Biol. Evol. 2015, 32, 1342–1353. [Google Scholar] [CrossRef]
- Poon, A.F.; Lewis, F.I.; Frost, S.D.; Kosakovsky Pond, S.L. Spidermonkey: Rapid detection of co-evolving sites using Bayesian graphical models. Bioinformatics 2008, 24, 1949–1950. [Google Scholar] [CrossRef]


| Species | Sex | Sample | Transcripts | SuperTranscripts | CDS |
|---|---|---|---|---|---|
| Anahita punctulata | male | 297 | 120,048 | 88,950 | 55,238 |
| Ctenus captiosus | female | 305 | 124,919 | 99,712 | 59,434 |
| Ctenus captiosus | female | 311 | 140,647 | 110,550 | 64,437 |
| Ctenus captiosus | male | 303 | 157,109 | 123,061 | 70,951 |
| Ctenus captiosus | male | 306 | 158,795 | 122,878 | 71,795 |
| Ctenus exlineae | female | 244 | 105,140 | 85,561 | 49,630 |
| Ctenus exlineae | female | 245 | 111,981 | 89,516 | 52,365 |
| Ctenus exlineae | female | 247 | 112,224 | 90,112 | 52,434 |
| Ctenus exlineae | male | 242 | 95,088 | 78,974 | 44,942 |
| Ctenus exlineae | male | 246 | 54,776 | 46,375 | 24,166 |
| Ctenus hibernalis | female | 91 | 194,576 | 148,947 | 83,512 |
| Ctenus hibernalis | female | 92 | 161,519 | 124,108 | 71,340 |
| Ctenus hibernalis | male | 148 | 202,764 | 157,422 | 81,381 |
| Leptoctenus byrrhus | female | 136 | 99,257 | 81,488 | 47,189 |
| Leptoctenus byrrhus | male | 213 | 108,687 | 88,723 | 50,313 |
| Leptoctenus byrrhus | male | 222 | 101,706 | 83,634 | 47,527 |
| Species | Sex | Sample | Peptides | ICKs | %ICK | Sum TPM | ICK TPM | %TPM |
|---|---|---|---|---|---|---|---|---|
| C. exlineae | male | 242 | 113 | 5 | 4.42% | 18,214.3 | 9670.5 | 53.1% |
| C. exlineae | female | 245 | 127 | 7 | 5.51% | 16,755.8 | 7002.0 | 41.8% |
| C. exlineae | male | 246 | 200 | 13 | 6.50% | 131,454 | 51,011.1 | 38.8% |
| C. exlineae | female | 244 | 339 | 5 | 1.47% | 42,546.4 | 12,227.6 | 28.7% |
| C. exlineae | female | 247 | 194 | 5 | 2.58% | 27,378.7 | 5436.3 | 19.9% |
| C. hibernalis | female | 4926 | 196 | 8 | 4.08% | 109,103 | 57,283.2 | 52.5% |
| C. hibernalis | female | 91 | 525 | 12 | 2.29% | 111,759 | 14,703.9 | 13.2% |
| C. hibernalis | male | 148 | 170 | 7 | 4.12% | 31,121.2 | 2581.5 | 8.3% |
| C. hibernalis | female | 92 | 176 | 5 | 2.84% | 37,128.6 | 2366.6 | 6.4% |
| Identifier | Dinez Numeral | Motif | Total |
|---|---|---|---|
| 6.0 | I | ---- | 123 |
| 8.0 | II | ---- | 538 |
| 10.0 | V | ----- | 117 |
| 10.1 | ----- | 23 | |
| 12.0 | VI | ------- | 100 |
| 12.1 | VII | ------ | 27 |
| 14.0 | VIII | --------- | 33 |
| Family | Species | 6.0 | 8.0 | 10.0 | 10.1 | 12.0 | 12.1 | 14.0 |
|---|---|---|---|---|---|---|---|---|
| Homalonychidae | Homalonychus theologus | 1 | 4 | 2 | 0 | 9 | 0 | 0 |
| Salticidae | Habronattus signatus | 6 | 13 | 2 | 1 | 1 | 0 | 0 |
| Xenoctenidae | Odo patricius | 2 | 21 | 2 | 0 | 3 | 0 | 0 |
| Anyphaenidae | Hibana sp. | 3 | 10 | 0 | 1 | 1 | 0 | 0 |
| Gnaphosidae | Sergiolus capulatus | 1 | 11 | 0 | 1 | 2 | 0 | 0 |
| Thomisidae | Thomisus spectabilis | 2 | 13 | 3 | 0 | 2 | 0 | 1 |
| Thomisidae | Misumenoides formosipes | 1 | 4 | 0 | 1 | 0 | 0 | 0 |
| Oxyopidae | Oxyopes sp. | 0 | 15 | 9 | 0 | 2 | 0 | 0 |
| Oxyopidae | Peucetia longipalpis | 1 | 11 | 4 | 1 | 0 | 0 | 0 |
| Lycosidae | Hippasa holmerae | 1 | 11 | 5 | 1 | 3 | 0 | 0 |
| Lycosidae | Pardosa pseudoannulata | 0 | 7 | 1 | 1 | 0 | 0 | 0 |
| Lycosidae | Schizocosa rovneri | 0 | 10 | 0 | 0 | 1 | 0 | 0 |
| Lycosidae | Sosippus placidus | 5 | 26 | 1 | 1 | 3 | 0 | 2 |
| Pisauridae | Nilus albocinctus | 1 | 8 | 2 | 1 | 3 | 0 | 0 |
| Pisauridae | Sphedanus quadrimaculatus | 1 | 6 | 2 | 1 | 4 | 0 | 0 |
| Pisauridae | Pisaurina mira | 0 | 1 | 0 | 1 | 1 | 0 | 0 |
| Pisauridae | Dolomedes triton | 1 | 11 | 0 | 0 | 10 | 0 | 1 |
| Psechridae | Fecenia protensa | 1 | 17 | 4 | 0 | 2 | 1 | 1 |
| Psechridae | Psechrus singaporensis | 0 | 13 | 3 | 1 | 2 | 1 | 0 |
| Ctenidae | Ctenus corniger | 11 | 19 | 5 | 1 | 3 | 1 | 1 |
| Ctenidae | Anahita punctulata | 2 | 15 | 1 | 0 | 2 | 1 | 1 |
| Ctenidae | Ctenus captiosus | 4 | 10 | 2 | 1 | 3 | 1 | 2 |
| Ctenidae | Ctenus exlineae | 2 | 9 | 2 | 1 | 2 | 1 | 1 |
| Ctenidae | Ctenus hibernalis | 2 | 10 | 3 | 1 | 3 | 1 | 1 |
| Ctenidae | Isoctenus sp. | 3 | 11 | 4 | 0 | 0 | 1 | 2 |
| Ctenidae | Leptoctenus byrrhus | 1 | 12 | 2 | 1 | 1 | 1 | 1 |
| Ctenidae | Phoneutria nigriventer | 5 | 12 | 3 | 0 | 1 | 1 | 2 |
| Class | Loop 1 | Loop 2 | Loop 3 | Loop 4 |
|---|---|---|---|---|
| C6.0 | - | - | - | - |
| C8.0 | - | - | - | - |
| C10.0 | - | - | - | - |
| C10.1 | - | - | -- | - |
| C12.0 | - | - | - | - |
| C12.1 | - | - | - | - |
| C14.0 | - | - | - | - |
| Model | log(likelihood) | Parameters | AICc | |||
|---|---|---|---|---|---|---|
| Unconstrained | −37,648.7 | 1169 | 77,691.4 | 0.06 | 0.09 | 3.35 |
| Constrained | −37,733.9 | 1168 | 77,859.6 | 0.03 | 0.03 | 1.00 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Brewer, M.S.; Cole, T.J. Killer Knots: Molecular Evolution of Inhibitor Cystine Knot Toxins in Wandering Spiders (Araneae: Ctenidae). Toxins 2023, 15, 112. https://doi.org/10.3390/toxins15020112
Brewer MS, Cole TJ. Killer Knots: Molecular Evolution of Inhibitor Cystine Knot Toxins in Wandering Spiders (Araneae: Ctenidae). Toxins. 2023; 15(2):112. https://doi.org/10.3390/toxins15020112
Chicago/Turabian StyleBrewer, Michael S., and T. Jeffrey Cole. 2023. "Killer Knots: Molecular Evolution of Inhibitor Cystine Knot Toxins in Wandering Spiders (Araneae: Ctenidae)" Toxins 15, no. 2: 112. https://doi.org/10.3390/toxins15020112
APA StyleBrewer, M. S., & Cole, T. J. (2023). Killer Knots: Molecular Evolution of Inhibitor Cystine Knot Toxins in Wandering Spiders (Araneae: Ctenidae). Toxins, 15(2), 112. https://doi.org/10.3390/toxins15020112





