The Gene vepN Regulated by Global Regulatory Factor veA That Affects Aflatoxin Production, Morphological Development and Pathogenicity in Aspergillus flavus
Abstract
:1. Introduction
2. Results
2.1. Analysis of CHIP-Seq Experimental Results
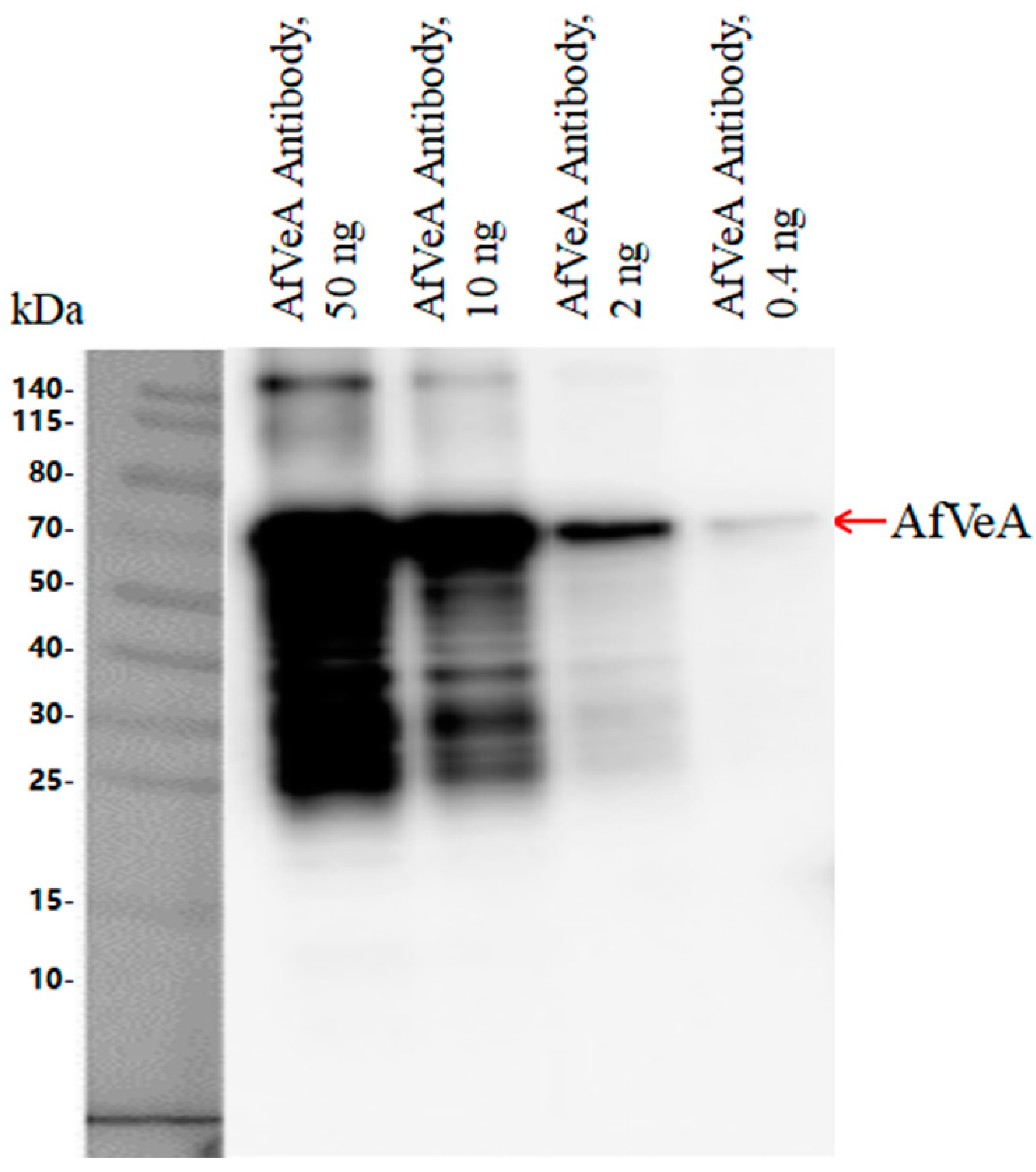
2.2. Identification A. flavus NRRL3357 Gene vepN
2.3. Construction of the Deletion Mutant (ΔvepN) and Overexpression (OE::vepN) Mutant
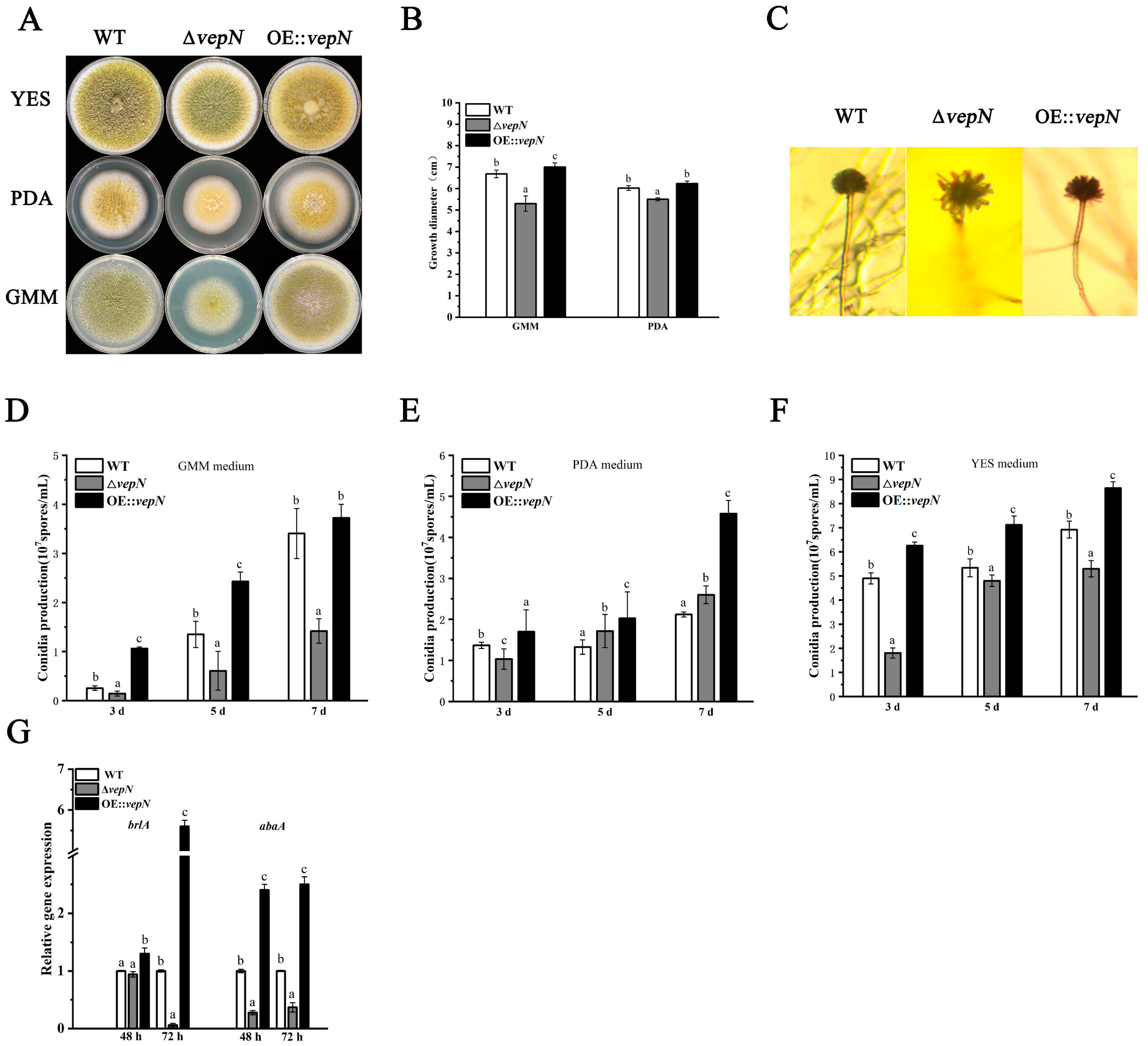
2.4. Effects of vepN on Morphology
2.5. Effects of vepN on Conidial Formation
2.6. Effects of vepN on Development of Sclerotia
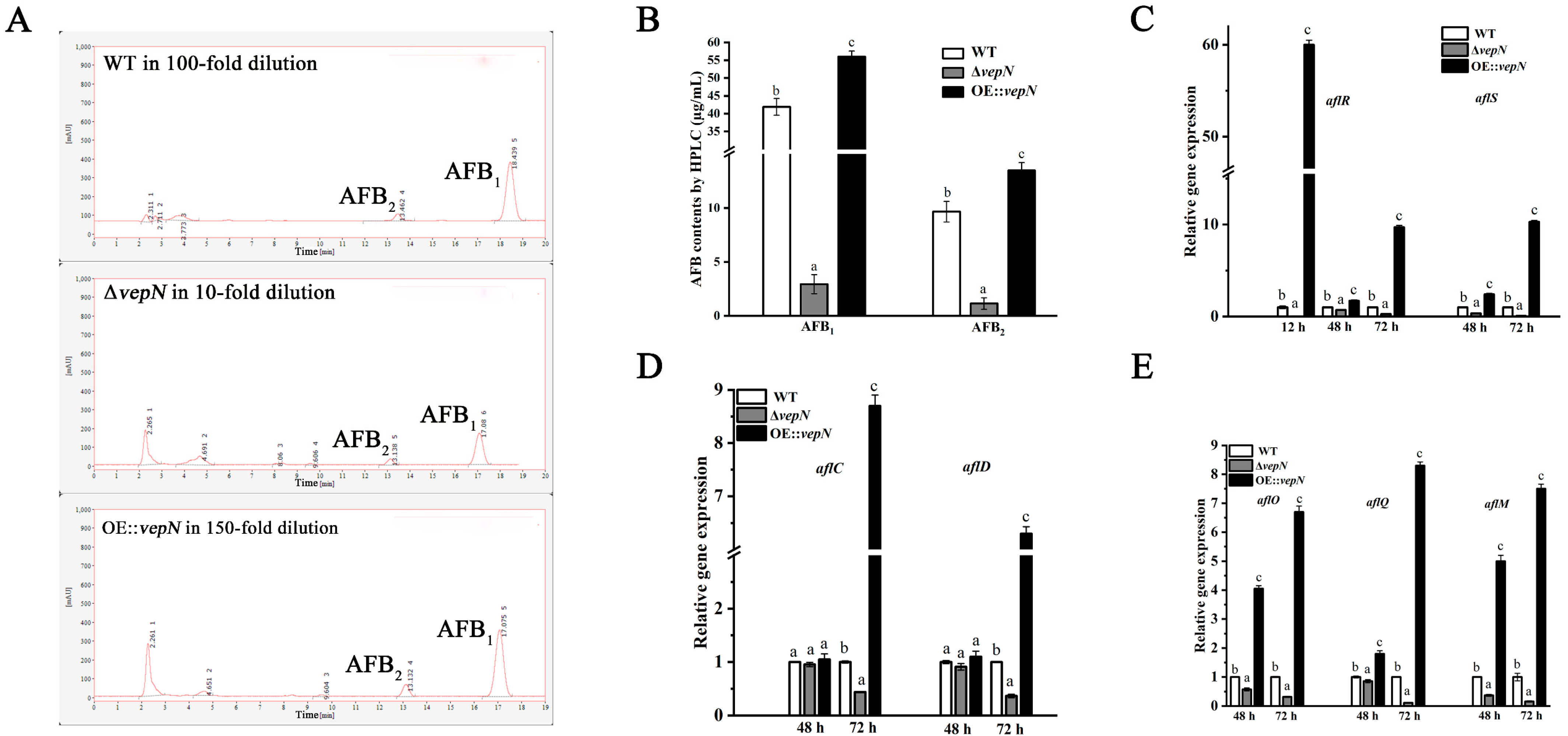
2.7. Effects of vepN on AFB
2.8. vepN Is Necessary for A. flavus Peanut Infections
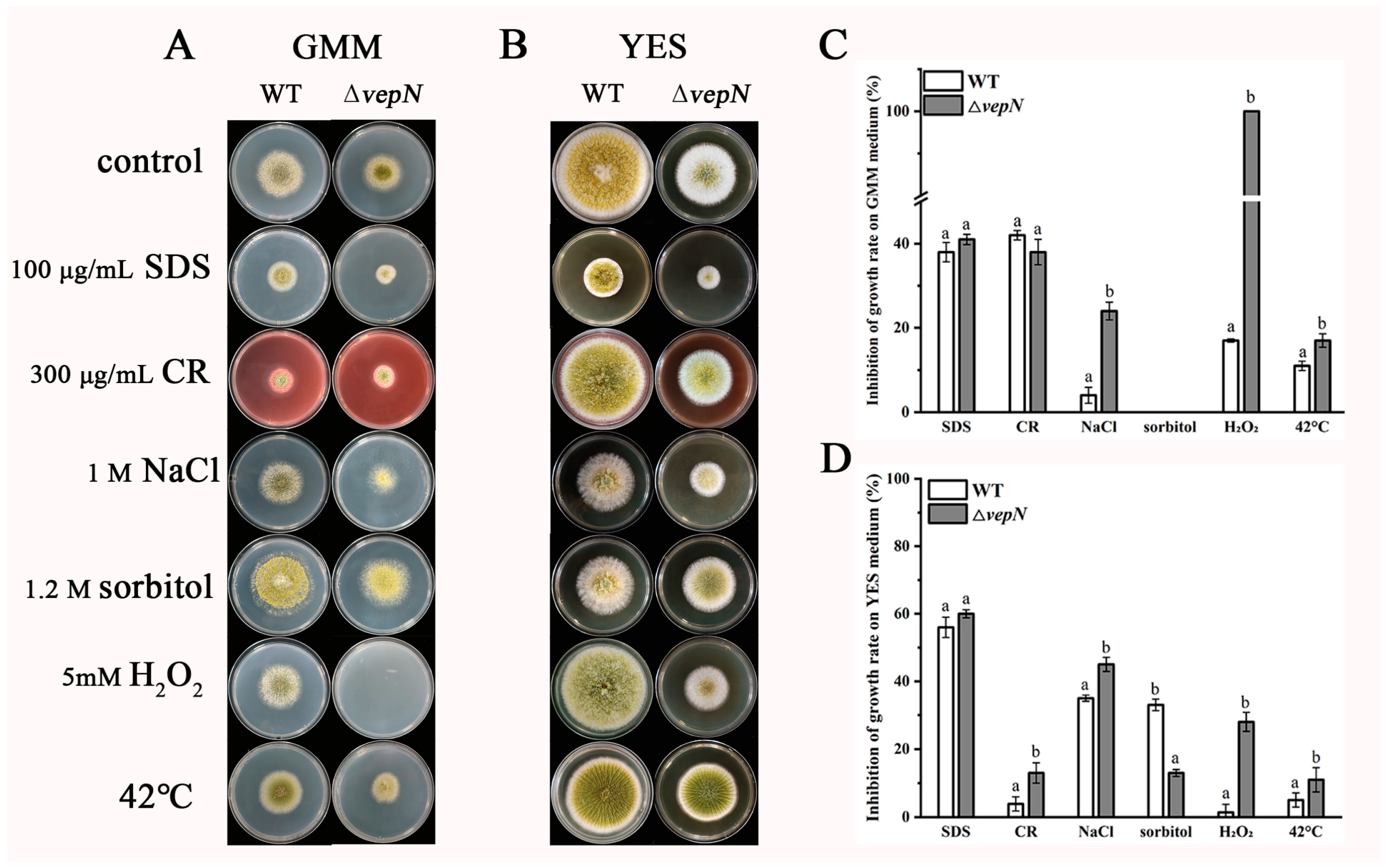
2.9. Responses to Different Stressors
2.10. Effects of vepN on the Cell Wall
3. Discussion
4. Materials and Methods
4.1. Strains, Media, and Growth Conditions
4.2. CHIP-Seq Experiment
4.3. Phylogenetic Analysis
4.4. Construction of Gene Knockout and Overexpression Strains
4.5. Observation of Conidia Morphology and Colonies
4.6. Conidial Formation
4.7. Sclerotial Production
4.8. Determination of Aflatoxin
4.9. Seed Infection
4.10. Stress Experiment
4.11. Transmission Electron Microscope (TEM)
4.12. RT-qPCR Analysis
4.13. Statistical Analysis
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Klich, M.A. Soil fungi of some low-altitude desert cotton fields and ability of their extracts to inhibit Aspergillus flavus. Mycopathologia 1998, 142, 97–100. [Google Scholar] [CrossRef] [PubMed]
- Ojeda-López, M.; Chen, W.; Eagle, C.E.; Gutiérrez, G.; Jia, W.L.; Swilaiman, S.S.; Huang, Z.; Park, H.S.; Yu, J.H.; Cánovas, D.; et al. Evolution of asexual and sexual reproduction in the aspergilli. Stud. Mycol. 2018, 91, 37–59. [Google Scholar] [CrossRef] [PubMed]
- Yu, J.; Cleveland, T.E.; Nierman, W.C.; Bennett, J.W. Aspergillus flavus genomics: Gateway to human and animal health, food safety, and crop resistance to diseases. Rev. Iberoam. Micol. 2005, 22, 194–202. [Google Scholar] [CrossRef] [PubMed]
- Denning, D.W.; Riniotis, K.; Dobrashian, R.; Sambatakou, H. Chronic cavitary and fibrosing pulmonary and pleural aspergillosis: Case series, proposed nomenclature change, and review. Clin. Infect. Dis. 2003, 37 (Suppl. 3), S265–S280. [Google Scholar] [CrossRef] [PubMed]
- Klich, M.A. Aspergillus flavus: The major producer of aflatoxin. Mol. Plant Pathol. 2007, 8, 713–722. [Google Scholar] [CrossRef] [PubMed]
- Frisvad, J.C.; Hubka, V.; Ezekiel, C.N.; Hong, S.B.; Nováková, A.; Chen, A.J.; Arzanlou, M.; Larsen, T.O.; Sklenář, F.; Mahakarnchanakul, W.; et al. Taxonomy of Aspergillus section Flavi and their production of aflatoxins, ochratoxins and other mycotoxins. Stud. Mycol. 2019, 93, 1–63. [Google Scholar] [CrossRef] [PubMed]
- Caceres, I.; Khoury, A.A.; Khoury, R.E.; Lorber, S.; Oswald, I.P.; Khoury, A.E.; Atoui, A.; Puel, O.; Bailly, J.D. Aflatoxin Biosynthesis and Genetic Regulation: A Review. Toxins 2020, 12, 150. [Google Scholar] [CrossRef] [PubMed]
- Yu, J.; Chang, P.K.; Ehrlich, K.C.; Cary, J.W.; Bhatnagar, D.; Cleveland, T.E.; Payne, G.A.; Linz, J.E.; Woloshuk, C.P.; Bennett, J.W. Clustered pathway genes in aflatoxin biosynthesis. Appl. Environ. Microbiol. 2004, 70, 1253–1262. [Google Scholar] [CrossRef]
- Bhatnagar, D.; Cary, J.W.; Ehrlich, K.; Yu, J.; Cleveland, T.E. Understanding the genetics of regulation of aflatoxin production and Aspergillus flavus development. Mycopathologia 2006, 162, 155–166. [Google Scholar] [CrossRef]
- Mooney, J.L.; Yager, L.N. Light is required for conidiation in Aspergillus nidulans. Genes. Dev. 1990, 4, 1473–1482. [Google Scholar] [CrossRef]
- Bayram, O.; Krappmann, S.; Ni, M.; Bok, J.W.; Helmstaedt, K.; Valerius, O.; Braus-Stromeyer, S.; Kwon, N.J.; Keller, N.P.; Yu, J.H.; et al. VelB/VeA/LaeA complex coordinates light signal with fungal development and secondary metabolism. Science 2008, 320, 1504–1506. [Google Scholar] [CrossRef] [PubMed]
- Amare, M.G.; Keller, N.P. Molecular mechanisms of Aspergillus flavus secondary metabolism and development. Fungal Genet. Biol. 2014, 66, 11–18. [Google Scholar] [CrossRef] [PubMed]
- Moon, H.; Lee, M.K.; Bok, I.; Bok, J.W.; Keller, N.P.; Yu, J.H. Unraveling the Gene Regulatory Networks of the Global Regulators VeA and LaeA in Aspergillus nidulans. Microbiol. Spectr. 2023, 11, e0016623. [Google Scholar] [CrossRef] [PubMed]
- Kim, H.; Han, K.; Kim, K.; Han, D.; Jahng, K.; Chae, K. The veA gene activates sexual development in Aspergillus nidulans. Fungal Genet. Biol. 2002, 37, 72–80. [Google Scholar] [CrossRef] [PubMed]
- Calvo, A.M.; Bok, J.; Brooks, W.; Keller, N.P. veA is required for toxin and sclerotial production in Aspergillus parasiticus. Appl. Environ. Microbiol. 2004, 70, 4733–4739. [Google Scholar] [CrossRef] [PubMed]
- Duran, R.M.; Cary, J.W.; Calvo, A.M. Production of cyclopiazonic acid, aflatrem, and aflatoxin by Aspergillus flavus is regulated by veA, a gene necessary for sclerotial formation. Appl. Microbiol. Biotechnol. 2007, 73, 1158–1168. [Google Scholar] [CrossRef] [PubMed]
- Kato, N.; Brooks, W.; Calvo, A.M. The expression of sterigmatocystin and penicillin genes in Aspergillus nidulans is controlled by veA, a gene required for sexual development. Eukaryot. Cell. 2003, 2, 1178–1186. [Google Scholar] [CrossRef] [PubMed]
- Wang, P.; Chang, P.K.; Kong, Q.; Shan, S.; Wei, Q. Comparison of aflatoxin production of Aspergillus flavus at different temperatures and media: Proteome analysis based on TMT. Int. J. Food Microbiol. 2019, 310, 108313. [Google Scholar] [CrossRef] [PubMed]
- Warenda, A.J.; Konopka, J.B. Septin function in Candida albicans morphogenesis. Mol. Biol. Cell. 2002, 13, 2732–2746. [Google Scholar] [CrossRef]
- Hernández-Rodríguez, Y.; Masuo, S.; Johnson, D.; Orlando, R.; Smith, A.; Couto-Rodriguez, M.; Momany, M. Distinct septin heteropolymers co-exist during multicellular development in the filamentous fungus Aspergillus nidulans. PLoS ONE. 2014, 9, e92819. [Google Scholar] [CrossRef]
- Tian, T.; Liu, Y.; Yan, H.; You, Q.; Yi, X.; Du, Z.; Xu, W.; Su, Z. agriGO v2.0: A GO analysis toolkit for the agricultural community, 2017 update. Nucleic Acids Res. 2017, 45, W122–W129. [Google Scholar] [CrossRef] [PubMed]
- Leipe, D.D.; Koonin, E.V.; Aravind, L. Evolution and classification of P-loop kinases and related proteins. J. Mol. Biol. 2003, 333, 781–815. [Google Scholar] [CrossRef]
- Milner-White, E.J.; Coggins, J.R.; Anton, I.A. Evidence for an ancestral core structure in nucleotide-binding proteins with the type A motif. J. Mol. Biol. 1991, 221, 751–754. [Google Scholar] [CrossRef] [PubMed]
- Gangwar, D.; Kalita, M.K.; Gupta, D.; Chauhan, V.S.; Mohmmed, A. A systematic classification of Plasmodium falciparum P-loop NTPases: Structural and functional correlation. Malar. J. 2009, 8, 69. [Google Scholar] [CrossRef]
- Wu, M.Y.; Mead, M.E.; Kim, S.C.; Rokas, A.; Yu, J.H. WetA bridges cellular and chemical development in Aspergillus flavus. PLoS ONE 2017, 12, e0179571. [Google Scholar] [CrossRef] [PubMed]
- Adams, T.H.; Boylan, M.T.; Timberlake, W.E. brlA is necessary and sufficient to direct conidiophore development in Aspergillus nidulans. Cell 1988, 54, 353–362. [Google Scholar] [CrossRef]
- Mead, M.E.; Borowsky, A.T.; Joehnk, B.; Steenwyk, J.L.; Shen, X.X.; Sil, A.; Rokas, A. Recurrent Loss of abaA, a Master Regulator of Asexual Development in Filamentous Fungi, Correlates with Changes in Genomic and Morphological Traits. Genome Biol. Evol. 2020, 12, 1119–1130. [Google Scholar] [CrossRef]
- Cho, H.J.; Son, S.H.; Chen, W.; Son, Y.E.; Lee, I.; Yu, J.H.; Park, H.S. Regulation of Conidiogenesis in Aspergillus flavus. Cells 2022, 11, 2796. [Google Scholar] [CrossRef] [PubMed]
- Wu, M.Y.; Mead, M.E.; Lee, M.K.; Ostrem Loss, E.M.; Kim, S.C.; Rokas, A.; Yu, J.H. Systematic Dissection of the Evolutionarily Conserved WetA Developmental Regulator across a Genus of Filamentous Fungi. mBio 2018, 9, e01130-18. [Google Scholar] [CrossRef]
- Cary, J.W.; Harris-Coward, P.Y.; Ehrlich, K.C.; Mack, B.M.; Kale, S.P.; Larey, C.; Calvo, A.M. NsdC and NsdD affect Aspergillus flavus morphogenesis and aflatoxin production. Eukaryot. Cell. 2012, 11, 1104–1111. [Google Scholar] [CrossRef]
- Chang, P.K.; Scharfenstein, L.L.; Li, R.W.; Arroyo-Manzanares, N.; De Saeger, S.; Diana Di Mavungu, J. Aspergillus flavus aswA, a gene homolog of Aspergillus nidulans oefC, regulates sclerotial development and biosynthesis of sclerotium-associated secondary metabolites. Fungal Genet. Biol. 2017, 104, 29–37. [Google Scholar] [CrossRef] [PubMed]
- Wada, R.; Jin, F.J.; Koyama, Y.; Maruyama, J.; Kitamoto, K. Efficient formation of heterokaryotic sclerotia in the filamentous fungus Aspergillus oryzae. Appl. Microbiol. Biotechnol. 2014, 98, 325–334. [Google Scholar] [CrossRef] [PubMed]
- Kale, S.P.; Milde, L.; Trapp, M.K.; Frisvad, J.C.; Keller, N.P.; Bok, J.W. Requirement of LaeA for secondary metabolism and sclerotial production in Aspergillus flavus. Fungal Genet. Biol. 2008, 45, 1422–1429. [Google Scholar] [CrossRef] [PubMed]
- Wang, P.; Xu, J.; Chang, P.K.; Liu, Z.; Kong, Q. New Insights of Transcriptional Regulator AflR in Aspergillus flavus Physiology. Microbiol. Spectr. 2022, 10, e0079121. [Google Scholar] [CrossRef]
- Chang, P.K.; Ehrlich, K.C.; Yu, J.; Bhatnagar, D.; Cleveland, T.E. Increased expression of Aspergillus parasiticus aflR, encoding a sequence-specific DNA-binding protein, relieves nitrate inhibition of aflatoxin biosynthesis. Appl. Environ. Microbiol. 1995, 61, 2372–2377. [Google Scholar] [CrossRef]
- Trail, F.; Mahanti, N.; Rarick, M.; Mehigh, R.; Liang, S.H.; Zhou, R.; Linz, J.E. Physical and transcriptional map of an aflatoxin gene cluster in Aspergillus parasiticus and functional disruption of a gene involved early in the aflatoxin pathway. Appl. Environ. Microbiol. 1995, 61, 2665–2673. [Google Scholar] [CrossRef] [PubMed]
- Chang, P.K.; Cary, J.W.; Yu, J.; Bhatnagar, D.; Cleveland, T.E. The Aspergillus parasiticus polyketide synthase gene pksA, a homolog of Aspergillus nidulans wA, is required for aflatoxin B1 biosynthesis. Mol. Gen. Genet. 1995, 248, 270–277. [Google Scholar] [CrossRef]
- Motomura, M.; Chihaya, N.; Shinozawa, T.; Hamasaki, T.; Yabe, K. Cloning and characterization of the O-methyltransferase I gene (dmtA) from Aspergillus parasiticus associated with the conversions of demethylsterigmatocystin to sterigmatocystin and dihydrodemethylsterigmatocystin to dihydrosterigmatocystin in aflatoxin biosynthesis. Appl. Environ. Microbiol. 1999, 65, 4987–4994. [Google Scholar] [PubMed]
- Yu, J.; Woloshuk, C.P.; Bhatnagar, D.; Cleveland, T.E. Cloning and characterization of avfA and omtB genes involved in aflatoxin biosynthesis in three Aspergillus species. Gene 2000, 248, 157–167. [Google Scholar] [CrossRef]
- Kong, Q.; Chang, P.K.; Li, C.; Hu, Z.; Zheng, M.; Sun, Q.; Shan, S. Identification of AflR Binding Sites in the Genome of Aspergillus flavus by ChIP-Seq. J. Fungi 2020, 6, 52. [Google Scholar] [CrossRef]
- He, Z.M.; Price, M.S.; Obrian, G.R.; Georgianna, D.R.; Payne, G.A. Improved protocols for functional analysis in the pathogenic fungus Aspergillus flavus. BMC Microbiol. 2007, 7, 104. [Google Scholar] [CrossRef] [PubMed]





| Strains | Description |
|---|---|
| WT | A. flavus NRRL3357 wild type |
| TJES19.1 | ΔKu70 and ΔpyrG |
| ΔvepN | ΔKu70, ΔvepN, and pyrG+ |
| OE::vepN | ΔKu70, ΔpyrG, vepN+, gpdA+, and SH ble+ |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Xu, J.; Jiang, M.; Wang, P.; Kong, Q. The Gene vepN Regulated by Global Regulatory Factor veA That Affects Aflatoxin Production, Morphological Development and Pathogenicity in Aspergillus flavus. Toxins 2024, 16, 174. https://doi.org/10.3390/toxins16040174
Xu J, Jiang M, Wang P, Kong Q. The Gene vepN Regulated by Global Regulatory Factor veA That Affects Aflatoxin Production, Morphological Development and Pathogenicity in Aspergillus flavus. Toxins. 2024; 16(4):174. https://doi.org/10.3390/toxins16040174
Chicago/Turabian StyleXu, Jia, Mengqi Jiang, Peng Wang, and Qing Kong. 2024. "The Gene vepN Regulated by Global Regulatory Factor veA That Affects Aflatoxin Production, Morphological Development and Pathogenicity in Aspergillus flavus" Toxins 16, no. 4: 174. https://doi.org/10.3390/toxins16040174





