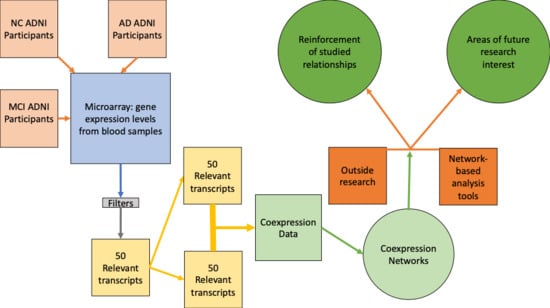
Network Medicine Approach for Analysis of Alzheimer’s Disease Gene Expression Data
Abstract
:1. Introduction
2. Results
3. Discussion
3.1. Network Medicine Applied to Gene Expression Data
3.2. Highly Disrupted Genes
3.3. Notable Connections and Clusters
3.3.1. Y-Linked Regulators
3.3.2. Immune-System Involved
3.3.3. Under Expressed Regulatory Elements
3.3.4. Folate Receptors
3.3.5. Unidentified Transcripts
4. Materials and Methods
4.1. Alzheimer’s Disease Neuroimaging Initiative
4.2. Network Medicine Applied to AD Gene Expression Data
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| AD | Alzheimer’s disease |
| ADNI | Alzheimer’s Disease Neuroimaging Initiative |
| CBS | Cystathionine beta synthase |
| FDR | False discovery rate |
| HP | Haptoglobin |
| lncRNA | Long non-coding RNA |
| MCI | Mild cognitive impairment |
| MHC | Major Histocompatibility Complex |
| NC | Normal condition |
| NM | Network medicine |
| PTGDS | Prostaglandin D2 synthase |
| XIST | X-inactive specific transcript |
References
- Alzheimer’s Association. 2016 Alzheimer’s disease facts and figures. Alzheimer’s Dement. 2016, 12, 459–509. [Google Scholar] [CrossRef]
- Hebert, L.E.; Weuve, J.; Scherr, P.A.; Evans, D.A. Alzheimer disease in the United States (2010–2050) estimated using the 2010 census. Neurology 2013, 80, 1778–1783. [Google Scholar] [CrossRef] [Green Version]
- Weiner, M.W.; Veitch, D.P.; Aisen, P.S.; Beckett, L.A.; Cairns, N.J.; Green, R.C.; Harvey, D.; Jack, C.R., Jr.; Jagust, W.; Morris, J.C.; et al. Recent publications from the Alzheimer’s Disease Neuroimaging Initiative: Reviewing progress toward improved AD clinical trials. Alzheimer’s Dement. 2017, 13, e1–e85. [Google Scholar] [CrossRef]
- Loscalzo, J.; Barabasi, A.L.; Silverman, E.K. Network Medicine: Complex. Systems in Human Disease and Therapeutics; Harvard University Press: Cambridge, MA, USA, 2017. [Google Scholar]
- Barabasi, A.L.; Gulbahce, N.; Loscalzo, J. Network medicine: A network-based approach to human disease. Nat. Rev. Genet. 2011, 12, 56–68. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Jarrell, J.T.; Gao, L.; Cohen, D.S.; Huang, X. Network Medicine for Alzheimer’s Disease and Traditional Chinese Medicine. Molecules 2018, 23, 1143. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hu, Y.S.; Xin, J.; Hu, Y.; Zhang, L.; Wang, J. Analyzing the genes related to Alzheimer’s disease via a network and pathway-based approach. Alzheimer’s Res. Ther. 2017, 9, 29. [Google Scholar] [CrossRef] [PubMed]
- Hu, W.; Lin, X.; Chen, K. Integrated analysis of differential gene expression profiles in hippocampi to identify candidate genes involved in Alzheimer’s disease. Mol. Med. Rep. 2015, 12, 6679–6687. [Google Scholar] [CrossRef] [PubMed]
- Silverman, E.K.; Loscalzo, J. Network medicine approaches to the genetics of complex diseases. Discov. Med. 2012, 14, 143–152. [Google Scholar]
- Hosp, F.; Vossfeldt, H.; Heinig, M.; Vasiljevic, D.; Arumughan, A.; Wyler, E.; Genetic and Environmental Risk for Alzheimer’s Disease GERAD1 Consortium; Landthaler, M.; Hubner, N.; Wanker, E.E.; et al. Quantitative interaction proteomics of neurodegenerative disease proteins. Cell Rep. 2015, 11, 1134–1146. [Google Scholar] [CrossRef]
- Kitsak, M.; Sharma, A.; Menche, J.; Guney, E.; Ghiassian, S.D.; Loscalzo, J.; Barabasi, A.L. Tissue Specificity of Human Disease Module. Sci. Rep. 2016, 6, 35241. [Google Scholar] [CrossRef]
- Siavelis, J.C.; Bourdakou, M.M.; Athanasiadis, E.I.; Spyrou, G.M.; Nikita, K.S. Bioinformatics methods in drug repurposing for Alzheimer’s disease. Brief. Bioinform. 2016, 17, 322–335. [Google Scholar] [CrossRef] [PubMed]
- Mostafavi, S.; Gaiteri, C.; Sullivan, S.E.; White, C.C.; Tasaki, S.; Xu, J.; Taga, M.; Klein, H.U.; Patrick, E.; Komashko, V.; et al. A molecular network of the aging human brain provides insights into the pathology and cognitive decline of Alzheimer’s disease. Nat. Neurosci. 2018, 21, 811–819. [Google Scholar] [CrossRef] [PubMed]
- Song, I.U.; Kim, Y.D.; Chung, S.W.; Cho, H.J. Association between serum haptoglobin and the pathogenesis of Alzheimer’s disease. Intern. Med. 2015, 54, 453–457. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhu, C.; Jiang, G.; Chen, J.; Zhou, Z.; Cheng, Q. Serum haptoglobin in Chinese patients with Alzheimer’s disease and mild cognitive impairment: A case-control study. Brain Res. Bull. 2018, 137, 301–305. [Google Scholar] [CrossRef]
- Yang, H.; Wang, H.; Wang, Y.; Addorisio, M.; Li, J.; Postiglione, M.J.; Chavan, S.S.; Al-Abed, Y.; Antoine, D.J.; Andersson, U.; et al. The haptoglobin beta subunit sequesters HMGB1 toxicity in sterile and infectious inflammation. J. Intern. Med. 2017, 282, 76–93. [Google Scholar] [CrossRef]
- Holmes, C. Review: Systemic inflammation and Alzheimer’s disease. Neuropathol. Appl. Neurobiol. 2013, 39, 51–68. [Google Scholar] [CrossRef]
- Corrada, M.M.; Kawas, C.H.; Hallfrisch, J.; Muller, D.; Brookmeyer, R. Reduced risk of Alzheimer’s disease with high folate intake: The Baltimore Longitudinal Study of Aging. Alzheimer’s Dement. 2005, 1, 11–18. [Google Scholar] [CrossRef] [Green Version]
- Sonkar, S.K.; Kumar, S.; Singh, N.K.; Tandon, R. Hyperhomocysteinemia induced locked-in syndrome in a young adult due to folic acid deficiency. Nutr. Neurosci. 2019. [Google Scholar] [CrossRef]
- Loera-Valencia, R.; Goikolea, J.; Parrado-Fernandez, C.; Merino-Serrais, P.; Maioli, S. Alterations in cholesterol metabolism as a risk factor for developing Alzheimer’s disease: Potential novel targets for treatment. J. Steroid Biochem. Mol. Biol. 2019, 190, 104–114. [Google Scholar] [CrossRef]
- Beyer, K.; Lao, J.I.; Carrato, C.; Rodriguez-Vila, A.; Latorre, P.; Mataro, M.; Llopis, M.A.; Mate, J.L.; Ariza, A. Cystathionine beta synthase as a risk factor for Alzheimer disease. Curr. Alzheimer Res. 2004, 1, 127–133. [Google Scholar] [CrossRef]
- McCarty, M.; O’Keefe, J.; DiNicolantonio, J. A diet rich in taurine, cysteine, folate B12 and betaine may lessen risk for Alzheimer’s disease by boosting brain synthesis of hydrogen sulfide. Med. Hypotheses 2019, 132, 109356. [Google Scholar] [CrossRef] [PubMed]
- Shi, M.T.; Zhang, Y.; Zhou, G.Q. The critical roles of TBC proteins in human diseases. Yi Chuan Hered. 2018, 40, 12–21. [Google Scholar]
- Tam, K.T.; Chan, P.K.; Zhang, W.; Law, P.P.; Tian, Z.; Fung Chan, G.C.; Philipsen, S.; Festenstein, R.; Tan-Un, K.C. Identification of a novel distal regulatory element of the human Neuroglobin gene by the chromosome conformation capture approach. Nucleic Acids Res. 2017, 45, 115–126. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gray, K.; Stefen, H.; Ly, T.; Keller, C.; Colpan, M.; Wayman, G.; Pate, E.; Fath, T.; Kostyukova, A. Tropomodulin’s actin-binding abilities are required to modulate dendrite development. Front. Mol. Neurosci. 2018, 11, 357. [Google Scholar] [CrossRef] [PubMed]
- Fagerberg, L.; Hallstrom, B.M.; Oksvold, P.; Kampf, C.; Djureinovic, D.; Odeberg, J.; Habuka, M.; Tahmasebpoor, S.; Danielsson, A.; Edlund, K.; et al. Analysis of the human tissue-specific expression by genome-wide integration of transcriptomics and antibody-based proteomics. Mol. Cell Proteom. 2014, 13, 397–406. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Luna, R.E.; Arthanari, H.; Hiraishi, H.; Akabayov, B.; Tang, L.; Cox, C.; Markus, M.A.; Luna, L.E.; Ikeda, Y.; Watanabe, R.; et al. The interaction between eukaryotic initiation factor 1A and eIF5 retains eIF1 within scanning preinitiation complexes. Biochemistry 2013, 52, 9510–9518. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rauschendorf, M.A.; Zimmer, J.; Hanstein, R.; Dickemann, C.; Vogt, P.H. Complex transcriptional control of the AZFa gene DDX3Y in human testis. Int. J. Androl. 2011, 34, 84–96. [Google Scholar] [CrossRef] [Green Version]
- Baarends, W.M.; van der Laan, R.; Grootegoed, J.A. Specific aspects of the ubiquitin system in spermatogenesis. J. Endocrinol. Investig. 2000, 23, 597–604. [Google Scholar] [CrossRef]
- Komura, K.; Yoshikawa, Y.; Shimamura, T.; Chakraborty, G.; Gerke, T.A.; Hinohara, K.; Chadalavada, K.; Jeong, S.H.; Armenia, J.; Du, S.Y.; et al. ATR inhibition controls aggressive prostate tumors deficient in Y-linked histone demethylase KDM5D. J. Clin. Investig. 2018, 128, 2979–2995. [Google Scholar] [CrossRef]
- Chen, X.; Tong, C.; Li, H.; Peng, W.; Li, R.; Luo, X.; Ge, H.; Ran, Y.; Li, Q.; Liu, Y.; et al. Dysregulated Expression of RPS4Y1 (Ribosomal Protein S4, Y-Linked 1) Impairs STAT3 (Signal Transducer and Activator of Transcription 3) Signaling to Suppress Trophoblast Cell Migration and Invasion in Preeclampsia. Hypertension 2018, 71, 481–490. [Google Scholar] [CrossRef]
- Forese, M.G.; Pellegatta, M.; Canevazzi, P.; Gullotta, G.S.; Podini, P.; Rivellini, C.; Previtali, S.C.; Bacigaluppi, M.; Quattrini, A.; Taveggia, C. Prostaglandin D2 synthase modulates macrophage activity and accumulation in injured peripheral nerves. Glia 2020, 68, 95–110. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bulow, S.; Zeller, L.; Werner, M.; Toelge, M.; Holzinger, J.; Entzian, C.; Shubert, T.; Waldow, F.; Gisch, N.; Hammerschmidt, S. Bactericidal/Permeability-increasing Protein is an Enhancer of Bacterial Lipoprotein Recognition. Front. Immunol. 2018, 9, 2768. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Brice, D.; Diamond, G. Antiviral Activities of Human Host Defense Peptides. Curr. Med. Chem. 2019. [Google Scholar] [CrossRef] [PubMed]
- Huntoon, K.; Wang, Y.; Eppolito, C.; Barbour, K.; Berger, F.; Shrikant, P.; HBaumann, H. The acute phase protein haptoglobin regulates host immunity. J. Leukoc. Biol. 2008, 84, 170–181. [Google Scholar] [CrossRef]
- McGeer, P.; Akiyama, H.; Itagaki, S.; McGeer, E. Immune system response in Alzheimer’s disease. Can. J. Neurol. Sci. 1989, 16, 516–527. [Google Scholar] [CrossRef] [Green Version]
- Jiang, J.; Natarajan, K.; Marguilies, D. MHC molecules, T cell receptors, natural killer cell receptors, and viron immunoevasins-key elements of adaptive and innate immunity. Adv. Exp. Med. Biol. 2019, 1172, 21–62. [Google Scholar]
- Loda, A.; Heard, E. Xist RNA in action: Past, present, and future. PLoS Genet. 2019, 15, e1008333. [Google Scholar] [CrossRef] [Green Version]
- Tang, L.; Liu, L.; Li, G.; Jiang, P.; Wang, Y.; Li, J. Expression Profiles of Long Noncoding RNAs in Intranasal LPS-Mediated Alzheimer’s Disease Model in Mice. Biomed. Res. Int. 2019, 2019, 9642589. [Google Scholar] [CrossRef] [Green Version]
- Rondinelli, B.; Rosano, D.; Antonini, E.; Frenquelli, M.; Montanini, L.; Huang, D.; Segalla, S.; Yoshihara, K.; Amin, S.; Lazarevic, D.; et al. Histone demethylase JARID1C inactivation triggers genomic instability in sporadic renal cancer. J. Clin. Investig. 2015, 125, 4625–4637. [Google Scholar] [CrossRef] [Green Version]
- LaSalle, J.M.; Powell, W.T.; Yasui, D.H. Epigenetic layers and players underlying neurodevelopment. Trends Neurosci. 2013, 36, 460–470. [Google Scholar] [CrossRef] [Green Version]
- Jack, C.R., Jr.; Bernstein, M.A.; Fox, N.C.; Thompson, P.; Alexander, G.; Harvey, D.; Borowski, B.; Britson, P.J.; Whitwell, J.L.; Ward, C.; et al. The Alzheimer’s Disease Neuroimaging Initiative (ADNI): MRI methods. J. Magn. Reson. Imaging 2008, 27, 685–691. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Weiner, M.W.; Veitch, D.P.; Aisen, P.S.; Beckett, L.A.; Cairns, N.J.; Cedarbaum, J.; Green, R.C.; Harvey, D.; Jack, C.R.; Jagust, W.; et al. 2014 Update of the Alzheimer’s Disease Neuroimaging Initiative: A review of papers published since its inception. Alzheimer’s Dement. 2015, 11, e1–e120. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Saykin, A.J.; Shen, L.; Yao, X.; Kim, S.; Nho, K.; Risacher, S.L.; Ramanan, V.K.; Foroud, T.M.; Faber, K.M.; Sarwar, N.; et al. Genetic studies of quantitative MCI and AD phenotypes in ADNI: Progress, opportunities, and plans. Alzheimer’s Dement. 2015, 11, 792–814. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mar, J.C.; Matigian, N.A.; Mackay-Sim, A.; Mellick, G.D.; Sue, C.M.; Silburn, P.A.; McGrath, J.J.; Quackenbush, J.; Wells, C.A. Variance of gene expression identifies altered network constraints in neurological disease. PLoS Genet. 2011, 7, e1002207. [Google Scholar] [CrossRef] [Green Version]
- Shannon, P.; Markiel, A.; Ozier, O.; Baliga, N.S.; Wang, J.T.; Ramage, D.; Amin, N.; Schwikowski, B.; Ideker, T. Cytoscape: A software environment for integrated models of biomolecular interaction networks. Genome Res. 2003, 13, 2498–2504. [Google Scholar] [CrossRef]
- Van Landeghem, S.; Van Parys, T.; Dubois, M.; Inze, D.; Van de Peer, Y. Diffany: An ontology-driven framework to infer, visualise and analyse differential molecular networks. BMC Bioinform. 2016, 17, 18. [Google Scholar] [CrossRef] [Green Version]
- Aguirre-Plans, J.; Pinero, J.; Menche, J.; Sanz, F.; Furlong, L.I.; Schmidt, H.; Oliva, B.; Guney, E. Proximal Pathway Enrichment Analysis for Targeting Comorbid Diseases via Network Endopharmacology. Pharmaceuticals 2018, 11, 61. [Google Scholar] [CrossRef] [Green Version]





| Key | Gene | Expression in MCI 2 | Expression in AD |
|---|---|---|---|
| 1 | [RPS4Y1] Ribosomal protein S4 Y-linked 1 Probe set: 11716411_x_at | Up | Up |
| 2 | [EIF1AY] eukaryotic translation initiation factor 1A Y-linked Probe set: 11720807_x_at | Up | Up |
| 3 | [GATA2] GATA binding protein 2 Probe set: 11722761_a_at | Down | Down p = 0.203 |
| 4 | [DDX3Y] DEAD (Asp-Glu-Ala-Asp) box helicase 3 Y-linked Probe set: 11724075_a_at | Up | Up |
| 5 | [HLA-DQA1] Major histocompatibility complex class II DQ alpha 1 Probe set: 11724799_x_at | Up | Up p = 0.694 |
| 6 | [USP9Y] ubiquitin specific peptidase 9 Y-linked Probe set: 11725294_at | Up | Up p = 0.194 |
| 7 | [KDM5D] lysine (K)-specific demethylase 5D Probe set: 11726813_a_at | Up | Up |
| 8 | [KDM5D] lysine (K)-specific demethylase 5D Probe set: 11726814_x_at | Up | Up |
| 9 | [TBC1D22B] TBC1 domain family member 22B Probe set: 11728078_a_at | Up p = 0.670 | Down |
| 10 | [BPI] Bactericidal/permeability-increasing protein Probe set: 11729344_at | Up p = 0.225 | Up |
| 11 | [ANKRD22] Ankyrin repeat domain 22 Probe set: 11732425_at | Up p = 0.334 | Up |
| 12 | [TMOD1] Tropomodulin 1 Probe set: 11732501_a_at | Down p = 0.587 | Down |
| 13 | [DEFA4] Defensin alpha 4 corticostatin Probe set:11732546_at | Up p = 0.293 | Up |
| 14 | [HP] Haptoglobin Probe set: 11733829_x_at | - p = 0.991 | Up |
| 15 | [DDX3Y] DEAD (Asp-Glu-Ala-Asp) box helicase 3 Y-linked Probe set: 11734664_x_at | Up | Up |
| 16 | [OSBP2] Oxysterol binding protein 2 Probe set: 11736205_a_at | Down p = 0.473 | Down |
| 17 | [FCRL1] Fc receptor-like 1 Probe set: 11736882_a_at | Down p = 0.641 | Down |
| 18 | [FCRL1] Fc receptor-like 1 Probe set: 11736883_x_at | Down p = 0.694 | Down |
| 19 | [FAM46C] Family with sequence similarity 46 member C Probe set: 11739338_at | Down p = 0.531 | Down |
| 20 | [OR2W3] (Locus via Non-standard RNA) olfactory receptor family 2 subfamily W member 3 Probe set: 11741636_at | - p = 0.843 | Down |
| 21 | [CACNG6] Calcium channel voltage-dependent gamma subunit 6 Probe set: 11742124_a_atex | Down | Down p = 0.616 |
| 22 | [FOLR3] folate receptor 3 (gamma) Probe set: 11744140_a_at | Up p = 0.645 | Up p = 0.163 |
| 23 | [FOLR3] folate receptor 3 (gamma) Probe set: 11744141_x_at | Up p = 0.582 | Up p = 0.137 |
| 24 | [CBS] Cystathionine-beta-synthase Probe set: 11744286_s_at | - p = 0.995 | Up |
| 25 | [HP] Haptoglobin Probe set: 11744649_x_at | - p = 0.76 | Up p = 0.141 |
| 26 | [CBS] Cystathionine-beta-synthase Probe set: 11744835_s_at | - p = 0.995 | Up |
| 27 | [KDM5D] lysine (K)-specific demethylase 5D Probe set: 11745012_a_at | Up | Up |
| 28 | [HLA-DQB1] Major histocompatibility complex class II DQ beta 1 Probe set: 11746804_x_at | Up | Up p = 0.46 |
| 29 | [DDX3Y] DEAD (Asp-Glu-Ala-Asp) box helicase 3 Y-linked Probe set: 11748424_x_at | Up | Up |
| 30 | [DDX3Y] DEAD (Asp-Glu-Ala-Asp) box helicase 3 Y-linked Probe set: 11749841_x_at | Up | Up |
| 31 | [HLA-DQA1] Major histocompatibility complex class II DQ alpha 1 Probe set: 11750528_x_at | Up | - p = 0.89 |
| 32 | [ENSG00000211625 || ENSG00000239951] (Matches 2 Loci; Matches Ensembl Gene) Putative uncharacterized protein ENSP00000374805 [Source:UniProtKB/TrEMBL;Acc:A6NLY3] || Ig kappa chain V-III region HAH Precursor [Source:UniProtKB/Swiss-Prot;Acc:P18135] Probe set: 11753832_x_at | Down p = 0.123 | Down |
| 33 | [XIST] X inactive specific transcript (non-protein coding) Probe set: 11754194_s_at | Down | Down |
| 34 | [EGR1] Early growth response 1 Probe set: 11754334_s_at | Down | Down |
| 35 | [NUDT4 || NUDT4P2 || NUDT4P1] (Matches 3 Loci) Nudix (Nucleoside diphosphate linked moiety X)-type motif 4 || nudix (nucleoside diphosphate linked moiety X)-type motif 4 pseudogene 2 || Nudix (nucleoside diphosphate linked moiety X)-type motif 4 pseudogene 1 Probe set: 11754453_s_at | - p = 0.846 | Down |
| 36 | [SHISA4] shisa family member 4 Probe set: 11756240_a_at | Down p = 0.641 | Down |
| 37 | [PTGDS] prostaglandin D2 synthase 21kDa (brain) Probe set: 11756587_a_at | - p = 0.853 | Up |
| 38 | [HP] Haptoglobin Probe set: 11757277_x_at | - p = 0.932 | Up |
| 39 | [XIST] X inactive specific transcript (non-protein coding) Probe set: 11757733_s_at | Down | Down |
| 40 | [XIST] X inactive specific transcript (non-protein coding) Probe set: 11757857_s_at | Down | Down |
| 41 | [TRIM10] tripartite motif containing 10 Probe set: 11758611_s_at | Up p = 0.647 | Down |
| 42 | (Matches Non-standard RNA) JARID1C protein (JARID1C) mRNA complete cds alternatively spliced Probe set: 11761133_at | Down | Down |
| 43 | [HLA-DQB1] (POOR HIT 44%) Major histocompatibility complex class II DQ beta 1 Probe set: 11762641_x_at | Up | Up p = 0.56 |
| 44 | (DEPRECATED TARGET; Matches RefSeq) (Deprecated) PREDICTED: Homo sapiens similar to hCG2042707 (LOC650405) || (Deprecated) PREDICTED: Homo sapiens similar to pre-B lymphocyte gene 1 (LOC652493) || (Deprecated) PREDICTED: Homo sapiens similar to hCG26659 (LOC100291464) Probe set: 11763222_x_at | Down p = 0.316 | Down |
| 45 | [ENSG00000211663] (Matches Ensembl Gene) V2-13 protein Fragment [Source:UniProtKB/TrEMBL; Acc:Q5NV73] Probe set: 11763229_x_at | Down p = 0.331 | Down |
| 46 | [ENSG00000242534 || ENSG00000244116] (Matches 2 Loci; Matches Ensembl Gene) immunoglobulin kappa variable 2D-28 [ENST00000453166 ENST00000558026] || immunoglobulin kappa variable 2-28 [ENST00000482769] | Up p = 0.614 | Down p = 0.122 |
| 47 | [ENSG00000211663] (Matches Ensembl Gene) V2-13 protein Fragment [Source:UniProtKB/TrEMBL;Acc:Q5NV73] Probe set: 11763255_x_at | Down p = 0.279 | Down |
| 48 | [ENSG00000211663] (Matches Ensembl Gene) V2-13 protein Fragment [Source:UniProtKB/TrEMBL;Acc:Q5NV73] Probe set: 11763551_x_at | Down p = 0.332 | Down |
| 49 | (Matches Non-standard RNA) mRNA; cDNA DKFZp686L12190 (from clone DKFZp686L12190): Probe set: 11763837_s_at | Up | Up |
| 50 | [TXLNG2P] (Matches Ensembl Gene) Uncharacterized protein CYorf15B (Lipopolysaccaride-specific response 5-like protein) [Source:UniProtKB/Swiss-Prot;Acc:Q9BZA4] Probe set: 11764064_s_at | Up | Up |
| Gene | DyNet Rewiring Score |
|---|---|
| [HP] Haptoglobin Probe set: 11757277_x_at | 8.33 |
| [FOLR3] folate receptor 3 (gamma) Probe set: 11744141_x_at | 8.00 |
| [FOLR3] folate receptor 3 (gamma) Probe set: 11744140_a_at | 8.00 |
| [NUDT4 || NUDT4P2 || NUDT4P1] (Matches 3 Loci) Nudix (Nucleoside diphosphate linked moiety X)-type motif 4 || nudix (nucleoside diphosphate linked moiety X)-type motif 4 pseudogene 2 || Nudix (nucleoside diphosphate linked moiety X)-type motif 4 pseudogene 1HP Probe set: 11754453_s_at | 7.00 |
| [HP] Haptoglobin Probe set: 11744649_x_at | 7.00 |
| [HP] Haptoglobin Probe set: 11733829_x_at | 7.00 |
| [SHISA4] shisa family member 4 Probe set: 11756240_a_at | 6.67 |
| [CBS] Cystathionine-beta-synthase Probe set: 11744835_s_at | 6.00 |
| [TBC1D22B] TBC1 domain family member 22B Probe set: 11728078_a_at | 5.67 |
| [CBS] Cystathionine-beta-synthase Probe set: 11744286_s_at | 5.67 |
| Gene | DyNet Rewiring Score |
|---|---|
| [GATA2] GATA binding protein 2 Probe set: 11722761_a_at | 8.00 |
| [PTGDS] prostaglandin D2 synthase 21kDa (brain) Probe set: 11756587_a_at | 6.33 |
| [ANKRD22] Ankyrin repeat domain 22 Probe set: 11732425_at | 6.33 |
| [SHISA4] shisa family member 4 Probe set: 11756240_a_at | 5.67 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Cohen, D.; Pilozzi, A.; Huang, X. Network Medicine Approach for Analysis of Alzheimer’s Disease Gene Expression Data. Int. J. Mol. Sci. 2020, 21, 332. https://doi.org/10.3390/ijms21010332
Cohen D, Pilozzi A, Huang X. Network Medicine Approach for Analysis of Alzheimer’s Disease Gene Expression Data. International Journal of Molecular Sciences. 2020; 21(1):332. https://doi.org/10.3390/ijms21010332
Chicago/Turabian StyleCohen, David, Alexander Pilozzi, and Xudong Huang. 2020. "Network Medicine Approach for Analysis of Alzheimer’s Disease Gene Expression Data" International Journal of Molecular Sciences 21, no. 1: 332. https://doi.org/10.3390/ijms21010332