A Novel Transcription Factor VPA0041 Was Identified to Regulate the Swarming Motility in Vibrio parahaemolyticus
Abstract
:1. Introduction
2. Results
2.1. VPA0041 Regulated the Swarming Motility of V. parahaemolyticus
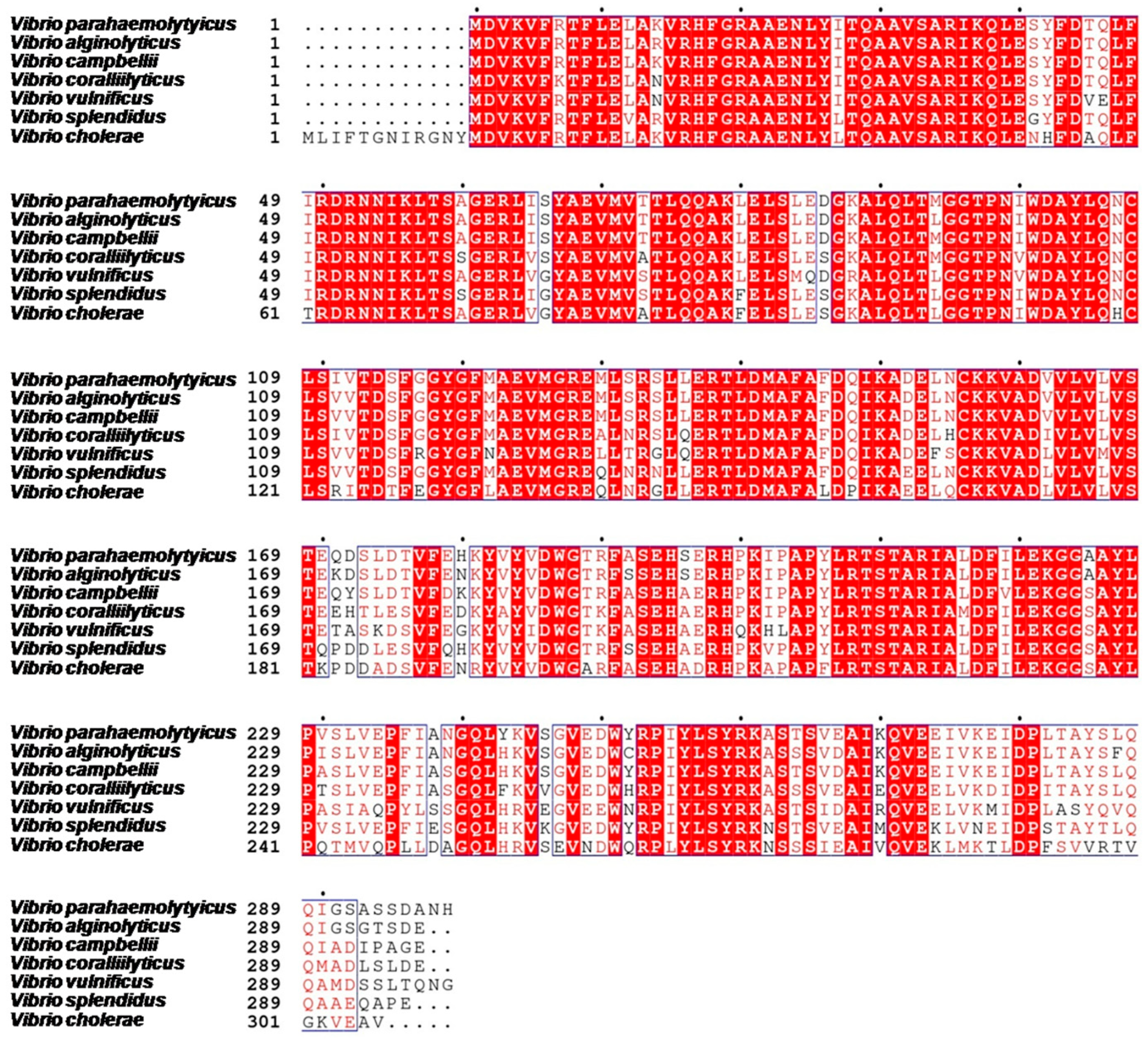
2.2. The Annotation of VPA0041 Protein in V. parahaemolyticus
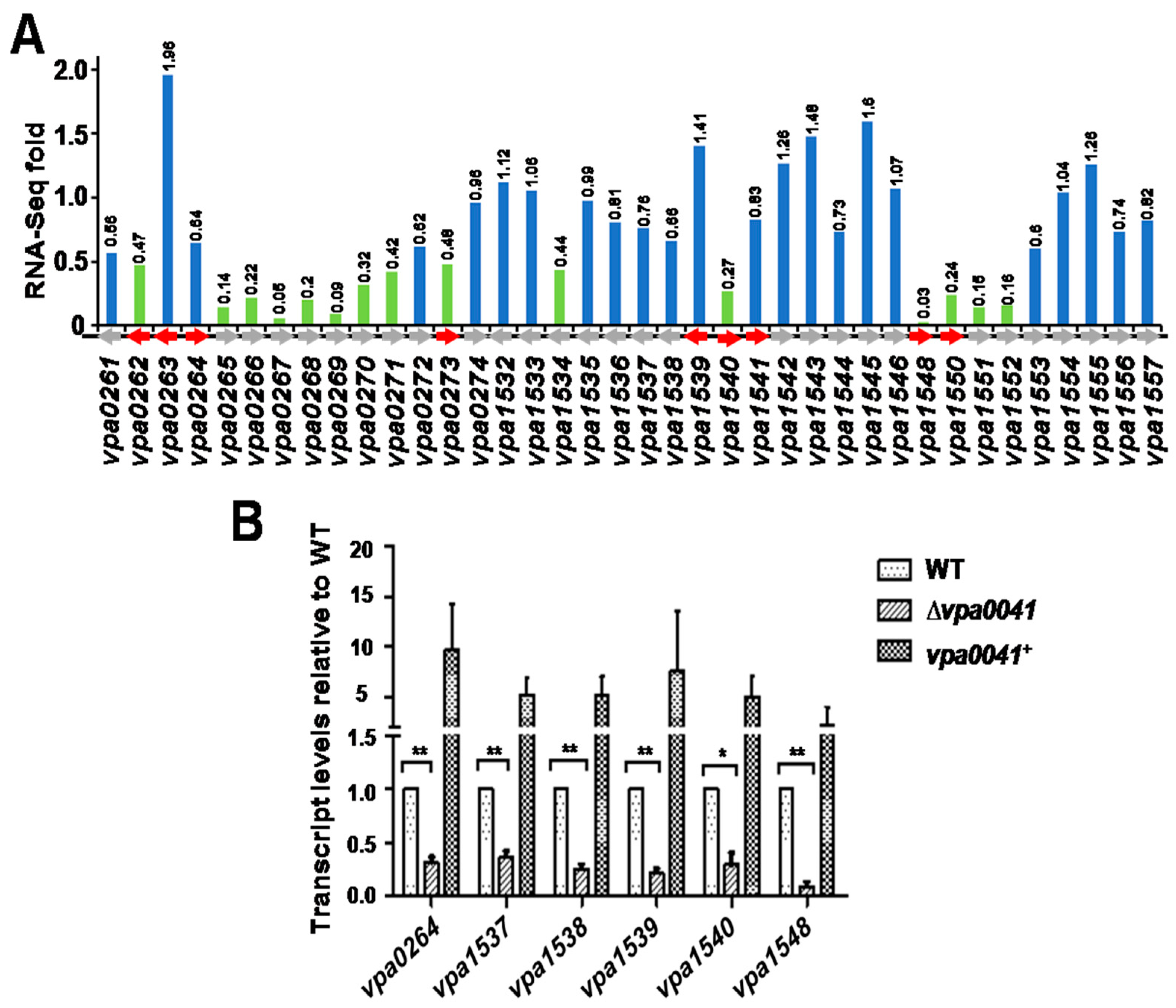
2.3. The Global Transcriptional Analysis of VPA0041 in V. parahaemolyticus
2.4. VPA0041 Regulates the Expression of the Lateral Flagellar Genes
2.5. VPA0041 Directly Bound to the Promoters of vpa0264, vpa1548, and vpa1550 to Activate the Lateral Flagellar System
3. Discussion
4. Materials and Methods
4.1. Bacterial Strains, Plasmids, and Growth Conditions
4.2. Construction of vpa0041 Mutant and Complemented Strains
4.3. The Motility Analysis
4.4. The Transmission Electron Microscope Analysis
4.5. qRT-PCR Analysis of the Lateral Flagellar Genes
4.6. RNA-Seq Analysis
4.7. Purification of VPA0041 Protein
4.8. Electrophoretic Mobility Shift Assay
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Austin, B. Vibrios as causal agents of zoonoses. Vet. Microbiol. 2010, 140, 310–317. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Johnson, C.N.; Bowers, J.C.; Griffitt, K.J.; Molina, V.; Clostio, R.W.; Pei, S. Ecology of Vibrio parahaemolyticus and Vibrio vulnificus in the coastal and estuarine waters of Louisiana, Maryland, Mississippi, and Washington (United States). Appl. Environ. Microbiol. 2012, 78, 7249–7257. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yang, C.; Zhang, X.; Fan, H.; Li, Y.; Hu, Q.; Yang, R.; Cui, Y. Genetic diversity, virulence factors and X farm-to-table spread pattern of Vibrio parahaemolyticus food-associated isolates. Food Microbiol. 2019, 84, 103270. [Google Scholar] [CrossRef] [PubMed]
- Chen, H.; Dong, S.; Yan, Y.; Zhan, L.; Zhang, J.; Chen, J.; Zhang, Z.; Zhang, Y.; Mei, L. Prevalence and Population Analysis of Vibrio parahaemolyticus Isolated from Freshwater Fish in Zhejiang Province, China. Foodborne Pathog. Dis. 2021, 18, 139–146. [Google Scholar] [CrossRef]
- Sar, N.; MrCarter, L.; Simon, M.; Silverman, M. Chemotactic control of the two flagellar systems of Vibrio parahaemolyticus. J. Bacteriol. 1990, 172, 334–341. [Google Scholar] [CrossRef] [Green Version]
- Harshey, R.M. Bacterial motility on a surface: Many ways to a common goal. Annu. Rev. Microbiol. 2003, 57, 249–273. [Google Scholar] [CrossRef]
- Freitas, C.; Glatter, T.; Ringgaard, S. The release of a distinct cell type from swarm colonies facilitates dissemination of Vibrio parahaemolyticus in the environment. ISME J. 2020, 14, 230–244. [Google Scholar] [CrossRef]
- McCarter, L.; Silverman, M. Surface-induced swarmer cell differentiation of Vibrio parahaemolyticus. Mol. Microbiol. 1990, 4, 1057–1062. [Google Scholar] [CrossRef]
- McCarter, L.L. Dual flagellar systems enable motility under different circumstances. J. Mol. Microbiol. Biotechnol. 2004, 7, 18–29. [Google Scholar] [CrossRef]
- Makino, K.; Oshima, K.; Kurokawa, K.; Yokoyama, K.; Uda, T.; Tagomori, K.; Iijima, Y.; Najima, M.; Nakano, M.; Yamashita, A.; et al. Genome sequence of Vibrio parahaemolyticus: A pathogenic mechanism distinct from that of Vibrio cholerae. Lancet 2003, 361, 743–749. [Google Scholar] [CrossRef]
- McCarter, L.L. Bacterial acrobatics on a surface: Swirling packs, collisions, and reversals during swarming. J. Bacteriol. 2010, 192, 3246–3248. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gode-Potratz, C.J.; Kustusch, R.J.; Breheny, P.J.; Weiss, D.S.; McCarter, L.L. Surface sensing in Vibrio parahaemolyticus triggers a programme of gene expression that promotes colonization and virulence. Mol. Microbiol. 2011, 79, 240–263. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Merino, S.; Shaw, J.G.; Tomás, J.M. Bacterial lateral flagella: An inducible flagella system. FEMS. Microbiol. Lett. 2016, 263, 127–135. [Google Scholar] [CrossRef] [PubMed]
- Kearns, D.B. A field guide to bacterial swarming motility. Nat. Rev. Microbiol. 2010, 8, 634–644. [Google Scholar] [CrossRef] [Green Version]
- Murray, T.S.; Ledizet, M.; Kazmierczak, B. Swarming motility, secretion of type 3 effectors and biofilm formation phenotypes exhibited within a large cohort of Pseudomonas aeruginosa clinical isolates. J. Med. Microbiol. 2010, 59, 511–520. [Google Scholar] [CrossRef]
- Khan, F.; Tabassum, N.; Anand, R.; Kim, Y.M. Motility of Vibrio spp.: Regulation and controlling strategies. Appl. Microbiol. Biotechnol. 2020, 104, 8187–8208. [Google Scholar] [CrossRef]
- McCarter, L.; Silverman, M. Iron regulation of swarmer cell differentiation of Vibrio parahaemolyticus. J. Bacteriol. 1989, 171, 731–736. [Google Scholar] [CrossRef] [Green Version]
- Kawagishi, I.; Imagawa, M.; Imae, Y.; McCarter, L.L.; Homma, M. The sodium-driven polar flagellar motor of marine Vibrio as the mechanosensor that regulates lateral flagellar expression. Mol. Microbiol. 1996, 20, 693–699. [Google Scholar] [CrossRef]
- Gode-Potratz, C.J.; Chodur, M.D.; McCarter, L.L. Calcium and iron regulate swarming and type III secretion in Vibrio parahaemolyticus. J. Bacteriol. 2010, 192, 6025–6038. [Google Scholar] [CrossRef] [Green Version]
- Jaques, S.; McCarter, L.L. Three new regulators of swarming in Vibrio parahaemolyticus. J. Bacteriol. 2006, 188, 2625–2635. [Google Scholar] [CrossRef] [Green Version]
- Gu, D.; Meng, H.; Li, Y.; Ge, H.; Jiao, X. A GntR family transcription factor (VPA1701) for swarming motility and colonization of Vibrio parahaemolyticus. Pathogens 2019, 8, 235. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhang, Y.; Liu, H.; Gu, D.; Lu, X.; Zhou, X.; Xia, X. Transcriptomic analysis of PhoR reveals its role in regulation of swarming motility and T3SS expression in Vibrio parahaemolyticus. Microbiol. Res. 2020, 235, 126448. [Google Scholar] [CrossRef] [PubMed]
- Hubbard, T.P.; Chao, M.C.; Abel, S.; Blondel, C.J.; Abel Zur Wiesch, P.; Zhou, X.; Davis, B.M.; Waldor, M.K. Genetic analysis of Vibrio parahaemolyticus intestinal colonization. Proc. Natl. Acad. Sci. USA 2016, 113, 6283–6288. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhang, N.; Zhang, S.; Ren, W.; Gong, X.; Long, H.; Zhang, X.; Cai, X.; Huang, A.; Xie, Z. Roles of rpoN in biofilm formation of Vibrio alginolyticus HN08155 at different cell densities. Microbiol. Res. 2021, 247, 126728. [Google Scholar] [CrossRef] [PubMed]
- Kim, Y.K.; McCarter, L.L. ScrG, a GGDEF-EAL protein, participates in regulating swarming and sticking in Vibrio parahaemolyticus. J. Bacteriol. 2007, 189, 4094–4107. [Google Scholar] [CrossRef] [Green Version]
- Ferreira, R.B.; Antunes, L.C.; Greenberg, E.P.; McCarter, L.L. Vibrio parahaemolyticus ScrC modulates cyclic dimeric GMP regulation of gene expression relevant to growth on surfaces. J. Bacteriol. 2008, 190, 851–860. [Google Scholar] [CrossRef] [Green Version]
- Phan, N.Q.; Uebanso, T.; Shimohata, T.; Nakahashi, M.; Mawatari, K.; Takahashi, A. DNA-binding protein HU coordinates pathogenicity in Vibrio parahaemolyticus. J. Bacteriol. 2015, 197, 2958–2964. [Google Scholar] [CrossRef] [Green Version]
- Chung, C.; Fen, S.; Yu, S.; Wong, H. Influence of oxyR on growth, biofilm formation, and mobility of Vibrio parahaemolyticus. Appl. Environ. Microbiol. 2016, 82, 788–796. [Google Scholar] [CrossRef] [Green Version]
- Chen, L.; Qiu, Y.; Tang, H.; Hu, L.; Yang, W.; Zhu, X.; Huang, X.; Wang, T.; Zhang, Y. ToxR is required for biofilm formation and motility of Vibrio parahaemolyticus. Biomed. Environ. Sci. 2018, 31, 848–850. [Google Scholar]
- Majander, F.; Korhonen, T.K.; Westerlund-Wikström, B. Simultaneous display of multiple foreign peptides in the FliD capping and FliC filament proteins of the Escherichia coli flagellum. Appl. Environ. Microbiol. 2005, 71, 4263–4268. [Google Scholar] [CrossRef] [Green Version]
- Maddocks, S.E.; Oyston, P.C.F. Structure and function of the LysR-type transcriptional regulator (LTTR) family proteins. Microbiology 2008, 154, 3609–3623. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rashid, M.M.; Ikawa, Y.; Tsuge, S. GamR, the LysR-type galactose metabolism regulator, regulates hrp gene expression via transcriptional activation of two key hrp regulators, HrpG and HrpX, in Xanthomonas oryzae pv. oryzae. Appl. Environ. Microbiol. 2016, 82, 3947–3958. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gao, X.; Wang, X.; Mao, Q.; Xu, R.; Zhou, X.; Ma, Y.; Liu, Q.; Zhang, Y.; Wang, Q. VqsA, a novel LysR-type transcriptional regulator, coordinates quorum sensing (QS) and is controlled by QS to regulate virulence in the pathogen Vibrio alginolyticus. Appl. Environ. Microbiol. 2018, 84, e00444-18. [Google Scholar] [CrossRef] [Green Version]
- Liu, Y.; Liu, B.; Yang, P.; Wang, T.; Chang, Z.; Wang, J.; Wang, Q.; Li, W.; Wu, J.; Huang, D.; et al. LysR-type transcriptional regulator OvrB encoded in O island 9 drives enterohemorrhagic Escherichia coli O157:H7 virulence. Virulence 2019, 10, 783–792. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ko, M.; Park, C. H-NS-Dependent regulation of flagellar synthesis is mediated by a LysR family protein. J. Bacteriol. 2000, 182, 4670–4672. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lehnen, D.; Blumer, C.; Polen, T.; Wackwitz, B.; Wendisch, V.F.; Unden, G. LrhA as a new transcriptional key regulator of flagella, motility and chemotaxis genes in Escherichia coli. Mol. Microbiol. 2002, 45, 521–532. [Google Scholar] [CrossRef]
- Heroven, A.K.; Dersch, P. RovM, a novel LysR-type regulator of the virulence activator gene rovA, controls cell invasion, virulence and motility of Yersinia pseudotuberculosis. Mol. Microbiol. 2006, 62, 1469–1483. [Google Scholar] [CrossRef]
- Harris, S.J.; Shih, Y.L.; Bentley, S.D.; Salmond, G.P. The hexA gene of Erwinia carotovora encodes a LysR homologue and regulates motility and the expression of multiple virulence determinants. Mol. Microbiol. 1998, 28, 705–717. [Google Scholar] [CrossRef] [Green Version]
- Zhang, M.; Kang, Z.; Guo, X.; Guo, S.; Xiao, D.; Liu, Y.; Ma, C.; Gao, C.; Xu, P. Regulation of glutarate catapolism by GntR family regulator CsiR and LysR family regulator GcdR in Pseudomonas putida KT2440. mBio 2019, 10, e01570-19. [Google Scholar] [CrossRef] [Green Version]
- Erwin, D.P.; Nydam, D.S.; Call, D.R. Vibrio parahaemolyticus ExsE is requisite for initial adhesion and subsequent type III secretion system1-dependent autophagy in HeLa cells. Microbiology 2012, 158, 2303–2314. [Google Scholar] [CrossRef] [Green Version]
- Cai, Q.; Zhang, Y. Structure, function and regulation of the thermostable direct hemolysin (TDH) in pandemic Vibrio parahaemolyticus. Microb. Pathog. 2018, 123, 242–245. [Google Scholar] [CrossRef] [PubMed]
- Ben-Yaakov, R.; Salomon, D. The regulatory network of Vibrio parahaemolyticus type VI secretion system 1. Environ. Microbiol. 2019, 21, 2248–2260. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Matsuda, S.; Okada, R.; Tandhavanant, S.; Hiyoshi, H.; Gotoh, K.; Iida, T.; Kodama, T. Export of a Vibrio parahaemolyticus toxin by the Sec and type III secretion machineries in tandem. Nat. Microbiol. 2019, 4, 781–788. [Google Scholar] [CrossRef]
- Liu, M.; Zhu, X.; Zhang, C.; Zhao, Z. LuxQ-LuxU-LuxO pathway regulates biofilm formation by Vibrio parahaemolyticus. Microbiol. Res. 2021, 250, 126791. [Google Scholar] [CrossRef] [PubMed]
- Lu, R.; Tang, H.; Qiu, Y.; Yang, W.; Yang, H.; Zhou, D.; Huang, X.; Hu, L.; Zhang, Y. Quorum sensing regulates the transcription of lateral flagellar genes in Vibrio parahaemolyticus. Future Microbiol. 2019, 14, 1043–1053. [Google Scholar] [CrossRef]
- Yin, W.; Zhang, N.; Xu, H.; Gong, X.; Long, H.; Ren, W.; Zhang, X.; Cai, X.; Huang, A.; Xie, Z. Stress adaptation and virulence in Vibrio alginolyticus is mediated by two (p)ppGpp synthetase genes, relA and spoT. Microbiol. Res. 2021, 253, 126883. [Google Scholar] [CrossRef]
- Mea, H.J.; Yong, P.V.C.; Wong, E.H. An overview of Acinetobacter baumannii pathogenesis: Motility, adherence and biofilm formation. Microbiol. Res. 2021, 247, 126722. [Google Scholar] [CrossRef]
- Gautam, L.K.; Sharma, P.; Capalash, N. Attenuation of Acinetobacter baumannii virulence by inhibition of polyphosphate kinase 1 with repurposed drugs. Microbiol. Res. 2021, 242, 126627. [Google Scholar] [CrossRef]
- Brescia, F.; Marchetti-Deschmann, M.; Musetti, R.; Perazzolli, M.; Pertot, I.; Puopolo, G. The rhizosphere signature on the cell motility, biofilm formation and secondary metabolite production of a plant-associated Lysobacter strain. Microbiol. Res. 2020, 234, 126424. [Google Scholar] [CrossRef]
- Salomon, D.; Gonzalez, H.; Updegraff, B.L.; Orth, K. Vibrio parahaemolyticus type VI secretion system 1 is activated in marine conditions to target bacteria, and is differentially regulated from system 2. PLoS ONE 2013, 8, e61086. [Google Scholar] [CrossRef]
- Calder, T.; de Souza Santos, M.; Attah, V.; Klimko, J.; Fernandez, J.; Salomon, D.; Krachler, A.M.; Orth, K. Structural and regulatory mutations in Vibrio parahaemolyticus type III secretion system display variable effects in virulence. FEMS Microbiol. Lett. 2014, 361, 107–114. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lin, I.T.; Tulman, E.R.; Geary, S.J.; Zhou, X. A gatekeeper protein contributes to T3SS2 function via interaction with an ATPase in Vibrio parahaemolyticus. Microbiol. Res. 2021, 252, 126857. [Google Scholar] [CrossRef] [PubMed]
- Wang, Q.; Millet, Y.A.; Chao, M.C.; Sasabe, J.; Davis, B.M.; Waldor, M.K. A genome-wide screen reveals that the Vibrio cholerae phosphoenolpyruvate phosphotransferase system modulates virulence gene expression. Infect. Immun. 2015, 83, 3381–3395. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhou, X.; Konkel, M.E.; Call, D.R. Vp1659 is a Vibrio parahaemolyticus type III secretion system 1 protein that contributes to translocation of effector proteins needed to induce cytolysis, autophagy, and disruption of actin structure in HeLa cells. J. Bacteriol. 2010, 192, 3491–3502. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Liang, W.; Wang, S.; Yu, F.; Zhang, L.; Qi, G.; Liu, Y.; Gao, S.; Kan, B. Construction and evaluation of a safe, live, oral Vibrio cholerae vaccine candidate, IEM108. Infect. Immun. 2003, 71, 5498–5504. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Morales, V.M.; Bäckman, A.; Bagdasarian, M. A series of wide-host range low-copy-number vectors that allow direct screening for recombinants. Gene 1991, 97, 39–47. [Google Scholar] [CrossRef]
- Wang, S.Y.; Lauritz, J.; Jass, J.; Milton, D.L. A ToxR homolog from Vibrio anguillarum serotype O1 regulates its own production, bile resistance, and biofilm formation. J. Bacteriol. 2002, 184, 1630–1639. [Google Scholar] [CrossRef] [Green Version]
- Tjaden, B. De novo assembly of bacterial transcriptomes from RNA-seq data. Genome Biol. 2015, 16, 1. [Google Scholar] [CrossRef] [Green Version]
- McClure, R.; Balasubramanian, D.; Sun, Y.; Bobrovskyy, M.; Sumby, P.; Genco, C.A.; Vanderpool, C.K.; Tjaden, B. Computational analysis of bacterial RNA-seq data. Nucleic Acids Res. 2013, 41, e140. [Google Scholar] [CrossRef] [Green Version]
- Kanehisa, M.; Goto, S. KEGG: Kyoto Encyclopedia of Genes and Genomes. Nucleic Acids Res. 2000, 28, 27–30. [Google Scholar] [CrossRef]
- Gu, D.; Guo, M.; Yang, M.; Zhang, Y.; Zhou, X.; Wang, Q. A σE-mediated temperature gauge controls a switch from LuxR-mediated virulence gene expression to thermal stress adaptation in Vibrio alginolyticus. PLoS Pathog. 2016, 12, e1005645. [Google Scholar] [CrossRef] [PubMed] [Green Version]




| Strain or Plasmid | Relevant Characteristics | Reference |
|---|---|---|
| E. coli | ||
| DH5α λpir | Host for π requiring plasmids | [53] |
| SM10 λpir | Host for π requiring plasmids, conjugal donor | [54] |
| BL21(DE3) | Host strain for protein expression | Novagen |
| V. parahaemolyticus | ||
| RIMD 2210633 | Clinical isolate. Carbr | [23] |
| ∆vpa0041 | RIMD 2210633, in-frame deletion in vpa0041, Carbr | This study |
| vpa0041+ | ∆vpa0041, pMMB207 expressing the vpa0041-his gene, Carbr, Cmr | This study |
| Plasmids | ||
| pDM4 | Suicide vector, pir dependent, R6K, SacBR, Cmr | [55] |
| pMMB207 | IncQ lacIq Δbla Ptac-lac lacZa, Cmr | [56] |
| pET30a | Expressing vector, Kmr | Novagen |
| vpa0041::pDM4 | vpa0041 up and down sequences clones into pDM4, Cmr | This study |
| vpa0041::pMMB207 | RBS and vpa0041-his sequences clones into pMMB207, Cmr | This study |
| vpa0041::pET30a | vpa0041 ORF clones into pET30a, Kmr | This study |
| Primer Name | Primer Sequence (5′ to 3′) | Target |
|---|---|---|
| vpa0041-up-F | GATAACAATTTGTGGAATCCCGGGAAGAAGAAATGGGTCAGAAGCGTT | vpa0041 deletion mutant |
| vpa0041-up-R | CGTCCGATGACTTAACATCCATAAACCAACTCCTG | vpa0041 deletion mutant |
| vpa0041-down-F | GGATGTTAAGTCATCGGACGCAAACCATTAAC | vpa0041 deletion mutant |
| vpa0041-down-R | AGTGTATATCAAGCTTATCGATACCCGCACAAAGACAGTGAAGGCAAT | vpa0041 deletion mutant |
| vpa0041-out-F | TTGTTGCGACCGTTTAGCGTATG | vpa0041 deletion mutant |
| vpa0041-out-R | CGCACAAAGACAGTGAAGGCAAT | vpa0041 deletion mutant |
| vpa0041-in-F | TACGCAAGCAGCGGTCAGTG | vpa0041 deletion mutant |
| vpa0041-in-R | GCTCAGAAGCGAAGCGTGTT | vpa0041 deletion mutant |
| pDM4-F | GGTGCTCCAGTGGCTTCTGTTTCTA | vpa0041 deletion mutant |
| pDM4-R | CAGCAACTTAAATAGCCTCTAAT | vpa0041 deletion mutant |
| vpa0041-F | CCGCGAGCTCTAAGGAGGTAGGATAATAATGGATGTTAAGGTCTTTAGAACAT | vpa0041+ |
| vpa0041-R | GCGTCGACTTAGTGATGATGATGATGATGATGGTTTGCGTCCGATGATGC | vpa0041+ |
| pMMB207-F | CTCCCGTTCTGGATAATGTTTTTTG | vpa0041+ |
| pMMB207-R | TCTTCTCTCATCCGCCAAAACAGCC | vpa0041+ |
| vpa0041-his-F | GGAATTCCATATGATGGATGTTAAGGTCTTTAGAAC | VPA0041 expression |
| vpa0041-his-R | GGAATTCCATATGATGGATGTTAAGGTCTTTAGAAC | VPA0041 expression |
| pET30a-F | TAGTTATTGCTCAGCGGTGGC | VPA0041 expression |
| pET30a-R | ACGATGCGTCCGGCGTAGAG | VPA0041 expression |
| gyrb-RT-F | TTACCGTCATGGTGAGCCTG | qRT-PCR |
| gyrb-RT-R | CACGCAGACGTTTTGCTAGG | qRT-PCR |
| vpa0264-RT-F | GCAGGTTCAGGCCCAGTATT | qRT-PCR |
| vpa0264-RT-R | TCATGTTGAGAAACGTCAGGCT | qRT-PCR |
| vpa1537-RT-F | CGCTTGAGAAAACGACAGTGG | qRT-PCR |
| vpa1537-RT-R | CCTACTAATGCGGTCTCGGC | qRT-PCR |
| vpa1538-RT-F | CACGTACGCACATATCCGGT | qRT-PCR |
| vpa1538-RT-R | ACGAACACCTTGCTCAACCT | qRT-PCR |
| vpa1539-RT-F | ATTAGTGAGGGTGCGCCTTT | qRT-PCR |
| vpa1539-RT-R | GGTGAAGGGAAGGAATGGCA | qRT-PCR |
| vpa1540-RT-F | CAACGCCAGTTCGTCTTAACG | qRT-PCR |
| vpa1540-RT-R | ACGGCCAGTAAAGAGAGGTTG | qRT-PCR |
| vpa1548-RT-F | GCTGGTGGCCTTATCGAAGA | qRT-PCR |
| vpa1548-RT-R | TACTGCGAAGTCTGCATCCAT | qRT-PCR |
| FAM-F | FAM-TGCCTGCAGGTCGACGAT | EMSA |
| gyrB-EMSA-F | TGCCTGCAGGTCGACGATTGCCTGCAGGTCGACGATGCGCGCG | EMSA |
| gyrB-EMSA-R | TGCCAGCGCACCGCTGACCGCAG | EMSA |
| vpa0264-EMSA-1F | TGCCTGCAGGTCGACGATGTTTGTCCTGTCGAAAGAATTC | EMSA |
| vpa0264-EMSA-1R | CATGCATCTTTCCTTACAGTCGGCT | EMSA |
| vpa0264-EMSA-2F | TGCCTGCAGGTCGACGATTAGAGTTTTCCCCCTAATTTT | EMSA |
| vpa0264-EMSA-3F | TGCCTGCAGGTCGACGATTCCACTCTTGTTTGTAAGTCAT | EMSA |
| vpa0264-EMSA-4F | TGCCTGCAGGTCGACGATGTATCTTGTTTGTATCTTGGCG | EMSA |
| vpa1548-EMSA-1F | TGCCTGCAGGTCGACGATGCCGATCAAAGCACATCGGAA | EMSA |
| vpa1548-EMSA-1R | CATCTTAGTCTCCTTAGTTTATCAC | EMSA |
| vpa1548-EMSA-2F | TGCCTGCAGGTCGACGATCTTAGTGGAATGCAAGTCACT | EMSA |
| vpa1548-EMSA-3F | TGCCTGCAGGTCGACGATAAATTTTTAATTTTCAAATTA | EMSA |
| vpa1548-EMSA-4F | TGCCTGCAGGTCGACGATAGTGACTAGGGAATATCCCAAG | EMSA |
| vpa1550-EMSA-1F | TGCCTGCAGGTCGACGATTTAGTCGCCTTGTCTTATAGGGA | EMSA |
| vpa1550-EMSA-1R | CACGAGCTTACTCCCTCTCTCATTG | EMSA |
| vpa1550-EMSA-2F | TGCCTGCAGGTCGACGATGGAACACATAAGGTGGAAAATAC | EMSA |
| vpa1550-EMSA-3F | TGCCTGCAGGTCGACGATCAAGTCAATGTTTTAAAAGAAT | EMSA |
| vpa1550-EMSA-4F | TGCCTGCAGGTCGACGATAGACATACTTTCAAGGCATAGAG | EMSA |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Li, M.; Meng, H.; Li, Y.; Gu, D. A Novel Transcription Factor VPA0041 Was Identified to Regulate the Swarming Motility in Vibrio parahaemolyticus. Pathogens 2022, 11, 453. https://doi.org/10.3390/pathogens11040453
Li M, Meng H, Li Y, Gu D. A Novel Transcription Factor VPA0041 Was Identified to Regulate the Swarming Motility in Vibrio parahaemolyticus. Pathogens. 2022; 11(4):453. https://doi.org/10.3390/pathogens11040453
Chicago/Turabian StyleLi, Mingzhu, Hongmei Meng, Yang Li, and Dan Gu. 2022. "A Novel Transcription Factor VPA0041 Was Identified to Regulate the Swarming Motility in Vibrio parahaemolyticus" Pathogens 11, no. 4: 453. https://doi.org/10.3390/pathogens11040453




