Synthesis, Spectroscopic Characterization, Molecular Docking and Biological Activity of Novel Secnidazole Metal Complexes
Abstract
:1. Introduction
2. Results and Discussion
2.1. Synthesis of Secnidazole Complexes
2.2. Elemental Analysis and Physical Properties
2.3. Conductance Measurements
2.4. Infrared Spectra
2.5. Electronic Absorption Spectra
2.6. H-NMR Spectroscopy
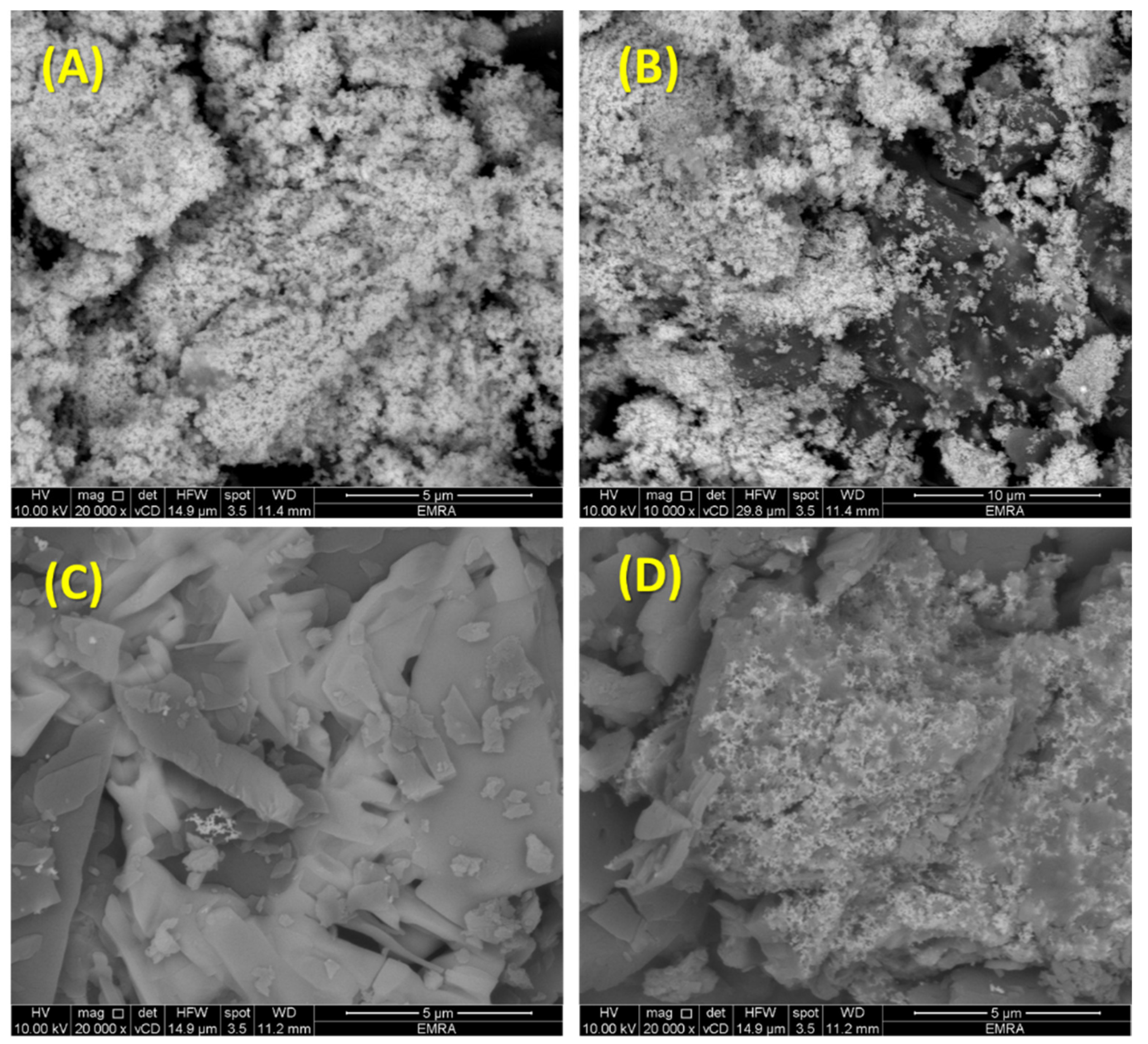
2.7. Scanning Electron Microscope (SEM)
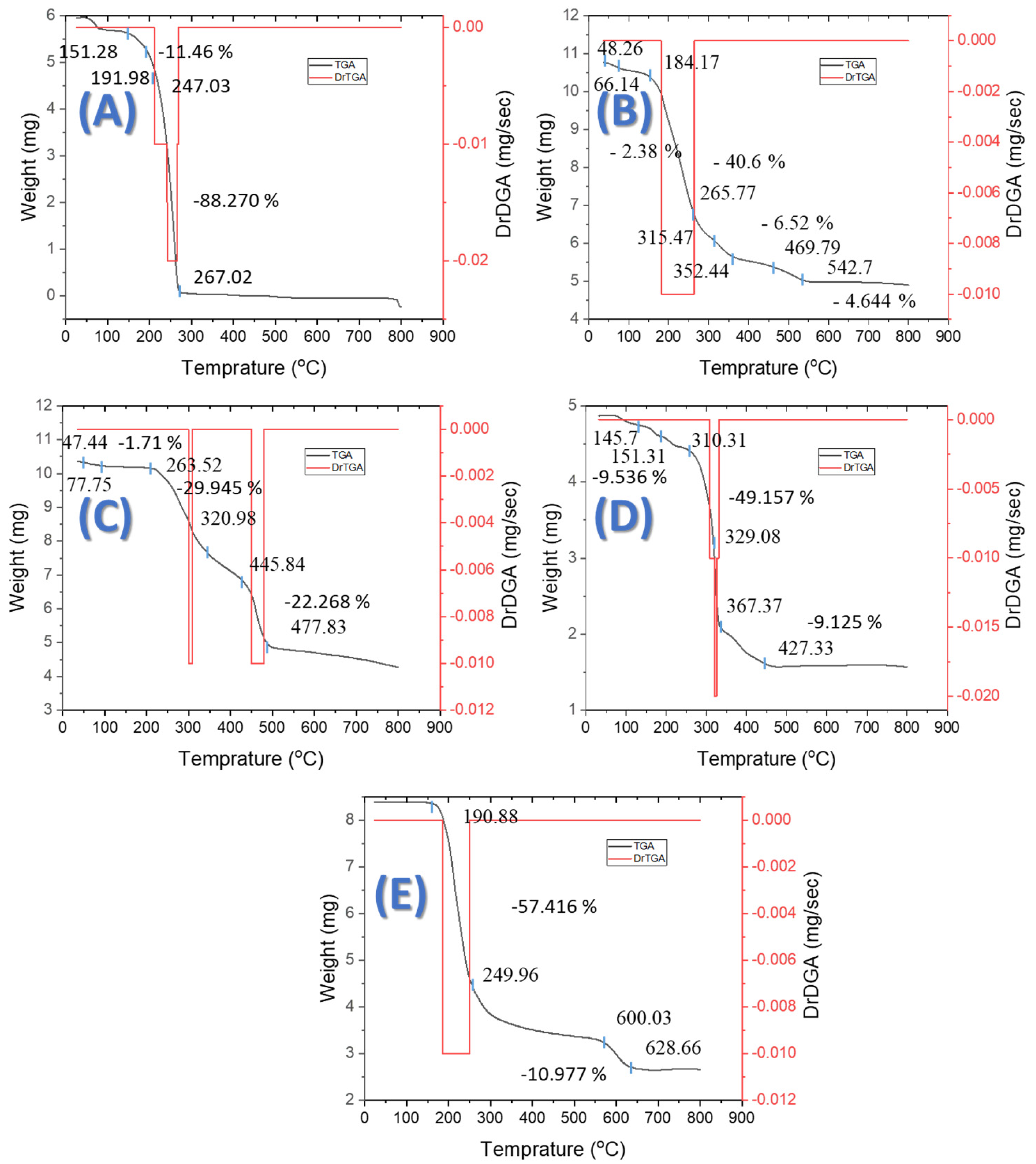
2.8. Thermogravimetric Analysis
3. Biological Activity
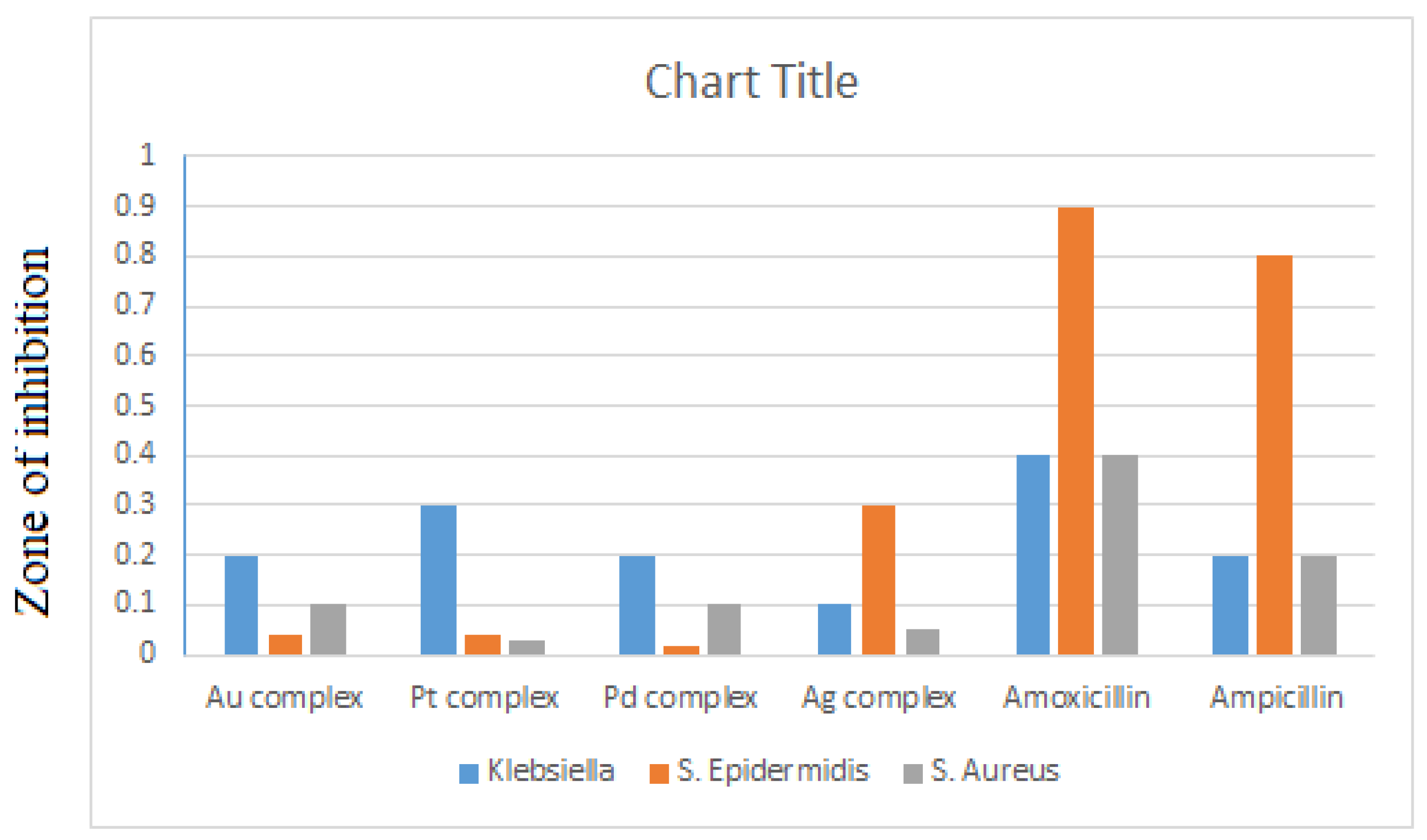
3.1. Antimicrobial Activity
3.2. Anticancer Activities
4. Docking Studies
5. Materials and Methods
5.1. Reagents
5.2. General Procedure for the Preparation of Secnidazole Complexes
5.2.1. [Au(sec)2Cl] (1)
5.2.2. [Pt(sec)2] (2)
5.2.3. [Pd(sec)2] (3)
5.2.4. [Ag(sec)] (4)
5.3. Spectroscopic Measurements
5.4. Antimicrobial Activity
5.5. Cytotoxicity Assay
6. Conclusions
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Ang, C.W.; Jarrad, A.M.; Cooper, M.A.; Blaskovich, M.A. Nitroimidazoles: Molecular fireworks that com-bat a broad spectrum of infectious diseases. J. Med. Chem. 2017, 60, 7636–7657. [Google Scholar] [CrossRef] [PubMed]
- Soledad, B.L.; Isabel, G.M.; Pilar, G.M.; Marcos, F.; Marcos, F.À; Noráh, B.B. Synthesis, characterization, and biological activity of cobalt(II), nickel(II), copper(II), and zinc(II) complexes of secnidazole. Inorg. Chim. Acta 2013, 397, 94–100. [Google Scholar]
- Ling, Z.; Xin, M.P.; Guri, L.V.D.; Rong, X.G.; Cheng, H.Z. Comprehensive review in current devel-opments of imidazole-based medicinal chemistry. Med. Res. Rev. 2014, 34, 340–437. [Google Scholar]
- Zhang, H.Z.; Gan, L.L.; Wang, H.; Zhou, C.H. New Progress in Azole Compounds as Antimicrobial Agents. Mini-Rev. Med. Chem. 2017, 17, 122–166. [Google Scholar] [CrossRef] [PubMed]
- Zhen, Z.L.; Lavanya, G.; Vijai, K.; Reddy, T.; Wei, W.G.; Cheng, H.Z. Discovery of novel nitroimid-azole enols as Pseudomonas aeruginosa DNA cleavage agents. Bioorg. Med. Chem. 2017, 25, 6511–6522. [Google Scholar]
- Huang, X.S.; Wang, L.S.; Yin, Y.; Li, W.M.; Duan, M.; Ran, W.; Zhu, H.L. Synthesis, characterization and bioactivity research of a derivative of secnidazole: 1-(2-chloropropyl)-2-methyl-5-nitro-1H-imidazole. J. Chem. Crystallogr. 2011, 41, 1360–1364. [Google Scholar] [CrossRef]
- López, S.H.; Londoño, L.M.E.; Garza, V.R.; Poblano, M.I.I.; Granada, M.P.; Gracia, M.I.; Barba, B.N. Synthesis, structure and biological activities of cobalt(II) and zinc(II) coordination compounds with 2-benzimidazole derivatives. J. Inorg. Biochem. 2008, 102, 1267. [Google Scholar] [CrossRef] [PubMed]
- Clarke, M.J.; Zhu, F.; Frasca, D.R. Non-platinum chemotherapeutic metallopharmaceuticals. Chem. Rev. 1999, 99, 2511. [Google Scholar] [CrossRef] [PubMed]
- Selwin, J.R.; Sivasankaran, N.M. Synthesis, characterization and antimicrobial activity of transition metal complexes with the schiff base derived from imidazole-2-carboxaldehyde and glycylglycine. J. Coord. Chem. 2009, 62, 319–327. [Google Scholar] [CrossRef]
- Geary, W.J. The use of conductivity measurements in organic solvents for the characterization of coor-dination compounds. Coord. Chem. Rev. 1971, 7, 81–122. [Google Scholar] [CrossRef]
- El-Bindary, A.A.; El-Desouky, M.G.; El-Afify, M.A.M. Thermal and Spectroscopic Studies of Some Prepared Metal Complexes and Investigation of their Potential Anticancer and Antiviral Drug Activity against SARS-CoV-2 by Molecular Docking Simulation. Biointerface Res. Appl. Chem. 2021, 12, 1053–1075. [Google Scholar]
- Bauer, A.W.; Kirby, W.A.; Sherris, C.; Turck, M. Antibiotic susceptibility testing by a standardized single disc method. Am. J. Clin. Pathol. 1996, 45, 493–496. [Google Scholar] [CrossRef]
- Repetto, G.; del Peso, A.; Zurita, J.L. Neutral red uptake assay for the estimation of cell viabil-ity/cytotoxicity. Nat. Protoc. 2008, 3, 1125–1131. [Google Scholar] [CrossRef] [PubMed]
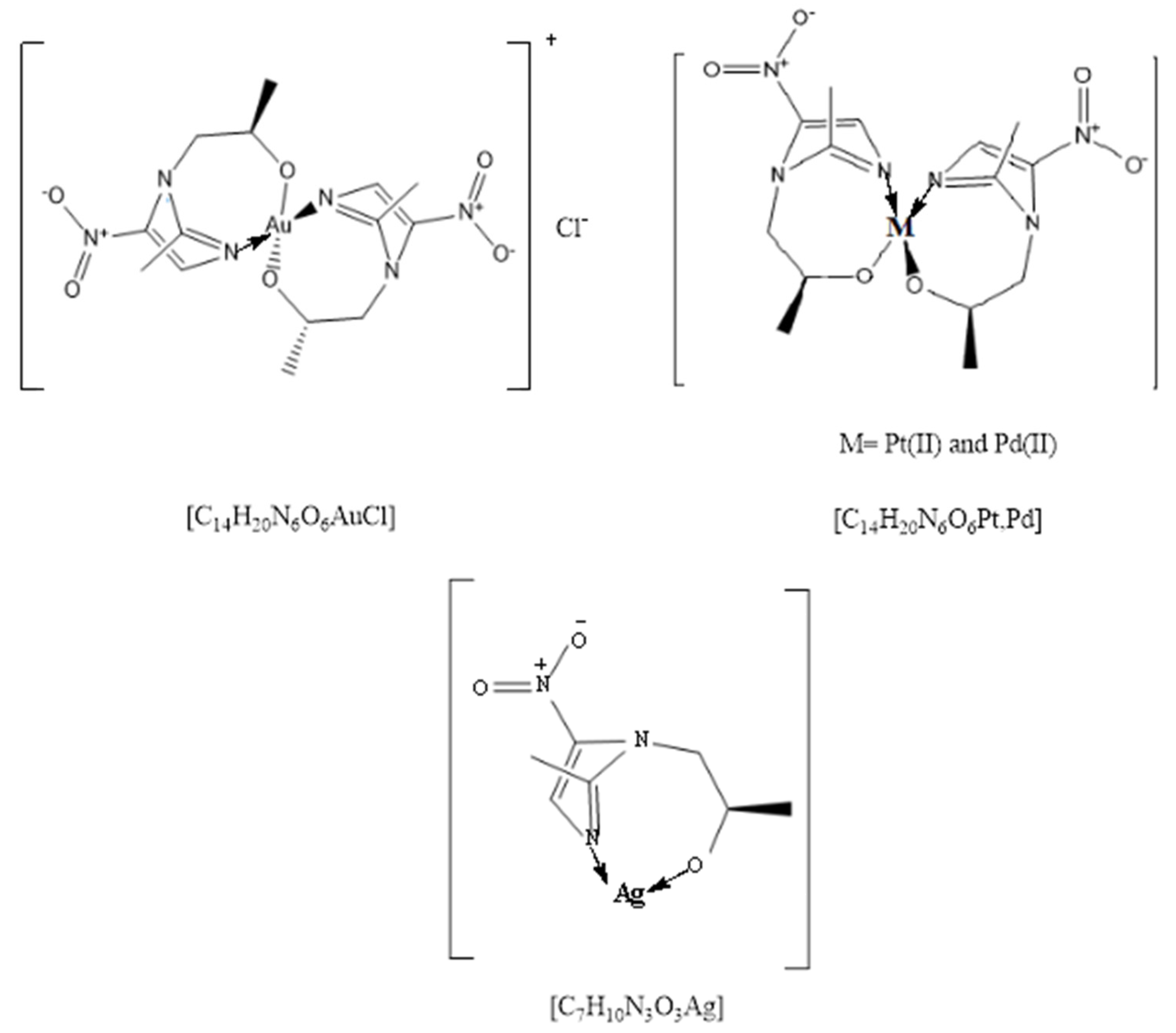


| No. | Complexes Formula | Λ (Ohm−1·cm2mol−1) | Color | Yield% | m.p. °C | CHN (Found) | ||
|---|---|---|---|---|---|---|---|---|
| %C | %H | %N | ||||||
| 1 | C14H20N6O6AuCl | 110.20 | Light brown | 91 | 350 | 27.96 (28.02) | 3.32 (3.29) | 13.98 (13.81) |
| 2 | C14H20N6O6Pt | 13.10 | black | 74 | 260 | 29.81 (29.75) | 3.54 (3.50) | 14.90 (15.08) |
| 3 | C14H20N6O6Pd | 11.60 | Gray | 80 | 300 | 35.38 (35.22) | 4.21 (4.35) | 17.69 (17.58) |
| 4 | C7H10N3O3Ag | 16.80 | Light gray | 66 | 200 | 28.76 (28.85) | 3.42 (3.38) | 14.38 (14.26) |
| Compound | OH | C-O | C=N | C-N | M-O | M-N |
|---|---|---|---|---|---|---|
| Secnidazole | 3500 | 1171 | 1658 | 1278 | - | - |
| Au Complex | - | 1150 | 1580 | 1264 | 521 | 423 |
| Pt Complex | - | 1158 | 1578 | 1260 | 535 | 485 |
| Pd Complex | - | 1148 | 1585 | 1258 | 544 | 470 |
| Ag Complex | - | 1160 | 1583 | 1262 | 562 | 447 |
| Compound | λmax (nm) | Assignment |
|---|---|---|
| Secnidazole | 295 324 | π➝π * transition n➝π * transition |
| Au Complex | 319 | n➝π * transition |
| Pt Complex | 330 377 | π➝π * transition n➝π * transition |
| Pd Complex | 285 322 | π➝π * transition n➝π * transition |
| Ag Complex | 355 385 376 | π➝π * transition π➝π * transition n➝π * transition |
| Sample | Klebsiella | E. Coli | Staphylococcus Epidermidis | Staphylococcus aureus |
|---|---|---|---|---|
| DMSO | 0.0 | 0.0 | 0.0 | 0.0 |
| Amoxicillin | 0.4 | 0.3 | 0.9 | 0.4 |
| Ampicillin | 0.2 | 0.1 | 0.8 | 0.2 |
| Secnidazole | 0.0 | 0.0 | 0.0 | 0.0 |
| Au Complex | 0.2 | 0.0 | 0.1> | 0.1 |
| Pt Complex | 0.3 | 0.0 | 0.1> | 0.1> |
| Pd Complex | 0.2 | 0.0 | 0.1> | 0.1 |
| Ag Complex | 0.1 | 0.0 | 0.3 | 0.1> |
| Complexes | MCF-7 | HepG-2 |
|---|---|---|
| Doxorubicin HCl | 8.50 | 4.7 |
| Secnidazole | - | - |
| Au Complex | 21.6 | >100 |
| Pt Complex | 11.3 | - |
| Pd Complex | 13.5 | >100 |
| Ag Complex | 7.4 | - |
| Protein | Receptor | Interaction | Distance | E (kcal/mol) |
|---|---|---|---|---|
| Breast cancer ID: 3hb5 | VAL 188 | H-acceptor | 3 | −2.6 |
| Protein | Receptor | Interaction | Distance | E (kcal/mol) |
|---|---|---|---|---|
| Breast cancer ID: 3hb5 | SER 12 | H-donor | 3.41 | −0.7 |
| THR 190 | H-donor | 3.21 | −1 | |
| SER 12 | H-acceptor | 3.41 | −0.6 | |
| GLY 15 | H-acceptor | 2.98 | −3 | |
| ASN 90 | H-acceptor | 2.91 | −3.1 |
| Protein | Receptor | Interaction | Distance | E (kcal/mol) |
|---|---|---|---|---|
| Breast cancer ID: 3hb5 | GLY 92 | H-acceptor | 3.2 | −1.3 |
| LYS 159 | H-acceptor | 3.15 | −2 | |
| GLY 92 | H-acceptor | 3.44 | −1.2 |
| Protein | Receptor | Interaction | Distance | E (kcal/mol) |
|---|---|---|---|---|
| Breast cancer ID: 3hb5 | SER 142 | H-donor | 2.91 | −4.3 |
| SER 142 | H-acceptor | 3.23 | −0.8 | |
| THR 190 | H-acceptor | 3.05 | −1.7 |
| Protein | Receptor | Interaction | Distance | E (kcal/mol) |
|---|---|---|---|---|
| Breast cancer ID: 3hb5 | VAL 188 | H-acceptor | 3.3 | −0.6 |
| SER 142 | H-acceptor | 2.7 | −1.1 | |
| MET 193 | H-acceptor | 3.7 | −0.7 |
| Compound | S | rmsd_Refine | E_Conf | E_Place | E_Score1 | E-Refine | E_Score2 |
|---|---|---|---|---|---|---|---|
| Secnidazole | −5.11 | 1.48 | 6.27 | −55.58 | −8.23 | −17.35 | −5.11 |
| −5.07 | 1.61 | 7.34 | −57.05 | −8.06 | −26.67 | −5.07 | |
| −5.01 | 1.89 | 5.44 | −51.78 | −8.04 | −23.10 | −5.01 | |
| −5.00 | 0.94 | 7.24 | −49.75 | −8.46 | −17.36 | −5.00 | |
| −4.95 | 0.84 | 5.87 | −51.82 | −8.20 | −20.08 | −4.95 | |
| Au complex | −6.87 | 1.32 | 216.30 | −52.50 | −9.90 | −25.52 | −6.87 |
| −4.77 | 0.76 | 214.28 | −76.74 | −10.78 | −26.23 | −4.77 | |
| −4.76 | 2.19 | 227.67 | −47.30 | −10.66 | −33.73 | −4.76 | |
| −4.74 | 1.65 | 214.07 | −53.80 | −10.04 | −21.61 | −4.74 | |
| −4.70 | 1.06 | 215.14 | −61.43 | −10.15 | −18.83 | −4.70 | |
| Pt complex | −7.48 | 2.49 | 149.63 | −55.58 | −10.83 | −37.00 | −7.48 |
| −7.03 | 1.94 | 147.23 | −69.33 | −9.63 | −34.12 | −7.03 | |
| −6.91 | 0.99 | 147.06 | −56.01 | −9.56 | −23.91 | −6.91 | |
| −6.79 | 1.39 | 157.33 | −43.52 | −10.23 | −25.46 | −6.79 | |
| −6.78 | 2.19 | 158.43 | −54.43 | −10.47 | −23.45 | −6.78 | |
| Pd complex | −7.43 | 1.58 | 134.52 | −60.31 | −10.19 | −35.53 | −7.43 |
| −6.84 | 2.18 | 154.23 | −47.58 | −9.38 | −22.29 | −6.84 | |
| −6.63 | 1.68 | 123.10 | −54.16 | −9.68 | −31.50 | −6.63 | |
| −6.57 | 1.25 | 150.12 | −47.46 | −9.12 | −22.53 | −6.57 | |
| −6.52 | 1.81 | 132.06 | −65.60 | −10.14 | −33.36 | −6.52 | |
| Ag complex | −7.87 | 0.97 | 232.12 | −43.69 | −7.10 | −21.43 | −7.87 |
| −7.30 | 2.48 | 232.46 | −57.20 | −8.46 | −17.48 | −7.30 | |
| −7.06 | 1.17 | 233.05 | −52.75 | −6.99 | −13.68 | −7.06 | |
| −7.05 | 1.59 | 234.16 | −45.31 | −7.01 | −13.85 | −7.05 | |
| −7.02 | 0.67 | 232.78 | −57.80 | −7.61 | −15.75 | −7.02 |
| Secnidazole |  |
| Au complex | 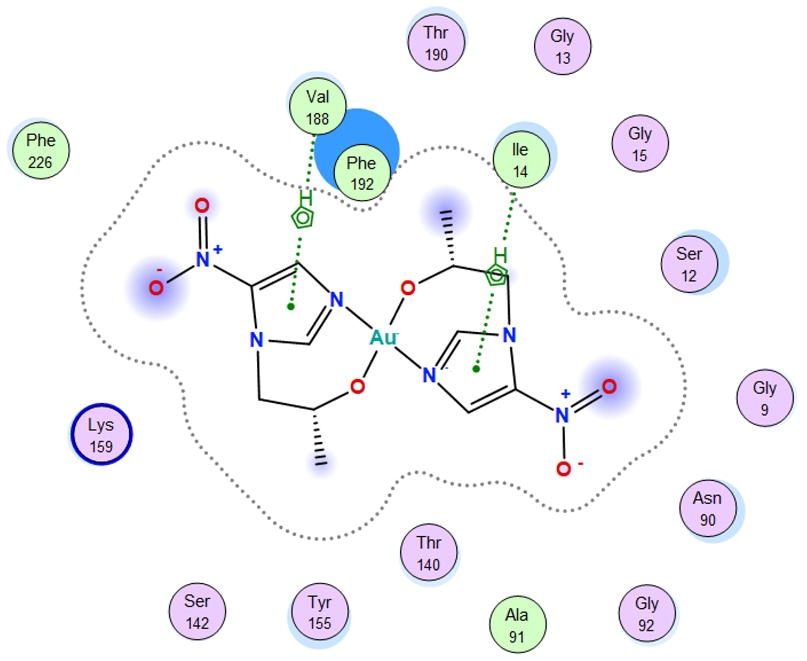 |
| Pt complex | 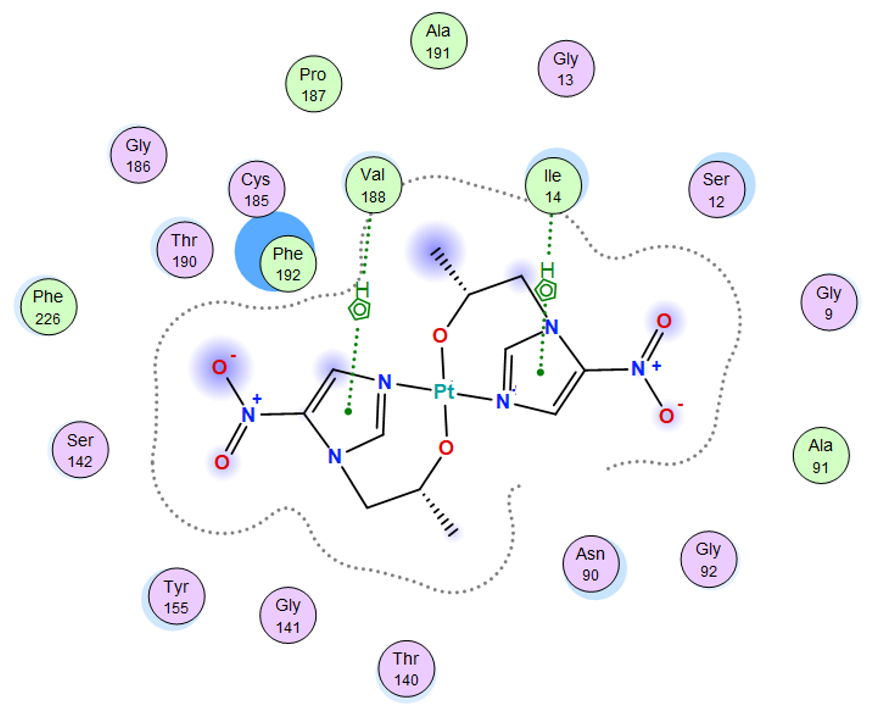 |
| Pd complex |  |
| Ag complex | 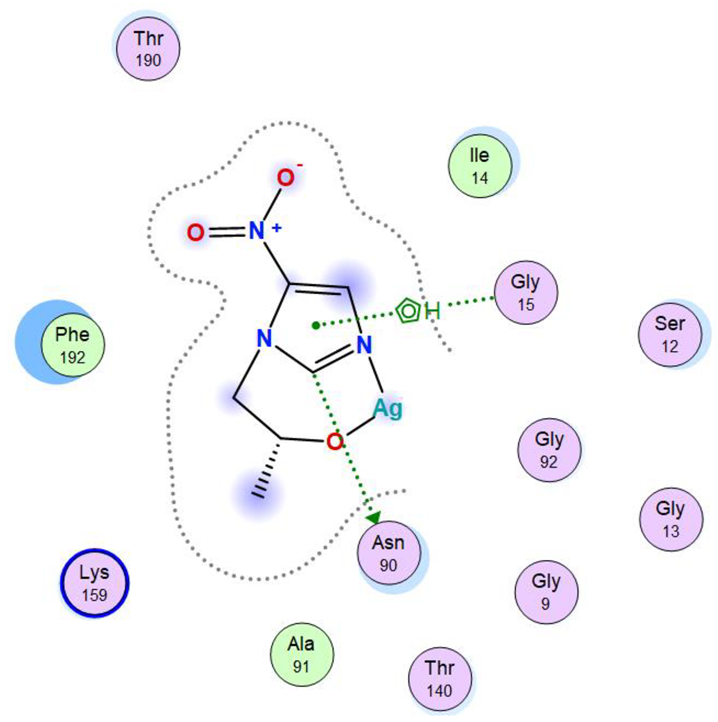 |
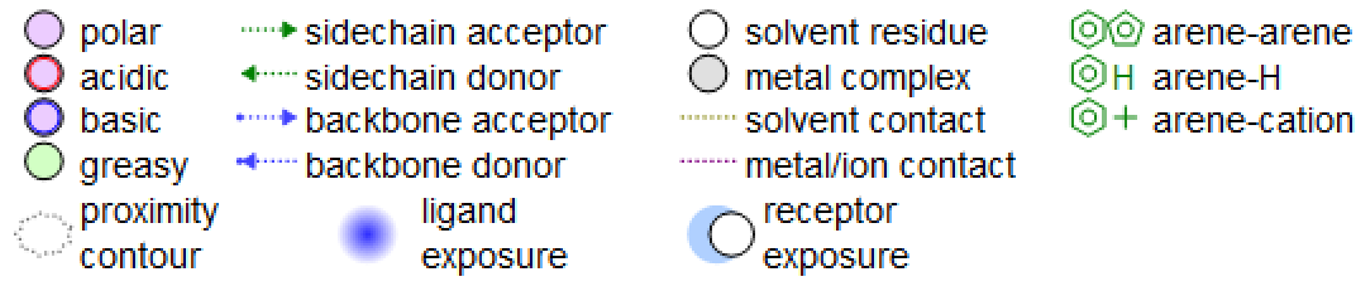 | |
| Secnidazole |  |  |
| Au complex | 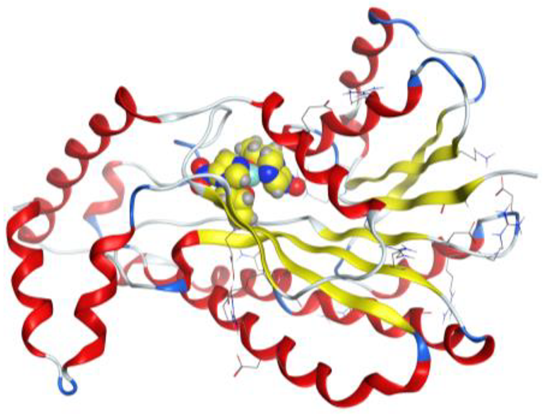 | 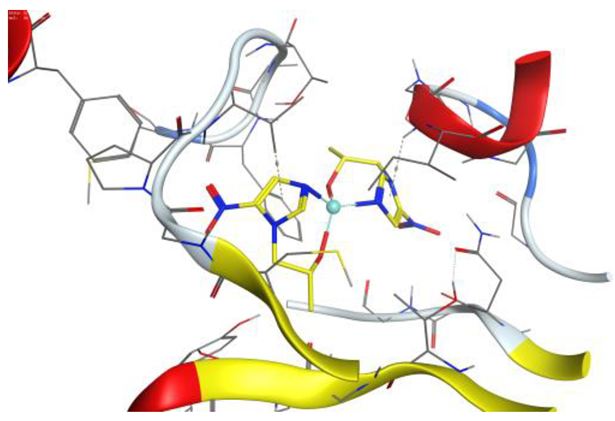 |
| Pt complex | 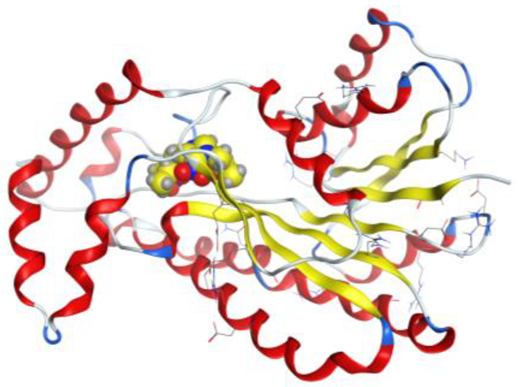 | 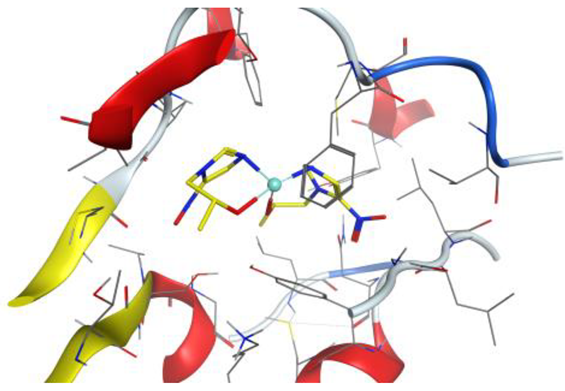 |
| Pd complex | 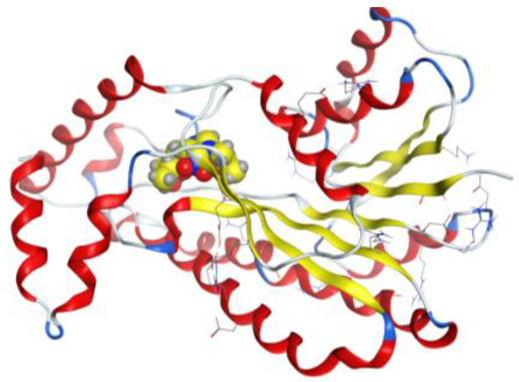 | 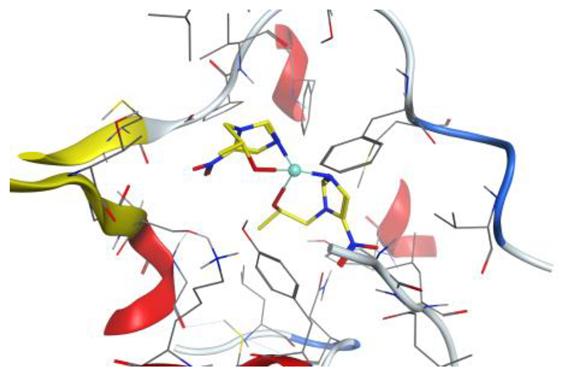 |
| Ag complex | 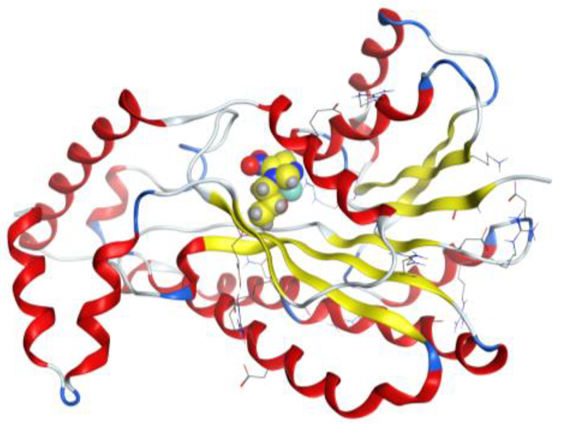 | 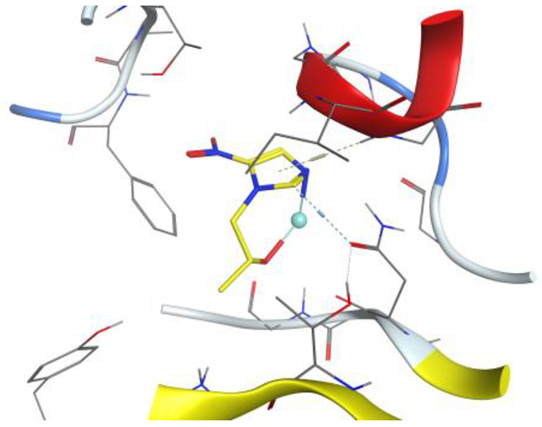 |
| Compound | Ligand | Receptor | Interaction | Distance | E (kcal/mol) |
|---|---|---|---|---|---|
| Secnidazole | 5-ring | CB VAL 188 (X) | pi-H | 4.30 | −0.5 |
| Au complex | 5-ring | N ILE 14 (X) | pi-H | 3.97 | −1.1 |
| 5-ring | CB VAL 188 (X) | pi-H | 4.01 | −0.6 | |
| Pt complex | 5-ring | N ILE 14 (X) | pi-H | 4.26 | −1.0 |
| 5-ring | CB VAL 188 (X) | pi-H | 4.14 | −0.8 | |
| Pd complex | O 37 | N GLY 92 (X) | H-acceptor | 2.87 | −1.2 |
| O 40 | N VAL 188 (X) | H-acceptor | 2.98 | −1.2 | |
| Ag complex | C 5 | OD1 ASN 90 (X) | H-donor | 3.40 | −1.5 |
| 5-ring | N GLY 15 (X) | pi-H | 4.48 | −4.1 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Abd El-Fattah, W.; Elamin, N.Y. Synthesis, Spectroscopic Characterization, Molecular Docking and Biological Activity of Novel Secnidazole Metal Complexes. Inorganics 2022, 10, 156. https://doi.org/10.3390/inorganics10100156
Abd El-Fattah W, Elamin NY. Synthesis, Spectroscopic Characterization, Molecular Docking and Biological Activity of Novel Secnidazole Metal Complexes. Inorganics. 2022; 10(10):156. https://doi.org/10.3390/inorganics10100156
Chicago/Turabian StyleAbd El-Fattah, Wesam, and Nuha Y. Elamin. 2022. "Synthesis, Spectroscopic Characterization, Molecular Docking and Biological Activity of Novel Secnidazole Metal Complexes" Inorganics 10, no. 10: 156. https://doi.org/10.3390/inorganics10100156







