Abstract
With the increasing understanding of the molecular mechanism of the microRNAs (miRNAs) in prostate cancer (PCa), the predictive potential of miRNAs has received more attention by clinicians and laboratory scientists. Compared with the traditional prognostic tools based on clinicopathological variables, including the prostate-specific antigen, miRNAs may be helpful novel molecular biomarkers of biochemical recurrence for a more accurate risk stratification of PCa patients after radical prostatectomy and may contribute to personalized treatment. Tissue samples from prostatectomy specimens are easily available for miRNA isolation. Numerous studies from different countries have investigated the role of tissue-miRNAs as independent predictors of disease recurrence, either alone or in combination with other clinicopathological factors. For this purpose, a PubMed search was performed for articles published between 2008 and 2017. We compiled a profile of dysregulated miRNAs as potential predictors of biochemical recurrence and discussed their current clinical relevance. Because of differences in analytics, insufficient power and the heterogeneity of studies, and different statistical evaluation methods, limited consistency in results was obvious. Prospective multi-institutional studies with larger sample sizes, harmonized analytics, well-structured external validations, and reasonable study designs are necessary to assess the real prognostic information of miRNAs, in combination with conventional clinicopathological factors, as predictors of biochemical recurrence.
1. Introduction
Prostate cancer (PCa) is the second most common cancer among men worldwide. It encompasses 15% of all diagnosed male malignancies every year, with an estimated 1.112 million new cases and 307,000 deaths according to the last global cancer statistics in 2012 [1].
Since the mid 1980s, the widespread use of the prostate-specific antigen (PSA) has substantially shaped the management of this cancer, but its overdiagnosis and overtreatment has gained increasing attention after a controversial debate on the PSA-based early detection and screening approach [2,3]. This is a result of the heterogeneous behavior of the disease from the entirely indolent to the extremely aggressive tumor. Numerous pre- and post-treatment nomograms based on well-established clinicopathological factors, such as clinical and pathological tumor stage, bioptic and pathological histological grading systems according to Gleason, and PSA values, have been used to estimate the individual risk of the disease course. This particularly refers to the prediction of different clinical end points like biochemical recurrence (BCR), occurrence of metastases, cancer-specific death, and overall survival [4,5,6]. However, the achieved accuracy of the outcome prediction using these nomograms is generally limited, resulting in an agreement between predicted and observed outcomes of only 70–80%. Thus, the identification of prognostic factors and the elucidation of the underlying molecular mechanisms that determine the course of the disease are essential future tasks for improving the cancer decision-making process [7,8]. This is true both for the risk estimation after PCa detection and for the follow-up after treatment.
Radical prostatectomy is the surgically preferred treatment option with curative intention of clinically significant PCa. Molecular markers of genomic, transcriptomic, proteomic or metabolomic nature are capable of enhancing the prediction accuracy if they are included in prediction tools that are based on only clinicopathological factors [9,10,11,12]. For such an approach, microRNAs (miRNAs), as decisive regulators of the cellular processes, are also candidate biomarkers [13,14,15]. miRNAs can function both as tumor suppressors or oncogenes in urological tumors as described in several recent reviews [16,17,18,19,20,21]. More detailed information regarding their special role in cancerogenesis and the progression of these tumors as well as their biogenesis and general function can be found in these reviews.
The expression of miRNAs can be specifically quantified in prostatectomy tissue samples. These analytes have been suggested in previous studies as promising prognostic markers to improve the prediction of the biochemical recurrence of PSA as the first alarming sign of cancer relapse after prostatectomy [22,23,24,25]. Approximately 15–30% of patients suffer from a biochemical recurrence after radical prostatectomy [26,27]. Thus, an early and reliable detection of these PCa patients at risk after radical prostatectomy would improve the decision-making for the initiation of adjuvant therapy and for the selection of patients who need a more frequent monitoring during follow-up.
Therefore, in the present review, we aimed (a) to compile the relevant data of existing miRNA-based studies, (b) to identify the most promising miRNAs as potential predictors of biochemical recurrence proven in several independent studies, (c) to critically assess the real benefit of these new markers compared or in combination with the conventional parameters and (d) to formulate preconditions for robust assays to translate validated results into clinical practice.
2. Literature Search Strategy
2.1. Medical Subject Heading (MeSH) Terms and Keywords
For this review, a PubMed search was performed for original articles in the database from 2008 to May 2017. The search strategy included the followings terms: the MeSH term “MicroRNAs” combined with the search string [“microRNAs” OR “microRNA” OR “micro-RNA” OR “micro-RNAs” OR “miRNAs”], the MeSH term “prostatic neoplasms” linked with the keyword “prostate cancer” using the Boolean operator “OR” and always connected with the search strings [“biochemical recurrence” OR “recurrence” OR “biochemical relapse” OR “biochemical failure”], and “radical prostatectomy” using the Boolean operator AND. Furthermore, references in the identified articles and reviews were considered to detect additional relevant articles. Publications were included in this review only if (a) they were peer-reviewed and supplied with full text in English, (b) the sample resources were tissue specimens, either fresh-frozen or formalin-fixed, paraffin-embedded (FFPE) tissue, whereas articles pertaining to miRNAs from blood, urine, cell lines, and animal models were disregarded, and (c) study objects were single miRNAs, patterns of various miRNAs, or miRNAs combined with clinicopathological variables, resulting in potential prognostic value for BCR.
2.2. Defining BCR as the Clinical Endpoint
BCR refers to the occurrence of increasing PSA value after its decline due to treatment. Radical prostatectomy and radiotherapy are the two main curative options for treatment of prostate cancer. Here, we exclusively refer to the surgical option as only this treatment mode allows the investigation of tissue samples of the removed cancer.
After radical prostatectomy, circulating PSA rapidly declines in a biphasic elimination, with a half-life of approximately one to three days [28]. Thus, patients with a preoperative PSA value of 20 µg/L generally reach a PSA level of <0.1 µg/L after 10 to 20 days, but not later than four weeks after successful operation [28]. According to the guidelines of the European Association of Urology (EAU), a PSA value <0.1 µg/L after radical prostatectomy is considered as undetectable. In this case, the definition of BCR is based on a renewed PSA increase to >0.2 µg/L that is confirmed by two consecutive elevated values [29]. Patients with increasing PSA values before the PSA nadir is reached should not be included in this biochemical recurrence group as the clinical outcome of patients with such a persistent PSA value after radical prostatectomy is generally poor. The 0.2 µg/L PSA cutoff also corresponds to the definition of BCR recommended by the American Urological Association (AUA) Prostate Guideline Update Panel [30]. However, it should be pointed out that this panel registered in their literature search of 145 studies more than 53 varying different definitions of biochemical recurrence after radical prostatectomy. The improved detection limit and analytical accuracy to measure low PSA values contributed to the recommendation to use the 0.2 µg/L PSA cutoff instead of 0.4 µg/L as previously suggested [31].
Thus, various factors summarized in Table 1 influence BCR directly, such as adverse tumor characteristics, or indirectly, such as different PSA analytics, varying definitions of BCR, and the clinician’s judgment of BCR. Within a selected definition of BCR, the tumor characteristics of the individual patient are the most important factor that determine the occurrence of BCR [5,6,32,33,34]. The numerous pre- and postoperative nomograms predicting BCR-free probability after radical prostatectomy are based on these clinicopathological data [4,5,6,35,36]. Thus, the clinical usability of all additional classifiers, in our case miRNAs, as potential more informative decision-making tools or adjunctive parameters have to be validated in relation to these conventional clinicopathological data in multivariate statistical models. Only their additional diagnostic benefit or cost-efficiency in comparison to conventional tools would justify the introduction in clinical practice. In the present review, we focused on the assessment of this aspect in the studies. Despite the controversial discussions regarding a standardized definition of the PSA cutoff of BCR and its use as a surrogate for the clinical outcome in these patients [37,38,39,40], an increasing PSA concentration after radical prostatectomy is considered by the clinician to be the first sign of potential later cancer metastasis [41]. It is obvious that BCR is not equal to clinical relapse, but elevated postoperative serum PSA levels enable the isolation of patients with high risk of true disease recurrence [41]. Therefore, in our tabulated summary reports, we included the specific cutoffs of BCR used in the particular studies.

Table 1.
Factors influencing the “biochemical recurrence” diagnosis after radical prostatectomy.
3. Overview of the Evaluated Studies
3.1. Number of Annual Publications and Type of Tissue Samples Used in the Studies
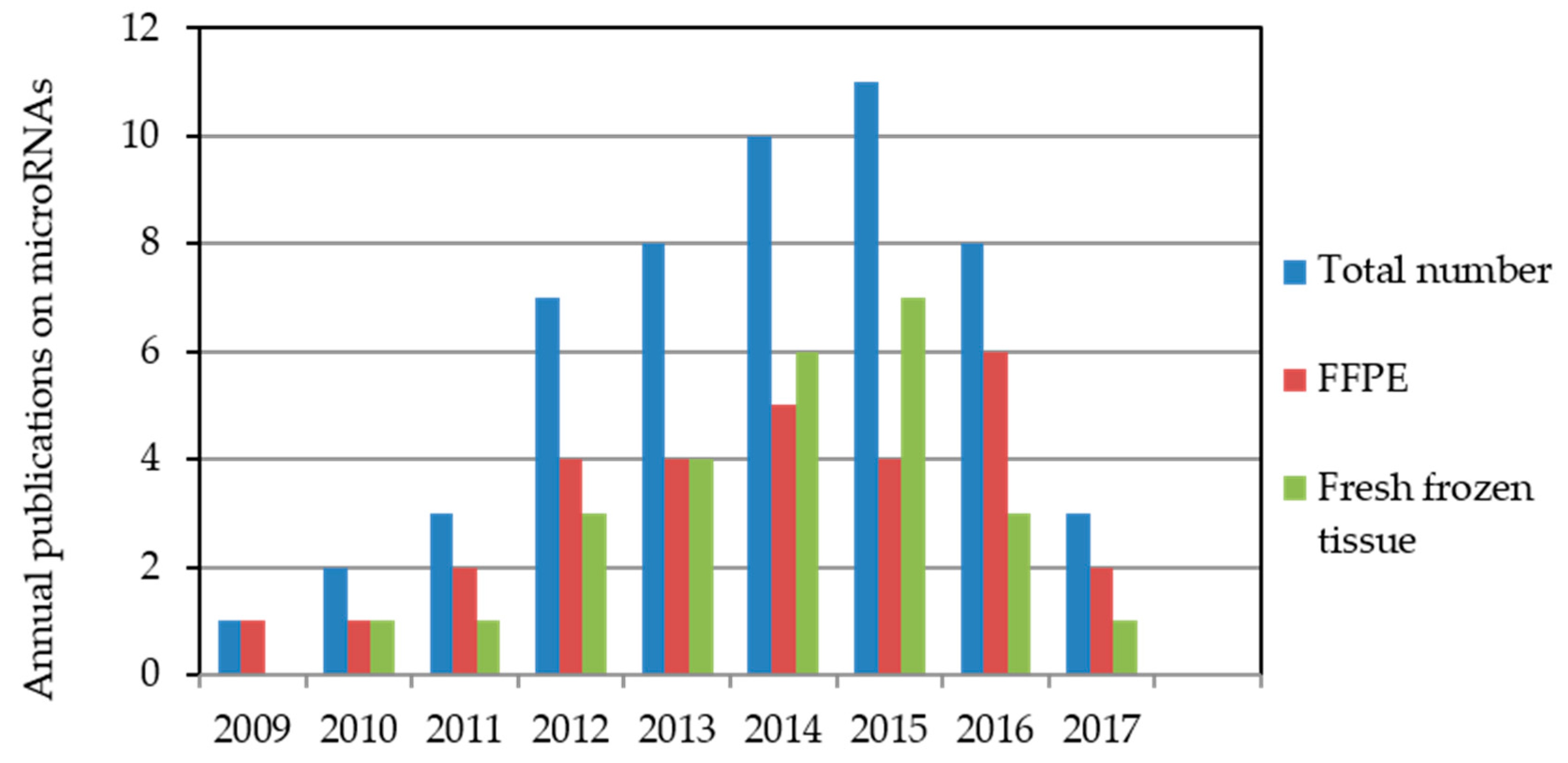
After preliminary screening of 148 papers, we identified 53 publications that complied with the described inclusion criteria. Forty-nine of these 53 articles were published in the past seven years. Only three papers appeared before 2011 [22,23,24]. Details can be seen in Figure 1.
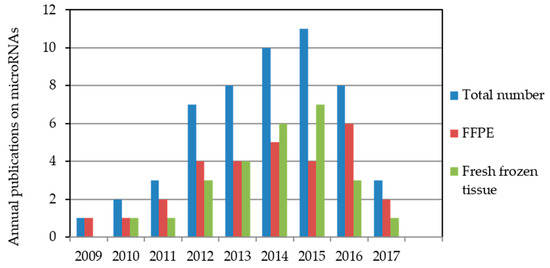
Figure 1.
Annual microRNA publications indexed in the PubMed database relating to biochemical recurrence after radical prostatectomy. The literature search was performed for the period from October 2008 to May 2017 with miRNA measurements in formalin-fixed, paraffin-embedded (FFPE) or fresh-frozen tissue samples. Two studies used both FFPE and fresh-frozen tissue samples [49,50].
Interest in the prognostic value of miRNAs in PCa has been reflected in the increasing number of publications. In 2009, Tong et al. [22] presented the first relevant study on the prognostic potential of miRNAs in PCa tissue; miR-135b and miR-194 were proven to reflect a tendency for early PCa relapse by comparing patients with early and late BCR. The results summarized in this review of 53 studies are based on data from 29 and 26 studies that analyzed FFPE and fresh-frozen tissue samples, respectively. In two studies, both FFPE and fresh-frozen tissue samples were used [49,50].
3.2. Characteristics of the Studies Evaluated in This Review
Biomarker studies with the intention to develop a robust assay for clinical practice must successfully undergo various phases of testing. Simply speaking, a discovery phase with the identification or selection of potential candidate biomarkers based on different principles for the intended objective should be distinguished from validation processes [51]. This classification with their subdivided characteristics is helpful to facilitate the assessment of studies and has therefore been adapted with regard to the use of miRNAs as BCR biomarkers in Table 2. On this basis, essential data and results of every study of the 53 evaluated studies including our own assessment have been compiled in Table 3. For the sake of clarity and facilitating the later discussion, the studies are listed by year of publication and are numbered accordingly.

Table 2.
Development phases to use miRNAs as predictors of biochemical recurrence after radical prostatectomy.

Table 3.
Studies regarding tissue miRNAs as predictive markers for biochemical recurrence in prostate cancer after radical prostatectomy.
3.2.1. Dysregulated miRNAs with Association to Biochemical Recurrence
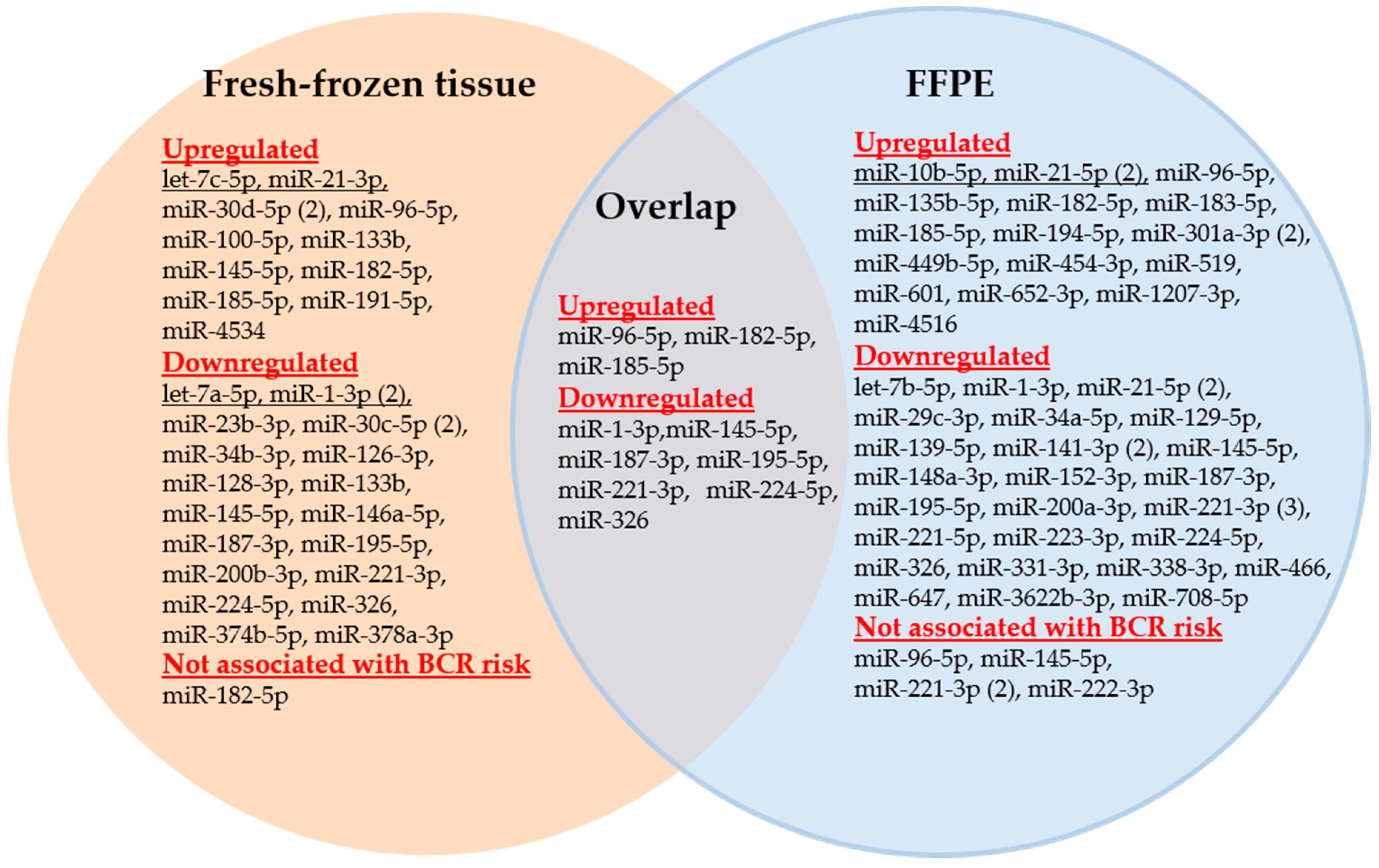
The differentially expressed miRNAs in prostatectomy tissue samples that have been proven to be potentially predictive BCR markers in the 53 evaluated studies are represented in Figure 2 as a Venn diagram.
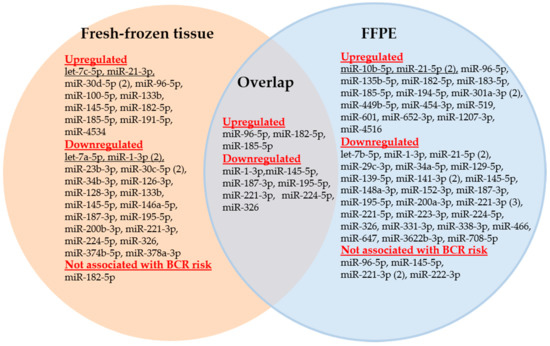
Figure 2.
Venn diagram of the miRNAs analyzed in FFPE and fresh-frozen tissue samples of studies examining the predictive capability of miRNAs for biochemical recurrence. Numbers in parentheses indicate the number of studies that examined the respective miRNA.
As previously mentioned, both samples from fresh-frozen tissue and FFPE archived tissue blocks were used for analysis of miRNAs in these studies. In contrast to fresh-frozen tissue, FFPE tissue samples are easily available as they are generally used in the tissue-based diagnostic routine process and do not require time-consuming workflow in comparison to fresh-frozen tissue samples. FFPE blocks are archived in repositories of the pathological institutes along with all clinical and pathological information. In contrast to the non-stability of mRNAs in FFPE tissue, miRNAs were found to be congruently expressed in fresh-frozen and FFPE tissue samples including prostate cancer [108,109,110,111]. Because of their small size and association with protectively acting macromolecules, miRNAs are obviously more robust molecules and are less affected by degradation processes than mRNAs. This was also demonstrated in model experiments of RNA degradation [112]. Li et al. [113] showed comparable miRNA profiles between FFPE and paired snap-frozen materials with R2 > 0.95. Moreover, this observation is consistent with the results of Casanova-Salas et al. [49] and Kristensen et al. [50], who used both FFPE and fresh-frozen tissue samples in their BCR studies (see Table 3, Study nos. 22 and 46). However, there are conflicting data on the stability of miRNAs in FFPE tissue blocks stored for more than ten years [108,109,111,114]. Two studies recently proved the differential long-term stability of various miRNAs in FFPE samples over ten years [114,115], probably depending on the different GC contents in the distinct miRNAs [115]. This issue needs to be controlled in studies using long-term archived samples to consider this possible storage effect for a correct assessment of analytical data [114].
In reviewing the 53 studies, 41 distinct miRNAs were described in FFPE and 27 miRNAs in fresh-frozen tissue samples as significant miRNAs (Figure 2). Moreover, only 10 miRNAs were simultaneously detected in both sample types as shown in the overlap section of Figure 2. As miR-21-5p, miR-133b, and miR-145-5p were found to be both up- and downregulated in various studies, a total of 58 distinct miRNAs were used as potential BCR markers. Of these 58 miRNAs, only 15 miRNAs were examined in at least two studies, whereas 43 miRNAs were determined in only one study (Table 4, Table S2). The direction of the dysregulation of the miRNAs is indicated by arrows in Table 4.

Table 4.
Distinct miRNAs analyzed in at least two studies for predicting biochemical recurrence.
3.2.2. miR-221-3p, miR-21-5p, miR-145-5p, miR-1-3p, and miR-96-5p, the Most Frequently Analyzed miRNA-Based BCR Markers
The miRNAs miR-221-3p, miR-21-5p, miR-145-5p, miR-1-3p, and miR-96-5p were found to be the most frequently analyzed miRNAs in the reviewed BCR studies. Their results are of particular interest as they allow some general conclusions with regard to the potential predictive BCR capability of miRNAs but also to future research requirements. In referring to the direction of the dysregulation of these miRNAs in the corresponding studies as indicated in Table 4 and subsequently mentioned using the list number from Table 3, the following short comments should summarize the situation.
- miR-221-3p. Three of the four studies confirmed the downregulated expression of miR-221 as a useful BCR predictor and independent factor in multivariate analyses with the standard clinicopathological variables (Study nos. 3, 31, and 46; [24,50,83]). Kristensen et al. [50] (Study 46) validated miR-221 in two independent BCR cohorts and an additional external validation using a publicly available data set as part of their 3-miRNA signature while Spahn et al. [24] (Study 3) proved the usefulness of this miRNA especially in high-risk PCa patients. Thus, these studies can be assessed as successful approaches from the discovery phase to validation by clinical assessment with the aim to develop a potential clinical tool as suggested in Table 1. The miRNA tool miQ that was primarily developed for diagnostic purposes included the also downregulated 5p strand of miR-221 in predicting BCR (Study 17, [67]). Strong correlations were observed in these studies between the increased expression of miR-221 and the tumor stage, Gleason score, and the pre-operative PSA level. In contrast, these correlations were not found in Study 9 with the missing predictor evidence of miR-221 [58]. However, this failure could also be caused by the short follow-up period of less than two years in this study.
- miR-21-5p. Increased and decreased expression of this miRNA was suggested as a potential BCR predictor in two studies (Table 4). Correlations were described between the increased expression of miR-21-5p as a BCR predictor and the standard clinicopathological variables (Study nos. 11 and 29; [60,81] while these data were not reported in the controversial studies with the decreased miRNA expression (Study nos. 14 and 31; [64,83]). After adjustment with clinicopathological factors, decreased miRNA expression failed to be an independent BCR risk factor (Study 31, [83]) or was only appropriate in obese patients (Study 14, [64]). Only one of the three studies with upregulated expression in tumor tissue clearly proved miR-21 as an independent factor for shorter BCR-free survival in multivariate analysis (Study 11, [60]).
- miR-145-5p. Both a study with increased (Study 5, [52]) and two studies with decreased expression of miR-145 estimated this miRNA as a potential BCR predictor or part of a significant prediction signature (Study nos. 15 and 17; [65,67]). It cannot be excluded that these discrepant findings were caused by analytical reasons, as two studies calculated the expression of miR-145 with normalizers (RNU43 and SNORD48) that were criticized regarding their suitability as reference genes [116]. Another study with decreased miR-145-5p expression (Study 9, [58]) was not able to confirm miR-145-5p as a BCR predictor in Kaplan-Meier analysis. However, it should be noted that the above-mentioned very short follow-up period in that study makes a true assessment difficult.
- miR-1-3p. Three studies examined the potential BCR capability of downregulated miR-1. Two studies (Study nos. 8 and 53; [55,107]) identified miR-1 as an independent BCR predictor after adjustment with the conventional clinicopathological factors. However, the additional benefit was not demonstrated when miR-1-3p was included in the model based only on clinicopathological factors. miR-1 was also demonstrated to be a successful BCR predictor in the third study (Study 24, [74]), but its clinical accuracy was exceeded by the pre-operative PSA value. The inconsistent documentation of clinicopathological variables in these studies makes it impossible to attribute this uniform BCR predictor result to congruent clinical characteristics between the studies.
- miR-96-5p. In two studies (Study nos. 2 and 17; [23,67]), increased levels of this miRNA in PCa tissue were successfully identified as a single BCR predictor or part of a BCR predictor combination. A third study (Study 9, [58]) did not confirm an association of the recurrence-free survival and the miR-9-5p expression level.
The heterogeneity of results of the particular miRNAs in these multiple studies also reflects the situation of the other miRNAs with only two studies available (Table 4). For example, opposite expression data were reported for miR-133b, but both studies suggested this miRNA as a potential BCR predictor despite their discordant expression data (Study nos. 24 and 26; [74,76]). Moreover, studies that partly use data from publicly available databases or from previous studies may lack clearly defined characteristics complicate objective assessment. This applies to miR-30c-5p (Study nos. 28 and 47; [78,99]) and miR-301-3p (Study nos. 37 and 48; [89,100]).
3.2.3. Multiple miRNAs as Signatures or in Combination with Other Analytes
In the discovery phase of the development of a tissue-based miRNA assay for predicting BCR, highthroughput “-omics” approaches like microarrays or sequencing technologies provide extensive data sets with numerous candidate miRNAs to meet this pursued objective. One approach is to search this pattern of analytes and to select the most effective miRNAs for the validation of BCR prediction in the subsequent development phases. However, there is now a great interest in using this wealth of information not only for selecting single markers but also for combining multiple markers into a specific panel or signature together with clinicopathological data [117,118]. Particular attention should be paid to implement orthogonal markers in such a signature [119]. Orthogonal markers are uncorrelated among each other and to the conventional clinicopathological factors. This uncorrelated particularity is an essential precondition to improve the predictive significance of the signature due to the additional information achieved by these independent factors. For miRNAs, this orthogonal aspect could be demonstrated for the miR-29c-3p, miR-34a-5p, miR-141-3p, and miR-148a-3p that were not associated with tumor size and pathological stage but were inversely correlated with Gleason grades [88]. The Decipher genomic classifier using a 22-gene signature for post-prostatectomy risk stratification or other similar approaches has shown the potential usefulness of such multi-analyte tools [120,121,122].
In this review, the studies by Nam et al. [89] (Study 37: 5-miR signature with miR-139-5p, miR-223-3p, miR-301a-3p, miR-454-3p, and miR-652-3p) and Kristensen et al. [50] (Study 46: 3-miRNA prognostic classifier with miR-185-5p, miR-221-3p, and miR-326) support these ideas. Based on the multiple-miRNA approach as a signature combined with rigorous validation processes (three validations in Study 46) or a high sample size and a high number of BCR events (n = 491, 167 BCRs in Study 37), the two studies yielded promising results. Both studies are among the most convincing studies evaluated in this review and can be considered future-oriented examples. Nam et al. [100] focused in a subsequent study (Study 48) on the predictive validity of the single miR-301a-3p from the above-mentioned 5-miRNA signature. The authors also described a good BCR prediction rate using only this single miRNA, but they did not compare the results of the two approaches. Bell et al. [84] published a further BCR prediction study based on a multiple miRNA signature (Study 32). A panel of 88 miRNA was required for a reliable BCR prediction within 3 years after surgery. However, the inclusion of only miR-4516 and miR-601 in a model with Gleason score and lymph node status alone improved the BCR prediction accuracy after salvage radiation treatment from 0.66 to 0.83 of the area under the receiver operating characteristic (ROC) curves. The previously discussed 4-miRNA tool miQ by Larne et al. [67] also proved that the integrated implementation of several differentially regulated miRNAs with orthogonal characteristics improved decision making in the management of PCa patients both in diagnosis and prognosis. Lichner et al. [69] developed three statistical models based on 2 to 3 miRNAs (Study 18: miR-331-3p + miR152-3p, miR-331-3p + miR-152-3p + miR135a-5p, and miR-148a-3p + miR-429) that were verified by internal validation and on an independent cohort. The authors achieved a correct classification rate of 92 to 100% in predicting patients with a high risk of BCR.
The combined use of panels with miRNA and mRNAs is also noteworthy. The mRNAs were either targets of the accompanying miRNAs or independently selected BCR markers, such as those in Study 6 [53] with miR-519, miR-647, and 10 mRNAs, in Study 41 [94] with miR-224-5p and its target APLN or in Study 52 [105] with miR-30d-5p and its target Protein phosphatase 1 regulatory subunit 12A (official symbol: PPP1R12A)(MYPT1).
4. Critical Assessment of the Recent Situation of miRNA-Based BCR Prediction
4.1. Analytical Considerations
In a previous review on circulating miRNAs in patients suffering from urological tumors, we discussed the typical influential and interfering factors that determine the results of miRNA measurements [51]. These are variables in the collection, further processing and storage of samples in the preanalytical phase, the various isolation and quantification methods based on different principles and technologies in the actual analytical phase and the different normalization strategies in the postanalytical phase. For more details of all these aspects, we refer the interested reader to the overview of Pritchard et al. [123]. In particular, different miRNA extraction procedures for fresh-frozen or FFPE samples and different measurement platforms showed qualitative and quantitative miRNA differences depending on the determination [124,125,126,127]. These differences might especially attribute to the lack of comparability of miRNA profiling data between studies that applied different analytical techniques. On the other hand, this effect needs a strict method harmonization in multi-institutional studies if the analytics are separately performed in every center. New comparative analyses recommended the Qiagen miRNeasy FFPE kit to be the best kit for miRNA isolation from FFPE samples and the new TaqMan advanced miRNA assays as the quantification method of superior sensitivity and specificity in comparison to competitor products [126,128].
All these issues also apply to the evaluated studies in this review and therefore do not need to be discussed again in detail. However, as a concrete example (Table 3, column “Methodology”), it is remarkable that confirmed stably expressed miRNAs for normalizing the expression results were only used in six (12%) of the 53 studies. In contrast, in 33 (63%) of the reviewed studies, different small nuclear and nucleolar RNAs (U6 or RNU6 [official name: RNU6-1], RNU6B [RNU6-6P], RNU43 [SNORD43], RNU44 [SNORD44], RNU47 [SNORD47], RNU48 [SNORD48], and RNU66 [SNORD66]) were used as endogenous normalizers. This was done even though most of these small RNAs were found to be unstably expressed across non-malignant and malignant prostate tissue and therefore considered as less suitable normalizers [116]. The real suitability of RNA47, RNU48, and RNU66 as normalizers was only tested in one study [67]. Thus, the general neglect of analytical basics was obvious in several studies. This was particularly underlined by the fact that none of the reviewed articles referred to the “Minimum Information for Publication of Quantitative Real-Time PCR Experiments” (MIQE) guidelines [129]. These guidelines address the analytical essentials that have to be considered to assess the quality and potential traceability of reverse transcription-quantitative polymerase chain reaction (RT-qPCR) measurements in an extensive checklist. Our observation corresponds with results of a recent survey of over 1700 publications that criticized the frequently insufficient description of experimental details of RT-qPCR measurements in many articles [130]. The authors of that survey called upon journal editors and reviewers to draw more attention to this issue for improving the transparency and comparability of RT-qPCR data between studies. It might be a specific challenge for clinically oriented journals in publishing clinical studies based on modern molecular-biological methods as clinicians often do not place any great emphasis on analytical problems.
4.2. Study Design Considerations
Table 3, with the essential details of the evaluated recent miRNA-based BCR studies and our separate comments, illustrates the heterogeneity of the data situation in this field. Different starting points in the discovery phase and specific features in subsequent validation processes hamper a comparison of data between studies. However, to provide a more informative overview not only on the diversity of miRNAs examined but also on the fundamental characteristics between the various studies, we classified various study criteria into categories in Table 5. This facilitates the identification of protocol deficiencies of the particular studies according to the assessment criteria of the development phases for establishing a robust tool in clinical practice (Table 2). Some noteworthy points should be considered more closely in the following.

Table 5.
Characteristics of the 53 studies evaluated in this review.
The different definitions of the PSA cutoff as criterion for the biochemical recurrence were discussed in detail at the beginning of this review. This diversity of cutoffs was also reflected in our survey. Two-thirds of the studies used the cutoff of 0.2 µ/L recommended in the EAU and AUA guidelines [29,30]. However, 23% of the studies did not specify this threshold as a fundamental precondition of data comparability. We also noticed this essential lack of information with regard to the specification of the important risk variables “resection margin status” and “lymph node status” in 60 and 70% of the studies, respectively. In contrast, the pathological tumor stage and Gleason score were generally indicated. On the other hand, only 3% of the studies included PCa patients with PSA values below 10 µg/L. This indicates that few studies focused on low-risk PCa patients.
In addition to these clinicopathological characteristics of the study patients as one part of the study design, more or less formal conditions determine the implementation and, finally, the validity of clinical studies. These study specifications are listed in Table 5 under the category “Study design features”. The percentage data given for the respective items illustrate deficiencies and the limited validity deficiencies of several studies. Thus, studies with sample sizes of less than 50 patients, 10 to 20 BCRs, a mean follow-up period under 5 years, or evaluated only through univariate analysis remain questionable from the statistical and biological point of view. For example, in a multivariate Cox regression analysis as a standard statistical method for BCR analysis, at least ten events per predictor variable are necessary to obtain reliable results [133]. Because several clinicopathological factors have to be individually considered in such a model it is not surprising that a study cohort with 20 BRC events and the additional inclusion of miRNAs of interest can hardly meet a scientifically founded conclusion of clinical significance. Considering a proportion of one-third of patients with BCR as an example, cohorts of more than 150 patients would be advisable. In contrast to this, few (15%) of the studies reviewed here that were exclusively retrospective in nature included more than 150 patients, and those were mostly multi-institutionally implemented. In this regard, it is significant that power and sample size calculations were presented in only two studies (Study nos. 2 and 9; [23,58]). Furthermore, only eight studies (15%) performed an internal or external validation of data that was suggested as an important criterion of the development phase “Validation by clinical assessment” (Table 2; [51]). For example, Kristensen et al. (Study 46; [50]) confirmed the improved prognostic performance of their 3-miRNA prognostic classifier in comparison to the BCR prediction based on only clinicopathological factors in three independent PCa patient cohorts. A similar benefit, proved by increased C-indices, was shown by single miRNAs in two other studies (Study nos. 2 and 30; [23,82]). The proof of such an additional benefit by the inclusion of miRNAs in the conventional model has to be considered as a decisive criterion to proceed further with developing a new clinical decision-making tool. Therefore, it is striking that the authors of merely two studies (Study nos. 2 and 46; [23,50]) pointed out that their studies were performed according to the “Reporting Recommendations for Tumor Marker Prognostic Studies (REMARK)” [131]. The generally “benevolent” neglect of the guideline suggestions in performing the prognostic studies by principal investigators and in accepting final study reports as publications by the journal editors is consistent with the above-mentioned attitude of ignoring the analytical MIQE and “Standards for Reporting of Diagnostic Accuracy (STARD)” guidelines [129,132].
4.3. Divergences between BCR Outcome and the Functional Role of miRNAs
Divergent BCR outcome data between different studies contrast with the functional data of miRNAs. In addition to the apparent differences due to the previously discussed reasons of the heterogeneity of study results, real divergences seem to exist between the miRNA expression level as a BCR predictor and the potential functional role of the respective miRNA. It is therefore worth briefly mentioning this rarely considered aspect using the examples of let-7c-5p, miR-141-3p, miR-148a-3p, and miR-221.
Leite et al. [52] showed in Study 5 that increased let-7c-5p in the primary untreated PCa tissue was associated with a higher BCR risk. This seems to be in contrast to the generally decreased expression of let-7c-5p in PCa tissue compared with normal prostate tissue and its suppressive action of this miRNA on the androgen receptor [134,135]. However, it should be considered that in the assessment of BCR risk, the expression of let-7c-5p is evaluated only in tumor cells. The BCR indicator effect of the let-7c-5p expression disappeared as an independent factor in the multivariate analysis with all risk factors showing the complex interplay between clinicopathological variables and expression levels of markers [52]. A similar, but contrasting and also not plausibly explainable phenomenon applies to miR-141-3p (Study nos. 31 and 36; [83,88]) and miR-148a-3p (Study 36; [88]). Decreased levels of both miRNAs indicated a shorter recurrence-free survival in the here reviewed studies, whereas their upregulation was found to be increased in untreated PCa and castration-resistant PCa specimens, and these miRNAs enhanced the proliferation of PCa cell lines [134,136,137]. For miR-221-3p and miR-221-5p, decreased expression levels were characteristics of a shorter BCR-free period (Studies nos. 3, 17, 31, 46; [24,50,67,83]. This tumor-suppressive function corresponds with the expression levels and functional data observed in other studies [138,139,140,141]. However, increased expression of miR-221 in PCa metastases and PCa mouse models and an enhanced proliferation of PCa cell lines by this miRNA were also described [142,143,144]. It was recently postulated that this oncogenic role of miR-221 is likely transient, and the dual tumor-suppressive and oncogenic function of miR-221 probably reflects different phases of PCa progression [140]. In this context, the possible divergences between BCR as clinical endpoint and the development-dependent functions of miRNAs would be understandable.
5. Future Directions
Despite the discussed critical points and limitations of the reviewed studies, promising results provided by several studies can be considered as proof of the true potential of miRNAs as BCR predictors. It is the final aim of this review to learn from the deficiencies of the conducted studies hitherto and draw corresponding conclusions for future studies. Therefore, our overview of the published results and the background data of the 53 studies allows two essential conclusions:
- No study has thus been able to comply with the suggested requirements specified in the final development phase “Validation of clinical usability” (Table 2) to establish a robust BCR tool for clinical practice using miRNAs. In addition, few studies can be valued as successfully finished in the second development phase due to the lack of internal validation in most of the studies (Table 2 and Table 5).
- The evaluation and comparison of analytical and clinical conditions in the various studies provided a wealth of experience in the assessment of study design features. Based on these experiences, critical study deficiencies could be identified (see Section 4, comments to Table 5), and future directions could be elaborated to overcome these shortcomings. In the following, we focus on some essential issues.
The results of the various studies and their generalized assessment confirm once more the clear need of a good coordination between the intended study aims, all study design elements, and preanalytical and analytical conditions. The three guidelines MIQE, REMARK, and STARD should be strictly considered in future studies since they define the basic foundation for implementing a study under common clinical and analytical conditions [129,131,132]. For planned projects, especially prospective, multi-institutional studies, appropriate elements of these guidelines should be specified, and their compliance should be a subject of constant control to guarantee necessary preconditions for a reliable database. These guidelines not only allow the necessary transparency but also the harmonization and comparability of results between multi-institutional studies. However, because of numerous factors, such as different methods of RNA isolation, reverse transcription, and true miRNA measurements, as well as various platform applications that could influence RT-qPCR results, it is advisable to perform all analyses at one institution in early studies. This approach would a priori avoid misinterpretations, as errors can be excluded due to missing traceability between results obtained through different methods. The issue of analytical differences could be solved later in a second step of method harmonization. The same applies to retrospective studies.
While BCR does not equal clinical relapse, elevated postoperative serum PSA levels make it possible to filter patients with a high risk of true disease recurrence [41]. Therefore, future studies should additionally focus on the predictive capability of miRNAs with regard to the clinical endpoints of distant metastasis, cancer-specific death, and response rate to drugs. Some studies (Study nos. 3, 14, 32, 53; [24,64,84,107]) have already considered these endpoints. However, the “mixed” use of these endpoints should be avoided in future studies because the distinct time difference between the endpoints could result in a systematic bias. In this respect, the study design should also clearly address the specific need for different patient groups. For example, the predictive tools using miRNAs differ between low-risk and high-risk patients following radical prostatectomy [69,71]. This result, which is also shown by using other genomic classifiers [122], should be considered in an adapted composition of the study groups according to the specific clinical objective. In addition, the Gleason-related association of miRNAs shown exemplarily by Lichner et al. [88] requires a re-assessment according to the new International Society of Urological Pathology (ISUP) Gleason group classification. Further multi-institutional studies are needed to validate the clinical usability of miRNA-based tools, either alone or combined with clinicopathological factors, for BCR prediction. The additional information provided by miRNAs in comparison to established BCR prediction tools [4,5,6] must be proven in these studies and should be demonstrated by decision curve analysis [145]. It is worth considering whether the above described 2–5 miRNA signatures could be confirmed in comparison to these clinically established tools in retrospective multi-institutional approaches as a validation step to initiate prospective studies.
6. Conclusions
In summary, miRNAs were shown in several studies of this review as promising marker candidates and miRNA signatures for predicting BCR after radical prostatectomy. However, the general non-consideration of the MIQE, REMARK, and STARD guidelines in most studies resulted in study design deficiencies, primarily a lack of internal validation of data. The unequivocal evidence of additional information through miRNAs in comparison to the conventional approaches of BCR has not been proven thus far. Further studies are needed to address these deficiencies both in retrospective and prospective multi-institutional studies to validate the clinical usability and benefit of miRNA-based BCR tools in combination with the conventional clinicopathological variables.
Supplementary Materials
Supplementary materials can be found at www.mdpi.com/1422-0067/18/10/2023/s1.
Acknowledgments
The authors thank Wilhelm Sander-Stiftung, SONNENFELD-Stiftung, and Stiftung Urologische Forschung for their support of the research work.
Conflicts of Interest
The authors declare no conflict of interest. The founding sponsors mentioned in the Acknowledgments had no role in the design of the study; in the collection, analyses, or interpretation of data; in the writing of the manuscript, and in the decision to publish the results.
Abbreviations
| APLN | Apelin |
| AUA | American Urological Association |
| AUC | Area under the ROC curve |
| BCL9 | B-cell CLL/lymphoma 9 |
| BCR | Biochemical recurrence |
| BPH | Benign prostatic hyperplasia |
| CoxM | Multivariate Cox regression analysis |
| CoxU | Univariate Cox regression analysis |
| EAU | European Association of Urology |
| FFPE | Formalin-fixed, paraffin-embedded |
| HR | Hazard ratio |
| ISUP | International Society of Urological Pathology |
| KMA | Kaplan-Meier analysis |
| MeSH | Medical Subject Heading of the U.S. National Library of Medicine |
| miQ | miRNA index quote |
| MIQE | Minimum information for publication of quantitative real-time PCR experiments |
| miRNA, miR | microRNA |
| MW | Mann-Whitney U-test |
| MYPT1 | Protein phosphatase 1 regulatory subunit 12A (official symbol: PPP1R12A) |
| PCa | Prostate carcinoma |
| PSA | Prostate-specific antigen |
| RB1CC1 | RB1 inducible coiled-coil 1 |
| REMARK | Reporting Recommendations for Tumor Marker Prognostic Studies |
| RFS | Biochemical recurrence-free survival |
| RM | Reference method, in general the reference gene |
| ROC | Receiver-operating characteristic curve |
| ROCK1 | Rho associated coiled-coil containing protein kinase 1 |
| RP | Radical prostatectomy |
| RT-qPCR | Reverse transcription-quantitative polymerase chain reaction |
| SOCS | Cytokine inducible SH2 containing protein (official symbol: CISH) |
| STARD | Standards for Reporting of Diagnostic Accuracy |
| TCGA | The Cancer Genome Atlas |
| TNM | Classification of malignant tumors describing the involment of the primary tumor, regional lymph nodes and the distant metastatic spread |
| TRIB1 | Tribbles pseudokinase 1 |
References
- Ferlay, J.; Soerjomataram, I.; Dikshit, R.; Eser, S.; Mathers, C.; Rebelo, M.; Parkin, D.M.; Forman, D.; Bray, F. Cancer incidence and mortality worldwide: Sources, methods and major patterns in GLOBOCAN 2012. Int. J. Cancer 2015, 136, E359–E386. [Google Scholar] [CrossRef] [PubMed]
- Andriole, G.L.; Crawford, E.D.; Grubb III, R.L.; Buys, S.S.; Chia, D.; Church, T.R.; Fouad, M.N.; Gelmann, E.P.; Kvale, P.A.; Reding, D.J.; et al. Mortality results from a randomized prostate-cancer screening trial. N. Engl. J. Med. 2009, 360, 1310–1319. [Google Scholar] [CrossRef] [PubMed]
- Schroder, F.H.; Hugosson, J.; Roobol, M.J.; Tammela, T.L.; Ciatto, S.; Nelen, V.; Kwiatkowski, M.; Lujan, M.; Lilja, H.; Zappa, M.; et al. Screening and prostate-cancer mortality in a randomized European study. N. Engl. J. Med. 2009, 360, 1320–1328. [Google Scholar] [CrossRef] [PubMed]
- Shariat, S.F.; Karakiewicz, P.I.; Roehrborn, C.G.; Kattan, M.W. An updated catalog of prostate cancer predictive tools. Cancer 2008, 113, 3075–3099. [Google Scholar] [CrossRef] [PubMed]
- Lughezzani, G.; Budaus, L.; Isbarn, H.; Sun, M.; Perrotte, P.; Haese, A.; Chun, F.K.; Schlomm, T.; Steuber, T.; Heinzer, H.; et al. Head-to-head comparison of the three most commonly used preoperative models for prediction of biochemical recurrence after radical prostatectomy. Eur. Urol. 2010, 57, 562–568. [Google Scholar] [CrossRef] [PubMed]
- Punnen, S.; Freedland, S.J.; Presti, J.C., Jr.; Aronson, W.J.; Terris, M.K.; Kane, C.J.; Amling, C.L.; Carroll, P.R.; Cooperberg, M.R. Multi-institutional validation of the CAPRA-S score to predict disease recurrence and mortality after radical prostatectomy. Eur. Urol. 2014, 65, 1171–1177. [Google Scholar] [CrossRef] [PubMed]
- Burke, H.B. Predicting clinical outcomes using molecular biomarkers. Biomark. Cancer 2016, 8, 89–99. [Google Scholar] [CrossRef] [PubMed]
- Reiter, R.E. Risk stratification of prostate cancer 2016. Scand. J. Clin. Lab. Investig. Suppl. 2016, 245, S54–S59. [Google Scholar] [CrossRef] [PubMed]
- Jung, K.; Reszka, R.; Kamlage, B.; Bethan, B.; Lein, M.; Stephan, C.; Kristiansen, G. Tissue metabolite profiling identifies differentiating and prognostic biomarkers for prostate carcinoma. Int. J. Cancer 2013, 133, 2914–2924. [Google Scholar] [CrossRef] [PubMed]
- Ross-Adams, H.; Lamb, A.D.; Dunning, M.J.; Halim, S.; Lindberg, J.; Massie, C.M.; Egevad, L.A.; Russell, R.; Ramos-Montoya, A.; Vowler, S.L.; et al. Integration of copy number and transcriptomics provides risk stratification in prostate cancer: A discovery and validation cohort study. EBioMedicine 2015, 2, 1133–1144. [Google Scholar] [CrossRef] [PubMed]
- Lalonde, E.; Alkallas, R.; Chua, M.L.; Fraser, M.; Haider, S.; Meng, A.; Zheng, J.; Yao, C.Q.; Picard, V.; Orain, M.; et al. Translating a prognostic DNA genomic classifier into the clinic: Retrospective validation in 563 localized prostate tumors. Eur. Urol. 2017, 72, 22–31. [Google Scholar] [CrossRef] [PubMed]
- Saad, F.; Latour, M.; Lattouf, J.B.; Widmer, H.; Zorn, K.C.; Mes-Masson, A.M.; Ouellet, V.; Saad, G.; Prakash, A.; Choudhury, S.; et al. Biopsy based proteomic assay predicts risk of biochemical recurrence after radical prostatectomy. J. Urol. 2017, 197, 1034–1040. [Google Scholar] [CrossRef] [PubMed]
- Filella, X.; Foj, L. Prostate cancer detection and prognosis: From prostate specific antigen (PSA) to exosomal biomarkers. Int. J. Mol. Sci. 2016, 17, 1784. [Google Scholar] [CrossRef] [PubMed]
- Luu, H.N.; Lin, H.Y.; Sorensen, K.D.; Ogunwobi, O.O.; Kumar, N.; Chornokur, G.; Phelan, C.; Jones, D.; Kidd, L.; Batra, J.; et al. miRNAs associated with prostate cancer risk and progression. BMC Urol. 2017, 17, 18. [Google Scholar] [CrossRef] [PubMed]
- Shukla, K.K.; Misra, S.; Pareek, P.; Mishra, V.; Singhal, B.; Sharma, P. Recent scenario of microRNA as diagnostic and prognostic biomarkers of prostate cancer. Urol. Oncol. 2017, 35, 92–101. [Google Scholar] [CrossRef] [PubMed]
- Fendler, A.; Stephan, C.; Yousef, G.M.; Jung, K. miRNAs as regulators of signal transduction in urological tumors. Clin. Chem. 2011, 57, 954–968. [Google Scholar] [CrossRef] [PubMed]
- Dietrich, D.; Meller, S.; Uhl, B.; Ralla, B.; Stephan, C.; Jung, K.; Ellinger, J.; Kristiansen, G. Nucleic acid-based tissue biomarkers of urologic malignancies. Crit. Rev. Clin. Lab. Sci. 2014, 51, 173–199. [Google Scholar] [CrossRef] [PubMed]
- Bertoli, G.; Cava, C.; Castiglioni, I. MicroRNAs as biomarkers for diagnosis, prognosis and theranostics in prostate cancer. Int. J. Mol. Sci. 2016, 17, 421. [Google Scholar] [CrossRef] [PubMed]
- Schubert, M.; Junker, K.; Heinzelmann, J. Prognostic and predictive miRNA biomarkers in bladder, kidney and prostate cancer: Where do we stand in biomarker development? J. Cancer Res. Clin. Oncol. 2016, 142, 1673–1695. [Google Scholar] [CrossRef] [PubMed]
- Gill, B.S.; Alex, J.M.; Navgeet; Kumar, S. Missing link between microRNA and prostate cancer. Tumour. Biol. 2016, 37, 5683–5704. [Google Scholar] [CrossRef] [PubMed]
- Kurozumi, A.; Goto, Y.; Okato, A.; Ichikawa, T.; Seki, N. Aberrantly expressed microRNAs in bladder cancer and renal cell carcinoma. J. Hum. Genet. 2017, 62, 49–56. [Google Scholar] [CrossRef] [PubMed]
- Tong, A.W.; Fulgham, P.; Jay, C.; Chen, P.; Khalil, I.; Liu, S.; Senzer, N.; Eklund, A.C.; Han, J.; Nemunaitis, J. MicroRNA profile analysis of human prostate cancers. Cancer Gene Ther. 2009, 16, 206–216. [Google Scholar] [CrossRef] [PubMed]
- Schaefer, A.; Jung, M.; Mollenkopf, H.J.; Wagner, I.; Stephan, C.; Jentzmik, F.; Miller, K.; Lein, M.; Kristiansen, G.; Jung, K. Diagnostic and prognostic implications of microRNA profiling in prostate carcinoma. Int. J. Cancer 2010, 126, 1166–1176. [Google Scholar] [CrossRef] [PubMed]
- Spahn, M.; Kneitz, S.; Scholz, C.J.; Stenger, N.; Rudiger, T.; Strobel, P.; Riedmiller, H.; Kneitz, B. Expression of microRNA-221 is progressively reduced in aggressive prostate cancer and metastasis and predicts clinical recurrence. Int. J. Cancer 2010, 127, 394–403. [Google Scholar] [PubMed]
- Fendler, A.; Jung, M.; Stephan, C.; Honey, R.J.; Stewart, R.J.; Pace, K.T.; Erbersdobler, A.; Samaan, S.; Jung, K.; Yousef, G.M. miRNAs can predict prostate cancer biochemical relapse and are involved in tumor progression. Int. J. Oncol. 2011, 39, 1183–1192. [Google Scholar] [PubMed]
- Amling, C.L.; Blute, M.L.; Bergstralh, E.J.; Seay, T.M.; Slezak, J.; Zincke, H. Long-term hazard of progression after radical prostatectomy for clinically localized prostate cancer: Continued risk of biochemical failure after 5 years. J. Urol. 2000, 164, 101–105. [Google Scholar] [CrossRef]
- Han, M.; Partin, A.W.; Zahurak, M.; Piantadosi, S.; Epstein, J.I.; Walsh, P.C. Biochemical (prostate specific antigen) recurrence probability following radical prostatectomy for clinically localized prostate cancer. J. Urol. 2003, 169, 517–523. [Google Scholar] [CrossRef]
- Lein, M.; Brux, B.; Jung, K.; Henke, W.; Koenig, F.; Stephan, C.; Schnorr, D.; Loening, S.A. Elimination of serum free and total prostate-specific antigen after radical retropubic prostatectomy. Eur. J. Clin. Chem. Clin. Biochem. 1997, 35, 591–595. [Google Scholar] [CrossRef] [PubMed]
- Cornford, P.; Bellmunt, J.; Bolla, M.; Briers, E.; de, S.M.; Gross, T.; Henry, A.M.; Joniau, S.; Lam, T.B.; Mason, M.D.; et al. EAU-ESTRO-SIOG Guidelines on Prostate Cancer. Part II: Treatment of relapsing, metastatic, and castration-resistant prostate cancer. Eur. Urol. 2017, 71, 630–642. [Google Scholar] [CrossRef] [PubMed]
- Cookson, M.S.; Aus, G.; Burnett, A.L.; Canby-Hagino, E.D.; D’Amico, A.V.; Dmochowski, R.R.; Eton, D.T.; Forman, J.D.; Goldenberg, S.L.; Hernandez, J.; et al. Variation in the definition of biochemical recurrence in patients treated for localized prostate cancer: The American Urological Association Prostate Guidelines for Localized Prostate Cancer Update Panel report and recommendations for a standard in the reporting of surgical outcomes. J. Urol. 2007, 177, 540–545. [Google Scholar] [PubMed]
- Amling, C.L.; Bergstralh, E.J.; Blute, M.L.; Slezak, J.M.; Zincke, H. Defining prostate specific antigen progression after radical prostatectomy: What is the most appropriate cut point? J. Urol. 2001, 165, 1146–1151. [Google Scholar] [CrossRef]
- Suardi, N.; Porter, C.R.; Reuther, A.M.; Walz, J.; Kodama, K.; Gibbons, R.P.; Correa, R.; Montorsi, F.; Graefen, M.; Huland, H.; et al. A nomogram predicting long-term biochemical recurrence after radical prostatectomy. Cancer 2008, 112, 1254–1263. [Google Scholar] [CrossRef] [PubMed]
- Walz, J.; Chun, F.K.; Klein, E.A.; Reuther, A.; Saad, F.; Graefen, M.; Huland, H.; Karakiewicz, P.I. Nomogram predicting the probability of early recurrence after radical prostatectomy for prostate cancer. J. Urol. 2009, 181, 601–607. [Google Scholar] [CrossRef] [PubMed]
- Aktas, B.K.; Ozden, C.; Bulut, S.; Tagci, S.; Erbay, G.; Gokkaya, C.S.; Baykam, M.M.; Memis, A. Evaluation of biochemical recurrence-free survival after radical prostatectomy by cancer of the prostate risk assessment post-surgical (CAPRA-S) score. Asian Pac. J. Cancer Prev. 2015, 16, 2527–2530. [Google Scholar] [CrossRef] [PubMed]
- Kang, M.; Jeong, C.W.; Choi, W.S.; Park, Y.H.; Cho, S.Y.; Lee, S.; Lee, S.B.; Ku, J.H.; Hong, S.K.; Byun, S.S.; et al. Pre- and post-operative nomograms to predict recurrence-free probability in korean men with clinically localized prostate cancer. PLoS ONE 2014, 9, e100053. [Google Scholar] [CrossRef] [PubMed]
- Hu, X.; Cammann, H.; Meyer, H.-A.; Jung, K.; Lu, H.; Leva, N.; Magheli, A.; Stephan, C.; Busch, J. Risk prediction models for biochemical recurrence after radical prostatectomy using prostate-specific anitigen and Gleason score. Asian J. Androl. 2014, 16, 897–901. [Google Scholar] [PubMed]
- Stephenson, A.J.; Kattan, M.W.; Eastham, J.A.; Dotan, Z.A.; Bianco, F.J., Jr.; Lilja, H.; Scardino, P.T. Defining biochemical recurrence of prostate cancer after radical prostatectomy: A proposal for a standardized definition. J. Clin. Oncol. 2006, 24, 3973–3978. [Google Scholar] [CrossRef] [PubMed]
- Mir, M.C.; Li, J.; Klink, J.C.; Kattan, M.W.; Klein, E.A.; Stephenson, A.J. Optimal definition of biochemical recurrence after radical prostatectomy depends on pathologic risk factors: Identifying candidates for early salvage therapy. Eur. Urol. 2014, 66, 204–210. [Google Scholar] [CrossRef] [PubMed]
- Tosoian, J.J.; Pierorazio, P.M. Optimizing use of serum prostate specific antigen to define biochemical recurrence—Is there a method to the madness? J. Urol. 2016, 195, 1648–1649. [Google Scholar] [CrossRef] [PubMed]
- Toussi, A.; Stewart-Merrill, S.B.; Boorjian, S.A.; Psutka, S.P.; Thompson, R.H.; Frank, I.; Tollefson, M.K.; Gettman, M.T.; Carlson, R.E.; Rangel, L.J.; et al. Standardizing the definition of biochemical recurrence after radical prostatectomy-what prostate specific antigen cut point best predicts a durable increase and subsequent systemic progression? J. Urol. 2016, 195, 1754–1759. [Google Scholar] [CrossRef] [PubMed]
- Pound, C.R.; Partin, A.W.; Eisenberger, M.A.; Chan, D.W.; Pearson, J.D.; Walsh, P.C. Natural history of progression after PSA elevation following radical prostatectomy. JAMA 1999, 281, 1591–1597. [Google Scholar] [CrossRef] [PubMed]
- D’Amico, A.V.; Whittington, R.; Malkowicz, S.B.; Weinstein, M.; Tomaszewski, J.E.; Schultz, D.; Rhude, M.; Rocha, S.; Wein, A.; Richie, J.P. Predicting prostate specific antigen outcome preoperatively in the prostate specific antigen era. J. Urol. 2001, 166, 2185–2188. [Google Scholar] [CrossRef]
- Freedland, S.J.; Sutter, M.E.; Dorey, F.; Aronson, W.J. Defining the ideal cutpoint for determining PSA recurrence after radical prostatectomy. Urology 2003, 61, 365–369. [Google Scholar] [CrossRef]
- Ward, J.F.; Moul, J.W. Biochemical recurrence after definitive prostate cancer therapy. Part I: Defining and localizing biochemical recurrence of prostate cancer. Curr. Opin. Urol. 2005, 15, 181–186. [Google Scholar] [CrossRef] [PubMed]
- Stephan, C.; Klaas, M.; Müller, C.; Schnorr, D.; Loening, S.A.; Jung, K. Interchangeability of measurements of total and free prostate-specific antigen in serum with 5 frequently used assay combinations: An update. Clin. Chem. 2006, 52, 59–64. [Google Scholar] [CrossRef] [PubMed]
- Stephan, C.; Kahrs, A.-M.; Klotzek, S.; Reiche, J.; Müller, C.; Lein, M.; Deger, S.; Miller, K.; Jung, K. Toward metrological traceability in the determination of prostate-specific antigen (PSA): Calibrating Beckman Coulter Hybritech Access PSA assays to WHO standards compared with the traditional Hybritech standards. Clin. Chem. Lab. Med. 2008, 46, 623–629. [Google Scholar] [CrossRef] [PubMed]
- Boorjian, S.A.; Thompson, R.H.; Tollefson, M.K.; Rangel, L.J.; Bergstralh, E.J.; Blute, M.L.; Karnes, R.J. Long-term risk of clinical progression after biochemical recurrence following radical prostatectomy: The impact of time from surgery to recurrence. Eur. Urol. 2011, 59, 893–899. [Google Scholar] [CrossRef] [PubMed]
- Herranz-Amo, F.; Molina-Escudero, R.; Ogaya-Pinies, G.; Ramirez-Martin, D.; Verdu-Tartajo, F.; Hernandez-Fernandez, C. Prediction of biochemical recurrence after radical prostatectomy. New tool for selecting candidates for adjuvant radiation therapy. Actas Urol. Esp. 2016, 40, 82–87. [Google Scholar] [CrossRef] [PubMed]
- Casanova-Salas, I.; Rubio-Briones, J.; Calatrava, A.; Mancarella, C.; Masia, E.; Casanova, J.; Fernandez-Serra, A.; Rubio, L.; Ramirez-Backhaus, M.; Arminan, A.; et al. Identification of miR-187 and miR-182 as biomarkers of early diagnosis and prognosis in patients with prostate cancer treated with radical prostatectomy. J. Urol. 2014, 192, 252–259. [Google Scholar] [CrossRef] [PubMed]
- Kristensen, H.; Thomsen, A.R.; Haldrup, C.; Dyrskjot, L.; Hoyer, S.; Borre, M.; Mouritzen, P.; Orntoft, T.F.; Sorensen, K.D. Novel diagnostic and prognostic classifiers for prostate cancer identified by genome-wide microRNA profiling. Oncotarget 2016, 7, 30760–30771. [Google Scholar] [CrossRef] [PubMed]
- Fendler, A.; Stephan, C.; Yousef, G.M.; Kristiansen, G.; Jung, K. The translational potential of microRNAs as biofluid markers of urologic tumors. Nat. Rev. Urol. 2016, 13, 734–752. [Google Scholar] [CrossRef] [PubMed]
- Leite, K.R.; Tomiyama, A.; Reis, S.T.; Sousa-Canavez, J.M.; Sanudo, A.; Dall'Oglio, M.F.; Camara-Lopes, L.H.; Srougi, M. MicroRNA-100 expression is independently related to biochemical recurrence of prostate cancer. J. Urol. 2011, 185, 1118–1122. [Google Scholar] [CrossRef] [PubMed]
- Long, Q.; Johnson, B.A.; Osunkoya, A.O.; Lai, Y.H.; Zhou, W.; Abramovitz, M.; Xia, M.; Bouzyk, M.B.; Nam, R.K.; Sugar, L.; et al. Protein-coding and microRNA biomarkers of recurrence of prostate cancer following radical prostatectomy. Am. J. Pathol. 2011, 179, 46–54. [Google Scholar] [CrossRef] [PubMed]
- Barron, N.; Keenan, J.; Gammell, P.; Martinez, V.G.; Freeman, A.; Masters, J.R.; Clynes, M. Biochemical relapse following radical prostatectomy and miR-200a levels in prostate cancer. Prostate 2012, 72, 1193–1199. [Google Scholar] [CrossRef] [PubMed]
- Hudson, R.S.; Yi, M.; Esposito, D.; Watkins, S.K.; Hurwitz, A.A.; Yfantis, H.G.; Lee, D.H.; Borin, J.F.; Naslund, M.J.; Alexander, R.B.; et al. MicroRNA-1 is a candidate tumor suppressor and prognostic marker in human prostate cancer. Nucleic Acids Res. 2012, 40, 3689–3703. [Google Scholar] [CrossRef] [PubMed]
- Ambs, S.; Prueitt, R.L.; Yi, M.; Hudson, R.S.; Howe, T.M.; Petrocca, F.; Wallace, T.A.; Liu, C.G.; Volinia, S.; Calin, G.A.; et al. Genomic profiling of microRNA and messenger RNA reveals deregulated microRNA expression in prostate cancer. Cancer Res. 2008, 68, 6162–6170. [Google Scholar] [CrossRef] [PubMed]
- Taylor, B.S.; Schultz, N.; Hieronymus, H.; Gopalan, A.; Xiao, Y.; Carver, B.S.; Arora, V.K.; Kaushik, P.; Cerami, E.; et al. Integrative genomic profiling of human prostate cancer. Cancer Cell 2010, 18, 11–22. [Google Scholar] [CrossRef] [PubMed]
- Kang, S.G.; Ha, Y.R.; Kim, S.J.; Kang, S.H.; Park, H.S.; Lee, J.G.; Cheon, J.; Kim, C.H. Do microRNA 96, 145 and 221 expressions really aid in the prognosis of prostate carcinoma? Asian J. Androl. 2012, 14, 752–757. [Google Scholar] [CrossRef] [PubMed]
- Kobayashi, N.; Uemura, H.; Nagahama, K.; Okudela, K.; Furuya, M.; Ino, Y.; Ito, Y.; Hirano, H.; Inayama, Y.; Aoki, I.; et al. Identification of miR-30d as a novel prognostic maker of prostate cancer. Oncotarget 2012, 3, 1455–1471. [Google Scholar] [CrossRef] [PubMed]
- Li, T.; Li, R.S.; Li, Y.H.; Zhong, S.; Chen, Y.Y.; Zhang, C.M.; Hu, M.M.; Shen, Z.J. miR-21 as an independent biochemical recurrence predictor and potential therapeutic target for prostate cancer. J. Urol. 2012, 187, 1466–1472. [Google Scholar] [CrossRef] [PubMed]
- Li, T.; Li, D.; Sha, J.; Sun, P.; Huang, Y. MicroRNA-21 directly targets MARCKS and promotes apoptosis resistance and invasion in prostate cancer cells. Biochem. Biophys. Res. Commun. 2009, 383, 280–285. [Google Scholar] [CrossRef] [PubMed]
- Majid, S.; Dar, A.A.; Saini, S.; Arora, S.; Shahryari, V.; Zaman, M.S.; Chang, I.; Yamamura, S.; Tanaka, Y.; Deng, G.; et al. miR-23b represses proto-oncogene Src kinase and functions as methylation-silenced tumor suppressor with diagnostic and prognostic significance in prostate cancer. Cancer Res. 2012, 72, 6435–6446. [Google Scholar] [CrossRef] [PubMed]
- Saini, S.; Majid, S.; Shahryari, V.; Arora, S.; Yamamura, S.; Chang, I.; Zaman, M.S.; Deng, G.; Tanaka, Y.; Dahiya, R. miRNA-708 control of CD44+ prostate cancer-initiating cells. Cancer Res. 2012, 72, 3618–3630. [Google Scholar] [CrossRef] [PubMed]
- Amankwah, E.K.; Anegbe, E.; Park, H.; Pow-Sang, J.; Hakam, A.; Park, J.Y. miR-21, miR-221 and miR-222 expression and prostate cancer recurrence among obese and non-obese cases. Asian J. Androl. 2013, 15, 226–230. [Google Scholar] [CrossRef] [PubMed]
- Avgeris, M.; Stravodimos, K.; Fragoulis, E.G.; Scorilas, A. The loss of the tumour-suppressor miR-145 results in the shorter disease-free survival of prostate cancer patients. Br. J. Cancer 2013, 108, 2573–2581. [Google Scholar] [CrossRef] [PubMed]
- He, H.C.; Han, Z.D.; Dai, Q.S.; Ling, X.H.; Fu, X.; Lin, Z.Y.; Deng, Y.H.; Qin, G.Q.; Cai, C.; Chen, J.H.; et al. Global analysis of the differentially expressed miRNAs of prostate cancer in Chinese patients. BMC Genomics 2013, 14, 757. [Google Scholar] [CrossRef] [PubMed]
- Larne, O.; Martens-Uzunova, E.; Hagman, Z.; Edsjo, A.; Lippolis, G.; den Berg, M.S.; Bjartell, A.; Jenster, G.; Ceder, Y. miQ—A novel microRNA based diagnostic and prognostic tool for prostate cancer. Int. J. Cancer 2013, 132, 2867–2875. [Google Scholar] [CrossRef] [PubMed]
- Martens-Uzunova, E.S.; Jalava, S.E.; Dits, N.F.; van Leenders, G.J.; Moller, S.; Trapman, J.; Bangma, C.H.; Litman, T.; Visakorpi, T.; Jenster, G. Diagnostic and prognostic signatures from the small non-coding RNA transcriptome in prostate cancer. Oncogene 2012, 31, 978–991. [Google Scholar] [CrossRef] [PubMed]
- Lichner, Z.; Fendler, A.; Saleh, C.; Nasser, A.N.; Boles, D.; Al-Haddad, S.; Kupchak, P.; Dharsee, M.; Nuin, P.S.; Evans, K.R.; et al. MicroRNA signature helps distinguish early from late biochemical failure in prostate cancer. Clin. Chem. 2013, 59, 1595–1603. [Google Scholar] [CrossRef] [PubMed]
- Majid, S.; Dar, A.A.; Saini, S.; Shahryari, V.; Arora, S.; Zaman, M.S.; Chang, I.; Yamamura, S.; Tanaka, Y.; Chiyomaru, T.; et al. miRNA-34b inhibits prostate cancer through demethylation, active chromatin modifications, and AKT pathways. Clin. Cancer Res. 2013, 19, 73–84. [Google Scholar] [CrossRef] [PubMed]
- Schubert, M.; Spahn, M.; Kneitz, S.; Scholz, C.J.; Joniau, S.; Stroebel, P.; Riedmiller, H.; Kneitz, B. Distinct microRNA expression profile in prostate cancer patients with early clinical failure and the impact of let-7 as prognostic marker in high-risk prostate cancer. PLoS ONE 2013, 8, e65064. [Google Scholar] [CrossRef] [PubMed]
- Sun, X.; Liu, Z.; Yang, Z.; Xiao, L.; Wang, F.; He, Y.; Su, P.; Wang, J.; Jing, B. Association of microRNA-126 expression with clinicopathological features and the risk of biochemical recurrence in prostate cancer patients undergoing radical prostatectomy. Diagn. Pathol. 2013, 8, 208. [Google Scholar] [CrossRef] [PubMed]
- Avgeris, M.; Stravodimos, K.; Scorilas, A. Loss of miR-378 in prostate cancer, a common regulator of KLK2 and KLK4, correlates with aggressive disease phenotype and predicts the short-term relapse of the patients. Biol. Chem. 2014, 395, 1095–1104. [Google Scholar] [CrossRef] [PubMed]
- Karatas, O.F.; Guzel, E.; Suer, I.; Ekici, I.D.; Caskurlu, T.; Creighton, C.J.; Ittmann, M.; Ozen, M. miR-1 and miR-133b are differentially expressed in patients with recurrent prostate cancer. PLoS ONE 2014, 9, e98675. [Google Scholar] [CrossRef] [PubMed]
- Katz, B.; Reis, S.T.; Viana, N.I.; Morais, D.R.; Moura, C.M.; Dip, N.; Silva, I.A.; Iscaife, A.; Srougi, M.; Leite, K.R. Comprehensive study of gene and microRNA expression related to epithelial-mesenchymal transition in prostate cancer. PLoS ONE 2014, 9, e113700. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Wan, X.; Chen, H.; Yang, S.; Liu, Y.; Mo, W.; Meng, D.; Du, W.; Huang, Y.; Wu, H.; et al. Identification of miR-133b and RB1CC1 as independent predictors for biochemical recurrence and potential therapeutic targets for prostate cancer. Clin. Cancer Res. 2014, 20, 2312–2325. [Google Scholar] [CrossRef] [PubMed]
- Lin, Z.Y.; Huang, Y.Q.; Zhang, Y.Q.; Han, Z.D.; He, H.C.; Ling, X.H.; Fu, X.; Dai, Q.S.; Cai, C.; Chen, J.H.; et al. MicroRNA-224 inhibits progression of human prostate cancer by downregulating TRIB1. Int. J. Cancer 2014, 135, 541–550. [Google Scholar] [CrossRef] [PubMed]
- Ling, X.H.; Han, Z.D.; Xia, D.; He, H.C.; Jiang, F.N.; Lin, Z.Y.; Fu, X.; Deng, Y.H.; Dai, Q.S.; Cai, C.; et al. MicroRNA-30c serves as an independent biochemical recurrence predictor and potential tumor suppressor for prostate cancer. Mol. Biol. Rep. 2014, 41, 2779–2788. [Google Scholar] [CrossRef] [PubMed]
- He, H.C.; Zhu, J.G.; Chen, X.B.; Chen, S.M.; Han, Z.D.; Dai, Q.S.; Ling, X.H.; Fu, X.; Lin, Z.Y.; Deng, Y.H.; et al. MicroRNA-23b downregulates peroxiredoxin III in human prostate cancer. FEBS Lett 2012, 586, 2451–2458. [Google Scholar] [CrossRef] [PubMed]
- Xiong, S.W.; Lin, T.X.; Xu, K.W.; Dong, W.; Ling, X.H.; Jiang, F.N.; Chen, G.; Zhong, W.D.; Huang, J. MicroRNA-335 acts as a candidate tumor suppressor in prostate cancer. Pathol. Oncol. Res. 2013, 19, 529–537. [Google Scholar] [CrossRef] [PubMed]
- Melbo-Jorgensen, C.; Ness, N.; Andersen, S.; Valkov, A.; Donnem, T.; Al-Saad, S.; Kiselev, Y.; Berg, T.; Nordby, Y.; Bremnes, R.M.; et al. Stromal expression of miR-21 predicts biochemical failure in prostate cancer patients with Gleason score 6. PLoS ONE 2014, 9, e113039. [Google Scholar] [CrossRef] [PubMed]
- Mortensen, M.M.; Hoyer, S.; Orntoft, T.F.; Sorensen, K.D.; Dyrskjot, L.; Borre, M. High miR-449b expression in prostate cancer is associated with biochemical recurrence after radical prostatectomy. BMC Cancer 2014, 14, 859. [Google Scholar] [CrossRef] [PubMed]
- Zheng, Q.; Peskoe, S.B.; Ribas, J.; Rafiqi, F.; Kudrolli, T.; Meeker, A.K.; de Marzo, A.M.; Platz, E.A.; Lupold, S.E. Investigation of miR-21, miR-141, and miR-221 expression levels in prostate adenocarcinoma for associated risk of recurrence after radical prostatectomy. Prostate 2014, 74, 1655–1662. [Google Scholar] [CrossRef] [PubMed]
- Bell, E.H.; Kirste, S.; Fleming, J.L.; Stegmaier, P.; Drendel, V.; Mo, X.; Ling, S.; Fabian, D.; Manring, I.; Jilg, C.A.; et al. A novel miRNA-based predictive model for biochemical failure following post-prostatectomy salvage radiation therapy. PLoS ONE 2015, 10, e0118745. [Google Scholar] [CrossRef] [PubMed]
- Cai, C.; Chen, Q.B.; Han, Z.D.; Zhang, Y.Q.; He, H.C.; Chen, J.H.; Chen, Y.R.; Yang, S.B.; Wu, Y.D.; Zeng, Y.R.; et al. miR-195 inhibits tumor progression by targeting RPS6KB1 in human prostate cancer. Clin. Cancer Res. 2015, 21, 4922–4934. [Google Scholar] [CrossRef] [PubMed]
- Guo, J.; Wang, M.; Liu, X. MicroRNA-195 suppresses tumor cell proliferation and metastasis by directly targeting BCOX1 in prostate carcinoma. J. Exp. Clin. Cancer Res. 2015, 34, 91. [Google Scholar] [CrossRef] [PubMed]
- Leite, K.R.; Reis, S.T.; Viana, N.; Morais, D.R.; Moura, C.M.; Silva, I.A.; Pontes, J., Jr.; Katz, B.; Srougi, M. Controlling RECK miR21 promotes tumor cell invasion and is related to biochemical recurrence in prostate cancer. J. Cancer 2015, 6, 292–301. [Google Scholar] [CrossRef] [PubMed]
- Lichner, Z.; Ding, Q.; Samaan, S.; Saleh, C.; Nasser, A.; Al-Haddad, S.; Samuel, J.N.; Fleshner, N.E.; Stephan, C.; Jung, K.; et al. miRNAs dysregulated in association with Gleason grade regulate extracellular matrix, cytoskeleton and androgen receptor pathways. J. Pathol. 2015, 237, 226–237. [Google Scholar] [CrossRef] [PubMed]
- Nam, R.K.; Amemiya, Y.; Benatar, T.; Wallis, C.J.; Stojcic-Bendavid, J.; Bacopulos, S.; Sherman, C.; Sugar, L.; Naeim, M.; Yang, W.; et al. Identification and validation of a five microrna signature predictive of prostate cancer recurrence and metastasis: A cohort study. J. Cancer 2015, 6, 1160–1171. [Google Scholar] [CrossRef] [PubMed]
- Sun, X.; Yang, Z.; Zhang, Y.; He, J.; Wang, F.; Su, P.; Han, J.; Song, Z.; Fei, Y. Prognostic implications of tissue and serum levels of microRNA-128 in human prostate cancer. Int. J. Clin. Exp. Pathol. 2015, 8, 8394–8401. [Google Scholar] [PubMed]
- Khan, A.P.; Poisson, L.M.; Bhat, V.B.; Fermin, D.; Zhao, R.; Kalyana-Sundaram, S.; Michailidis, G.; Nesvizhskii, A.I.; Omenn, G.S.; Chinnaiyan, A.M.; et al. Quantitative proteomic profiling of prostate cancer reveals a role for miR-128 in prostate cancer. Mol. Cell Proteom. 2010, 9, 298–312. [Google Scholar] [CrossRef] [PubMed]
- Tian, B.; Huo, N.; Li, M.; Li, Y.; He, Z. let-7a And its target, insulin-like growth factor 1 receptor, are differentially expressed in recurrent prostate cancer. Int. J. Mol. Med. 2015, 36, 1409–1416. [Google Scholar] [CrossRef] [PubMed]
- Wallis, C.J.; Gordanpour, A.; Bendavid, J.S.; Sugar, L.; Nam, R.K.; Seth, A. mir-182 is associated with growth, migration and invasion in prostate cancer via suppression of FOXO1. J. Cancer 2015, 6, 1295–1305. [Google Scholar] [CrossRef] [PubMed]
- Wan, Y.; Zeng, Z.C.; Xi, M.; Wan, S.; Hua, W.; Liu, Y.L.; Zhou, Y.L.; Luo, H.W.; Jiang, F.N.; Zhong, W.D. Dysregulated microRNA-224/apelin axis associated with aggressive progression and poor prognosis in patients with prostate cancer. Hum. Pathol. 2015, 46, 295–303. [Google Scholar] [CrossRef] [PubMed]
- Xu, B.; Huang, Y.; Niu, X.; Tao, T.; Jiang, L.; Tong, N.; Chen, S.; Liu, N.; Zhu, W.; Chen, M. Hsa-miR-146a-5p modulates androgen-independent prostate cancer cells apoptosis by targeting ROCK1. Prostate 2015, 75, 1896–1903. [Google Scholar] [CrossRef] [PubMed]
- Bakkar, A.; Alshalalfa, M.; Petersen, L.F.; Abou-Ouf, H.; Al-Mami, A.; Hegazy, S.A.; Feng, F.; Alhajj, R.; Bijian, K.; Alaoui-Jamali, M.A.; et al. microRNA 338-3p exhibits tumor suppressor role and its down-regulation is associated with adverse clinical outcome in prostate cancer patients. Mol. Biol. Rep. 2016, 43, 229–240. [Google Scholar] [CrossRef] [PubMed]
- Bucay, N.; Sekhon, K.; Majid, S.; Yamamura, S.; Shahryari, V.; Tabatabai, Z.L.; Greene, K.; Tanaka, Y.; Dahiya, R.; Deng, G.; et al. Novel tumor suppressor microRNA at frequently deleted chromosomal region 8p21 regulates epidermal growth factor receptor in prostate cancer. Oncotarget 2016, 7, 70388–70403. [Google Scholar] [CrossRef] [PubMed]
- Das, D.K.; Osborne, J.R.; Lin, H.Y.; Park, J.Y.; Ogunwobi, O.O. miR-1207-3p is a novel prognostic biomarker of prostate cancer. Transl. Oncol. 2016, 9, 236–241. [Google Scholar] [CrossRef] [PubMed]
- Ling, X.H.; Chen, Z.Y.; Luo, H.W.; Liu, Z.Z.; Liang, Y.K.; Chen, G.X.; Jiang, F.N.; Zhong, W.D. BCL9, a coactivator for Wnt/β-catenin transcription, is targeted by miR-30c and is associated with prostate cancer progression. Oncol. Lett. 2016, 11, 2001–2008. [Google Scholar] [CrossRef] [PubMed]
- Nam, R.K.; Benatar, T.; Wallis, C.J.; Amemiya, Y.; Yang, W.; Garbens, A.; Naeim, M.; Sherman, C.; Sugar, L.; Seth, A. miR-301a regulates E-cadherin expression and is predictive of prostate cancer recurrence. Prostate 2016, 76, 869–884. [Google Scholar] [CrossRef] [PubMed]
- Nip, H.; Dar, A.A.; Saini, S.; Colden, M.; Varahram, S.; Chowdhary, H.; Yamamura, S.; Mitsui, Y.; Tanaka, Y.; Kato, T.; et al. Oncogenic microRNA-4534 regulates PTEN pathway in prostate cancer. Oncotarget 2016, 7, 68371–68384. [Google Scholar] [CrossRef] [PubMed]
- Xu, S.; Yi, X.M.; Zhang, Z.Y.; Ge, J.P.; Zhou, W.Q. miR-129 predicts prognosis and inhibits cell growth in human prostate carcinoma. Mol. Med. Rep. 2016, 14, 5025–5032. [Google Scholar] [CrossRef] [PubMed]
- Zhai, J.; Qu, S.; Li, X.; Zhong, J.; Chen, X.; Qu, Z.; Wu, D. miR-129 suppresses tumor cell growth and invasion by targeting PAK5 in hepatocellular carcinoma. Biochem. Biophys. Res. Commun. 2015, 464, 161–167. [Google Scholar] [CrossRef] [PubMed]
- Colden, M.; Dar, A.A.; Saini, S.; Dahiya, P.V.; Shahryari, V.; Yamamura, S.; Tanaka, Y.; Stein, G.; Dahiya, R.; Majid, S. MicroRNA-466 inhibits tumor growth and bone metastasis in prostate cancer by direct regulation of osteogenic transcription factor RUNX2. Cell Death Dis. 2017, 8, e2572. [Google Scholar] [CrossRef] [PubMed]
- Lin, Z.Y.; Chen, G.; Zhang, Y.Q.; He, H.C.; Liang, Y.X.; Ye, J.H.; Liang, Y.K.; Mo, R.J.; Lu, J.M.; Zhuo, Y.J.; et al. MicroRNA-30d promotes angiogenesis and tumor growth via MYPT1/c-JUN/VEGFA pathway and predicts aggressive outcome in prostate cancer. Mol. Cancer 2017, 16, 48. [Google Scholar] [CrossRef] [PubMed]
- Xuan, H.; Xue, W.; Pan, J.; Sha, J.; Dong, B.; Huang, Y. Downregulation of miR-221, -30d, and -15a contributes to pathogenesis of prostate cancer by targeting BMI-1. Biochemistry 2015, 80, 276–283. [Google Scholar] [CrossRef] [PubMed]
- Wei, W.; Leng, J.; Shao, H.; Wang, W. miR-1, A potential predictive biomarker for recurrence in prostate cancer after radical prostatectomy. Am. J. Med. Sci. 2017, 353, 315–319. [Google Scholar] [CrossRef] [PubMed]
- Xi, Y.; Nakajima, G.; Gavin, E.; Morris, C.G.; Kudo, K.; Hayashi, K.; Ju, J. Systematic analysis of microRNA expression of RNA extracted from fresh frozen and formalin-fixed paraffin-embedded samples. RNA 2007, 13, 1668–1674. [Google Scholar] [CrossRef] [PubMed]
- Szafranska, A.E.; Davison, T.S.; Shingara, J.; Doleshal, M.; Riggenbach, J.A.; Morrison, C.D.; Jewell, S.; Labourier, E. Accurate molecular characterization of formalin-fixed, paraffin-embedded tissues by microRNA expression profiling. J. Mol. Diagn. 2008, 10, 415–423. [Google Scholar] [CrossRef] [PubMed]
- Leite, K.R.; Canavez, J.M.; Reis, S.T.; Tomiyama, A.H.; Piantino, C.B.; Sanudo, A.; Camara-Lopes, L.H.; Srougi, M. miRNA analysis of prostate cancer by quantitative real time PCR: Comparison between formalin-fixed paraffin embedded and fresh-frozen tissue. Urol. Oncol. 2011, 29, 533–537. [Google Scholar] [CrossRef] [PubMed]
- Liu, A.; Tetzlaff, M.T.; Vanbelle, P.; Elder, D.; Feldman, M.; Tobias, J.W.; Sepulveda, A.R.; Xu, X. MicroRNA expression profiling outperforms mRNA expression profiling in formalin-fixed paraffin-embedded tissues. Int. J. Clin. Exp. Pathol. 2009, 2, 519–527. [Google Scholar] [PubMed]
- Jung, M.; Schaefer, A.; Steiner, I.; Kempkensteffen, C.; Stephan, C.; Erbersdobler, A.; Jung, K. Robust microRNA stability in degraded RNA preparations from human tissue and cell samples. Clin. Chem. 2010, 56, 998–1006. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Smyth, P.; Flavin, R.; Cahill, S.; Denning, K.; Aherne, S.; Guenther, S.M.; O’Leary, J.J.; Sheils, O. Comparison of miRNA expression patterns using total RNA extracted from matched samples of formalin-fixed paraffin-embedded (FFPE) cells and snap frozen cells. BMC Biotechnol. 2007, 7, 36. [Google Scholar] [CrossRef] [PubMed]
- Peskoe, S.B.; Barber, J.R.; Zheng, Q.; Meeker, A.K.; de Marzo, A.M.; Platz, E.A.; Lupold, S.E. Differential long-term stability of microRNAs and RNU6B snRNA in 12–20 year old archived formalin-fixed paraffin-embedded specimens. BMC Cancer 2017, 17, 32. [Google Scholar] [CrossRef] [PubMed]
- Kakimoto, Y.; Tanaka, M.; Kamiguchi, H.; Ochiai, E.; Osawa, M. MicroRNA stability in FFPE tissue samples: Dependence on GC content. PLoS ONE 2016, 11, e0163125. [Google Scholar] [CrossRef] [PubMed]
- Carlsson, J.; Helenius, G.; Karlsson, M.; Lubovac, Z.; Andren, O.; Olsson, B.; Klinga-Levan, K. Validation of suitable endogenous control genes for expression studies of miRNA in prostate cancer tissues. Cancer Genet. Cytogenet. 2010, 202, 71–75. [Google Scholar] [CrossRef] [PubMed]
- Bovelstad, H.M.; Nygard, S.; Borgan, O. Survival prediction from clinico-genomic models—A comparative study. BMC Bioinform. 2009, 10, 413. [Google Scholar] [CrossRef] [PubMed]
- Klein, E.A.; Cooperberg, M.R.; Magi-Galluzzi, C.; Simko, J.P.; Falzarano, S.M.; Maddala, T.; Chan, J.M.; Li, J.; Cowan, J.E.; Tsiatis, A.C.; et al. A 17-gene assay to predict prostate cancer aggressiveness in the context of Gleason grade heterogeneity, tumor multifocality, and biopsy undersampling. Eur. Urol. 2014, 66, 550–560. [Google Scholar] [CrossRef] [PubMed]
- Gerszten, R.E.; Wang, T.J. The search for new cardiovascular biomarkers. Nature 2008, 451, 949–952. [Google Scholar] [CrossRef] [PubMed]
- Klein, M.E.; Dabbs, D.J.; Shuai, Y.; Brufsky, A.M.; Jankowitz, R.; Puhalla, S.L.; Bhargava, R. Prediction of the Oncotype DX recurrence score: Use of pathology-generated equations derived by linear regression analysis. Mod. Pathol. 2013, 26, 658–664. [Google Scholar] [CrossRef] [PubMed]
- Xiao, K.; Guo, J.; Zhang, X.; Feng, X.; Zhang, H.; Cheng, Z.; Johnson, H.; Persson, J.L.; Chen, L. Use of two gene panels for prostate cancer diagnosis and patient risk stratification. Tumour. Biol 2016, 37, 10115–10122. [Google Scholar] [CrossRef] [PubMed]
- Ross, A.E.; Johnson, M.H.; Yousefi, K.; Davicioni, E.; Netto, G.J.; Marchionni, L.; Fedor, H.L.; Glavaris, S.; Choeurng, V.; Buerki, C.; et al. Tissue-based genomics augments post-prostatectomy risk stratification in a natural history cohort of intermediate- and high-risk men. Eur. Urol. 2016, 69, 157–165. [Google Scholar] [CrossRef] [PubMed]
- Pritchard, C.C.; Cheng, H.H.; Tewari, M. MicroRNA profiling: Approaches and considerations. Nat. Rev. Genet. 2012, 13, 358–369. [Google Scholar] [CrossRef] [PubMed]
- Meng, W.; McElroy, J.P.; Volinia, S.; Palatini, J.; Warner, S.; Ayers, L.W.; Palanichamy, K.; Chakravarti, A.; Lautenschlaeger, T. Comparison of microRNA deep sequencing of matched formalin-fixed paraffin-embedded and fresh frozen cancer tissues. PLoS ONE 2013, 8, e64393. [Google Scholar]
- Nagy, Z.B.; Wichmann, B.; Kalmar, A.; Bartak, B.K.; Tulassay, Z.; Molnar, B. miRNA isolation from FFPET specimen: A technical comparison of miRNA and total RNA isolation methods. Pathol. Oncol. Res. 2016, 22, 505–513. [Google Scholar] [CrossRef] [PubMed]
- Howe, K. Extraction of miRNAs from formalin-fixed paraffin-embedded (FFPE) tissues. Methods Mol. Biol. 2017, 1509, 17–24. [Google Scholar] [PubMed]
- Doleshal, M.; Magotra, A.A.; Choudhury, B.; Cannon, B.D.; Labourier, E.; Szafranska, A.E. Evaluation and validation of total RNA extraction methods for microRNA expression analyses in formalin-fixed, paraffin-embedded tissues. J. Mol. Diagn. 2008, 10, 203–211. [Google Scholar] [CrossRef] [PubMed]
- ThermoFisher Scientific. TaqMan Advanced miRNA Assays—Superior Performance for miRNA Detection and Quantification. Available online: https://www.thermofisher.com/content/dam/LifeTech/Documents/PDFs/TaqMan-Advanced-miRNA-Performance-White-Paper.pdf (accessed on 20 September 2017).
- Bustin, S.A.; Benes, V.; Garson, J.A.; Hellemans, J.; Huggett, J.; Kubista, M.; Mueller, R.; Nolan, T.; Pfaffl, M.W.; Shipley, G.L.; et al. The MIQE guidelines: Minimum information for publication of quantitative real-time PCR experiments. Clin. Chem. 2009, 55, 611–622. [Google Scholar] [CrossRef] [PubMed]
- Bustin, S.A.; Benes, V.; Garson, J.; Hellemans, J.; Huggett, J.; Kubista, M.; Mueller, R.; Nolan, T.; Pfaffl, M.W.; Shipley, G.; et al. The need for transparency and good practices in the qPCR literature. Nat. Methods 2013, 10, 1063–1067. [Google Scholar] [CrossRef] [PubMed]
- Altman, D.G.; McShane, L.M.; Sauerbrei, W.; Taube, S.E. Reporting recommendations for tumor marker prognostic studies (REMARK): Explanation and elaboration. PLoS Med 2012, 9, e1001216. [Google Scholar] [CrossRef] [PubMed]
- Bossuyt, P.M.; Reitsma, J.B.; Bruns, D.E.; Gatsonis, C.A.; Glasziou, P.P.; Irwig, L.; Lijmer, J.G.; Moher, D.; Rennie, D.; de Vet, H.C.; et al. STARD 2015: An updated list of essential items for reporting diagnostic accuracy studies. Clin. Chem. 2015, 61, 1446–1452. [Google Scholar] [CrossRef] [PubMed]
- Peduzzi, P.; Concato, J.; Feinstein, A.R.; Holford, T.R. Importance of events per independent variable in proportional hazards regression analysis II. Accuracy and precision of regression estimates. J. Clin. Epidemiol. 1995, 48, 1503–1510. [Google Scholar] [CrossRef]
- Nadiminty, N.; Tummala, R.; Lou, W.; Zhu, Y.; Zhang, J.; Chen, X.; eVere White, R.W.; Kung, H.J.; Evans, C.P.; Gao, A.C. MicroRNA let-7c suppresses androgen receptor expression and activity via regulation of Myc expression in prostate cancer cells. J. Biol Chem. 2012, 287, 1527–1537. [Google Scholar] [CrossRef] [PubMed]
- Ozen, M.; Creighton, C.J.; Ozdemir, M.; Ittmann, M. Widespread deregulation of microRNA expression in human prostate cancer. Oncogene 2008, 27, 1788–1793. [Google Scholar] [CrossRef] [PubMed]
- Murata, T.; Takayama, K.; Katayama, S.; Urano, T.; Horie-Inoue, K.; Ikeda, K.; Takahashi, S.; Kawazu, C.; Hasegawa, A.; Ouchi, Y.; et al. miR-148a is an androgen-responsive microRNA that promotes LNCaP prostate cell growth by repressing its target CAND1 expression. Prostate Cancer Prostatic. Dis. 2010, 13, 356–361. [Google Scholar] [CrossRef] [PubMed]
- Waltering, K.K.; Porkka, K.P.; Jalava, S.E.; Urbanucci, A.; Kohonen, P.J.; Latonen, L.M.; Kallioniemi, O.P.; Jenster, G.; Visakorpi, T. Androgen regulation of micro-RNAs in prostate cancer. Prostate 2011, 71, 604–614. [Google Scholar] [CrossRef] [PubMed]
- Coarfa, C.; Fiskus, W.; Eedunuri, V.K.; Rajapakshe, K.; Foley, C.; Chew, S.A.; Shah, S.S.; Geng, C.; Shou, J.; Mohamed, J.S.; et al. Comprehensive proteomic profiling identifies the androgen receptor axis and other signaling pathways as targets of microRNAs suppressed in metastatic prostate cancer. Oncogene 2016, 35, 2345–2356. [Google Scholar] [CrossRef] [PubMed]
- Kneitz, B.; Krebs, M.; Kalogirou, C.; Schubert, M.; Joniau, S.; van, P.H.; Lerut, E.; Kneitz, S.; Scholz, C.J.; Strobel, P.; et al. Survival in patients with high-risk prostate cancer is predicted by miR-221, which regulates proliferation, apoptosis, and invasion of prostate cancer cells by inhibiting IRF2 and SOCS3. Cancer Res. 2014, 74, 2591–2603. [Google Scholar] [CrossRef] [PubMed]
- Gui, B.; Hsieh, C.L.; Kantoff, P.W.; Kibel, A.S.; Jia, L. Androgen receptor-mediated downregulation of microRNA-221 and -222 in castration-resistant prostate cancer. PLoS ONE 2017, 12, e0184166. [Google Scholar] [CrossRef] [PubMed]
- Goto, Y.; Kojima, S.; Nishikawa, R.; Kurozumi, A.; Kato, M.; Enokida, H.; Matsushita, R.; Yamazaki, K.; Ishida, Y.; Nakagawa, M.; et al. MicroRNA expression signature of castration-resistant prostate cancer: The microRNA-221/222 cluster functions as a tumour suppressor and disease progression marker. Br. J. Cancer 2015, 113, 1055–1065. [Google Scholar] [CrossRef] [PubMed]
- Sun, T.; Wang, Q.; Balk, S.; Brown, M.; Lee, G.S.; Kantoff, P. The role of microRNA-221 and microRNA-222 in androgen-independent prostate cancer cell lines. Cancer Res. 2009, 69, 3356–3363. [Google Scholar] [CrossRef] [PubMed]
- Sun, T.; Yang, M.; Chen, S.; Balk, S.; Pomerantz, M.; Hsieh, C.L.; Brown, M.; Lee, G. M.; Kantoff, P.W. The altered expression of miR-221/-222 and miR-23b/-27b is associated with the development of human castration resistant prostate cancer. Prostate 2012, 72, 1093–1103. [Google Scholar] [CrossRef] [PubMed]
- Sun, T.; Wang, X.; He, H.H.; Sweeney, C.J.; Liu, S.X.; Brown, M.; Balk, S.; Lee, G.S.; Kantoff, P.W. miR-221 promotes the development of androgen independence in prostate cancer cells via downregulation of HECTD2 and RAB1A. Oncogene 2014, 33, 2790–2800. [Google Scholar] [CrossRef] [PubMed]
- Vickers, A.J.; Elkin, E.B. Decision curve analysis: A novel method for evaluating prediction models. Med. Decis. Making 2006, 26, 565–574. [Google Scholar] [CrossRef] [PubMed]
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).